Abstract
Starch is the major constituent of rice grains determining eating, cooking, and processing properties. While rice endosperm mutants with modified starch characteristics have provided valuable materials for elucidating the gene mechanisms involved in starch biosynthesis and grain development, their practical use in crop improvement has been rarely documented. In this review, we focus on the use of floury endosperm mutants with reduced grain hardness to breed rice cultivars suitable for dry milling. We describe the identification of “Suweon542”, the original genetic source carrying flo4-4 [formerly flo7(t)] which has been used to develop Korean rice cultivars specialized for dry-milled flour production. Molecular breeding efforts are highlighted in the development of “Garumi1” and “Garumi2”, the floury endosperm rice cultivars with improved agronomic characteristics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice consumption in Korea has been sharply decreasing over the past four decades as the eating habits of the Korean people diversify with rapid economic growth. The per capita annual rice consumption in Korea recorded 61 kg in 2018, which is less than a half of 133 kg in 1980 (KOSIS 2019). As a result, overproduction of rice has been the major challenge in the Korean rice industry causing low market price, high storage cost, and stagnated income of rice growers. Despite the decrease in the total rice consumption, the amount of rice consumed as processed food (e.g., rice cake, bread, noodles, etc.) has been soaring in the recent years. While the total rice consumption in Korea decreased by 20% from 5.26 M tons in 2000 to 4.24 M tons in 2015, the amount consumed as processed food increased by 240% (from 180 to 610 thousand tons) during the same period (KOSIS 2019). Therefore, the government has been making policy and research efforts to invigorate the processed rice food industry and boost rice consumption in Korea.
The high cost of flour production is one of the main obstacles in scaling up processed rice food industry. Approximately 70% of rice used to manufacture processed food is utilized in the form of flour, of which the majority is produced using the wet milling technique (Kim 2013). To soften rice grains, wet milling involves the soaking and drying steps of rice before and after milling, consuming large amounts of water and energy (Chiang and Yeh 2002). While the dry milling technique mainly used for wheat flour production excludes the soaking and drying steps and can reduce the milling cost by 50–70%, the high grain hardness of regular rice cultivars causes high level of damaged starch in dry-milled flour and deteriorates end-use quality (Lee and Kim 2011; Leewatchararongjaroen and Anuntagool 2016; Kwak et al. 2017). This necessitates the development of rice cultivars with soft grain that pulverizes easily and is suitable for dry milling.
A number of endosperm mutants with modified starch structure have been identified and characterized in rice, providing valuable materials to elucidate the starch biosynthesis pathway (Nakamura 2018; Bao 2019). However, little information is available in terms of the practical use of the endosperm mutants in rice (Ashida 2014). In this mini review, we focus on the genetics and characteristics of the floury endosperm rice mutants, highlighting their utilization in molecular breeding to develop cultivars suitable for dry milling which would contribute to strengthening the processed rice food industry.
Floury endosperm mutants in rice
The term “floury” or “chalky” is generally used to describe soft, white non-translucent rice endosperm which breaks easily into powder due to the loosely packed starch granule structures (Satoh and Omura 1981; Kaushik and Khush 1991; Ashida et al. 2009). While some floury rice mutants referred to as “milky-white” exhibit the soft non-translucent characteristics in the entire endosperm area, others are partly floury and often classified as “white-core”, “white-back”, or “white-belly” that has the floury portion at the central, dorsal, or ventral area of the endosperm, respectively (Ashida et al. 2009). The non-translucent endosperm appearance of the floury mutants is similar to that of the “waxy” (glutinous) and “dull” (intermediate between waxy and non-waxy) mutants. However, when stained by I-KI solution, the endosperm of the floury mutants turns blue-black as regular translucent rice while the waxy mutants are not stained and the dull mutants show intermediate phenotype (Satoh and Omura 1981; Ashida et al. 2009). Also, the soft endosperm of the floury mutants is different from that of the waxy and dull mutants which shows high grain hardness as regular translucent rice (Satoh and Omura 1981). Recent reviews have provided extensive information on various types of rice endosperm mutants used for studying starch biosynthesis pathway (Nakamura 2018; Bao 2019). This review mainly focuses on the rice endosperm mutants classified as flo (floury), their characteristics, and potential use in breeding.
More than 16 mutants named flo have been reported in rice and 14 of them have been cloned (Table 1). Caution is required as some of the non-allelic mutants have been designated the same nomenclature, i.e., two non-allelic mutants each for flo6 (Zhang et al. 2012; Peng et al. 2014) and flo7 (Sheng et al. 2015; Zhang et al. 2016). Most of the flo mutants have been screened from the mutant populations induced by chemicals (MNU, EMS, SA), radiation (gamma ray), insertional mutagens (T-DNA, Tos17), or tissue culture owing to their abnormal grain appearance with white non-translucent endosperm which is different from the wild-type’s translucent endosperm. The I-KI staining method is often used to distinguish floury mutants from waxy and dull mutants (Mo et al. 2013).
While the flo mutants including flo(a), flo6, flo10, flo12, flo13, flo14 and flo16 exhibit the floury phenotype in most part of the endosperm except thin periphery, some are partially floury in the core (flo5, flo8, flo15) or peripheral (flo7) regions of the grain (Fig. 1). Interestingly, flo4-1 and flo4-4 [formerly flo7(t)], the allelic mutants of cyOsPPDK (Os05g0405000), exhibit different floury endosperm phenotype. While flo4-1 carrying a T-DNA insertion between exon 3 and exon 4 is floury only in the core endosperm region (Kang et al. 2005), flo4-4 with a missense mutation in exon 8 shows floury phenotype in most part of the endosperm except for the thin outer layer (Mo et al. 2013; Wang et al. 2018). Regardless of the extent of the floury phenotype within the endosperm, electron microscopy observations of the non-translucent endosperm area of the flo mutants reveal loosely packed starch granules with rounded shape that are different from the densely packed polyhedral starch granules observed in regular translucent endosperm (Fig. 2).
Transverse sections of the rice mutants with floury endosperm. Each figure was imported from the relevant reference with minor modification (i.e. cropping and rotation). The permission to reuse the images of flo4-1, flo5, flo6 (both Zhang et al. 2012 and Peng et al. 2014), flo8, flo10, flo11, flo12, flo13, and flo15 were obtained through Copyright Clearance Center (http://www.copyright.com/publishers/rightslink/). The images of flo7(t), flo7 (both Sheng et al. 2015 and Zhang et al. 2016), flo14, and flo16 were imported from the open access articles distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/). The permission to reuse the images of flo(a) and flo9 were obtained directly from the publisher (Korean Society for Molecular and Cellular Biology) and the corresponding author (Dr. Yihua Wang), respectively
Electron microscopy visualization of endosperm and brown rice appearance in floury endosperm rice cultivars. a Electron microscopy observation of endosperm in non-mutagenized Namil and its floury mutant line Suweon542. The pictures were imported from Mo et al. (2013) with minor modification under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses). b Brown rice and its transverse section of Suweon542, Jopyeong, and the two floury rice cultivars Garumi1 and Garumi2 derived from the cross between Suweon542 and Jopyeong
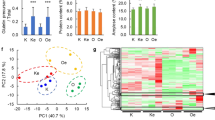
Major phenotype changes (e.g. grain weight, starch properties, etc.) observed in the flo mutants are summarized in Table 1. Most flo mutants exhibit significant decrease in grain weight mainly due to the altered endosperm starch structure with loosely packed granules that are different from wild-type’s densely packed starch granules. Only two flo mutants, namely flo5 (Ryoo et al. 2007) and flo8 (Long et al. 2017), have comparable grain weight relative to wild-type while none of the flo mutants show increase in grain weight. Most flo mutants exhibit significant reduction in amylose content while a few, namely flo6 (Zhang et al. 2012), flo11 (Zhu et al. 2018), flo13 (Hu et al. 2018) and flo14 (Xue et al. 2019), show no significant change. Two exceptions are flo5 (Ryoo et al. 2007) and flo4-4 (Mo et al 2013), which have amylose content significantly higher than wild-type. Changes in other phenotypes including amylopectin chain length distribution, protein and lipid content, and plant height vary among different flo mutants (Table 1). While these mutants have provided invaluable materials to study the gene mechanisms involved in starch biosynthesis and grain development, their potential use in practical breeding has been rarely explored.
“Suweon542”, a floury endosperm mutant suitable for dry milling
In Korea, “Suweon542” provided the original genetic source for developing rice cultivars suitable for producing dry-milled flour (Mo et al. 2013). Suweon542 is an elite rice line selected directly from a mutant population induced by sodium azide in the background of “Namil”, a high-yielding non-glutinous Korean japonica cultivar. From over 5,000 mutant lines at M8 generation, those with white non-translucent endosperm were visually screened. Waxy and dull mutants were excluded by I-KI staining. One line designated “Namil(SA)M2-1509-RGA-1-1-1-1” with floury appearance in the entire endosperm area was selected as a promising line, named Suweon542 and evaluated in yield trials and local adaptability tests. Compared to non-mutagenized Namil, Suweon542 was slightly taller and headed few days later. Although Suweon542 had significantly higher number of spikelets (155 per panicle) than Namil (117 per panicle), its brown rice yield was 5.67 t/ha, 6% lower than that of Namil (6.00 t/ha) mainly due to the low weight of the floury grains.
Compared to Namil, the grain hardness of Suweon542 was reduced by 56%, indicating that the soft grains of Suweon542 would be easily pulverized during dry milling and produce good quality flour (Mo et al. 2013). Dried-milled flour of Suweon542 showed damaged starch content of 4.9%, 47% lower than that of Namil (9.2%), and its particle size distribution was highly similar to commercial wheat flour produced by dry milling. End-use qualities of the dry-milled and wet-milled Suweon542 flour were compared for various processed food products including bread, noodles, beer, rice cake and confectionery. The panel tests by researchers and consumers reported no quality difference between the products made from dry-milled and wet-milled Suweon542 flour, demonstrating that Suweon542 would provide a promising genetic source to breed rice cultivars suitable for dry milling (KIPRIS 2012).
To identify the causal mutation responsible for Suweon542’s floury endosperm, genetic analysis was conducted using an F2 population derived from the cross between Suweon542 and Milyang23 (Mo et al. 2013). The causal gene was initially mapped at the 19.3–20.1 Mb region on chromosome 5 and temporarily designated flo7(t). Fine-mapping with the segregating F3:4 progenies further delimited the flo7(t) locus within the 33 kb interval containing four candidate genes (Wang et al. 2018). Sequencing of the candidate genes revealed a G-to-A SNP (Gly-to-Asp) in exon 8 of cyOsPPDK (Os05g0405000) which encodes a cytosolic pyruvate orthophosphate dikinase protein. As the three mutated alleles of cyOsPPDK (flo4-1, flo4-2 and flo4-3) had been reported previously (Kang et al. 2005), the novel allele from Suweon542 was named flo4-4 (Wang et al. 2018). To facilitate molecular breeding, PCR-based allele specific markers were developed for flo4-4 (KIPRIS 2019a).
Molecular breeding to improve rice cultivars for dry milling
Despite the novel starch properties suitable for dry milling, Suweon542 has two major drawbacks—the high susceptibility to bacterial blight, stripe virus and blast that are rampant in the Southern plain area of Korea, and the frequent occurrence of pre-harvest sprouting under the wet weather during grain filling. To overcome these limitations, we crossed Suweon542 with “Jopyeong”, an early-maturing Korean japonica rice cultivar carrying the Xa3 gene for bacterial blight resistance and the Stv-bi gene for stripe virus resistance (Nam et al. 2013). While the blast resistance gene carried by Jopyeong is unclear, pedigree information suggests that Jopyeong has a blast resistant allele at the Piz locus from a Korean rice cultivar “Jinbubyeo” (Jeung et al. 2007).
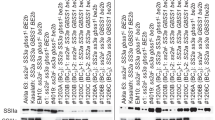
Marker assisted selection (MAS) was carried out from the F2 generation using the flo4-4 marker (KIPRIS 2019a) for floury endosperm, the 9643.T4 marker (Park et al. 2013) for Xa3, the InDel7 marker (KIPRIS 2013) for Stv-bi, and the 9871.T7E marker (Jeung et al. 2007) for the resistant allele at the Piz locus (Fig. 3). This enabled us an efficient breeding scheme as all the breeding lines evaluated in the field at F6 generation were fixed with the favorable alleles for floury endosperm and resistance to bacterial blight, stripe virus and blast. As MAS was not feasible for pre-harvest sprouting, phenotype selection was conducted by visually screening brown rice at F2–F4 generations and evaluating panicles for vivipary under saturated humidity at F5 generation. Two promising lines, “SR34136-B < fl-19-7-2-5” and “SR34136-B < fl-19-7-8-6”, showed pre-harvest sprouting resistance and were further evaluated in yield trials, demonstrating higher yield compared to Suweon542 under the late planting conditions (transplanting in late June–early July in Korea). These lines were subsequently registered as the new cultivars “Garumi1” and “Garumi2” (KIPRIS 2019b).
Enhanced disease resistance of Garumi1 and Garumi2 through molecular breeding. Disease resistance phenotype for bacterial blight (a) and blast (b). Marker genotypes of 9643.T4 for Xa3 (c), 9871.T7E for Piz (d) and InDel7 for Stv-bi (e). A and B indicate susceptible and resistant alleles, respectively, and M indicates the size marker
Perspectives
With improved resistance to diseases and pre-harvest sprouting, Garumi1 and Garumi2 provide useful materials as specialty cultivars for dry-milled rice flour production. High yield performance of Garumi1 and Garumi2 under the late planting condition is also valuable for promoting diverse double cropping systems in the rice paddies. Different dry milling techniques and conditions are currently being tested in collaboration with processed food manufacturers to optimize the flour characteristics of Garumi1 and Garumi2 for producing various food items such as bakery products, noodles, rice cake and liquor. Further breeding efforts are also underway to develop new floury endosperm cultivars with additional processing qualities such as color and aroma.
References
Ashida K (2014) Properties of floury rice mutant and its utilization for rice flour. Jpn Agr Res Q 48:51–56
Ashida K, Iida S, Yasui T (2009) Morphological, physical, and chemical properties of grain and flour from chalky rice mutants. Cereal Chem 86:225–231
Bao J (2019) Rice starch. In: Bao J (ed) Rice chemistry and technology, 4th edn. Woodhead Publishing, Duxford, pp 55–108
Chiang PY, Yeh AI (2002) Effect of soaking on wet-milling of rice. J Cereal Sci 35:85–94
Hu T, Tian Y, Zhu J, Wang Y, Jing R, Lei J, Sun Y, Yu Y, Li J, Chen X, Zhu X, Hao Y, Liu L, Wang Y, Wan J (2018) OsNDUFA9 encoding a mitochondrial complex I subunit is essential for embryo development and starch synthesis in rice. Plant Cell Rep 37:1667–1679
Jeung JU, Kim BR, Cho YC, Han SS, Moon HP, Lee YT, Jena KK (2007) A novel gene, Pi40(t), linked to the DNA markers derived from NBS-LRR motifs confers broad spectrum of blast resistance in rice. Theor Appl Genet 115:1163–1177
Kang HG, Park S, Matsuoka M, An G (2005) White-core endosperm floury endosperm-4 in rice is generated by knockout mutations in the C4-type pyruvate orthophosphate dikinase gene (OsPPDKB). Plant J 42:901–911
Kaushik RP, Khush GS (1991) Genetic analysis of endosperm mutants in rice Oryza sativa L. Theor Appl Genet 83:146–152
Kawasaki T, Mizuno K, Shimada H, Satoh H, Kishimoto N, Okumura S, Ichikawa N, Baba T (1996) Coordinated regulation of the genes participating in starch biosynthesis by the rice floury-2 locus. Plant Physiol 110:89–96
Kim MH (2013) Review on rice flour manufacturing and utilization. J Biosyst Eng 38:103–112
Kinoshita T, Takahashi M (1991) The one hundredth report of genetical studies on rice plant—linkage studies and future prospects. J Fac Agr Hokkaido Univ 65:1–61
KIPRIS (2012) A floury japonica rice line, Namil(SA)-flo2, suitable for dry milling process and the food compositions containing Namil(SA)-flo2 as an active ingredient (Registration No. 1011906420000). Korea Intellectual Property Rights Information Service. https://engportal.kipris.or.kr/engportal/search/total_search.do. Accessed 1 Nov 2019
KIPRIS (2013) Primer for selecting variety resistant to rice stripe disease containing Stv-bi gene and the selecting method thereof (Application No. 1020120012710). Korea Intellectual Property Rights Information Service. https://engportal.kipris.or.kr/engportal/search/total_search.do. Accessed 1 Nov 2019
KIPRIS (2019a) Markers for identifying floury endosperm characteristics and use thereof (Registration No. 1020108590000). Korea Intellectual Property Rights Information Service. https://engportal.kipris.or.kr/engportal/search/total_search.do. Accessed 4 Nov 2019
KIPRIS (2019b) New rice cultivar, Garumi1 and Garumi2 suitable for dry milling (Application No. 1020190101469). Korea Intellectual Property Rights Information Service. https://engportal.kipris.or.kr/engportal/search/total_search.do. Accessed 4 Nov 2019
KOSIS (2019) Korean Statistical Information Service. https://kosis.kr/eng/. Accessed 4 Nov 2019
Kwak J, Yoon MR, Lee JS, Lee JH, Ko S, Tai TH, Won YJ (2017) Morphological and starch characteristics of the Japonica rice mutant variety Seolgaeng for dry-milled flour. Food Sci Biotechnol 26:43–48
Lee YT, Kim YU (2011) Physicochemical properties of brown rice flours differing in amylose content prepared by different milling methods. J Korean Soc Food Sci Nutr 40:1797–1801
Leewatchararongjaroen J, Anuntagool J (2016) Effects of dry-milling and wet-milling on chemical, physical and gelatinization properties of rice flour. Rice Sci 23:274–281
Liu Y, Zhu X, Liu X, Tian Y, Liu S, Wang Y, Zhang W, Jiang L, Wang Y, Wan J (2018) Phenotyping and gene-mapping of a floury endosperm mutant flo9 in rice. J Nanjing Agr Univ 41:616–624
Long W, Dong B, Wang Y, Pan P, Wang Y, Liu L, Chen X, Liu X, Liu S, Tian Y, Chen L, Wan J (2017) FLOURY ENDOSPERM8, encoding the UDP-glucose pyrophosphorylase 1, affects the synthesis and structure of starch in rice endosperm. J Plant Biol 60:513–522
Maekawa M (1985) Location of a floury endosperm gene in the second linkage group. Rice Genet Newsl 2:57–58
Mo YJ, Jeung JU, Shin YS, Park CS, Kang KH, Kim BK (2013) Agronomic and genetic analysis of Suweon 542, a rice floury mutant line suitable for dry milling. Rice 6:37
Nakamura Y (2018) Rice starch biotechnology: rice endosperm as a model of cereal endosperms. Starch 70:1600375
Nam JK, Ko JK, Kim KY, Kim BK, Kim WJ, Ha KY, Shin MS, Ko JC, Kang HJ, Park HS, Baek MK, Shin WC, Mo YJ, Choi IB, Kim YD, Yang HS, Kim JG (2013) A new early-maturing rice with blast, bacterial blight, rice stripe virus resistant and high grain quality, ‘Jopyeong’. Korean J Breed Sci 45:177–182
Park HS, Shin MS, Kim KY, Noh TH, Baek SH, Lee JH, Ha KY, Baek MK, Kim WJ, Park JH, Yoo JS, Cho YC, Kim BK (2013) Reaction of single resistance genes and their pyramiding effects in indica and japonica rice against Xanthomonas oryzae pv. oryzae in Korea. Korean J Breed Sci 45:119–129
Peng C, Wang Y, Liu F, Ren Y, Zhou K, Lv J, Zheng M, Zhao S, Zhang L, Wang C, Jiang L, Zhang X, Guo X, Bao Y, Wan J (2014) FLOURY ENDOSPERM6 encodes a CBM48 domain-containing protein involved in compound granule formation and starch synthesis in rice endosperm. Plant J 77:917–930
Qiao Y, Lee SI, Piao R, Jiang W, Ham TH, Chin JH, Piao Z, Han L, Kang SY, Koh HJ (2010) Fine mapping and candidate gene analysis of the floury endosperm gene, FLO(a), in rice. Mol Cells 29:167–174
Ryoo N, Yu C, Park CS, Baik MY, Park IM, Cho MH, Bhoo SH, An G, Hahn TR, Jeon JS (2007) Knockout of a starch synthase gene OsSSIIIa/Flo5 causes white-core floury endosperm in rice (Oryza sativa L.). Plant Cell Rep 26:1083–1095
Satoh H, Omura T (1981) New endosperm mutations induced by chemical mutagens in rice, Oryza sativa L. Jpn J Breed 31:316–326
She KC, Kusano H, Koizumi K, Yamakawa H, Hakata M, Imamura T, Fukuda M, Naito N, Tsurumaki Y, Yaeshima M, Tsuge T, Matsumoto K, Kudoh M, Itoh E, Kikuchi S, Kishimoto N, Yazaki J, Ando T, Yano M, Aoyama T, Sasaki T, Satoh H, Shimada H (2010) A novel factor FLOURY ENDOSPERM2 is involved in regulation of rice grain size and starch quality. Plant Cell 22:3280–3294
Sheng Z, Fang P, Li S, Jiao G, Xie L, Hu P, Tang S, Wei X (2015) Phenotype of rice floury endosperm mutant flo7 and fine mapping of mutated gene. Rice Sci 22:162–170
Teng X, Zhong M, Zhu X, Wang C, Ren Y, Wang Y, Zhang H, Jiang L, Wang D, Hao Y, Wu M, Zhu J, Zhang X, Guo X, Wang Y, Wan J (2019) FLOURY ENDOSPERM16 encoding a NAD-dependent cytosolic malate dehydrogenase plays an important role in starch synthesis and seed development in rice. Plant Biotechnol J 17:1914–1927
Wang H, Mo YJ, Im DE, Jang SG, Ham TH, Lee J, Jeung JU, Kwon SW (2018) A new SNP in cyOsPPDK gene is associated with floury endosperm in Suweon 542. Mol Genet Genomics 293:1151–1158
Wu M, Ren Y, Cai M, Wang Y, Zhu S, Zhu J, Hao Y, Teng X, Zhu X, Jing R, Zhang H, Zhong M, Wang Y, Lei C, Zhang X, Guo X, Cheng Z, Lin Q, Wang J, Jiang L, Bao Y, Wang Y, Wan J (2019) Rice FLOURY ENDOSPERM10 encodes a pentatricopeptide repeat protein that is essential for the trans-splicing of mitochondrial nad1 intron 1 and endosperm development. New Phytol 223:736–750
Wu Y, Pu C, Lin H, Huang H, Huang Y, Hong C, Chang M, Lin Y (2015) Three novel alleles of FLOURY ENDOSPERM2 (FLO2) confer dull grains with low amylose content in rice. Plant Sci 233:44–52
You X, Zhang W, Hu J, Jing R, Cai Y, Feng Z, Kong F, Zhang J, Yan H, Chen W, Chen X, Ma J, Tang X, Wang P, Zhu S, Liu L, Jiang L, Wan J (2019) FLOURY ENDOSPERM15 encodes a glyoxalase I involved in compound granule formation and starch synthesis in rice endosperm. Plant Cell Rep 38:345–359
Xue M, Liu L, Yu Y, Zhu J, Gao H, Wang Y, Wan J (2019) Lose-of-function of a rice nucleolus-localized pentatricopeptide repeat protein is responsible for the floury endosperm14 mutant phenotypes. Rice 12:100
Zhang D, Wu J, Zhang Y, Shi C (2012) Phenotypic and candidate gene analysis of a new floury endosperm mutant (osagpl2-3) in rice. Plant Mol Biol Rep 30:1303–1312
Zhang L, Ren Y, Lu B, Yang C, Feng Z, Liu Z, Chen J, Ma W, Wang Y, Yu X, Wang Y, Zhang W, Wang Y, Liu S, Wu F, Zhang X, Guo X, Bao Y, Jiang L, Wan J (2016) FLOURY ENDOSPERM7 encodes a regulator of starch synthesis and amyloplast development essential for peripheral endosperm development in rice. J Exp Bot 67:633–647
Zhong M, Liu X, Liu F, Ren Y, Wang Y, Zhu J, Teng X, Duan E, Wang F, Zhang H, Wu M, Hao Y, Zhu X, Jing R, Guo X, Jiang L, Wang Y, Wan J (2019) FLOURY ENDOSPERM12 encoding alanine aminotransferase 1 regulates carbon and nitrogen metabolism in rice. J Plant Biol 62:61–73
Zhu X, Teng X, Wang Y, Hao Y, Jing R, Wang Y, Liu Y, Zhu J, Wu M, Zhong M, Chen X, Zhang Y, Zhang W, Wang C, Wang Y, Wan J (2018) FLOURY ENDOSPERM11 encoding a plastid heat shock protein 70 is essential for amyloplast development in rice. Plant Sci 277:89–99
Acknowledgements
This work was supported by the research grant (PJ01289003) from the Rural Development Administration (RDA), Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mo, Y., Jeung, JU. The use of floury endosperm mutants to develop rice cultivars suitable for dry milling. Plant Biotechnol Rep 14, 185–191 (2020). https://doi.org/10.1007/s11816-020-00604-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-020-00604-x