Abstract
Genome editing offers great advantages in identifying gene function and generating agronomical important mutations in crops. Here, we report the development of edited lines with reduced seed dormancy by knockout viviparous-1 (OsVP1) gene known as a transcription factor that regulates key aspects of plant seed development and ABA signaling in rice. Thirty-three genetic edited lines out of 55 T0 rice plants were generated using CRISPR/Cas9 system. Sequencing analysis showed that the plants had four different mutation types at the target site of OsVP1, the mutations were found to be transmitted to the succeeding generations. Stable transmission of CRISPR/Cas9-mediated mutant lines without the transferred DNA (T-DNA) was confirmed by segregation in the T1 generation. Regarding many investigated agronomic trait, there are no significant differences between homozygous mutants and wildtype plants under field’s growth conditions. Especially in RT-PCR analysis of ABA/GA signaling genes, the expression of OsNCED2, OsGA20ox1, OsGA20ox2, OsGA20ox3 genes in homozygous mutants was increased compared to wildtype plants. Results of this study exemplified the effectiveness of CRISPR/Cas9 as a gene editing tool in broke down the seed dormancy in rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Seed germination is an important process ensuring the continuity of life in plants that depend on it as exclusive mode of propagation (Bewley 1997). It is important to ensure that seeds have uniform and rapid germination after sowing for agricultural production. Thus many seed treatments, such as cold stratification and dry after-ripening, have been used to reduce or break seed dormancy after crop harvesting (Finch-Savage and Leubner-Metzger 2006). Seed dormancy is a condition that has not germinated during a specific period, even in environmental conditions that are prone to sprouting (Finch-Savage and Leubner-Metzger 2006; Graeber et al. 2012). These phenomena vary in proportion to the dry storage (after ripening) of the seeds and are genetically controlled by the plant genotypes. The dormancy imposed by the coat is enhanced by the tissue that covers the seed, i.e., glue and pale (or crust), pericarp and testis, and optionally endosperm (Bewley et al. 2013). Embryonic dormancy of seeds is finely controlled during development (Sugimoto et al. 2010). In rice, grains after harvest are dried under sunlight and are incubated at 55 ℃ for 3–5 days to break dormancy that will allow to germinate in rapid and uniform upon seed sowing. The mechanisms underlying the breaking of seed dormancy after-ripening treatment in rice is not fully well known. So far, many genetic researchers have reported that ABA plays an essential role in dormancy induction and maintenance (Kermode 2005). There are four major transcription factors discovered in A. thaliana that play an important role in seed maturation and dormancy, namely, ABSCISIC ACID INSENSITIVE 3 (ABI3), FUSCA 3 (FUS3), LEAFY COTYLEDON 1 (LEC1) and LEAFY COTYLEDON 1 (LEC2) (LeClere et al. 2002). All four abi3, lec1, lec2 and fus3 mutants are severely affected in seed maturation and share some common phenotypes, such as decreased dormancy at maturation (Raz et al. 2001) and reduced expression of seed storage proteins (Gutierrez et al. 2007). Also, the seed dormancy is a quantitative trait loci (QTL) controlled by multiple genetic and environmental factors (Du et al. 2015). So far, many QTLs for seed dormancy have been identified from rice, wheat and Arabidopsis (van der Schaar et al. 1997; Flintham et al. 2002; Dong et al. 2003). Unveiling of QTL genes and complex mechanisms underlying seed dormancy is accelerated by the rapid progress of crop genomics. To date, more than 40 QTLs related to seed dormancy and germination have been reported among cultivated, wild, and weedy rice (Cai and Morishima 2000; Dong et al. 2003; Gu et al. 2004; Li et al. 2006; Lu et al. 2011). In rice, qLTG3-1, the first clonal gene related to QTL limiting germination at low temperature, is preferentially expressed in sheath and epidermal blast cells (Fujino et al. 2008). It has been also reported that seed dormancy 4 (Sdr4) is one of the major factors determining seed dormancy in rice (Sugimoto et al. 2010). Sdr4 expression is positively regulated by rice viviparous-1 (OsVP1), a global regulator of seed maturation that is orthologous to maize (ZmVP1) and Arabidopsis (ABI3) (McCarty et al. 1991; Hattori et al. 1994; Sugimoto et al. 2010). Studies in Arabidopsis have revealed the framework of the regulatory network for seed maturation, which is controlled by several master transcription factors including ABI3, which is orthologous to ZmVP1 and OsVP1 (Koornneef et al. 1989; McCarty et al. 1991). Mutations in these genes profoundly affect the acquisition of seed dormancy. For OsVP1, studies on its knockdown rice plants using the gene silencing system reported that they showed enhanced germination speed of seeds than that of wild type plants (Gao et al. 2011). In recent years, genome editing techniques have been successfully applied to genetically modified plants. Among them, CRISPR/Cas9 system has revolutionized gene editing technology and is widely used because of its simplicity, efficiency, and versatility (Ma et al. 2015).
Here, we report the CRISPR/Cas9 target mutagenesis of OsVP1 in Dongjin, a Japonica rice cultivar. These homozygous mutant lines were significantly improved germination speed compared to wild type plants. The transgene-free homozygous mutants were efficiently selected in the T1 generation. Results of this study exemplified the effectiveness of CRISPR/Cas9 as a gene editing tool in breaking down the seed dormancy in rice.
Materials and methods
Plant materials and growth condition
Rice cultivar Dongjin (Oryza sativar L., ssp. Japonica) was used in this study. Rice plants were grown in the greenhouse facility, and in the paddy field of Hankyong National University in Korea. Harvested seeds were dried to ~ 14% moisture content and kept in dry conditions at 4 °C to retain dormancy.
Vector construction
The Cas9::sgRNA expression vector, pBAtC binary vector (Kim et al. 2016) was used to construct OsU3::pBAtC carrying the OsU3 promoter to express sgRNA designed from OsVP1 gene. The sgRNA expressing cassette sequences (Supplementary Fig. 1 and 2) with EcoRI and XhoI site between OsU3 promoter and sgRNA scaffold sequence were synthesized by Bioneer co., LTD (Dajeon, Korea). To amplify the OsU3 promoter, Phusion High-Fidelity DNA polymerase (Finnzymes, Thermo Scientific, Wlatham, MA, USA) was used in genomic DNA with the following primers: OsU3p-EcoRI-F, 5′-CTCGAATTCAAGGAATCTTTAAACATACGAACAG-3′ and OsU3p-XhoI-R, 5′-ACTCTCGAGACACCTGCGAGCTGCCACGGATCATCTGCACAACTCTTT-3′ (Supplementary Table 1). The purified PCR products were then digested using EcoRI and XhoI restriction sites, and subcloned into pBAtC vector, which was also digested with EcoRI and XhoI to replace AtU6 promoter (Cho et al. 2013). The full sequencing data of pBOsC vector was described in Supplementary Fig. 3.
Construction of the sgRNA expression vector
The OsU3::vp1-sgRNA/pBOsC vector expressing sgRNA from OsVP1 (vp1-sgRNA) was constructed according to the method previously described (Kim et al. 2016). Briefly, the target sequence for OsVP1 was designed by the CRISPR-RGEN Tools website (https://rgenome.ibs.re.kr/) (Park et al. 2015). To identify off-target sequences within coding region of the gene Cas-OFFinder (https://www.rgenome.net/cas-offinder/) (Bae et al. 2014) was used. Cleavage position, GC content and out-of-frame score of each target sites were considered for improve gene editing efficiency. These target sites which had potential off-target sites bearing 1–2 mismatches in rice genomic DNA were excluded to reduce the possibility of off-target mutations. The vp1-sgRNA templates were annealed using two primers, 5′ GGCAGCCGCCGCCTCGATGGTGGAT-3′ and 5′ AAACATCCACCATCGAGGCGGCGGC-3′, and cloned into AarI-digested OsU3::pBOsC binary vector. Construction of the sgRNA expression vector, OsU3::vp1-sgRNA/pBOsC, and its flanking sequences were confirmed by Sanger sequencing method.
Rice transformations
Rice transformations were carried out as previously described (Nishimura et al. 2006). Agrobacterium-tumefaciens strain EHA105 was used to infect callus tissue induced from Dongjin seeds. Transgenic plants were regenerated from transformed calli using selection medium containing 6 mg/L phosphinothricin and 250 mg/L cefotaxime before transplanting in soil.
Genomic DNA extraction and TaqMan copy number analysis
All transgenic plants were carefully cultivated in greenhouse and fields condition. Sampling of leaves was performed to isolate DNA via DNA Quick Plant Kit (Inclone, Korea). The transgenic plants were selected by PCR analysis using bar gene specific primers: Bar-F 5′-CGTCAACCACTACATCGAGA-3′, Bar-R 5′-TTGCGCGCTATATTTTGTTT-3′ (Supplementary Table 1). PCR reaction was performed an initial denaturation at 95 ℃ for 5 min, followed by 35 cycles of denaturation at 95 ℃ for 30 s, annealing at 56 ℃ for 30 s, extension at 72 ℃ for 1 min, and an extra extension at 72 ℃ for 7 min. To check transgene copy number in rice genome, we was performed on Applied Biosystems StepOnePlus™ (Applied Biosystems, Foster City, CA) with TaqManⓇ Gene Expression Master Mix (Applied Biosystems) kit. The rice tubulin alpha-1 chain gene (AK102560) were used with endogenous control for TaqManⓇ copy number assay using primers; NOS-F (5′-GCATGACGTTATTTATGAGATGGGTTT-3′), NOS-R (5′-TGCGCGCTATATTTTGTTTTCTATCG-3′), and NOS-probe (5ʹ-TAGAGTCCCGCAATTAT-3′) (Supplementary Table 1). TaqMan-PCR analysis was performed with the following parameters: enzyme activation at 95 ℃ for 10 min, followed by 40 cycles of denaturation at 95 ℃ for 15 s, annealing and extension at 60 ℃ for 1 min. The relative quantitative analysis was applied based on real-time PCR data using Applied Biosystems CopyCallerⓇ Software v2.0 (Applied Biosystems) according to the manufacturer’s instructions.
Targeted deep sequencing and mutation analysis
To determine the mutation at the target site, genomic DNA (50 ng) was used as template to perform PCR amplification using PCR Mastermix (Bioneer, Korea). The primers used to amplify the genomic region containing the CRISPR/Cas9 target site are listed in Supplementary Table 1. PCR amplicons were subjected to paired-end read sequencing using MiniSeq (Illumina, San Diego, CA, USA). The next generation sequencing data derived from MiniSeq were analyzed using Cas-Analyzer (https://www.rgenome.net/cas-analyzer) (Park et al. 2017). To minimize the errors in PCR handling and sequencing, reads that appears only once were excluded. Those insertion and deletion mutations found around the Cas9 cleavage site which was 3 bp upstream of the protospacer were considered as mutations induced by Cas9.
Selection of T-DNA free plants
To selection T-DNA-free lines, T1 generation seeds were obtained from self-pollination of T0 transgenic plants introduced single copy and edited with homo or bi allelic. T1 seeds were sown in pots, and a 1% (v/v) basta solution was embedded in gauze on a part of the seedling leaves grown to the 3-leaf stage, and observed the response in 3 days to distinguish between resistance and sensitivity. Also these plants were used to reconfirm whether they was resistance or sensitivity by bar-strip tests. T-DNA free plants were selected when the death of leaves by basta treatment and the absence of bands by bar-strip experiments.
Germination assay and α-amylase activity
To measure the germination rates, rice seeds were placed in a petri dish (9 cm in diameter) with two filter papers and 10 ml of distilled water. All petri dishes were placed in a dark incubator at 25 ± 1 °C for 12 h and then for 12 h with light/12 h of dark photoperiod. The seeds were considered as germinated when the seeds sprouted through the seed coat. α-amylase was extracted from ca. 0.5 g imbibed grains. The germination was observed daily to compute the germination rate using three replications. Activity was measured by digestion of soluble starch using a kit (Megazame co.) with a reaction mixture (0.6 ml) consisted of 0.5 ml substrate buffer and 0.1 ml of sample solution. The mixture was incubated at 37 °C for 7.5 min; 0.5 ml of iodine solution and 3.0 ml of distilled water were added. Soluble starch was determined by absorbance at 660 nm. One amylase unit (U) is defined as the amount of enzyme that can hydrolyze 10 mg of starch at 37 °C in 30 min. Three replications were used for this assay. Statistical analysis was performed using Tukey’s HSD test.
qRT-PCR analysis
Genes involved in the metabolism of ABA (OsNCED2) and GA (OsGA20ox1, OsGA20ox2, OsGA20ox3) were investigated for qRT-PCR analysis. The imbibed (24 and 48 h) embryos were cut quickly from seeds then stored in liquid nitrogen. Total RNA was isolated from ca. 80–100 mg of embryos using a RNeasy plant mini kit (Qiagen, www.qiagen.com). Total RNA was used to synthesize cDNA with random oligonucleotides using a reverse transcription system (Takara, www.takara-bio.com). The cocktail for quantitative RT-PCR was prepared using the cDNA, primers and SYBR Green Real-time PCR Master Mix (Toyobo, https://www.bio-toyobo.cn) in 20 μL reaction volume in three replicates. Light Cycler 480 device (Roche, https://www.roche-applied-science.com) was used with the following conditions: 95 ºC for 30 s, 40 cycles at 95 ºC for 5 s, 60 ºC for 35 s and 95 ºC for 15 s. The actin gene was used as a housekeeping gene. Relative quantification of transcript levels was performed using the comparative Ct method (Livak and Schmittgen 2001). Statistical analysis was performed using Tukey’s HSD test.
Results
CRISPR/Cas9 design for OsVP1 editing
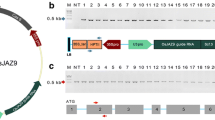
In monocot plants, the rice U3 small nuclear RNA promoter (OsU3) is generally used to express sgRNA (Belhaj et al. 2013). The OsU3 promoter was isolated by PCR amplification from Dongjin genome (Supplementary Fig. 1). The Arabidopsis U6 promoter in the CRISPR/Cas9 vector, pHAtC (Kim et al. 2016), was replaced with the Dongjin OsU3 promoter, and the resulting OsU3::pHAtC was used for rice CRISPR/Cas9-mediated target mutagenesis. To design a CRISPR/Cas9-OsVP1 that targets OsVP1 gene, a 20-bp nucleotide sequence (vp1) in the first exon of OsVP1 (Os01g68370) was chosen as the target site (Fig. 1a). The vp1 targeting sequence and protospacer adjacent motif (PAM) sequence are represented in red and in underlined lowercase letters, respectively. The predicted Cas9 cleavage site (vertical arrowhead) in the coding region of the gene was 632 bp downstream from the ATG initiation codon. The recombinant binary plasmid, OsU3::vp1-sgRNA/pHOsC, carrying vp1-sgRNA targeting OsVP1 gene under the control of OsU3 promoter, was then constructed based on OsU3::pHOsC (Fig. 1b).
Schematic representation of CRISPR/Cas9-mediated targeted mutagenesis in the rice OsVP1 gene. a Schematic diagram of OsVP1 gene and vp1 targeting sequence. Rice OsVP1 contains six exons, represented by blue rectangles, and the untranslated region portion, represented by blue lines. The 20-bp sgRNA targeting sequence (vp1) and protospacer adjacent motif (PAM) sequence are shown in red and in underlined lower-case letters, respectively. The vertical arrowhead indicates an expected cleavage site. b T-DNA region of the recombinant OsU3::vp1-sgRNA/pHOsC vector carrying vp1-sgRNA under the control of the OsU3 promoter. Expression of Cas9 is driven by the Cauliflower mosaic virus 35S (CaMV35S) promoter; expression of the vp1-sgRNA is driven by the OsU3 promoter; expression of phosphinothricin (PPT) is driven by the nopalin synthase (NOS) promoter; NLS: nuclear localization signal of Simian virus 40 (SV40) large T antigen; nos-t: gene terminator; LB and RB: left and right border, respectively. Primers used in the PCR are indicated by black arrows
CRISPR/Cas9 mediated targeted mutagenesis of OsVP1
The binary plasmid, OsU3::vp1-sgRNA/pHOsC, was then transformed into the WT calli via Agrobacterium-mediated transformation (Supplementary Fig. 4). There were 33 T0 transgenic plants that showed plant phenotype (Supplementary Fig. 5a). To verify the transgenic plants, PCR analysis with 33 plants was performed and the results showed positive amplification of bar genes (Supplementary Fig. 5b). To select single copy plants, TaqMan PCR was done using tubulin alpha-1 chain gene probe and NOS gene probe. Twenty-four transgenic plants showed copy number values ranging from 0.40 to 0.52, thus regarding as a single copy (Supplementary Fig. 5c). To further investigate CRISPR/Cas9 targeted mutagenesis of OsVP1, the targeting-containing amplicons obtained from all single copy lines were directly sequenced by MiniSeq and analyzed using Cas-Analyzer (https://www.rgenome.net/cas-analyzer/) (Park et al. 2017). Results showed that 18 independent plants have edited sequences near the protospacer adjacent motif (PAM) region. Sequence analyses detected 5 homozygous mutations, 3 heterozygous mutations, and 10 bi-allelic mutations (Table 1). Among allele mutation types, there were 69.4% (25/36) for deletions, 13.8% (5/36) for insertions, 5.5% (2/36) for substitutions, and 5.5% (2/36) for insertions/deletions, respectively (Table 1). The majority of mutations were short insertions and deletions (Indels). Every four base upstream of the protospacer adjacent motif (PAM) was mutated, and aligned perfectly with the expected Cas9 cleavage site, which is three base pairs downstream of the PAM sequence (Fig. 2a). The genotypes of mutants could be classified into three types: (1) putative homozygotes with both alleles containing the same mutations including lines, cr-vp1-4 and cr-vp1-8; (2) putative heterozygotes with only one allele mutated, such as line cv-vp1-11; (3) putative bi-allelic mutant with both copies of the target sequence mutated differently, including line cr-vp1-3 (Fig. 2a). Also, phenotypic analysis showed that there were no visible differences in plant growth and grain size between mutant lines and WT plants (Fig. 2b, c).
CRISPR/Cas9-induced mutations in the OsVP1 gene and phenotype of edited plants. a The mutant OsVP1 genotypes of representative T0 plants are identified by DNA sequencing and alignment. Deletions and insertions are indicated by dashes and red letters, respectively. The numbers on the right side show the sizes of the indels, with “−” and “+” indicating deletion and insertion of the nucleotides involved, respectively. The letters after the numbers represent different bases of the same length. b Plant morphologies of three mutant lines (cr-vp1-6, cr-vp1-8, cr-vp1-3) and WT plants were cultivated in the greenhouse for 2 months. c Appearance of mature seeds and brown rice of three mutant lines (cr-vp1-6, cr-vp1-8, cr-vp1-3) and WT plants
Obtaining transgene free lines in the T1 generation
To confirm the inheritance of mutation, four T0 plants and their progeny were investigated. For each T0 plants, 6–20 progeny were sampled randomly and genotyped individually at the target site (Table 2). As expected, all of the T0 putative homozygotes and their offspring had identical genotypes (cr-vp1-4, cr-vp1-8), suggesting that the mutations in these T0 plants were stable inherited to the next generation. For the T1 generation of putative bi-allelic mutant (cr-vp1-3), the genotype segregation ratio was consistent with Mendelian law, indicating that the targeted mutations in T0 plants were inherited normally. Thus, the results clearly showed that CRISPR/Cas9-induced gene mutations can enable stable inheritance to the subsequent generations. We also tracked the segregation of transgene (T-DNA) in the T1 population of the four T0 plants (Supplementary Fig. 6a, b) based on a PCR analysis and Bar-strip assay of bar gene, which are the elements of T-DNA (Supplementary Fig. 6c, d). Transgene-free plants were selected in the progeny of all detected T0 plants, with the proportion ranging from 16.1 to 27.6% (Supplementary Fig. 6e). These results indicated that transgene-free homozygous mutants could be easily obtained in the T1 generation, as the inheritance of T-DNA and the targeted gene was relatively independent. We selected four transgene-free homozygous knockout lines, including cr-vp1-6-4, cr-vp1-8-3, cr-vp1-3-6 and cr-vp1-11-3, which have coding frame shifts and premature translational stops, in the T1 generation (Fig. 3).
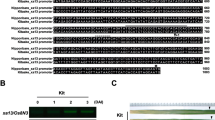
Transgene-free homozygous mutant lines induced by CRISPR/Cas9. a DNA sequence alignments for the four homozygous mutants identified in the T1 generation, together with a wild-type (WT) control. The numbers on the right side are the sizes of the indels, with “−” and “+” showing deletion and insertion of nucleotides involved, respectively. b Alignments of deduced amino acid sequence for the four homozygous mutants and WT. Each of the mutant alleles codes for truncated and disrupted OsVP1 proteins
Agronomic characterization of the transgene-free knockout lines
To assess the germination ability in the transgene-free knockout mutants, the progeny of cr-vp1-6-4, cr-vp1-8-3, cr-vp1-3-6, and cr-vp1-11-3 were tested along with WT. The WT seeds started to germinate in 2 days after sowing and reached the plateau in 7 days. However, cr-vp1-6-4, cr-vp1-8-3, cr-vp1-3-6, and cr-vp1-11-3, germination began as early as in 1 day after sowing and reached its peak in 3 days (Fig. 4). These results showed that the seed germination of these mutant lines was faster than that of WT, implying that knockout of the OsVP1 broke the dormancy and promoted germination of rice seeds. Moreover, mutant lines exhibited no significant differences in grain yield, straw weight, grain quality, and other main agronomic traits (Fig. 5a, b, Supplementary Fig. 7), compared with WT plants. These results demonstrate that transgene-free mutant lines could have negligible effects on growth, biomass, and main agronomic traits under normal cultivation conditions after gene editing process. Generally, GA induces synthesis of α-amylase in the aleurone layer, which then diffuses into the starchy endosperm where it initiates hydrolysis of starch (Bewley et al. 2013). To investigate the α-amylase activity changes in the seed germination process, cr-vp1-6-4 and the WT seeds were soaked in water for 48 h and α-amylase activity was monitored. The α-amylase activity of cr-vp1-6-4 line seeds was increased two times more than WT suggesting that knockout of OsVP1 increased α-amylase activity and enhanced the hydrolysis of endosperm starch and then promoted seed germination (Fig. 6).
Freshly harvested seeds from transgene-free homozygous mutant lines (cr-vp1-6, cr-vp1-8, cr-vp1-11, cr-vp1-3) showed improved germination percentage compared with those of WT. Germination assays were conducted in a petri dish (9 cm in diameter) with two filter papers at 25 ± 1 °C for 12 h (day) and for 12 h (night). Error bars indicate SD; n = 3
Agronomic characters of transgene-free homozygous mutant lines. Two mutant lines (cr-vp1-6-4, cr-vp1-8-3) and WT plants were cultivated in the experimental field. a Grain yield. b Straw dry weight. There is no significant difference between mutant lines and WT in grain yield and straw dry weight. Error bars indicate SD; n = 3
Expression of ABA/GA biosynthesis-related genes is affected in cr-vp1-6-4 line
Together with ABA and GA is associated with the regulation of seed dormancy and germination (Ogawa et al. 2003; Liu et al. 2010). To further evaluate the impact of seed dormancy, we investigated the expression of ABA/GA signaling genes, including OsNCED2, OsGA20ox1, OsGA20ox2 and OsGA20ox3. In cr-vp1-6-4 line, the transcripts of OsNCED2, OsGA20ox1, OsGA20ox2 and OsGA20ox3 were significantly increased more than the wild type. These data suggested that the increased expression of ABA/GA signaling genes could partially reduce the acquisition of seed dormancy of cr-vp1-6-4 line (Fig. 7).
Discussion
CRISPR/Cas9 system is a highly specific and an efficient technique for genome editing in all organisms. CRISPR/Cas9 technology has been widely used to improve major crops, such as grape, corn, rice, and soybean (Bortesi and Fischer 2015). Recently, several studies have attempted to further improve superior varieties using CRISPR/Cas9 technology to edit genes involved in agriculturally important traits especially in elite rice varieties (Li et al. 2016; Han et al. 2018; Martín-Pizarro et al. 2018). Knockout of thermo-sensitive genic male-sterile (TGMS) gene, tms5 in 11 fertile elite cultivars produced successful TGMS lines with good agronomic traits (Zhou et al. 2016).
In this study, OsU3::pBOsC, which replaced the rice U3 promoter in the pBOsC vector (Kim et al. 2016) with the OsU3 promoter, was constructed for rice CRISPR/Cas9 mediated target mutagenesis (Fig. 1). Using the OsU3::pBOsC, targeted mutagenesis in the viviparous-1 gene, OsVP1, was generated. We generated 18 edited plants out of 33 T0 transgenic plants (Table 1). With them, we selected four transgene-free homozygous knockout lines including cr-vp1-6-4, cr-vp1-8-3, cr-vp1-3-6 and cr-vp1-11-3, which showed coding frame shifts and premature translational stops, in the T1 generation (Fig. 3b). The evaluation of the uniform and rapid germination after sowing showed that the cr-vp1-6-4, cr-vp1-8-3, cr-vp1-3-6 and cr-vp1-11-3 lines were significantly enhanced compared to that of WT plants (Fig. 4). Furthermore, the result of field trials showed no significant differences between T2 homozygous mutant lines and WT plants in terms of agronomic traits under normal field conditions (Fig. 5, Supplementary Fig. 7). This study provides a successful case for improving germination speed based on gene editing using CRISPR/Cas9 system. The knockout of OsVP1 demonstrated speeding up the germination of seeds and then as a result, in reducing the seed dormancy. Generally, GA induces synthesis of α-amylase in the aleurone layer, which then diffuses into the starchy endosperm where it initiates hydrolysis of starch (Bewley et al. 2013). In our experiments, the α-amylase activity of cr-vp1-6-4 line was clearly increased compared to WT (Fig. 6). These results suggest that knockout of the OsVP1 increased α-amylase activity and enhanced the hydrolysis of endosperm starch and promoted seed germination. A previous work (Sugimoto et al. 2010) on OsVP1 was reported that a 32-bp deletion in the first exon resulted in a regulation of seed dormancy and further domestication of rice. This mutant line showed an improved dormancy release and germination speed of matured seeds (Du et al. 2015). Also together with ABA, GA is associated with the regulation of seed dormancy and germination (Ogawa et al. 2003; Liu et al.2010; Nambara et al. 2010; Graeber et al. 2012). GA enhances germination by weakening the endosperm, increasing embryo growth potential and activating hydrolytic enzymes such as α -amylase in the seed. These results indicate that the starch preserved by α-amylase during the germination of cr-vp1-6-4 seeds was exposed to the amylolytic breakdown more rapidly than the WT (Fig. 6). In cereals like wheat, corn and rice, α-amylase plays an important role in hydrolyzing endosperm starch into metabolism able sugars for a vigorous seedling growth, and its complex synthesis is positively and negatively regulated by ABA/GA signaling, respectively (Bewley et al. 2013). Liu et al. (2011) revealed that ABA/GA signaling genes were five GA2ox, five NCED, four GA20ox and three CYP707A gene family members in rice. To further evaluate the impact of seed dormancy, we investigated the expression of ABA/GA signaling genes, including OsNCED2, OsGA20ox1, OsGA20ox2 and OsGA20ox3. In cr-vp1-6-4 line, the transcripts of OsNCED2, OsGA20ox1, OsGA20ox2 and OsGA20ox3 were significantly increased compare to the wild type. These data suggested that increasing expression of ABA/GA signaling genes could partially reduce the acquisition of seed dormancy of cr-vp1-6-4 line. Therefore, cr-vp1-6-4 line not only affects seed germination but also reduced the acquisition of seed dormancy.
Lastly, breeding strategy using CRISPR/Cas9 system by knockout of rice transcription factor has been demonstrated here and this can be an alternative approach for genetic improvement of rice. And this strategy could allow escaping the rigorous biosafety regulation done in transgenic plants generated with gene transfer.
References
Bae S, Park J, Kim JS (2014) Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30:1473–1475. https://doi.org/10.1093/bioinformatics/btu048
Belhaj K, Chaparro-Garcia A, Kamoun S, Nekrasov V (2013) Plant genome editing made easy: targeted mutagenesis in model and crop plants using the CRISPR/Cas system. Plant Methods 9:39. https://doi.org/10.1186/1746-4811-9-39
Bewley JD (1997) Seed germination and dormancy. Plant Cell 9:1055–1066
Bewley JD, Bradford KJ, Hilhorst HWM, NonogakiH (2013) Seeds: physiology of development, germination and dormancy, 3rd edn. Springer, Heidelberg, pp 249–250. https://doi.org/10.1007/978-1-4614-4693-4https://doi.org/10.1007/978-1-4614-4693-4
Bortesi L, Fischer R (2015) The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol Adv 33:41–52. https://doi.org/10.1016/j.biotechadv.2014.12.006
Cai HW, Morishima H (2000) Genomic regions affecting seed shattering and seed dormancy in rice. Theor Appl Genet 100:840–846. https://doi.org/10.1007/s001220051360
Cho SW, Kim S, Kim JM, Kim JS (2013) Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat Biotechnol 31:230–232. https://doi.org/10.1038/nbt.2507
Dong Y, Tsuzuki E, Kamiunten H, Terao H, Lin D, Matsuo M, Zheng Y (2003) Identification of quantitative trait loci associated with pre-harvest sprouting resistance in rice (Oryza sativa L.). Field Crop Res 81:133–139. https://doi.org/10.1016/S0378-4290(02)00217-4
Du W, Cheng J, Cheng Y, Wang L, He Y, Wang Z, Zhang H (2015) Physiological characteristics and related gene expression of after-ripening on seed dormancy release in rice. Plant Biol 17:1156–1164. https://doi.org/10.1111/plb.12371
Finch-Savage WE, Leubner-Metzger G (2006) Seed dormancy and the control of germination. New Phytol 171:501–523. https://doi.org/10.1111/j.1469-8137.2006.01787.x
Flintham J, Adlam R, Bassoi M, Holdsworth M, Gale M (2002) Mapping genes for resistance to sprouting damage in wheat. Euphytica 126:39–45. https://doi.org/10.1023/A:1019632008244
Fujino K, Sekiguchi H, Matsuda Y, Sugimoto K, Ono K, Yano M (2008) Molecular identification of a major quantitative trait locus, qLTG3–1, controlling low-temperature germinability in rice. Proc Natl Acad Sci USA 105:12623–12628. https://doi.org/10.1073/pnas.0805303105
Gao Y, Liu J, Fan J (2011) Construction and transformation of RNAi vector of OsVP1 for a regulatory gene of pre-harvest sprouting in Oryza sativa. AGRIS 43:1321–1327
Graeber KAI, Nakabayashi K, Miatton E, Leubner-Metzger GERHARD, Soppe WJ (2012) Molecular mechanisms of seed dormancy. Plant Cell Environ 3:1769–1786. https://doi.org/10.1111/j.1365-3040.2012.02542.x
Gu XY, Kianian SF, Foley ME (2004) Multiple loci and epistases control genetic variation for seed dormancy in weedy rice (Oryza sativa). Genetics 166:1503–1516. https://doi.org/10.1534/genetics.166.3.1503
Gutierrez L, Van Wuytswinkel O, Castelain M, Bellini C (2007) Combined networks regulating seed maturation. Trends Plant Sci 12:294–300. https://doi.org/10.1016/j.tplants.2007.06.003
Han Y, Luo D, Usman B, Nawaz G, Zhao N, Liu F, Li R (2018) Development of high yielding glutinous cytoplasmic male sterile rice (Oryza sativa L.) lines through CRISPR/Cas9 based mutagenesis of Wx and TGW6 and proteomic analysis of anther. Agronomy 8:290. https://doi.org/10.3390/agronomy8120290
Hattori M, Adachi H, Tsujimoto M, Arai H, Inoue K (1994) Miller-Dieker lissencephaly gene encodes a subunit of brain platelet-activating factor. Nature 370:216. https://doi.org/10.1038/370216a0
Kermode AR (2005) Role of abscisic acid in seed dormancy. J Plant Growth Regul 24:319–344. https://doi.org/10.1007/s00344-005-0110-2
Kim H, Kim ST, Ryu J, Choi MK, Kweon J, Kang BC, Kim SG (2016) A simple, flexible and high-throughput cloning system for plant genome editing via CRISPR-Cas system. J Integr Plant Biol 58:705–712. https://doi.org/10.1111/jipb.12474
Koornneef M, Hanhart CJ, Hilhorst HW, Karssen CM (1989) In vivo inhibition of seed development and reserve protein accumulation in recombinants of abscisic acid biosynthesis and responsiveness mutants in Arabidopsis thaliana. Plant Physiol 90:463–469. https://doi.org/10.1104/pp.90.2.463
LeClere S, Tellez R, Rampey RA, Matsuda SPT, Bartel B (2002) Characterization of a family of IAA amino acid conjugate hydrolases from Arabidopsis. J Biol Chem 277:20446–20452. https://doi.org/10.1074/jbc.M111955200
Li C, Zhou A, Sang T (2006) Rice domestication by reducing shattering. Science 311:1936–1939. https://doi.org/10.1126/science.1123604
Li M, Li X, Zhou Z, Wu P, Fang M, Pan X, Li H (2016) Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas9 system. Front Plant Sci 7:377. https://doi.org/10.3389/fpls.2016.00377
Liu F, Marquardt S, Lister C, Swiezewski S, Dean C (2010) Targeted 3′ processing of antisense transcripts triggers Arabidopsis FLC chromatin silencing. Science 327:94–97. https://doi.org/10.1126/science.1180278
Liu Y, Xu Y, Xiao J, Ma Q, Li D, Xue Z, Chong K (2011) OsDOG, a gibberellin-induced A20/AN1 zinc-finger protein, negatively regulates gibberellin-mediated cell elongation in rice. J Plant Physiol 168:1098–1105. https://doi.org/10.1016/j.jplph.2010.12.013
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lu B, Xie K, Yang C, Wang S, Liu X, Zhang L, Wan J (2011) Mapping two major effect grain dormancy QTL in rice. Mol Breed 28:453–462. https://doi.org/10.1007/s11032-010-9495-0
Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R, Xie Y (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8:1274–1284. https://doi.org/10.1016/j.molp.2015.04.007
Martín-Pizarro C, Triviño JC, Posé D (2018) Functional analysis of the TM6 MADS-box gene in the octoploid strawberry by CRISPR/Cas9-directed mutagenesis. J Exp Bot 70:885–895. https://doi.org/10.1093/jxb/ery400
McCarty DR, Hattori T, Carson CB, Vasil V, Lazar M, Vasil IK (1991) The Viviparous-1 developmental gene of maize encodes a novel transcriptional activator. Cell 66:895–905. https://doi.org/10.1016/0092-8674(91)90436-3
Nambara E, Okamoto M, Tatematsu K, Yano R, Seo M, Kamiya Y (2010) Abscisic acid and the control of seed dormancy and germination. Seed Sci Res 20:55–67. https://doi.org/10.1017/S0960258510000012
Nishimura A, Aichi I, Matsuoka M (2006) A protocol for Agrobacterium-mediated transformation in rice. Nat Protoc 1:2796. https://doi.org/10.1038/nprot.2006.469
Ogawa M, Hanada A, Yamauchi Y, Kuwahara A, Kamiya Y, Yamaguchi S (2003) Gibberellin biosynthesis and response during Arabidopsis seed germination. Plant Cell 15:1591–1604. https://doi.org/10.1105/tpc.011650
Park J, Bae S, Kim JS (2015) Cas-Designer: a web-based tool for choice of CRISPR-Cas9 target sites. Bioinformatics 31:4014–4016. https://doi.org/10.1093/bioinformatics/btv537
Park J, Lim K, Kim JS, Bae S (2017) Cas-analyzer: an online tool for assessing genome editing results using NGS data. Bioinformatics 33:286–288. https://doi.org/10.1093/bioinformatics/btw561
Raz V, Bergervoet JH, Koornneef M (2001) Sequential steps for developmental arrest in Arabidopsis seeds. Development 128:243–252
Sugimoto K, Takeuchi Y, Ebana K, Miyao A, Hirochika H, Hara N, Yano M (2010) Molecular cloning of Sdr4, a regulator involved in seed dormancy and domestication of rice. Proc Natl Acad Sci USA 107:5792–5797. https://doi.org/10.1073/pnas.0911965107
Van Der Schaar W, Alonso-Blanco C, Léon-Kloosterziel KM, Jansen RC, Van Ooijen JW, Koornneef M (1997) QTL analysis of seed dormancy in Arabidopsis using recombinant inbred lines and MQM mapping. Heredity 79:190. https://doi.org/10.1038/hdy.1997.142
Zhou H, He M, Li J, Chen L, Huang Z, Zheng S, Zhuang C (2016) Development of commercial thermo-sensitive genic male sterile rice accelerates hybrid rice breeding using the CRISPR/Cas9-mediated TMS5 editing system. Sci Rep-UK 6:37395. https://doi.org/10.1038/srep37395
Acknowledgements
This research was supported by a Grant from the Next-Generation BioGreen 21 Program (Project no. PJ01368902), Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jung, Y.J., Lee, H.J., Bae, S. et al. Acquisition of seed dormancy breaking in rice (Oryza sativa L.) via CRISPR/Cas9-targeted mutagenesis of OsVP1 gene. Plant Biotechnol Rep 13, 511–520 (2019). https://doi.org/10.1007/s11816-019-00580-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-019-00580-x