Abstract
The B-box proteins are a class of zinc finger transcription factors and play important roles in regulating plant growth, development, and stress response. However, their origin and expansion model in plants have been very vague. In this study, 208 B-box genes were identified in 18 plant species, and phylogenetic analyses divided them into five structure groups. Subsequently, the sequence analysis including theoretical isoelectric point, instability index, and aliphatic index showed the wide variation of B-box gene in different species. Our multispecies genome-wide analysis reveals that the structure groups III and IV in the plant BBX gene family have the earliest origin (Rhodophyta) and are extensively expanded in land plants, while the other three structure groups (I, II, and V) seem to originate at least in the last common ancestor of land plants. Furthermore, whole genome duplication (WGD) was the main driver (28 gene pairs, 65.12%) of the B-box gene family expansion, followed by segmental duplication, which tend to have more introns and are subject to more intense purification selections. We also analyzed the sequence differences between B-box domains to propose a new evolutionary model of B-box domain. These analyses provide new insights for understanding the origin and evolution of the B-box gene family.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The B-box (BBX) proteins belong to zinc finger transcription factors and have been reported to participate in regulating plant growth and development. Specifically, B-box (BBX) protein regulation of plant development is mostly related to photoperiod control in plants (Onouchi et al. 2000; Samach 2000). For example, CO/AtBBX1, COL3/AtBBX4, COL9/AtBBX7 with double B-box, and CCT domains in Arabidopsis have been proved to control and regulate the flowering process by participating in the photoperiod (Crocco and Botto 2013). The mutant of CO/AtBBX1 only flowered in long days, while Arabidopsis thaliana flowered in both long and short days when it was overexpressed (Suárez-López et al. 2001). In addition, AtBBX proteins with only the B-box domains also participates in the control of plant growth through light factors. Some BBX proteins also act as assistants to other members, such as AtBBX32, which has been shown to be a transcriptional regulator of STH/AtBBX21 in the light signaling pathway (Huang et al. 2012). Furthermore, BBX genes in Arabidopsis and multiple species have been reported to play an important role in carotenoid biosynthesis (Xiong et al. 2019), biological, and abiotic stress response (Ding et al. 2018; Soitamo et al. 2008).
Despite the BBX genes have been characterized in many plants, their origin and evolution in the plant kingdom have not been thoroughly studied. The article of Crocco and Botto put a spotlight on a recent study that reported the identification of the BBX gene in green algae and believed that the earliest BBX protein in photosynthetic organisms originated about 1 billion years ago (Crocco and Botto 2013). The BBX gene family in red algae has not been characterized, possibly due to the lack of genomic data. The availability of more plant genome data provides the possibility to further study the origin of this gene family. Besides, gene duplication usually plays an important role in the expansion of gene families (Wang et al. 2018b). As previously reported, tandem duplication made important contributions to the expansion of gene families in Arabidopsis, tomato, rice and other species (Huang et al. 2015; Liu et al. 2018; Liu and Widmer 2014; Quan et al. 2019). However, the driving force for the expansion of the BBX gene family is still very vague.
In the current study, we investigated and analyzed BBX proteins from 18 representative plants (including red algae, green algae, moss, ferns, gymnosperm, and angiosperms) to understand the origin and evolution of the BBX gene family in the plant kingdom. Furthermore, the huge differences in the gene structure and molecular properties of BBX genes indicate the wide diversity of members of this gene family. We also analyzed the duplication models of the BBX gene through collinearity analysis within the genome to determine the main driving force for its expansion, as well as the gene structure and selection pressure characteristics of different duplication models. Finally, through the analysis and comparison of the B-box domain sequences of the BBX genes, a possible evolution model for them is proposed. In particular, the analysis of sea buckthorn BBX gene family in this article was first reported. This study provides new insights for further clarifying the origin and expansion of the BBX gene family in plants.
Methods
Gene identification
The whole genome data of 18 species were downloaded from the corresponding database or website (Table S1), and the protein sequence data were processed to only retain the longest transcript of each gene. The protein sequences of the 32 BBX gene family members of Arabidopsis have been reported and they were used as BLAST queries to search against the protein sequences of 17 species using default parameters (Khanna et al. 2009). Then, the Hidden Markov Model (HMM) file (PF00643) (http://pfam.sanger.ac.uk/) of B-box domain was used to search against the all proteins of 17 species with default parameters using HMMER 3.0. We also used the NCBI Batch CD-Search tools to check all members containing the BBX core domain. Those with exactly the same sequence in the same species are considered the same gene. Finally, a total of 208 BBX gene family members including the 32 BBX gene family members of Arabidopsis were identified after comprehensive curation. The three-dimensional structure of some BBX proteins were predicted to more rigorously speculate its origin. Specifically, the predicted protein structure models of A. thaliana protein were download from SWISS-MODEL Repository (SMR). The construction of homologs structure model in other plants selected the template used in A. thaliana protein homology prediction as template. The three-dimensional structure models were verified using the Qualitative Model Energy Analysis (QMEAN) and VERIFY 3D programs available from the Structural Analysis and Verification Sever v 6.0 (SAVES) (https://saves.mbi.ucla.edu/SAVES) (Table S2). We used the tools from ExPasy website (http://web.expasy.org/protparam/) to obtain protein characteristic information such as isoelectric point (pI), aliphatic index, instability index, grand average of hydropathicity (GRAVY), and the subcellular localization (Table S3).
Phylogenetic and sequence analysis
The BBX proteins from 18 plants were used for multiple alignments by ClustalW program. All gene accession of BBX genes used in this study are listed in Table S3. We used neighbor-joining method of MEGA7.0 to construct unrooted phylogenetic tree. The MEME online program (http://meme.nbcr.net/meme/intro.html) was used to determine the conserved motifs in BBX proteins. We obtained the exon–intron organization of BBX genes from the corresponding Generic Feature Format Version 3 (GFF3) files of each species using TBtools (Chen et al. 2020). As some BBX proteins have a double B-box domain, we used the following criteria to designate them: the first B-box that appeared within the protein in the N-terminal position was called B1 and the second B-box termed B2 (Crocco and Botto 2013).
Renaming of BBX gene family members and gene duplication
We obtain the position of each BBX proteins on the chromosome or scaffold from the gff3 file of each species, and rename them according to its position order (Table S3). Multiple Collinearity Scan toolkit (MCScanX) (Wang et al. 2012) was adopted to analyze the synteny and the dot-plot of collinear homologous genes in the genomes of seven species is drawn with Python program, which was used to distinguish the duplication models of BBX members (Wang et al. 2012). Collinear blocks formed by WGD were generally large fragments and were complementary to each other. In addition, the median of the Ks value of the anchor gene pairs on the collinearity blocks formed by a certain WGD is generally close to form the Ks peak corresponding to the WGD. Based on the above characteristics, we confirmed the collinearity blocks formed by WGD, and the B-box genes located on these blocks were naturally formed by WGD. Furthermore, the results of collinearity analysis performed by MCScanX clarified the tandem duplication genes. We also used the downstream analysis program (add_ka_and_ks_to_collinearity.pl) of MCScanX to calculate the non-synonymous (Ka) and synonymous (Ks) substitutions of each duplicated gene pair (Wang et al. 2012).
Results
Genome-wide identification and global phylogenetic analysis of BBX proteins
We obtained 208 no-redundant BBX genes in 18 plants. Based on their location on the scaffolds or chromosomes, we renamed the BBX genes (Table S3). The molecular mass and length of BBX proteins varied greatly, with MWs ranging from 7.16 to 154.93 KDa and length from 66 to 1493 aa. The theoretical pI was ranging from 4.22 to 9.36, showing greater variation. The instability index of most (192 of 208) BBX proteins were more than 40, and they were considered unstable. Only two BBX proteins were considered non-hydrophilic because the high GRAVY value (> 0) and others were considered hydrophilic. The aliphatic indices of BBX proteins range from 47.66 to 88.79. Overall, the huge differences in molecular properties of BBX genes indicated the wide diversity of members of this gene family.
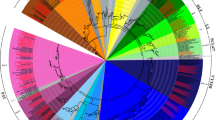
The full-length BBX proteins of 18 plants were used to perform phylogenetic analysis to investigate the classifications of BBX proteins. According to the topology structure of the phylogenetic tree and classification of AtBBX in a previous research, all BBX proteins were classified to five structure groups (Fig. 1). Structure groups I and II comprised a total of 64 BBX proteins, and most of them with a CCT domain and two B-box domains. A total of 38 BBX proteins were classified into structure group III and usually contained a single B-box and CCT domains. Structure group IV contained the largest number of BBX proteins (80), most of which had two B-box domains. Only 26 BBX proteins were classified into structure group V, and they tend to contain a single B-box domain (B1). The classification of BBX proteins through phylogenetic analysis was not completely consistent with their domains. For example, both OsBBX2 (from Oryza sativa) and PpBBX13 (from Physcomitrella patens) had only one B-box domain, although they were classified into structure group II. AzBBX3 (from Azolla filiculoides) was classified into structure group IV, but contains a single B-box domain. These results suggested that BBX gene family members belonging to a structure group may have lost a B-box domain during evolutionary events.
Generally, highly similar sequences often have highly similar structures and functions. We speculated that members with the same domain and classified into a structure group are more likely to have similar functions. For example, AtBBX1 is a photoperiodic flowering-time protein in which the overexpression induces early flowering, and mutations delay flowering (Crocco and Botto 2013), so we think that HrBBX21 may play a role in the flowering photoperiod of sea buckthorn.
Origin and evolution of BBX gene family in plant
We analyzed the collected dataset of the 18 species to gain a comprehensive insight into the evolutionary history of BBX genes. Among the 208 non-redundant BBX genes identified, the most primitive member appeared in the red algae C. crispus (Fig. 2a). The BBX genes showed a large increase in the number of land plants, and all five structure group appeared in land plants (Fig. 2b), and this may be related to the fact that BBX participates in the control of plant growth and development by regulating the photoperiod, which has been widely reported. Since the BBX gene was not identified in two red algae C. merolae and G. sulphuraria, and the CcBBX1/CcBBX2 identified in C. crispus have significant sequence and structure similarities with the AtBBX11/AtBBX4 in A. thaliana (Back bone: Root Mean Squared Deviations [RMSD] = 0.09/1.58 Å; All atoms: RMSD = 0.65/1.82 Å) (Fig. 2c), it is reasonable to speculate that the BBX gene family originated from the Rhodophyta (Fig. 2a). In fact, the above results also indicated that structures groups III and IV originated from the Rhodophyta. In P. patens and M. polymorpha, we identified PpBBX8 belonging to the structure group I and MpBBX2 belonging to the structure group V. The predicted three-dimensional structure of PpBBX8 was significantly similar to the predicted three-dimensional structure of AtBBX30 belonging to the structure group I, and this result also appeared in MpBBX2 and AtBBX31 belonging to the V structure group. The liverwort M. polymorpha was used as the most basal plant linage, because liverworts are the sister to all other land plants and represent the earliest branching land plant lineage (Qiu et al. 2006; Wang et al. 2015). These lines of evidence suggested that structure group V was likely to originate at least in the last common ancestor of land plants, and structure group I may originated in moss (Fig. 2a). For structure group II, the CbBBX3 identified in C. braunii had poor similarity with the members of this group in A. thaliana in the predicted three-dimensional structure. Then, the MpBBX5 we identified in M. polymorpha showed high structural similarity with the MpBBX5 of the group of A. thaliana, which suggested that this structure group may originate from moss.
The origin of BBX genes in plants. a Distribution of the number of BBX genes in 18 plants. b Predicted three-dimensional structure of BBX protein in A. thaliana and some primitive plants. The similarities between the related proteins and the corresponding proteins in Arabidopsis are annotated with Root Mean Squared Deviations (RMSD of back bone and all atoms)
Gene structure and conserved motif analysis
The genetic structure of BBX genes was examined to gain a deeper understanding of its evolution (Fig. S1). The BBX genes in structure group II, structure group III, and structure group V had less introns (no more than four introns), while that in structure group I and IV had more introns. All BBX proteins had one or two B-box domains and some members had CCT conserved domain. There were extremely long intron regions in some BBX genes. The BBX genes in different species showed strong conservation in the domains, but there were obvious variations in gene length and gene structure.
The conserved motif composition of all BBX proteins was displayed based on the motif analysis from MEME (Fig. S2). Similar motif compositions were detected in the same group, and some motifs were distributed in all groups. For example, each member of structure group I had motif 3 and there was no motif 2 in each members of structure group I and III. In addition, motifs 1 almost distributed in all BBX genes. The compositions of these motifs were generally consistent with the results of phylogenetic analysis, but their roles in plant growth and development need to be further studied.
BBX gene duplication models and characteristics analysis
To determine the main driving force of BBX gene expansion, the duplication model of BBX genes was analyzed. Using collinearity analysis in the genomes of seven representative species (genome assembly at the chromosome level) to identify the duplication model of the BBX genes (Figs. 3, S3–S8). We marked the median value of the synonymous substitution site (Ks) of the collinear blocks on the homologous dot-plot, and analyzed the complementary features of them to identify the collinear region originating from WGD. We finally identified the duplication models of BBX genes (Table S4), which showing that WGD (28 gene pairs, 45 genes) made important contributions to the expansion of BBX genes than segmental duplication (15 gene pairs, 26 genes), and no tandem duplication was found in 43 gene pairs (Fig. 4a). These results reasonably suggested that WGD was the main driving force to BBX gene family expansion.
Collinear homologous collinear dot-plot within sea buckthorn genome. The B-box gene names are displayed at the edge. The collinearity blocks formed by the WGD are in green or red boxes in the figure, and the median value of Ks or the range of the median value of Ks of the collinearity blocks are marked
The variations in gene structure, selection pressure of different duplication models. a Number and percentage of different duplication models of BBX genes. b Distribution of duplication genes in structure groups. c Comparison of the number of introns in genes with WGD and segmental duplication models. d Comparison of the Ka/Ks n genes with WGD and segmental duplication models
Among the BBX genes identified to be duplicated by WGD or Segmental, 20 (31.25%) belonged to structure group IV, suggesting that gene duplication had a greater contribution to the expansion of this structure group (Fig. 4b). The above results were consistent with structure group IV with the most BBX gene members (Fig. 1). Of note, many members of this structure group (such as AtBBX1 and AtBBX21) influenced the growth and development of plants by participating in photoperiod regulation, and this structure group was significantly expanded in land plants (Crocco et al. 2010; Onouchi et al. 2000; Samach 2000).
We further paid attention to the variation in gene structure and selection pressure of genes with different duplication models. Surprisingly, genes with segmental duplication had a higher average number of introns than genes with WGD (Fig. 4c). In addition, most gene pairs (41 of 43) with duplication were subject to purification selection (Ka/Ks ratios less than 1) (Table S4), and among them, the gene pairs with segmental duplication has smaller Ka/Ks ratios showing that they were more affected by selection (Fig. 4d). Genes with segmental duplication had more introns and more potential transcripts to perform more functions, which may make them more susceptible to environmental selection to show smaller Ka/Ks.
Evolutionary analysis of BBX and CCT domains
The BBX and CCT domain sequence of all BBX proteins were extracted for a multiple sequence alignment, and some BBX and CCT domain were incomplete (Figs. S9 and S10).{Sakai, 2015 #96}{Sakai, 2015 #96} To analyze the evolution of BBX and CCT more rigorously, we only retained the complete members of the domain sequence and removed the amino acid residues outside the topological structure that had been reported for phylogenetic analysis (Crocco and Botto 2013). Phylogenetic analysis suggested that most of the B-box1/B-box2 domain sequence were very clearly separated but some B-box1 domains were in the clade of B-box2, such as AcBBX12 B-box1 (Fig. 5a). More interestingly, the B-box2 domain sequences were divided into two main clades (I and III) (Fig. 5a), and this implied that the B-box2 domain sequences from the two clades may evolved independently. Multiple sequence alignments of the three clades (I, II, and III) based on the phylogenetic tree were displayed, respectively (Figs. S11–S13). The topological structures of the three clades were different, and the number of conserved sites in the three clades was as follows: III > II > I (Fig. 5b). At the same time, the clades with more conservative sites completely retain the conservative sites with fewer conservative sites clades. These results were consistent with the following ideas: The domain in clade I (still called B2) has an earlier evolutionary origin than the domain in clade II (called B1), and the domain in clade II has an earlier evolutionary origin than the domain in clade III (called B2’), which was also supported by the phylogenetic genetic tree of B-box domains (Fig. 5a). No matter which clade they belong to, the topological structure of the domain generated by cysteine was retained, which indicated that the conservation of the topological structure was extremely important for the molecular functions of this domain. We removed the incomplete CCT domain sequence, and removed the amino acid residues outside the CCT topology for multiple sequence alignment, and found that the core sequence of the CCT domain was roughly divided into three clades (Fig. S14).
Discussion
As the genomes of more and more species are sequenced, it is possible to study the origin and evolution of gene families with important functions. BBX transcription factors play important roles in plant growth and development, such as controlling photomorphogenesis, and have been found in many species (Crocco et al. 2011; Lin et al. 2018; Wang et al. 2013). In the present work, we identified this important transcription factor family in 18 species to explore its origin and evolution.
Significant increase in the number of members of the BBX gene family and all structure groups of the gene family in land plants was identified, which indicated that lots of BBX genes were duplicated and it may play important physiological roles during adaptation of land plants (Crocco and Botto 2013). Gene duplication is an important reason for gene family expansion (Liu et al. 2018). Actually, potential segmental duplication and internal deletion events result in the differences of the consensus sequences and inter space of the zinc-binding residues in the two B-box domains (Crocco and Botto 2013; Massiah et al. 2007). Tandem duplication can be easily identified, but segmental duplication and WGD are extremely difficult to distinguish, as the fragmentation and fusion of chromosomes after WGD. The method of collinearity analysis has been used to analyze the genome duplication of various plants, and this method has been widely used and recognized (Tang et al. 2008; Wang et al. 2017, 2018a; b; Yu et al. 2021). Our comparative genome results suggested that the main driving force for the expansion of the BBX gene family was WGD, even although they may occur millions of years ago. In previous studies, it was found that WGD had made a great contribution to the expansion of kiwi ascorbic acid synthesis-related gene family (Wang et al. 2018b). In contrast, celery and other Apiaceae plants lost a large number of NBS genes after WGD, and had a huge impact on their gene regulatory network (Song et al. 2020). The more particular impact of WGD on the members of the BBX gene family needs to be identified and analyzed from multiple perspectives in more species.
In addition, we identified the BBX genes from structure groups III and IV in C. crispus, but did not identify the BBX genes in the other two red algae, which suggests that the BBX genes in plants may originate from red algae. The three structure groups (I, II, and V) originated from the last common ancestor of terrestrial plants, which is consistent with previous findings that some members of these structure groups participate in the control of plant growth through light and other abiotic stress factors. In fact, the significant increase in the number of BBX gene family members and all structure groups of the gene family in terrestrial plants indicates that a large number of BBX genes are replicated and may play an important physiological role in the adaptation process of terrestrial plants (Gangappa and Botto 2014; Kenrick and Crane 1997).
According to previous reports, the B-box domains of BBX genes in plant are divided into two classes, known as B-box1 (B1) and B-box2 (B2) domains that are recognized on their consensus sequence and the distance between the zinc-binding residues(Khanna et al. 2009). However, previous studies mostly used the following criteria to define B-box1 and B-box2: the first B-box that appears at the N-terminal position in the protein is called B-box1 (B1), and the second B-box is called B-box2 (B2). In other words, they performed evolutionary analysis after subjective classification based on the relative position of the B-box domain rather than the sequence of all B-box domains. In this study, we extracted the topological structure sequence of all complete B-box domains for phylogenetic analysis to further analyze its evolution. Specifically, we removed the incomplete B-box domain sequence and the extra amino acid defects on both sides of the consensus type of B-box domain to avoid their influence. Even so, the B-box1 and B-box2 sequences were not clearly distinguished in phylogenetic tree. Some B-box1 domains were in the clades of B-box2 and some B-box2 domains were in the clades of B-box1. This result required us to further consider the accuracy of the hypothetical model on the evolutionary trajectory of the B-box domain (Crocco and Botto 2013).
Here, we proposed a new hypothetical model of the evolutionary trajectory of the BBX domain in plants (Fig. 6). The early plant BBX protein initially had only one B-box domain, and the B-box domain occurred duplication, mutation, and deletion formed several types of a–h (Fig. 6). Some of these types had added CCT domains in the evolution to correspond to the structure group I–V categories of the Crocco and Botto hypothetical models (Crocco and Botto 2013). Interestingly, three types of conserved domains of B2, B1, and B2' were found in algae, indicating that the B-box domain duplication and mutation happened before plants colonized the land (Table S5). Admittedly, a complete and convincing hypothesis about the evolution of the BBX gene still requires more researchers' efforts.
A hypothetical model of B-box domain evolution. a The new hypothetical model of B-box domain evolution proposed in this article. a–h respectively, represents the categories in the newly proposed B-box domain evolution hypothesis model (the CCT domain is not considered). b B-box gene classification (structure groups I–V) according to the evolutionary hypothesis model proposed by Crocco and Botto
Availability of data and materials
All data analyzed during this study are included in this article and its Online Resource files.
Code availability
Not applicable.
References
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Crocco CD, Botto JF (2013) BBX proteins in green plants: insights into their evolution, structure, feature and functional diversification. Gene 531:44–52. https://doi.org/10.1016/j.gene.2013.08.037
Crocco C, Holm M, Yanovsky MJ, Botto JF (2010) AtBBX21 and COP1 genetically interact in the regulation of shade avoidance. Plant J 64:551–562
Crocco CD, Holm M, Yanovsky MJ, Botto JF (2011) Function of B-BOX under shade. Plant Signal Behav 6:101–104. https://doi.org/10.4161/psb.6.1.14185
Ding L et al (2018) Two B-box domain proteins, BBX18 and BBX23, interact with ELF3 and regulate thermomorphogenesis in arabidopsis. Cell Rep 25:1718-1728 e1714. https://doi.org/10.1016/j.celrep.2018.10.060
Gangappa SN, Botto JF (2014) The BBX family of plant transcription factors. Trends Plant Sci 19:460–470
Huang J, Zhao X, Weng X, Wang L, Xie W, Phan LS (2012) The Rice B-box zinc finger gene family: genomic identification, characterization, expression profiling and diurnal analysis. PLoS ONE 7:e48242
Huang W, Xian Z, Kang X, Tang N, Li Z (2015) Genome-wide identification, phylogeny and expression analysis of GRAS gene family in tomato. BMC Plant Biol 15:209
Kenrick P, Crane PR (1997) The origin and early evolution of plants on land. Nature 389:33–39
Khanna R, Kronmiller B, Maszle DR, Coupland G, Holm M, Mizuno T, Wu SH (2009) The Arabidopsis B-box zinc finger family. Plant Cell 21:3416–3420. https://doi.org/10.1105/tpc.109.069088
Lin F et al (2018) B-BOX DOMAIN PROTEIN28 negatively regulates photomorphogenesis by repressing the activity of transcription factor HY5 and undergoes COP1-mediated degradation. Plant Cell 30:2006–2019. https://doi.org/10.1105/tpc.18.00226
Liu X, Widmer A (2014) Genome-wide comparative analysis of the gras gene family in populus arabidopsis and rice. Plant Mol Biol Rep 32:1129–1145. https://doi.org/10.1007/s11105-014-0721-5
Liu B, Sun Y, Xue J, Jia X, Li R (2018) Genome-wide characterization and expression analysis of GRAS gene family in pepper (Capsicum annuum L.). PeerJ 6:e4796. https://doi.org/10.7717/peerj.4796
Massiah MA, Matts JA, Short KM, Simmons BN, Singireddy S, Yi Z, Cox TC (2007) Solution structure of the MID1 B-box2 CHC(D/C)C(2)H(2) zinc-binding domain: insights into an evolutionarily conserved RING fold. J Mol Biol 369:1–10. https://doi.org/10.1016/j.jmb.2007.03.017
Onouchi H, Igeño MI, Périlleux C, Coupland GG (2000) Mutagenesis of plants overexpressing CONSTANS demonstrates novel interactions among Arabidopsis flowering-time genes. Plant Cell Online 12:885–900
Qiu Y-L et al (2006) The deepest divergences in land plants inferred from phylogenomic evidence. Proc Natl Acad Sci USA 103:15511–15516
Quan SW et al (2019) Genome-wide identification classification, expression and duplication analysis of GRAS Family Genes in Juglans Regia L. Sci Rep 9:11643–11643
Samach A (2000) Distinct roles of CONSTANS target genes in reproductive development of arabidopsis. Science 288:1613–1616
Soitamo AJ, Piippo M, Allahverdiyeva Y, Battchikova N, Aro EM (2008) Light has a specific role in modulating Arabidopsis gene expression at low temperature. BMC Plant Biol 8:13. https://doi.org/10.1186/1471-2229-8-13
Song X et al (2020) The celery genome sequence reveals sequential paleo-polyploidizations, karyotype evolution and resistance gene reduction in apiales. Plant Biotechnol J. https://doi.org/10.1111/pbi.13499
Suárez-López P, Wheatley K, Robson F, Onouchi H, Coupland G (2001) CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nature 410:1116–1120
Tang H, Bowers JE, Wang X, Ming R, Alam M, Paterson AH (2008) Synteny and collinearity in plant genomes. Science 320:486–488. https://doi.org/10.1126/science.1153917
Wang Y et al (2012) MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res 40:e49. https://doi.org/10.1093/nar/gkr1293
Wang Q, Tu X, Zhang J, Chen X, Rao L (2013) Heat stress-induced BBX18 negatively regulates the thermotolerance in Arabidopsis. Mol Biol Rep 40:2679–2688. https://doi.org/10.1007/s11033-012-2354-9
Wang CY, Liu SS, Han GZ (2015) Insights into the origin and evolution of the plant hormone signaling machinery. Plant Physiol 167(3):872
Wang J et al (2017) Hierarchically aligning 10 legume genomes establishes a family-level genomics platform. Plant Physiol 174:284–300. https://doi.org/10.1104/pp.16.01981
Wang J et al (2018a) An overlooked paleotetraploidization in cucurbitaceae. Mol Biol Evol 35:16–26. https://doi.org/10.1093/molbev/msx242
Wang JP et al (2018b) Two Likely auto-tetraploidization events shaped kiwifruit genome and contributed to establishment of the Actinidiaceae family. iScience 7:230–240. https://doi.org/10.1016/j.isci.2018.08.003
Xiong C et al (2019) A tomato B-box protein SlBBX20 modulates carotenoid biosynthesis by directly activating PHYTOENE SYNTHASE 1, and is targeted for 26S proteasome-mediated degradation. New Phytol 221:279–294. https://doi.org/10.1111/nph.15373
Yu L, Zhang G, Lyu Z, He C, Zhang J (2021) Genome-wide analysis of the GRAS gene family exhibited expansion model and functional differentiation in sea buckthorn (Hippophae rhamnoides L.). Plant Biotechnol Rep. https://doi.org/10.1007/s11816-021-00694-1
Acknowledgements
We would like to thank Mrs. H.M. Luo for their assistance with sample collection. We would also like to thank everyone who contributed to this manuscript in various capacities.
Funding
This research was funded by the National Natural Science Foundation of China [U2003116] and the Fundamental Research Funds for the Central Non-profit Research Institution of Chinese Academy of Forestry [No. ZDRIF2019; No. CAFYBB2020SZ001-2].
Author information
Authors and Affiliations
Contributions
CYH and JGZ supervised the research. LYY performed the experiments, analyzed the data, prepared figures and captions, and wrote the manuscript. GYZ and HL collected field samples. ZRL analyzed RNA-seq data. All of the authors have read and approve of the submitted manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11816_2022_745_MOESM6_ESM.pdf
Figure S1 Gene structure of B-box genes. a The phylogenetic tree was constructed based on the full-length sequences of B-box proteins using MEGA 7 software. Details of clusters are shown in different colors. b Gene structure of B-box genes. Black line indicates intron, boxes of different colors are used to represent UTR, CDS, B-box domain, and CCT domain. Y-axis represents the subfamily name of each B-box genes. The lengths of the elements were drawn to scale
11816_2022_745_MOESM7_ESM.pdf
Figure S2 Distribution of conserved motifs in B-box proteins. a The phylogenetic tree was constructed based on the full-length sequences of B-box proteins using MEGA 7 software. Details of clusters are shown in different colors. b The motif composition of B-box proteins. The motifs, numbers 1–10, are displayed in different-colored boxes. The length of protein can be estimated using the scale at the bottom
11816_2022_745_MOESM8_ESM.pdf
Figure S3 Homologous collinear dot-plot within pineapple genome. The ID of the B-box gene is displayed at the edge. The collinearity blocks formed by the WGD are in green boxes in the figure, and the median value of Ks or the range of the median value of Ks of the collinearity blocks are marked
11816_2022_745_MOESM9_ESM.pdf
Figure S4 Homologous collinear dot-plot within Ostreococcus lucimarinus genome. The ID of the B-box gene is displayed at the edge
11816_2022_745_MOESM10_ESM.pdf
Figure S5 Homologous collinear dot-plot within Arabidopsis thaliana genome. The ID of the B-box gene is displayed at the edge. The collinearity blocks formed by the WGD are in green boxes in the figure, and the median value of Ks or the range of the median value of Ks of the collinearity blocks are marked
11816_2022_745_MOESM11_ESM.pdf
Figure S6 Homologous collinear dot-plot within Grape genome. The ID of the B-box gene is displayed at the edge. The collinearity blocks formed by the WGD are in green boxes in the figure, and the median value of Ks or the range of the median value of Ks of the collinearity blocks are marked
11816_2022_745_MOESM12_ESM.pdf
Figure S7 Homologous collinear dot-plot within Rice genome. The ID of the B-box gene is displayed at the edge. The collinearity blocks formed by the WGD are in green boxes in the figure, and the median value of Ks or the range of the median value of Ks of the collinearity blocks are marked
11816_2022_745_MOESM13_ESM.pdf
Figure S8 Homologous collinear dot-plot within Physcomitrella patens genome. The ID of the B-box gene is displayed at the edge. The collinearity blocks formed by the WGD are in green boxes in the figure, and the median value of Ks or the range of the median value of Ks of the collinearity blocks are marked
Rights and permissions
About this article
Cite this article
Yu, L., Lyu, Z., Liu, H. et al. Insights into the evolutionary origin and expansion of the BBX gene family. Plant Biotechnol Rep 16, 205–214 (2022). https://doi.org/10.1007/s11816-022-00745-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-022-00745-1