Abstract
Phosphate (Pi) is one of the major nutrients for crop growth and yield. Although several studies have revealed a global view of Pi starvation responses in rice, the detailed features were not well-addressed. To identify differentially expressed genes associated with phosphate starvation on a genome scale, we analyzed transcriptome data of roots from 2-week old seedlings that were grown on Pi-sufficient or -deficient media for 7 and 21 days. Using publicly available RNA-sequencing data, we subsequently identified 820 up-regulated genes in roots under Pi starvation. Gene ontology enrichment analysis of these genes indicated that secondary metabolic processes are most significantly enriched under Pi starvation, and Pi transport and defense to biotic stress also play significant roles in response against Pi deficiency. Functional classification analysis using MapMan emphasizes the significance of transcription factors, such as MYB, WRKY, and bHLH, various transporters, and genes in secondary metabolic processes. Use of promoter trap lines or transgenic plants expressing the GUS reporter gene under the control of Pi starvation-inducible gene promoters confirmed the meta-expression patterns of two genes stimulated by Pi starvation, suggesting novel promoters for enhancing Pi use efficiency. In addition to the identification of two novel promoters for Pi starvation response, cis-regulatory elements for the regulation of Pi starvation are suggested. Overall, our study provides a global view of Pi starvation response based on transcriptome data and novel tools for improving PUE and Pi uptake in rice, a model crop plant.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phosphate (Pi) is one of the major nutrients of growth, development, and reproduction in crop plants and functions in energy metabolism and signal transduction cascades and regulates enzymatic activities (Schachtman et al. 1998). Root growth plasticity determines the survival of plants in continuously changing environmental conditions (Koevoets et al. 2016). Most rice ecosystems frequently suffer from Pi deficiency. Pi fertilizer is a limited and unrenewable resource that is supplied by only a few countries. In addition, Pi as a fertilizer is rapidly immobilized owing to fixation and microbial activity. Only approximately 20–30% of the Pi applied is utilized by plants, and the rest is lost (Cordell et al. 2011; Schroder et al. 2011). An increase in the efficiency of Pi use is a very important target for current and future crop breeding.
In rice, a model crop plant, the functions of 18 genes relating to Pi use efficiency (PUE) have been reported. The genes include five Pi transporters, two auxin response factors, three SYG1/Pho81/XPR1 (SPX) domain-containing proteins, two MYB-like transcription factors (TFs), an MYB TF, and a phosphatase (Yamamoto et al. 2012). Interestingly, most of them are significantly stimulated in conditions of Pi starvation. Transcriptome data is an effective way to address limitations in current understanding, quickly expand current knowledge, and provide an improved strategy to enhance PUE in crops.
Pi starvation transcriptomes in rice cultivars were revealed through microarray or RNA-seq technologies. Transcriptomes in rice under low Pi stress were analyzed using a BGI 60 K rice array (Li et al. 2009). RNA-seq-based mRNA transcriptome analysis was performed using rice seedling roots and shoots under Pi starvation and excess Pi conditions (Oono et al. 2011). Later, mRNA sequencing of four rice cultivars with different growth and physiological responses to Pi starvation were produced, and the potential for the development of novel strategies for improving tolerance to Pi starvation in rice was suggested (Oono et al. 2013). More intensive transcriptomes under Pi starvation using RNA-seq technologies have been produced. The expression profiles for 126 samples were analyzed and revealed a very complex diversity of transcriptomes including short-, middle-, and long-term Pi starvation responses (Secco et al. 2013). Although several studies have reported global transcriptome responses to Pi starvation in rice, the functional analysis of candidate genes in roots that show primary responses to Pi starvation has been less intensively studied. The functions of 47 genes obtained through genetic and molecular studies have been reported and are well-summarized in the Overview of functionally characterized Genes in Rice Online (OGRO) database (http://qtaro.abr.affrc.go.jp/ogro) (Yamamoto et al. 2012). The functions of the remaining genes could be studied with gene indexed mutants produced by the use of T-DNA or transposons, such as Tos17 or Ac/Ds, or ds/espm insertion and then used for further functional genomic studies. At present, 65% of Rice Genome Annotation Project (RGAP) non-transposable element (TE) genes have gene indexed mutants (Chandran et al. 2016). Categorization of candidate genes from previous studies can be improved through an advanced integrating omics analysis.
In this study, we focus on 820 genes with a significant up-regulation in rice roots under long-term Pi starvation and the recovery process obtained through the reanalysis of previous global transcriptome data produced by Secco et al. (2013). MapMan analysis was used to categorize our candidate genes in terms of metabolic and regulatory pathways and transporting activities, which may assist researchers in prioritizing the candidate genes for further studies. In addition, we validated the expression patterns of two candidate genes showing up-regulation under Pi starvation through a GUS reporter system/promoter-GUS transgenic line and RT-PCR analyses, providing a new tool to optimize the function of candidate genes for enhanced PUE. Analysis of functionally characterized candidate genes identified in this study support the significance of selected candidate genes for further study to enhance PUE. Detailed data analyses and discussion are presented.
Materials and methods
Plant growth
The experiment was conducted using rice cultivar Dongjin seedlings that were first germinated for 2 weeks in Yoshida solution and then grown for 7 or 21 days in Pi-sufficient (0.320 mM Pi) or -deficient Yoshida solution (Yoshida et al. 1976). The pH of the culture solution was adjusted to 5.5 using 1 M NaOH. The Yoshida solution was replaced with new solution every 3 days. In all the hydroponic experiments, seedlings were directly grown in each culture solution (8 L) with an 8-h light (28°C)/16-h dark (22 °C) photoperiod. To verify that the samples responded to the Pi starvation treatment, we assessed the expression patterns of two Pi starvation marker genes, SULFOQUINOVOSYLDIACYLGLYCEROL 2 (OsSQD2, LOC_Os01g04920) and PHOSPHATE TRANSPORTER 6 (OsPT6, LOC_Os08g45000), through RT-PCR and qRT-PCR analyses.
Global identification of rice genes stimulated under phosphate starvation using RNA-sequencing data
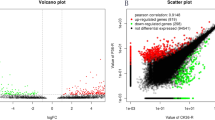
We used RNA-seq data obtained under Pi starvation conditions (Secco et al. 2013). Fragments per kilobase of transcript per million mapped reads (FPKM) values from whole samples used in this study were downloaded from the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/sra) under accession number SRA097415. We then generated fold changes of roots under Pi starvation for 1 h, 6 h, 24 h, 3 days, 7 days, 21 days, + 1 h, 21 days + 6 h, and 21 days + 24 h with roots under normal conditions. In addition, fold changes were generated to compare roots under recovery conditions (+ 1, + 6, + 24 h) after Pi starvation for 21 days over roots under Pi starvation for 21 days + 1 h, 21 days + 6 h, and 21 days + 24 h. With this fold change data, we carried out k-means clustering analysis and produced 10 clusters with differential expression patterns. As a result, we identified 820 candidate genes showing more than a significant log2 2.5-fold change up-regulation at 21 days under Pi starvation from three clusters (Fig. 2).
Analysis of phosphate starvation-inducible genes with known function
To determine the functional features of our candidate genes associated with Pi starvation, we searched the OGRO database, which contains summaries of rice genes with known functions obtained by genetic and molecular studies (Yamamoto et al. 2012). Of our 820 candidate genes, the functions of 47 have been reported and are summarized in Table 1.
MapMan analysis
Rice MapMan classification covers 36 BINs, and these BINs can be extended in a hierarchical manner into subBINs (Usadel et al. 2005; Urbanczyk-Wochniak et al. 2006). Using diverse MapMan tools, significant gene lists selected from high-throughput data analysis can be integrated into diverse overviews. To do this, we generated a dataset carrying locus IDs from the Rice Genome Annotation Project and the average log2 fold-change data of Pi starvation/control. For the functional classification of 820 genes up-regulated in Pi starvation conditions, we used Metabolism_overview (Fig. 3a), Transporter_overview (Fig. 3b), Transcription_overview (Fig. 3c), and Regulation_overview (Fig. 3d). The detailed information is presented in Table S1.
Gene ontology enrichment analysis
The gene ontology (GO) enrichment tool installed in the Rice Oligonucleotide Array Database (ROAD, http://www.ricearray.org/analysis/go_enrichment.shtml) (Cao et al. 2012) was used to determine the biological roles of high-throughput candidate genes (Jung et al. 2008). We applied GO enrichment tools for 820 genes up-regulated in rice root after 21 days of Pi starvation. Based on a p value < 0.05 and fold enrichment value > 2, we selected 36 significant GO terms (Fig. 4 and Table S2).
Construction of promoter-GUS vectors and GUS assay
The promoter region (− 1 to − 2411 bp from the initiation ATG codon) of LOC_Os02g44654/Cytochrome p450 was amplified with the 5′-GGTACCTAGTAGCCGTATCACATTAC-3′ (Kpn I)/5′-TCTAGAGG GTCTGGTCTACCTCAAGGCG-3′ (Xba I) primer set through genomic DNA PCR analysis. The resulting promoter DNA fragment was placed upstream of the beta-glucuronidase (GUS) reporter gene located in binary vector pGA3519 (Kim et al. 2009). Transgenic plants harboring the above construct were obtained in the japonica cultivar Dongjin background through the Agrobacterium tumefaciens mediating transformation method (Lee et al. 1999). To analyze GUS expression patterns using the GUS reporter system, we grew the promoter-GUS transgenic line for 21 days on Pi-sufficient or -deficient media after growing germinated plants for 2 weeks using Yoshida solution (Yoshida et al. 1976). Whole seedlings were soaked for 1 h in GUS staining solution. GUS staining assay was performed as previously described (Jefferson et al. 1987) on 21-day-old seedlings of one transgenic line for LOC_Os02g44654 encoding cytochrome p450 (Fig. 5a–c) carrying promoter-GUS vector and promoter trap lines for LOC_Os01g52560 (1D-014-17) (Fig. 5d–f). Photographs of GUS assayed whole plants were produced using an EOS 560 digital camera (Cannon, Tokyo, Japan), and those of roots were obtained using an Olympus microscope BX61 (Olympus, Tokyo, Japan).
Cis-acting element analysis
To identify conserved cis-acting elements (CREs) in the promoters of LOC_Os01g52560, LOC_Os02g44654, and LOC_Os02g44654 genes showing up-regulation under Pi starvation, we extracted 2-kb sequences upstream of ATG of these genes from PLANTPAN (http://plantpan2.itps.ncku.edu.tw/) (Chow et al. 2016), and CREs in the promoters were analyzed using OSIRIS (Morris et al. 2008) and PLACE (Higo et al. 1998). CREs in the promoters were aligned with the Motif Alignment and Search Tool (MAST) (Bailey et al. 2015). The results are displayed in Fig. 6, and known target motifs with a p value ≤ 0.05 were selected for further analysis.
Results
Morphological features of rice seedlings grown under low phosphate conditions
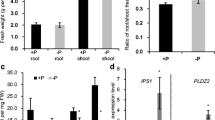
We identified the morphological features of rice cultivar Dongjin seedlings grown for 7 days (Fig. 1a) or 21 days (Fig. 1b) on Pi-sufficient or -deficient media. It was shown that Pi plays particularly important roles in early root development of rice (Fig. 1a). The numbers of leaves and tillers are reduced due to low Pi supplement. The length of shoots is also shorter than that of rice grown under normal conditions. Young leaves appear to be healthy, but older leaves turn brown (Fig. 1a). For evaluating the quality of these samples, we assessed the expression patterns of two Pi starvation-inducible marker genes (Fig. 1c, d). As a result, we found that two marker genes OsSQD2 (LOC_Os01g04920) (Yu et al. 2002) and OsPT6 (LOC_Os08g45000) (Ai et al. 2009) are significantly up-regulated under Pi starvation of 7 and 21 days. This result indicates that root samples grown under Pi starvation for 7 and 21 days are relevant for analyzing the physiological response.
Morphological appearance under phosphate starvation and schematic diagram of samples used in transcriptome analysis. Morphological appearance of Rice cultivar Dongjin seedling grown for 7 days (a) or 21 days (b) on Pi-sufficient or -deficient media after germination for 2 weeks using Yoshida solution. Preferential expression patterns based on qRT-PCR (c) and RT-PCR analysis (d) for two marker genes: OsSQD2 (LOC_Os01g04920) and OsPT6 (LOC_Os08g45000). OsUBI5 (LOC_Os01g22490) was used as internal control for both analyses. Bar = 1 cm. *P value < 0.05; **P value < 0.01; ***P value < 0.001
Identification of phosphate starvation-responsive genes using RNA-seq data
Transcriptome analysis was conducted with publicly available RNA-sequencing data using the roots of 2-week old seedlings exposed to Pi starvation for 7 and 21 days in Pi-sufficient or -deficient conditions or when Pi is re-supplemented (additional 0.320 mM Pi at 1, 6, or 24 h after 21 days). The RNA-seq data were used under the criteria of more than 5 FPKM values in Pi starvation, less than 0.5 p value, and more than 2.5 (log2 scale) fold up-regulation in response compared to the control (rice roots under sufficient Pi conditions). Subsequently, we identified 820 up-regulated genes in rice root under conditions of Pi starvation (Fig. 2a).
Functional evaluation of phosphate starvation-inducible genes using literature searches
To evaluate the functional significance of our candidate genes, we searched the literature to determine whether their functions have been reported previously. Out of 820 up-regulated genes under Pi deficiency in rice roots, we identified 47 with known functions through analysis using the OGRO database, which contains functionally characterized rice genes obtained through genetic and molecular studies (Yamamoto et al. 2012). Of the 47 genes with known functions, 10 are involved in the efficiency of Pi use and include OsPT2 (Zhang et al. 2014), OsPT6 (Ai et al. 2009), OsPT8 (Jia et al. 2011), OsPT9, OsPT10 (Lapis-Gaza et al. 2014), OsSPX1, OsSPX3 (Wang et al. 2009), OsSPX2 (Wang et al. 2014), and Purple acid phosphatase 10 a (OsPap10a) (Wang and Liu 2012). The functions of nine genes are reported to be associated with root growth and development, and they include rice OsDwarf10 (OsD10) (Arite et al. 2007), rice cis-Zeatin-O-glucosyltransferase (OscZOGT2) (Kudo et al. 2012), rice REPETITIVE PROLINE-RICH PROTEIN 1.1 (OsRePRP1.1), OsRePRP1.2 (Tseng et al. 2013), rice ATP-binding cassette transporter5 (OsABCG5) (Shiono et al. 2014), rice Ca2+/calmodulin-dependent protein kinase (OsCCaMK) (Bao et al. 2014), OsPT2 (Zhang et al. 2014), OsPT9, and OsPT10 (Lapis-Gaza et al. 2014). OsD10 is a rice ortholog of MAX4/RMS1/DAD1 that encodes carotenoid cleavage dioxygenase 8 (CCD8) and is believed to be involved in the synthesis of an unidentified inhibitor of shoot branching by strigolactones (SL) (Arite et al. 2007). Recently, osd10 mutants were found to have a loss of sensitivity in root responses under Pi deficiency (Sun et al. 2014; Xi et al. 2015). Mutations in rice regulators of SL, such as NODULATION SIGNALING PATHWAY 1 (OsNSP1) and OsNSP2, induced an accumulation of SL during Pi starvation (Yokota et al. 2010; Liu et al. 2011). Moreover, SLs are mediators of root growth under low Pi conditions (Koltai and Kapulnik 2011) and play an important role in the Pi uptake of plants. The functions of 18 of the 47 genes, including OsD10, OsCPS1, and OsKS1, are related to length growth/elongation of the plant (Table 1). These results indicate that Pi starvation-inducible genes might also have roles in regulating morphological or physiological traits. Additionally, five genes, including OsJAZ8 and OsWRKY76, are related to pathogen resistance (Table 1). Taken together, these findings appear to have an important role in root development, length growth, and biotic stress tolerance in association with the efficiency of Pi use, and there is the possibility of crosstalk between Pi uptake and these agronomic traits.
MapMan analysis of phosphate deficiency-related genes in rice roots
For the functional classification of 820 genes up-regulated under Pi starvation, we used the MapMan toolkit. Among them, 141 elements were assigned to the ‘miscellaneous function’ category, 81 to ‘protein’, 72 to ‘secondary metabolism’, 67 to ‘transporter’, 64 to ‘stress’, 58 to ‘RNA-regulation’, 41 to ‘lipid metabolism’, 40 to ‘signaling’, 32 to ‘hormone’, and the rest to other functional groups (Table S1). We selected four MapMan overviews for further analysis: metabolism_overview (Fig. 3a), transporter_overview (Fig. 3b), transcription_overview (Fig. 3c), and regulation_overview (Fig. 3d). With regard to metabolism overview, we found that the secondary metabolism category (72 elements) has the closest links with up-regulation under Pi starvation in rice root. From the transport overview, we found that genes involved in phosphate transporters and ATP-binding cassette (ABC) transporters are more abundant than others. From the cellular response overview, we found that the Pi starvation response is likely to be associated with the biotic stress response. In the ‘RNA-regulation’ category, we found that MYB, WRKY, and bHLH TFs are more important than others in response to Pi starvation. The remaining 273 genes did not have assigned MapMan terms. As in the GO enrichment analysis above, the functional category ‘transport’, including ‘ion transport’, ‘cation transport’, and ‘metal ion transport’, was more frequent under Pi starvation conditions.
MapMan analysis of genes associated with phosphate starvation in rice root. Secondary metabolism overview (a), Transporter (b), Transcription (c), and Regulation (d) were mapped with selected phosphate starvation-inducible genes in rice root. Red boxes indicate groups of genes up-regulated by phosphate starvation. Details are presented in Table S1
Gene ontology enrichment analysis reveals the significance of 36 biological processes
To determine the biological functions of Pi deficiency-inducible genes in rice root, we performed GO term enrichment analysis of 820 genes up-regulated by Pi deficiency in root. GO term enrichment analysis suggests that there were significant GO terms in the biological process categories associated with Pi starvation (Table S2). Through the use of GO enrichment analysis data with a criteria of ≥ twofold enrichment value and ≤ 0.05 hyper-geometric p value, we showed that secondary metabolic processes such as ‘Diterpene phytoalexin precursor biosynthetic process pathway’ (26.4-fold enrichment value), ‘l-phenylalanine catabolic process’ (24.0), and ‘Phenylpropanoid metabolic process’ (22.6); biological processes relating to defense response such as ‘defense response to fungus’ (17.59), ‘response to biotic stimulus’ (13.3), and ‘defense response to bacterium’ (11.7); and biological processes related to various transports such as ‘phosphate transport’ (22.6), ‘tetracycline transport’ (3.9), ‘metal ion transport’ (3.7), and ‘lipid transport’ (3.0) are highly enriched in Pi starvation-induced genes (Fig. 4).
Validation of phosphate deficiency-inducible genes in rice roots using the GUS reporter system
The GUS reporter gene system has been used to identify in vivo activity of promoters for the regulation of gene expression (Jefferson 1989). From RNA-seq data of samples under Pi starvation for 21 days compared to Pi-sufficient conditions (Control), we identified 40 genes showing the highest significant up-regulation under Pi starvation. Then, we screened a GUS reporter system-based promoter trap lines of 32 genes and carried out co-segregation analysis of the T-DNA insertion and GUS expression in T2 seedlings of these lines. As a result, the promoter trap lines showed the co-segregation of Pi starvation-induced GUS expression patterns and T-DNA insertion: Line 1D-014-17, a promoter trap line for LOC_Os01g52560, displayed GUS expression at the elongated region in seedling roots under Pi starvation for 3 weeks (Fig. 5 and Figure S1). In addition, we produced a promoter-GUS transgenic line for LOC_Os02g44654 encoding CytochromeP450 and found that this line showed a similar expression pattern with the line 1D-014-17 under Pi starvation. Interestingly, application of the promoter trap system or promoter-GUS system combined with qualified genome-wide transcriptome data is a very effective way to quickly identify the activity of a novel promoter.
Validation of expression patterns of two phosphate starvation-inducible genes using a promoter trap system and a promoter-GUS transgenic line. Expression patterns of promoter-GUS transgenic line for LOC_Os02g44654 (a–c) and a promoter trap line (1D-014-17, D-F) for LOC_Os01g52560 were analyzed under Pi starvation for 3 weeks a. Roots (a, b, d, e) and leaf (c, f) were then incubated for 3 h in GUS staining solution. Additional data of the validation using GUS reporter system for expression patterns of two genes under Pi-sufficient or deficient condition are presented in Figure S1. Validation of expression patterns using qRT-PCR analysis under phosphate starvation for LOC_Os02g44654 and LOC_Os01g52560 was prepared, and OsSQD2 (LOC_Os01g04920) was used as a positive control for inducible expression under phosphate starvation (g). OsUBI5 (Os01g22490) was used as an internal control for qRT-PCR analysis. *p value < 0.05; **p value < 0.01; ***p value < 0.001
Cis-acting element analysis of three phosphate starvation-inducible promoters confirmed by the GUS reporter system
To find CREs for triggering Pi deficiency response in rice roots, we used promoter regions of LOC_Os02g44654 and LOC_Os01g52560 that showed Pi starvation-inducible expression patterns confirmed using GUS reporter systems. The promoters were analyzed using the PLANTPAN 2.0 database (http://plantpan2.itps.ncku.edu.tw) (Chow et al. 2016), PLACE (Higo et al. 1998), and MEME tools (Bailey et al. 2015). As a result, we found 65 common CREs between them; from these, we obtained five consensus CREs in the promoters of the genes as candidate CREs responsible for Pi starvation-inducible expression: P1BS/GNATATNC, WBBOXPCWRKY1/TTTGACY, MYB2CONSENSUSAT/YAACKG, Helix-loop-helix DNA-binding domain/NSCACGTGSN, and IRO2OS/CACGTGG. Of these CREs, P1BS was the most frequently identified in all promoter regions (Fig. 6). P1BS is the binding site of an MYB domain transcription factor, PHR1. P1BS is present in the promoters of many crucial Pi-responsive genes (Rubio et al. 2001; Schunmann et al. 2004; Bustos et al. 2010; Nilsson et al. 2010; Oropeza-Aburto et al. 2012). WBBOXPCWRKY1 is one of the WRKY box (W-box) elements commonly found in the promoters of genes relevant for Pi-retranslocation and scavenging including PT1, PT3, PT4, PT5, PT7, INDUCED BY PHOSPHATE STARVATION 1 (IPS1), ACID PHOSPHATASE GENES2 (PS2), Purple Acid Phosphatase (PAP11), and PHOSPHATE STARVATION RESPONSE 1 (PHR1) (Devaiah et al. 2007). MYB2CONSENSUSAT is a MYB recognition site found in the promoters of the dehydration-responsive gene in Arabidopsis. The bHLH transcription factor-binding domain (NSCACGTGSN) is the most similar to E-box (CANNTG) and is found in all promoters of OsPT genes except OsPT1, OsPT6, and OsPT8 (Hatorangan et al. 2009) and phosphate starvation-induced bHLH transcription factor (OsPTF1) (Yi et al. 2005). In addition, IRO2OS is also an OsIRO2-binding core sequence that is an essential regulatory element of the genes involved in Fe uptake under Fe-deficient conditions (Ogo et al. 2006). Recent studies in response to Pi starvation revealed its relationship with the accumulation of iron (Hirsch et al. 2006; Misson et al. 2005). Based on these findings, P1BS and WBBOXPCWRKY1 might be related to Pi starvation-inducible expression, and bHLH transcription factor-binding domain and MYB2CONSENSUSAT might be associated with crosstalk between various TFs and Pi starvation-inducible expression. The other CREs not mentioned here could have novel roles in driving Pi starvation-inducible expression. Future experiments will be necessary to confirm our predictions (Table S3).
Identification of conserved cis-acting elements of phosphate starvation-inducible genes. Conserved CRE was indicated by node color red nodes indicate P1BS motif, WBBOXPCWRKY1 motif; light green nodes, MYB2CONSENSUSAT motif; purple nodes, Helix-loop-helix DNA-binding domain motif; gray nodes, IRO2OS motif; and orange nodes. Detailed information is presented in Table S3
Analyses of predicted protein–protein interactions associated with phosphate starvation
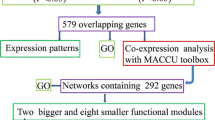
Regulatory genes are primary targets when investigating diverse stress responses and developmental processes. Among the 820 genes that were up-regulated under Pi starvation, we identified 22 TFs, 43 protein degradation and modification pathways, 16 transporters, and 47 functionally characterized genes (Fig. 7, Table S4, and Table S5). Understanding the regulatory relationships among them might improve our ability to develop novel strategies for enhancing PUE. To do this, we utilized the Rice Interactions Viewer to generate a hypothetical protein–protein interaction network associated with the up-regulated genes mentioned above. The network was further refined by highlighting genes in two categories: 60 functionally characterized genes and 52 up-regulated genes under low Pi in rice root. Of the latter, 31 genes have been functionally characterized. We also highlighted elements with multiple interactions among these regulatory genes or functionally characterized genes. This network data will be a useful resource in designing the hypothetical molecular model for detailed regulatory pathways in response to low Pi stress. Details on locus IDs, gene names, and putative functions of elements used in this refined network are provided in Table S5.
Construction of a regulatory network associated with genes up-regulated in root under phosphate starvation. Predicted protein–protein interaction analysis using the Rice interaction viewer and Cytoscape 3.2.1 is prepared. We queried predicted protein to protein interaction network associated with functionally characterized 47 genes (large hexagons) from the OGRO database and 59 up-regulated regulatory genes under low Pi in rice root (red dotted circles or red dotted hexagons). In total, 197 interactions with 209 proteins including 21 up-regulated uncharacterized genes under low Pi in rice root and 60 functionally characterized genes were visualized. Functional classifications using MapMan annotation were indicated by 13 node colors Detailed information of the network is presented in Table S4 and S5
Discussion
Pi is a major nutrient with unequivocal importance for plant development such as root development, shoot length, and the number of tillers. Our data has shown that root development is enhanced for the first 7 days under low Pi conditions, but after 21 days, low Pi causes growth retardation and brown roots. The global understanding on the molecular mechanism of rice roots in response to low Pi stress will be very important in studies to enhance PUE. MapMan and GO enrichment analyses suggest that the Pi starvation response primary affect root development, secondary metabolism process, biotic stress response, and various transporter activities.
Of all the enriched GO terms, ‘diterpene metabolic process’ was the most significantly enriched by Pi starvation in rice roots. Rice produces diterpene phytoalexins, such as momilactones, oryzalexins, and phytocassanes. It was also reported that the production and secretion of momilactone A and B, which act as potent phytoalexins and allelochemicals, are increased in rice roots under various stress conditions (Kato-Noguchi 2011; Zhao et al. 2005; Kong et al. 2006). Recently, it was reported that rice exhibited high allelopathic activity when incubated under Pi- or nitrogen-limited conditions (Shen and Lin 2007; Song et al. 2008), indicating that the allelopathic activity of rice may be increased by nutrient starvation. Kim et al. (2005) also reported that nutrient competitive conditions with barnyard grass increased allelopathic activity in aqueous extracts of rice plants. These results suggest that phytoalexin diterpene probably led to allelopathic activity in rice under nutrient competitive conditions. Strigolactone (SL) resulting from terpenoid biosynthetic processes accumulates in Arabidopsis and rice roots under Pi starvation and is known to suppress shoot branching by inhibiting the outgrowth of axillary buds (Gomez-Roldan et al. 2008) and to promote the elongation of seminal/primary roots and adventitious roots (ARs) (Kapulnik et al. 2011Arite et al. 2012). The elevated endogenous SL concentration by Pi deficiency contributes to inhibition of tiller bud outgrowth (Umehara et al. 2010) and enhances root development in rice (Sun et al. 2014; Kumar et al. 2015). Elevated endogenous SL-accumulating phenotypes are consistent with the major phenotypes seen in Pi deficiency. In addition, a protein, PLEIOTROPIC DRUG RESISTANCE 1 (PDR1), belonging to the ABC transporters is known to be involved in the long-distance transport of SLs from root to shoot and also in root tissues (Kretzschmar et al. 2012).
Furthermore, changes of the secondary metabolites in response to low Pi stress resulted in accumulation of anthocyanin and total phenolic compounds in leaves under long-term Pi deficiency (Misson et al. 2005). Similar results were observed in Zea mays, with genes related to the phenylpropanoid pathway identified as up-regulated or down-regulated under Pi starvation (Juszczuk et al. 2004; Calderon-Vazquez et al. 2008). Our data suggest that Pi deficiency stress may affect the accumulation of anthocyanin as well as lignin development in rice roots. In addition, secondary metabolites in rice may play an important role in enhancing survival rate under nutrient-deficient conditions.
From both MapMan and GO enrichment analyses, phosphate transporters were significantly identified in genes up-regulated under Pi starvation conditions, supporting the quality of samples used for transcriptome analysis. Interestingly, biotic stress or defense-related genes are also closely associated with Pi deficiency (Fig. 3e). We also identified other significant GO terms such as ‘DNA catabolic process’ and ‘respiratory gaseous exchange’ (Fig. 4). Therefore, the biological processes identified as being closely associated with Pi-limited conditions in rice root might be novel resources for improving our understanding of the molecular mechanism and PUE through further applications.
Under Pi starvation, the expression of the rice Phosphate transporter 1 (PHT1) gene group belonging to high-affinity transporters is strongly induced to increase the ability of the roots in acquiring Pi from soils and remobilizing Pi within plants (Mudge et al. 2003; Raghothama and Karthikeyan 2005). Most transporters in the PHT1 group are known to be strongly expressed in roots of Arabidopsis, Zea mays, and rice (Muchhal et al. 1996; Koyama et al. 2005; Nagy et al. 2006). For regulation of genes in the PHT1 group, TFs might have important roles. From the transcription overview installed in the MapMan toolkit, we found that MYB, WRKY, and bHLH TFs are more important than others. Previous studies have shown that MYB, WRKY, and bHLH TFs were involved in a regulation system under Pi starvation. PHR1, a MYB TF (Rubio et al. 2001); WRKY75, a WRKY family TF (Devaiah et al. 2007) in Arabidopsis; another WRKY family TF named OsWRKY74; and a bHLH transcription factor named OsPTF1 have been reported to play a role in providing tolerance against Pi starvation stress by improving root system architecture in rice (Dai et al. 2016; Yi et al. 2005). Other TFs than those mentioned might have important roles in regulating Pi starvation responses in rice.
Our study provides a global view of Pi starvation response based on transcriptome data and novel promoters for improving PUE and Pi uptake. This strategy is very promising for future applications because recent studies using both ubiquitous and root-preferred promoters for drought stress tolerance clarified that root-preferred promoters are more effective at enhancing OsNAC6, which is involved in the drought tolerance regulatory pathway (Lee et al. 2017). Histochemical GUS assays were carried out on 21-day-old seedlings under Pi starvation and Pi-sufficient conditions using Yoshida solution. Interestingly, a GUS staining pattern was observed only in initiated or elongated areas of crown roots and lateral roots under Pi starvation and was not identified in any tissue under sufficient Pi conditions (Fig. 4d). These expression patterns are consistent with physiological changes in the rice root under Pi deficiency. In addition, the promoter of OsPTF1, a known TF involved in tolerance to Pi starvation in rice, was expressed in all cells of newly developed lateral roots and in the elongation zone of primary roots in the Pi starvation condition (Yi et al. 2005). We anticipate that the use of a Pi starvation-inducible promoter will more effectively improve root quality and Pi uptake in a Pi-limited field than that of ubiquitous promoters.
References
Ai P, Sun S, Zhao J, Fan X, Xin W, Guo Q, Yu L, Shen Q, Wu P, Miller AJ, Xu G (2009) Two rice phosphate transporters, OsPht1;2 and OsPht1;6, have different functions and kinetic properties in uptake and translocation. Plant J 57:798–809
Arite T, Iwata H, Ohshima K, Maekawa M, Nakajima M, Kojima M, Sakakibara H, Kyozuka J (2007) DWARF10, an RMS1/MAX4/DAD1 ortholog, controls lateral bud outgrowth in rice. Plant J 51:1019–1029
Arite T, Kameoka H, Kyozuka J (2012) Strigolactone positively controls crown root elongation in rice. J Plant Growth Regul 31:165
Bailey TL, Johnson J, Grant CE, Noble WS (2015) The MEME suite. Nucleic Acids Res 43:e39–e49
Bao Z, Watanabe A, Sasaki K, Okubo T, Tokida T, Liu D, Ikeda S, Imaizumi-Anraku H, Asakawa S, Sato T, Mitsui H, Minamisawa K (2014) A rice gene for microbial symbiosis, Oryza sativa CCaMK, reduces CH4 Flux in a paddy field with low nitrogen input. Appl Environ Microbiol 80:995–2003
Bustos R, Castrillo G, Linhares F, Puga MI, Rubio V, Perez-Perez J, Solano R, Leyva A, Paz-Ares J (2010) A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis. PLoS Genet 6:e1001102
Calderon-Vazquez C, Ibarra-Laclette E, Caballero-Perez J, Herrera-Estrella L (2008) Transcript profiling of Zea mays roots reveals gene responses to phosphate deficiency at the plant- and species-specific levels. J Exp Bot 59:2479–2497
Cao P, Jung K-H, Choi D, Hwang D, Zhu J, Ronald PC (2012) The rice oligonucleotide array database: an atlas of rice gene expression. Rice 5:17
Chandran AKN, Lee GS, Yoo YH, Yoon UH, Ahn BO, Yun DW, Kim JH, Choi HK, An G, Kim TH, Jung KH (2016) Functional classification of rice flanking sequence tagged genes using MapMan terms and global understanding on metabolic and regulatory pathways affected by dxr mutant having defects in light response. Rice 9:17
Chow CN, Zheng HQ, Wu NY, Chien CH, Huang HD, Lee TY, Chiang-Hsieh YF, Hou PF, Yang TY, Chang WC (2016) PlantPAN 2.0: an update of plant promoter analysis navigator for reconstructing transcriptional regulatory networks in plants. Nucleic Acids Res 44:D1154–D1160
Cordell D, Rosemarin A, Schroder JJ, Smit AL (2011) Towards global phosphorus security: a systems framework for phosphorus recovery and reuse options. Chemosphere 84:747–758
Dai X, Wang Y, Zhang WH (2016) OsWRKY74, a WRKY transcription factor, modulates tolerance to phosphate starvation in rice. J Exp Bot 67:947–960
Devaiah BN, Karthikeyan AS, Raghothama KG (2007) WRKY75 transcription factor is a modulator of phosphate acquisition and root development in Arabidopsis. Plant Physiol 143:1789–1801
Gomez-Roldan V, Fermas S, Brewer PB, Puech-Pages V, Dun EA, Pillot JP, Letisse F, Matusova R, Danoun S, Portais JC, Bouwmeester H, Becard G, Beveridge CA, Rameau C, Rochange SF (2008) Strigolactone inhibition of shoot branching. Nature 455:189–194
Hatorangan MR, Sentausa E, Wijaya GY (2009) In silico identification of cis-regulatory elements of phosphate transporter genes in rice (Oryza sativa L.). J Crop Sci Biotechnol 12:25–30
Higo K, Ugawa Y, Iwamoto M, Higo H (1998) PLACE: a database of plant cis-acting regulatory DNA elements. Nucleic Acids Res 26:358–359
Hirsch J, Marin E, Floriani M, Chiarenza S, Richaud P, Nussaume L, Thibaud MC (2006) Phosphate deficiency promotes modification of iron distribution in Arabidopsis plants. Biochimie 88:1767–1771
Jefferson RA (1989) The GUS reporter gene system. Nature 342:837–838
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Jia H, Ren H, Gu M, Zhao J, Sun S, Zhang X, Chen J, Wu P, Xu G (2011) The phosphate transporter gene OsPht1;8 is involved in phosphate homeostasis in rice. Plant Physiol 156:1164–1175
Jung KH, Dardick C, Bartley LE, Cao P, Phetsom J, Canlas P, Seo YS, Shultz M, Ouyang S, Yuan Q, Frank BC, Ly E, Zheng L, Jia Y, Hsia AP, An K, Chou HH, Rocke D, Lee GC, Schnable PS, An G, Buell CR, Ronald PC (2008) Refinement of light-responsive transcript lists using rice oligonucleotide arrays: evaluation of gene-redundancy. PLoS One. https://doi.org/10.1371/journal.pone.0003337
Juszczuk IM, Wiktorowska A, Malusá E, Rychter AM (2004) Changes in the concentration of phenolic compounds and exudation induced by phosphate deficiency in bean plants (Phaseolus vulgaris L.). Plant Soil 267:41–49
Kapulnik Y, Delaux PM, Resnick N, Mayzlish-Gati E, Wininger S, Bhattacharya C, Sejalon-Delmas N, Combier JP, Becard G, Belausov E, Beeckman T, Dor E, Hershenhorn J, Koltai H (2011) Strigolactones affect lateral root formation and root-hair elongation in Arabidopsis. Planta 233:209–216
Kato-Noguchi H (2011) Convergent or parallel molecular evolution of momilactone A and B: potent allelochemicals, momilactones have been found only in rice and the moss Hypnum plumaeforme. J Plant Physiol 168:1511–1516
Kim SY, Madrid AV, Park ST, Yang SJ, Olofsdotter M (2005) Evaluation of rice allelopathy inhydroponics. Weed Res 45:74–79
Kim SR, Lee DY, Yang JI, Moon SO, An G (2009) Cloning vectors for rice. J Plant Biol 52:73
Koevoets IT, Venema JH, Elzenga JT, Testerink C (2016) Roots withstanding their environment: exploiting root system architecture responses to abiotic stress to improve crop tolerance. Front Plant Sci 7:1335
Koltai H, Kapulnik Y (2011) Strigolactones as mediators of plant growth responses to environmental conditions. Plant Signal Behav 6:37–41
Kong CH, Li HB, Hu F, Xu XH, Wang P (2006) Allelochemicals released by rice roots and residues in soil. Plant Soil 288:47–56
Koyama T, Ono T, Shimizu M, Jinbo T, Mizuno R, Tomita K, Mitsukawa N, Kawazu T, Kimura T, Ohmiya K, Sakka K (2005) Promoter of Arabidopsis thaliana phosphate transporter gene drives root-specific expression of transgene in rice. J Biosci Bioeng 99:38–42
Kretzschmar T, Kohlen W, Sasse J, Borghi L, Schlegel M, Bachelier JB, Reinhardt D, Bours R, Bouwmeester HJ, Martinoia E (2012) A petunia ABC protein controls strigolactone-dependent symbiotic signalling and branching. Nature 483:341–344
Kudo T, Makita N, Kojima M, Tokunaga H, Sakakibara H (2012) Cytokinin activity of cis-zeatin and phenotypic alterations induced by overexpression of putative cis-Zeatin-O-glucosyltransferase in rice. Plant Physiol 160:319–331
Kumar M, Pandya-Kumar N, Kapulnik Y, Koltai H (2015) Strigolactone signaling in root development and phosphate starvation. Plant Signal Behav 10:e1045174
Lapis-Gaza HR, Jost R, Finnegan PM (2014) Arabidopsis PHOSPHATE TRANSPORTER1 genes PHT1;8 and PHT1;9 are involved in root-to-shoot translocation of orthophosphate. BMC Plant Biol 14:334
Lee SH, Shon YG, Lee SI, Kim CY, Koo JC, Lim CO, Choi YJ, Han CD, Chung CH, Choe ZR, Cho MJ (1999) Cultivar variability in the Agrobacterium-rice cell interaction and plant regeneration. Physiol Plant 107:338–340
Lee DK, Chung PJ, Jeong JS, Jang G, Bang SW, Jung H, Kim YS, Ha SH, Choi YD, Kim JK (2017) The rice OsNAC6 transcription factor orchestrates multiple molecular mechanisms involving root structural adaptions and nicotianamine biosynthesis for drought tolerance. Plant Biotechnol J 15:754–764
Li L, Qiu X, Li X, Wang S, Lian X (2009) The expression profile of genes in rice roots under low phosphorus stress. Sci China C Life Sci 52:1055–1064
Liu W, Kohlen W, Lillo A, Op den Camp R, Ivanov S, Hartog M, Limpens E, Jamil M, Smaczniak C, Kaufmann K, Yang WC, Hooiveld GJ, Charnikhova T, Bouwmeester HJ, Bisseling T, Geurts R (2011) Strigolactone biosynthesis in Medicago truncatula and rice requires the symbiotic GRAS-type transcription factors NSP1 and NSP2. Plant Cell 23:3853–3865
Misson J, Raghothama KG, Jain A, Jouhet J, Block MA, Bligny R, Ortet P, Creff A, Somerville S, Rolland N, Doumas P, Nacry P, Herrerra-Estrella L, Nussaume L, Thibaud MC (2005) A genome-wide transcriptional analysis using Arabidopsis thaliana Affymetrix gene chips determined plant responses to phosphate deprivation. PNAS 102:11934–11939
Morris RT, O’Connor TR, Wyrick JJ (2008) Osiris: an integrated promoter database for Oryza sativa L. Bioinformatics 24:2915–2917
Muchhal US, Pardo JM, Raghothama KG (1996) Phosphate transporters from the higher plant Arabidopsis thaliana. PNAS 93:10519–10523
Mudge SR, Smith FW, Richardson AE (2003) Root-specific and phosphate-regulated expression of phytase under the control of a phosphate transporter promoter enables Arabidopsis to grow on phytate as a sole P source. Plant Sci 165:871–878
Nagy R, Vasconcelos MJ, Zhao S, McElver J, Bruce W, Amrhein N, Raghothama KG, Bucher M (2006) Differential regulation of five Pht1 phosphate transporters from maize (Zea mays L.). Plant Biol (Stuttg) 8:186–197
Nilsson L, Muller R, Nielsen TH (2010) Dissecting the plant transcriptome and the regulatory responses to phosphate deprivation. Physiol Plant 139:129–143
Ogo Y, Itai RN, Nakanishi H, Inoue H, Kobayashi T, Suzuki M, Takahashi M, Mori S, Nishizawa NK (2006) Isolation and characterization of IRO2, a novel iron-regulated bHLH transcription factor in graminaceous plants. J Exp Bot 57:2867–2878
Oono Y, Kawahara Y, Kanamori H, Mizuno H, Yamagata H, Yamamoto M, Hosokawa S, Ikawa H, Akahane I, Zhu Z, Wu J, Itoh T, Matsumoto T (2011) mRNA-Seq reveals a comprehensive transcriptome profile of rice under phosphate stress. Rice 4:50–65
Oono Y, Kawahara Y, Yazawa T, Kanamori H, Kuramata M, Yamagata H, Hosokawa S, Minami H, Ishikawa S, Wu J, Antonio B, Handa H, Itoh T, Matsumoto T (2013) Diversity in the complexity of phosphate starvation transcriptomes among rice cultivars based on RNA-Seq profiles. Plant Mol Biol 83:523–537
Oropeza-Aburto A, Cruz-Ramirez A, Acevedo-Hernandez GJ, Perez-Torres CA, Caballero-Perez J, Herrera-Estrella L (2012) Functional analysis of the Arabidopsis PLDZ2 promoter reveals an evolutionarily conserved low-Pi-responsive transcriptional enhancer element. J Exp Bot 63:2189–2202
Raghothama KG, Karthikeyan AS (2005) Phosphate acquisition. Plant Soil 274:37
Rubio V, Linhares F, Solano R, Martin AC, Iglesias J, Leyva A, Paz-Ares J (2001) A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev 15:2122–2133
Schachtman DP, Reid RJ, Ayling SM (1998) Phosphorus uptake by plants: from soil to cell. Plant Physiol 116(2):447–453
Schroder JJ, Smit AL, Cordell D, Rosemarin A (2011) Improved phosphorus use efficiency in agriculture: a key requirement for its sustainable use. Chemosphere 84:822–831
Schunmann PH, Richardson AE, Vickers CE, Delhaize E (2004) Promoter analysis of the barley Pht1;1 phosphate transporter gene identifies regions controlling root expression and responsiveness to phosphate deprivation. Plant Physiol 136:4205–4214
Secco D, Jabnoune M, Walker H, Shou H, Wu P, Poirier Y, Whelan J (2013) Spatio-temporal transcript profiling of rice roots and shoots in response to phosphate starvation and recovery. Plant Cell 25:4285–4304
Shen L, Lin W (2007) Effects of phosphorus levels on allelopathic potential of rice cocultured with barnyardgrass. Allelopathy J 19:393–402
Shiono K, Ando M, Nishiuchi S, Takahashi H, Watanabe K, Nakamura M, Matsuo Y, Yasuno N, Yamanouchi U, Fujimoto M, Takanashi H, Ranathunge K, Franke RB, Shitan N, Nishizawa NK, Takamure I, Yano M, Tsutsumi N, Schreiber L, Yazaki K, Nakazono M, Kato K (2014) RCN1/OsABCG5, an ATP-binding cassette (ABC) transporter, is required for hypodermal suberization of roots in rice (Oryza sativa). Plant J 80:40–51
Song B, Xiong J, Fang C, Qiu L, Lin R, Liang Y (2008) Allelopathic enhancement and differential gene expression in rice under low nitrogen treatment. J Chem Ecol 34:688–695
Sun H, Tao J, Liu S, Huang S, Chen S, Xie X, Yoneyama K, Zhang Y, Xu G (2014) Strigolactones are involved in phosphate- and nitrate-deficiency-induced root development and auxin transport in rice. J Exp Bot 65:6735–6746
Tseng IC, Hong CY, Yu SM, Ho TH (2013) Abscisic acid- and stress-induced highly proline-rich glycoproteins regulate root growth in rice. Plant Physiol 163:118–134
Umehara M, Hanada A, Magome H, Takeda-Kamiya N, Yamaguchi S (2010) Contribution of strigolactones to the inhibition of tiller bud outgrowth under phosphate deficiency in rice. Plant Cell Physiol 51:1118–1126
Urbanczyk-Wochniak E, Usadel B, Thimm O, Nunes-Nesi A, Carrari F, Davy M, Blasing O, Kowalczyk M, Weicht D, Polinceusz A, Meyer S, Stitt M, Fernie AR (2006) Conversion of MapMan to allow the analysis of transcript data from Solanaceous species: effects of genetic and environmental alterations in energy metabolism in the leaf. Plant Mol Biol 60:773–792
Usadel B, Nagel A, Thimm O, Redestig H, Blaesing OE, Palacios-Rojas N, Selbig J, Hannemann J, Piques MC, Steinhauser D, Scheible WR, Gibon Y, Morcuende R, Weicht D, Meyer S, Stitt M (2005) Extension of the visualization tool MapMan to allow statistical analysis of arrays, display of corresponding genes, and comparison with known responses. Plant Physiol 138:1195–1204
Wang L, Liu D (2012) Arabidopsis purple acid phosphatase 10 is a component of plant adaptive mechanism to phosphate limitation. Plant Signal Behav 7:306–310
Wang Z, Hu H, Huang H, Duan K, Wu Z, Wu P (2009) Regulation of OsSPX1 and OsSPX3 on expression of OsSPX domain genes and Pi-starvation signaling in rice. J Integr Plant Biol 51:663–674
Wang Z, Ruan W, Shi J, Zhang L, Xiang D, Yang C, Li C, Wu Z, Liu Y, Yu Y, Shou H, Mo X, Mao C, Wu P (2014) Rice SPX1 and SPX2 inhibit phosphate starvation responses through interacting with PHR2 in a phosphate-dependent manner. PNAS 111:14953–14958
Xi L, Wen C, Fang S, Chen X, Nie J, Chu J, Yuan C, Yan C, Ma N, Zhao L (2015) Impacts of strigolactone on shoot branching under phosphate starvation in chrysanthemum (Dendranthema grandiflorum cv. Jinba). Front Plant Sci 6:694
Yamamoto E, Yonemaru J, Yamamoto T, Yano M (2012) OGRO: the overview of functionally characterized Genes in Rice online database. Rice (N Y) 5:26
Yi K, Wu Z, Zhou J, Du L, Guo L, Wu Y, Wu P (2005) OsPTF1, a novel transcription factor involved in tolerance to phosphate starvation in rice. Plant Physiol 138:2087–2096
Yokota K, Soyano T, Kouchi H, Hayashi M (2010) Function of GRAS proteins in root nodule symbiosis is retained in homologs of a non-legume, rice. Plant Cell Physiol 51:1436–1442
Yoshida S, Forno DA, Cock JH, Gomez KA (1976) Laboratory manual for physiological studies of rice, vol 3. IRRI, Los Banos, pp 27–34
Yu B, Xu C, Benning C (2002) Arabidopsis disrupted in SQD2 encoding sulfolipid synthase is impaired in phosphate-limited growth. PNAS 99:5732–5737
Zhang L, Hu B, Li W, Che R, Deng K, Li H, Yu F, Ling H, Li Y, Chu C (2014) OsPT2, a phosphate transporter, is involved in the active uptake of selenite in rice. New Phytol 201:1183–1191
Zhao H, Li H, Kong C, Xu X, Liang W (2005) Chemical response of allelopathic rice seedlings under varying environmental conditions. Allelopathy J 15:105–110
Acknowledgements
This work was supported by grants from the Next-Generation BioGreen 21 Program (PJ01100401 to KHJ), and Research Program for Agriculture Science & Technology Development (Project title: Construction of forage rice genetic population for adaptation to reclaimed land, Project No. PJ01358001 to HMP), Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gho, YS., An, G., Park, HM. et al. A systemic view of phosphate starvation-responsive genes in rice roots to enhance phosphate use efficiency in rice. Plant Biotechnol Rep 12, 249–264 (2018). https://doi.org/10.1007/s11816-018-0490-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-018-0490-y