Abstract
ASYMMERTIC LEAVES1 (AS1) and AS2 in Arabidopsis are essential in determining leaf cell fates of adaxial axis. Here, we report ASYMMERTIC LEAVES2-LIKE38 (ASL38/LBD41), is an important gene of the LATERAL ORGAN BOUNDARY DOMAIN family. ASL38/LBD41 was detected in the adaxial and internal domain between the ab-adaxial domains of leaves. For explaining ASL38/LBD41 role in Arabidopsis development, a construct of the sense-expressing ASL38/LBD41 was transformed to Arabidopsis (Col-0); thus gained both overexpressing and cosuppressing 35S:ASL38/LBD41 plants. Rosette leaves of these overexpressing plants showed narrow (Class I) to radial symmetric needle-like patterns (Class II), compared with those of wild-type control, exhibiting adaxilized defect. However, rosette leaves of these cosuppressing plants showed narrow (Class I) to radial symmetric needle-like patterns (Class II), in contrast to those of wild-type (Col-0), exhibiting abaxilized defect. Furthermore, phenotypes observed in asl38-1 mutants suggest a redundant function for ASL38/LBD41 in boosting adaxial and/or inhibiting cell fate in abaxial axis. Together, our data suggest that ASL38/LBD41 might play a role of specialization in adaxial cell fate in Arabidopsis lateral organs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The primordium of leaf blades is as a result of the peripheral section in shoot apical meristem (SAM). When the primordia of leaves form, it is necessary to build the proximodistal, mediolateral, and ab/adaxial axes (Hudson 2000). The ab/adaxial factor might be most crucial of all among these axes (Bowman et al. 2002).
Recently, a lot of researches have revealed a few genes that function in building ad-abaxial axes of leaf blades. In this homeodomain/Leu zipper (HD-ZIP) (class III) family, five members function in special adaxiality of lateral organs (Emery et al. 2003; Prigge et al. 2005; McConnell et al. 2001; McConnell and Barton 1998). With miR165 and miR166 targeting the mRNAs of this family genes; in abaxial cells, the down-regulation is determined (Williams et al. 2005; Kim et al. 2005; Bao et al. 2004; Tang et al. 2003). Under the Ler (Landsberg erecta) background, as2 mutant shows that the pattern of adaxial cells is transformed into the pattern of abaxial cells (Xu et al. 2003). However, its overexpressing lines (35S:AS2) exhibit obviously adaxial and radial leaf blades (Xu et al. 2003; Lin et al. 2003). In the AS2/LOB domains, the LOB domain is essential, and it cannot be shifted via that of other members in the family (Matsumura et al. 2009). bop1 bop2 double mutants exhibit that adaxial cell pattern is transformed into abaxial cell pattern (Ha et al. 2007). On the other hand, in KANADI (KAN) and YABBY (YAB) families, some members exhibit abaxial pattern (Eshed et al. 1999; Sawa et al. 1999; Siegfried et al. 1999; Kerstetter et al. 2001; Emery et al. 2003; Eshed et al. 2004; Candela et al. 2008).
Here, we report that ASL38/LBD41 gene is one member of the Arabidopsis LATERAL ORGAN BOUNDARY DOMAIN family, and studies its mode in expression level. Our data indicate that during Arabidopsis leaf growth and development, such expression mode is essential. We constructed sense 35S:ASL38/LBD41 to transform Arabidopsis, and attained 35S:ASL38/LBD41 plants that were both overexpression and cosuppression mutants. By studying these mutants, our findings indicate that ASL38 function is specification of adaxial cell fates. Furthermore, phenotypes observed in asl38-1 mutant suggested that ASL38/LBD41 might play a character of specialization of adaxial cell fate in lateral organs of Arabidopsis.
Materials and methods
Arabidopsis material, constructing vector and genetic transformation
Arabidopsis plants were sown in MS medium for 7 days, and then transferred to green room and grown under 21 ± 2 °C (Meng 2015; Meng et al. 2015).
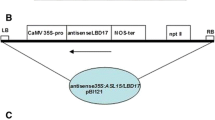
The ASL38/LBD41 (At3g02550) codes a zone, which is a DNA fragment (214–1005 in the full-length Arabidopsis ASL38/LBD41). This DNA fragment is enlarged derived from 20-day-old plant cDNA of Arabidopsis wild-type (Col-0). And then the DNA was built in the sense direction and seated in downstream of the 35S promoter (Fig. 2a). By sequencing, we confirm the 35S:ASL38/LBD41 construct.
The 35S:ASL38/LBD41 plants were produced via vacuum infiltration of Arabidopsis Col-0 by usualizing GV3101 strain (Meng and Yao 2015). All 35S:ASL38/LBD41 transgenic lines (T2) were confirmed through PCR via usualizing an nptII primer (Meng et al. 2009a, b).
Molecular biological experiments
Total RNA was secluded via usualizing Trizol. To perform RT-PCR research, we synthesized cDNA by 1 μg of total RNA by usualizing an oligo dT 18 primers and MMuLV Reverse Transcriptase. To perform PCR experiments, in each cDNA, 1/4 volume was selected, and usualized them as a template.
To perform ASL38/LBD41 and ASL37/LBD40 PCR amplification, conditions below is used, that is, at 94 °C for 4 min (for denaturation), and then for 40 cycles (for cosupression of ASL38/LBD41) and 33 cycles (for overexpression of ASL38/LBD41) of 94 °C for 45 s, 60 °C for 45 s, 72 °C for 1 min, at last at 72 °C for 7 min. The PCR primers: ASL38/LBD41-F (5′-tcggaaagggtgtagtgag-3′) and ASL38/LBD41-R (5′-aggacgaaggtgattgggac-3′); ASL37/LBD40-F (5′-tacgaaaaggctgcagtgaa-3′) and ASL38/LBD41-R (5′-ggtaccaccacgtgatttcc-3′) (Shuai et al. 2002).
For assaying the expressing change of abaxial polarity genes in 35S:ASL38/LBD41 lines, RT-PCR was implemented, its conditions are below in details: at 94 °C for 4 min (for denaturation), for 35 (for FIL, YAB and PHB) cycles of 94 °C for 40 s, 58 °C for 40 s, and 72 °C for 1 min, and at last, a incubation at 72 °C for 7 min. Gene-specific primers used were described by Lin et al. (2003).
Morphological analyses
We performed morphological experiments, which refer to methods in Fuchs (1963) and Baum and Rost (1996). Also we completed experiments of scanning electron microscopy using Quanta 200 FEG SEM (Alvarez et al. 1992). FIL probe was produced via cDNA and synthesized by using DIG-labeled antisense RNA via usualizing T7 RNA polymerase. FIL probes were produced, which refer to methods in Eshed et al. (2001), and ASL38/LBD41 probes were produced via usualizing ASL38/LBD41 F (5′-tcggaaagggtgtagtgag-3′) and R (5′-aggacgaaggtgattgggac-3′).
Results
Site accumulation of ASL38/LBD41 transcripts during vegetative growth
In the wild-type Columbia ecotype, similar to previous report (Shuai et al. 2002), ASL38/LBD41 gene transcripts can be detected, at variable levels, in all tissues examined, for example, cauline leaves, roots, rosette leaves, inflorescence stems and blossom flowers (Fig. 1a), suggesting ASL38/LBD41 was expressed throughout Arabidopsis tissues. Furthermore, for better detecting ASL38/LBD41 transcripts in Arabidopsis, in situ hybridization of ASL38/LBD41 was constructed. In transverse sections of 10-day-old Arabidopsis (Col-0), relatively weak signals were detected in terms of ASL38/LBD41 transcripts in this zone of the expected primordium on leaf blades (P0) and early primordia (P1 and P2) can be seen. With primordia developed, stronger signals in adaxial region were detected. In general, with leaf blades developed, the relative signals became weaker (Fig. 1b). These data suggest that ASL38/LBD41 might regulate adaxial leaf blade of Arabidopsis.
Expression of some genes in wild-type and 35S: ASL38/LBD41 plants. a In rosette leaves, cauline leaves, roots, inflorescence stems and blossom flowers, ASL38/LBD41 can all be detected, at variable levels. RL rosette leaves, CL cauline leaves, RT roots, IS inflorescence stems, BF blossom flowers. b in situ hybridization with an ASL38/LBD41 probe in wild-type. a–f are serial sections of a vegetative meristem with 10 um interval. Bars 100 um. c ASL38/LBD41 mRNA levels are completely inhibited in sense 35S:ASL38 plants using primers of coding regions, in contrast to those of wild-type (Col-0). RNA was used as a control
Identification of 35S:ASL38/LBD41 Arabidopsis seedlings
For explaining the function of ASL38/LBD41, we performed the sense- expressing status of the ASL38/LBD41 in the 35S promoter, which is a strong promoter (Fig. 2b). The 35S:ASL38/LBD41 transgenic plants were made. Using PCR assay, seventy-two transgenic plants containing the nptII reporter gene were proved (Meng et al. 2009a, b). In these identified seedlings, 30 with narrow rosette leaves (Class I) and 22 with radially- symmetric needle-like leaves (Class II) were found. While residual 20 identified seedlings are proved via PCR, they were not obviously diffinct with Col-0 (data not shown). RT-PCR findings in these 52 (52/72) transgenic plants with aberrant phenotypes were performed. In 12 with narrow rosette leaves (Class I) and 9 with radially-symmetric needle-like leaves (Class II) (Fig. 3a–d, g, h; arrow), RT-PCR findings indicated that both transgenic ASL38/LBD41 and endogenous were entirely inhibited, for example, not any expression of endogenous ASL38/LBD41 mRNA compared with Col-0 was observed (Fig. 1c). These findings indicate that in the ASL38/LBD41 plants, the cosuppression is observed. These identified plants (9 + 12/30) of RT-PCR analysis were selected for further analysis. Besides these, 18 with narrow rosette leaves and 13 with radially-symmetric needle-like leaves (Fig. 3l, m; arrow) showed overexpression of ASL38/LBD41 by RT-PCR analysis (Fig. 2a). In this same construct, independent transgenic seedlings always fluctuate >100-fold in transgene expression level and cosupression frequently evokes gene silencing (Chen et al. 2007; Holtorf et al. 1995; Jones et al. 1985). Thus, it was not surprised that both cosupression and overexpression were triggered in 35S:ASL38/LBD41 seedlings.
Expression of some genes in 35S: ASL38/LBD41 plants. a ASL38 expression levels are enhanced in overexpressing 35S:ASL38 plants, in contrast to those of wild-type. b Scheme of the sense 35S:ASL38/LBD constructs. The vector used for the introduction of the 793-bp Arabidopsis 35:ASL38/LBD41cDNA sense orientation into Arabidopsis. The sense ASL38/LBD41cDNA insert is flanked by the cauliflower mosaic virus 35S promoter at the 5′ end and by the transcriptional terminator at the 3′ end. LB left border, RB right border, nptII neomycin phosphotransferase gene whose expression confers plant resistance to kanamycin. c In overexpression of ASL38/LBD41, expression of FIL and YAB3 genes is decreased, whereas expression of PHB genes is enhanced, in contrast to that of wild-type (Col-0). d ASL37 expression leaves of cosupressing 35S:ASL38/LBD41 plants were compared with those of wild-type. e 35S:ASL38/LBD41 cosupression enhances expression of FIL and YAB3 genes, whereas it decreases expression of PHB genes, in contrast to those of wild-type (Col-0). Levels of RNA of genes above are determined by RT-PCR analysis using 1 ug of total RNA from each sample. RNA was used as a control
Phenotypes of 35S:ASL38/LBD41 plants and asl38-1 mutants. a, d A 14-day-old cosupressing 35S:ASL38/LBD41 plant with a narrow rosette leaf. Arrow indicates a narrow rosette leaf. b, c A cosupressing 35S:ASL38/LBD41 narrow rosette leaf grown for 14 and 30-day-old, respectively. Arrow indicates a narrow rosette leaf. e Vascular-pattern of 35S:ASL38/LBD41 leaf. f Arabidopsis thaliana ecotype Columbia wild-type. g Cosupressing 35S:ASL38/LBD41 plant with the radial symmetric needle-like leaf. h The radial symmetric needle-like leaf (box) of g shows a higher magnification. i Cauline leaves of cosupressing 35S:ASL38/LBD41 plants curled downward. j Vascular-pattern of wild-type leaf. k Wild-type grown in flowerpot. l Overexpressing 35S:ASL38 plant with the radial symmetric needle-like leaf. m Overexpressing 35S:ASL38 plant with a narrow rosette leaf. Arrow indicates a narrow rosette leaf. n, o asl38-1 mutants with narrow rosette leaves. Arrow indicates a narrow rosette leaf. Bars 1.0 mm
Overexpression and cosuppression of ASL38/LBD41 show Ab-adaxial polarity defects
The vasculature of rosette leaves displayed a reticulate pattern (Fig. 3j) in wild-type plants, whereas midvein often disappeared and secondary vein drastically simplified in the cosupressing 35S:ASL38/LBD41 rosette leaves (Fig. 3e). Since the establishment of a vascular system requires the existence of different ab/adaxial axes (Lin et al. 2003), these alternations of the vasculature of the cosupressing 35S:ASL38/LBD41 leaves may result from the loss of polar gene expression. For further explaining the function of ASL38/LBD41, we analyzed the dissecting characteristics of these 35S:ASL38/LBD41 rosette leaves. In rosette leaves of wild-type, an obvious ab-adaxial polarity is visible. Tightly packed, extended palisade mesophyll cells locate at the adaxial side, while irregular or round spongy mesophyll cells, loosely packed, at the abaxial side (Fig. 4a). However, in overexpressing and cosuppressing 35S:ASL38/LBD41 radially-symmetric needle-like leaves (Class II), the polarity was disoriented; as the palisade mesophyll cells were well developed on abaxial side as well as adaxial side in overespressing 35S:ASL38/LBD41 leaf blades (Fig. 4n), and the spongy mesophyll cells were well developed on adaxiality except as on abaxiality in cosuppressing 35S:ASL38/LBD41 leaves (Fig. 4d).
Cross-section of 35S:ASL38/LBD41 plants and asl38-1 mutants. a Cross-section through the rosette leaf blade of a wild-type. b Cross-section through the rosette leaf petiole of a wild- type. c is the magnified views of the regions boxed in b. d, i Cross-section through the radial symmetric needle-like rosette leaves of cosupressing 35S:ASL38/LBD41 plants. h, j are the magnified views of the regions boxed in d and i, respectively. e Cross-section through the narrow rosette leaves of cosupressing 35S:ASL38 plants. f the magnified views of the big box regions in e. g Cross-section through the narrow rosette leaf blade of asl38-1 mutants. k Cross-section through the narrow rosette leaf petiole of asl38-1 mutants. l is the magnified views of the regions boxed in k. m, o Cross-section through the narrow and radial symmetric needle-like rosette leaves of overexpressing 35S:ASL38/LBD41 plant, respectively. n, p The magnified views of the box regions in m, o, respectively; Bars 50 um in a, b, d, e, g, i, k, m and o; Bars 20 um in c, f, h, j, l, n and p. ab abaxial, ad adaxial, p palisade mesophyll, s spongy mesophyll, ph phloem, x xylem
In the elementary vascular tracts of Arabidopsis Col-0 leaf blades, ab/adaxial antagonism was obvious via the positioning of distinct tissues. Xylem located adaxially, while phloem positioned abaxially (Fig. 4b, c). In a transverse section of overexpressing and cosuppressing 35S:ASL38/LBD41 rosette leaves, regardless of narrow leaf blades (Class I) or radially-symmetric needle-like leaves (Class II), the primary tracts showed alterable xylem/phloem array, for example, xylems twined by phloems of cosuppressing 35S:ASL38/LBD41 leaves (Fig. 4d, h, e, f, i, j), and phloems situated amid xylems of overexpressing 35S:ASL38/LBD41 leaves (Fig. 4m–p).
Like the internal tissues, polar deficiency was also obvious in the epidermal cells of 35S:ASL38/LBD41 leaf blades. By scanning electron microscopy analysis, the wild-type abaxial epidermis was arranged via a wavelike surface containing no uniform measured cells, which contains long cells (Fig. 5a; arrow), whereas the control (Col-0) adaxial surface of leaf blades was arranged through a flat epidermis containing consistently measured cells (Fig. 5b). In comparison, the abaxial epidermises of overexpressing 35S:ASL38/LBD41 narrow rosette leaves (Class I) reveal a compound of ab/adaxial leaf blades (Fig. 5l). Their adaxial surface was not distinct from that of control (Col-0) leaves (data not shown). Moreover, the epidermis of the radially-symmetric needle-like leaves (class II) reveal adaxial blade traits, as the epidermis containing consistently measured cells can be observed (Fig. 5h). On the contrary, the adaxial epidermises of cosuppressing 35S:ASL38/LBD41 narrow rosette leaves (Class I) reveal an undulating surface resembling those of wild-type abaxial blade; and on this side, long cells were well observed (Fig. 5c, g; arrow). Their abaxial epidermises were not distinct from that of control (Col-0) leaves (data not shown). Moreover, the distal, middle and proximal epidermis in the radially- symmetric needle-like leaves (class II) reveal abaxial blade traits in cosuppressing 35S:ASL38/LBD41 narrow rosette leaves, as a lot of long cells can be observed (Fig. 5i–k; arrow).
Epidermal Surface of 35S:ASL38/LBD41 Plants and asl38-1 mutants. a Abaxial epidermis of a wild-type (wt) rosette leaf showing an undulating surface composed of irregular cell size. Arrow indicates elongated cell. b Adaxial epidermal cells of a wild-type (wt) rosette leaf showing uniform cell size. e, f The adaxial epidermises of the asl38-1 narrow rosette leaf blades resembling a wild-type abaxial epidermis. Arrow indicates elongated cell. c, g The adaxial epidermises of the cosupressing 35S:ASL38/LBD41 narrow rosette leaf blades resembling a wild-type abaxial epidermis. Arrow indicates elongated cell. i–k The adaxial tip, middle and proximal epidermises of the cosupressing 35S:ASL38 radially-symmetric, needle-like leaf (arrowhead) resembling a wild-type abaxial epidermis. Arrow indicates elongated cell. d Abaxial epidermis of a wild-type (wt) rosette leaf with three to four branches. h The epidermises of the overexpressing 35S:ASL38/LBD41 radially-symmetric, needle-like leaf resembling a wild-type adaxial epidermis. l The abaxial epidermises of the overexpressing 35S:ASL38/LBD41 narrow leaf resembling a wild-type adaxial epidermis. Bars 50 um in a, b, f, g, h, j, k, and l; Bars 100.0 um in c–e and i
To obtain molecular evidence for the ASL38/LBD41 function, by utilizing RNA in situ hybridization, we detected the expressing traits of the FIL polar gene. In Col-0 seedlings, transcripts of the FIL gene store up in younger leaves and their expressions decrease on the adaxial side with younger leaf growing (Sawa et al. 1999; Siegfried et al. 1999) (Fig. 6a; arrow). In cosuppressing 35S:ASL38/LBD41 plants with narrow (Class I) and needle-like rosette leaves (Class II), FIL expression was entirely detected in the ab/adaxial side of the developing leaf blades and the apex meristem (Fig. 6c, d; arrow), suggesting lateral organ of these 35S:ASL38/LBD41 plants was abaxialized patterning. However, in overexpressing 35S:ASL38/LBD41 plants with narrow (Class I) and needle-like rosette leaves (Class II), FIL was not detected in the ab-adaxial side of the developing leaf and the apex meristem (Fig. 5e, f; arrow), suggesting lateral organ of these 35S:ASL38/LBD41 plants was adaxialized patterning. In general, our data lead to the conclusion that the cosuppressing 35S:ASL38/LBD41 leaf blades closely link to the loss of adaxial polarity, but overexpressing 35S:ASL38/LBD41 leaf blades are closely relative to the loss of abaxial polarity.
RNA in situ hybridization analysis of FIL expression in 35S:ASL38/LBD41 Plants and asl38-1 mutants. In 12-day-old Col (a); asl38-1 mutants (b); cosupressing 35S:ASL38/LBD41 Class I seedlings (c); cosupressing 35S:ASL38 Class II seedlings (d); overexpressing 35S:ASL38 Class I seedlings (e); overexpressing 35S:ASL38 Class II seedlings (f). Arrows indicate expression of FIL in (a). Arrows indicate ectopic expression of FIL in b–d. Bars 100 um
Aberrant phenotype of the overexpressing 35S:ASL38/LBD41 trichomes indicate that ASL38/LBD41 is multiple functions. There was a decrease in the divergent section of trichomes on the 35S:ASL38/LBD41 radial needle-like leaves (class II), to two (Fig. 5h) rather than the three to four branches of wild- type (Fig. 5d). Also, on 35S:ASL38/LBD41 leaves, both the size and the number of the trichome support cells were increased, and protruded from the leaf blade surface (Fig. 5h); in contrast to those of wild-type Col-0 (Fig. 5d).
ASL37 expression analysis in cosuppressing 35S:ASL38/LBD41 seedlings
While our data indicated that ASL38/LBD41 was probably involved in ab-adaxil polarity, it is reasonable that our findings may be produced via an artifact through the influence of the outcome of RNAs of ASL38/LBD41, influencing levels of undisclosed genes of the AS2/LOB family, which contains 42 members and unidentified members. In LBDs gene family, ASL37/LBD40 is the most similar to ASL38/LBD41. Since ASL38/LBD41 is highly homologous to ASL37 both in the AS2/LOB domains (58 % identical amino acids) and the C-terminal halves (38 % identical amino acids), it is possible that ASL38 and ASL37 would have the overlapping functions. Thus, RT-PCR analysis was performed for examining ASL37 expression levels in cosuppressing 35S:ASL38/LBD41 plants. Our data indicated that ASL37 expression levels in cosuppressing 35S:ASL38/LBD41 plants were consistent with those in control plants (Fig. 2d), suggesting that these aberrant phenotypes of cosuppressing 35S:ASL38/LBD41 plants were triggered by endogenous RNAs of suppressing ASL38/LBD41 other than those of suppressing ASL37 or ASL38 and ASL37. Previous reports (McHale and Koning 2004; Andersson et al. 2001; Oeller et al. 1991; Hamilton et al. 1990; Ecker and Davis 1986; Rothstein et al. 1987; Smith et al. 1988; Van der Krol et al. 1988) have proved that antisense mRNA and sense has been proved to be a helpful tool for suppressing the expression of particular genes; more importantly, the mRNA level of this requested protein in this gene family can be only altered, on the other hand, other genes in this family keep up no influence (Chen et al. 2007; Andersson et al. 2001; Ganeteg et al. 2001; Zhang et al. 1997).
T-DNA insertions of ASL38/LBD41 expression
In order to further assure that these aberrant phenotypes of 35S:ASL38/LBD41 plants do result from ASL38/LBD41 overexpression and cosuppression, we gained a asl38 mutant of T-DNA insertion, which is from the Salk collection (SALK-090708, T4; asl38-1). In 22 asl38-1 mutants, 5 with narrow rosette leaves were acquired; suggesting that ASL38/LBD41 might be participated in leaf enlargement (Fig. 3n, o). Remaining 17 asl38-1 mutants (17/22) displayed to be analogical leaf enlargement pattern to control (Col-0) plants at either younger or older stages (data not shown). Phenotypes analysis indicated that the polarity was disturbed in the subepidermis of the asl38-1 narrow rosette leaves, which indicates on the both abaxial and adaxial side, there are spongy mesophyll cells (Fig. 4g). Moreover, in arrangement of the primary vascular bundles, partially abaxialized rosette leaves appeared in asl38-1 plants, namely, leafstalks showed vasculature with half-moon-traits; where xylem was observed on the interior and phloem was observed on the external (Fig. 4k, l). This is interpreted as partially abaxialized vasculature (Ha et al. 2007). By scanning electron microscopy analysis, the adaxial epidermises of asl38-1 narrow rosette leaf blades showed an undulating surface resembling those of wild-type abaxial blade; and on this side, long cells were well observed (Fig. 5e, f; arrow). Their abaxial surface was not distinct from that of control (Col-0) leaves (data not shown). Furthermore, in asl38-1 with narrow rosette leaf blades, by utilizing RNA in situ hybridization, we detected the status of the FIL expression. In control (Col-0) seedlings, the FIL transcripts were seen in younger leaves, and when younger leaf grows their expressions were decreased on the adaxial surface (Siegfried et al. 1999; Sawa et al. 1999) (Fig. 6a; arrow), whereas in asl38-1 plants with narrow rosette leaves, FIL expression was entirely detected in the ab-adaxial side of the developing leaves and the apex meristems, which was never found in that of wild-type (Fig. 6b; arrow). At adult stages, asl38-1 plants displayed to be analogical to control (Col-0) seedlings (data not shown). These partially abaxilized phenotypes seen in asl38-1 mutant might be as a result of the redundant function in LBD/ASL family.
Discussion
LOB is considered as playing an important character in building boundary of plant lateral organ development. When the meristem and lateral organs initiate, LOB can establish boundary or communicate signaling (Shuai et al. 2002). LOB is known as a nuclear protein, which participates in regulation of transcriptional level (Husbands et al. 2007). Although we have well understood their some functions, such as AS2 (Xu et al. 2003; Lin et al. 2003), in this family, many genes are not fully studied.
Aberrant phenotypes of some transgenic plants are triggered by ASL38/LBD41 cosupression
Independent transformed plants containing the identical construct always fluctuate >100-fold in transgene expression level and cosupression frequently evokes gene silencing (Chen et al. 2007; Holtorf et al. 1995; Peach and Velten 1991; Jones et al. 1985). In our work, wild-type exhibits evident ASL38/LBD41 transcripts by RT-PCR analysis, whereas 21 (9 + 12) 35S:ASL38/LBD41 transgenic plants with similar traits yield no signal in the code regions corresponding to endogenous ASL38/LBD41 mRNA, strongly suggesting that these 35S:ASL38/LBD41 plants are co-suppressed (Chen et al. 2007; Dougherty and Parks 1995; van der Krol et al. 1990; Napoli et al. 1990). Interestingly, the 21 35S:ASL38/LBD41 plants show similar traits that xylems are surrounded by phloems in xylem/phloem arrangements (traits of abaxially lateral organ), suggesting cosuppression, but not overexpression, of sense 35S:ASL38/LBD41. Furthermore, had sense 35S:ASL38/LBD41 in transgenic plants overexpressed, xylem/phloem arrangements would have been different or even reversed (Lin et al. 2003; McConnell and Barton 1998; McConnell et al. 2001). Sense or antisense suppression is a helping instrument for building Arabidopsis plants which knocks out specific protein, more significantly, specific in the sense transgenic plants in which the mRNA expression of the requested protein is influenced, whereas other genes in this family are not influenced (Andersson et al. 2001; Ganeteg et al. 2001; Zhang et al. 1997). Sense cosupression can be induced by divergent mechanism that is, at present, still poorly understood. Chen et al. (2007) thought that RNA silencing was triggered by sense cosupression evoked by siRNAs. The mechanism of ASL38/LBD41 cosupression remains to be elucidated.
ASL38/LBD41 specifies adaxial cell fate
A lot of studies imply there is a link between cosuppressing 35S:ASL38/LBD41 and fate of abaxial cells or overexpressing 35S:ASL38/LBD41 and fate of adaxial cells. Cauline leaves of cosuppressing 35S:ASL38/LBD41 plants curled downward, toward the abaxial side, suggesting that these 35S:ASL38/LBD41 plants may be involved in abaxialized defect. More significantly, blade expansion of rosette leaves of 35S:ASL38/LBD41 overexpression and cosuppression were radically limited. The similar narrow leaf blades have been reported in 35S:AS2 overexpression, curling upward, hence are believed to be adaxialized organs (Lin et al. 2003). Mutations that related to ab-adaxial axes induce decreased blade elognation (Bowman et al. 2002), which agrees with the hypothesis that juxtaposition of ab/adaxial and abaxial regions is needed to leaf growth (Waites and Hudson 1995). In extreme cases, 35S:ASL38/LBD41 rosette leaves exhibit radially-symmetric, needle-like appearance. Similar to these extreme leaves of these 35S:ASL38/LBD41 plants, in the Antirrhinum, mutants of PHAN lead to the radial and symmetric needle-like leaf blades, which are thought to be the results of abaxialization (Waites and Hudson 1995), and loss-of-function mutant of phb-phv-rev triple exhibits a single radial cotyledon and hypocotyl, which are believed to be abaxialized organs (Emery et al. 2003). ae3-1 leaves have a radially-symmetric, needle-like look, which is considered resulting from the defective adaxial identity (Huang et al. 2006). These radial cotyledon, hypocotyl, and rosette leaves all show very similar phenotypes, namely, the radially-symmetric, needle-like traits are always closely related to ab-adaxial polarity, suggesting that the needle-like leaves of 35S:ASL38/LBD41 plants may be the adaxial-abaxial defective organs. The development with needle-like leaves cannot further produce laminae (Xu et al. 2003). Together, added losses with asymmetric growth are kept company via decreased blade expansion (Eshed et al. 2004). This phenomenon accords the classic theory that the build of ab/adaxial axis is necessary to laminae development (Sussex 1954, 1955).
The examination of the internal tissues reveals that adaxial side of the rosette leaves of cosupressing 35S:ASL38/LBD41 plants are spongy mesophyll, which is different from that extended cells of the palisade mesophyll locate at the adaxial surface in wild-type; whereas abaxial side of overexpressing 35S:ASL38/LBD41 rosette leaves are palisade mesophyll. In a vascular-pattern of Arabidopsis leaf blades, ab-adaxial axis is organized via arranging distinct tissues. Xylem is positioned adaxially, whereas phloem abaxially. However, in a transverse section of cosuppressing 35S:ASL38/LBD41 rosette leaf blades, most primary bundles display distinct xylem/phloem arrangements, i.e., they typically show phloem-surrounding-xylem patterns; whereas overexpressing 35S:ASL38/LBD41 rosette leaves reveal xylem-surrounding-phloem patterns. In previous reports, a link has been found, for example, in kan1-kan2 or bop1-bop2 double mutant, phb-phv-rev triple mutant and phan mutant leaf blade growth (Ha et al. 2007; Huang et al. 2006; Eshed et al. 2004; Emery et al. 2003; Waites and Hudson 1995), vascular-pattern presents a pattern of xylem surrounded by phloem, and the vascular-shape deficiencies are always closely relative to the missing of adaxial axis. For another, in adaxial phb-1d/+ leaf blades, amphivasal positioning is observed, in which the tissue of phloem is enclosed via xylem (McConnell and Barton 1998); and in leaf blades of adaxial 35S:AS2 lines, phloem is enclosed via xylem tissue (Lin et al. 2003). The above observations demonstrate that there is a close linkage between the vascular patterning and ab/adaxial axis, and that the cosuppressing 35S:ASL38/LBD41 leaf blades closely link to the loss of adaxial polarity and overexpressing 35S:ASL38/LBD41 leaf blades closely link to the loss of abaxial polarity. By scanning electron microscopy analysis, the abaxial epidermises of overexpressing 35S:ASL38/LBD41 reveal adaxial blade traits of wild-type, as a even face made of uniformly sized cells can be observed. On the contrary, the adaxial epidermises of cosuppressing 35S:ASL38/LBD41 narrow rosette leaves show an undulating surface resembling those of wild-type abaxial blade; and on this side, long cells were well observed.
Phenotypic analyses of overexpression and cosupression of ASL38/LBD41 reveal that the ASL38/LBD41 is necessary to specify the adaxial position in leaf blades. To better understand if overexpression and cosuppression of ASL38/LBD41 is closely related to ab-adaxial later lateral organs of shoots, the molecular evidence of ASL38/LBD41 function was attained by examining the expressing modes of the FIL via utilizing RNA in situ hybridization. Our data showed that FIL was entirely detected in the abaxial and adaxial surfaces of the developing leaf blades and apex meristems; whereas FIL was not detected in the abaxial and adaxial surfaces of the developing leaves and the apex meristems in overexpressing 35S:ASL38/LBD41 plants, which was never found in that of wild-type.
Author contribution statement
Z. B Wang designed the research. J. P Song and Y. B Wang performed the research. Z. B Wang wrote the article.
References
Alvarez J, Guli CL, Yu XH, Smyth DR (1992) Terminal flower: a gene affecting inflorescence development in Arabidopsis thaliana. Plant J 2:103–116
Andersson J, Walters RG, Horton P, Jansson S (2001) Antisense inhibition of the photosynthetic antenna proteins CP29 and CP26: implications for the mechanism of protective energy dissipation. Plant Cell 13:1193–1204
Bao N, Lye KW, Barton MK (2004) MicroRNA binding sites in Arabidopsis class III HD-ZIP mRNAs are required for methylation of the template chromosome. Dev Cell 7:653–662
Baum SF, Rost TL (1996) Root apical organization in Arabidopsis thaliana: 1. Root cap and protoderm. Protoplasma 192:178–188
Bowman JL, Eshed Y, Baum SF (2002) Establishment of polarity in angiosperm lateral organs. Trends Genet 18:134–141
Candela H, Johnston R, Gerhold A, Foster T, Hake S (2008) The milkweed pod1 Gene Encodes a KANADI Protein that is required for Abaxial/Adaxial Patterning in Maize Leaves. Plant Cell 20:2073–2087
Chen RZ, Zhao X, Shao Z, Wei Z, Wang YY, Zhu LL, Zhao J, Sun MX, He RF, He GC (2007) Rice UDP-glucose pyrophosphorylase1 is essential for pollen callose deposition and its cosuppression results in a new type of thermosensitive genic male sterility. Plant Cell 19:847–861
Dougherty WG, Parks TD (1995) Transgenes and gene supression: telling us something new? Curr Opin Cell Biol 7:399–405
Ecker JR, Davis RW (1986) Inhibition of gene expression in plant cells by expression of antisense RNA. Proc Natl Acad Sci USA 83(5):372–5376
Emery JF, Floyd SK, Alvarez J, Eshed Y, Hawker NP, Izhaki A, Baum SF, Bowman JL (2003) Radial patterning of Arabidopsis shoots by Class III HD-ZIP and KANADI genes. Curr Biol 13:1768–1774
Eshed Y, Baum SF, Bowman JL (1999) Distinct mechanisms promote polarity establishment in carpels of Arabidopsis. Cell 99:199–209
Eshed Y, Baum SF, Perea JV, Bowman JL (2001) Establishment of polarity in lateral organs of plants. Curr Biol 11:1251–1260
Eshed Y, Izhaki A, Baum SF, Floyd SK, Bowman JL (2004) Asymmetric leaf development and blade expansion in Arabidopsis are mediated by KANADI and YABBY activities. Development 131:2997–3006
Fuchs C (1963) Fuchsin staining with NaOH clearing for lignified elements of whole plants or plant organs. Stain Technol 38:141–144
Ganeteg U, Strand Å, Gustafsson P, Jansson S (2001) Theroperties of the chlorophyll a/b-binding proteins Lhca2 and Lhca3 studied in vivo using antisense inhibition. Plant Physiol 127:150–158
Ha CM, Jun JH, Nam HG, Fletcher JC (2007) BLADE-ON-PETIOLE1 and 2 control Arabidopsis lateral organ fate through regulation of LOB domain and adaxial-abaxial polarity genes. Plant Cell 19:1809–1825
Hamilton AJ, Lycett GW, Grierson D (1990) Antisense gene that inhibits synthesis of the hormone ethylene in transgenic plants. Nature 346:284–287
Holtorf S, Apel K, Bohlmann H (1995) Comparison of different constitutive and inducible promoters for the overexpression of transgenes in Arabidopsis thaliana. Plant Mol Biol 29:637–646
Huang W, Pi L, Liang W, Xu B, Wang H, Cai R, Huang H (2006) The proteolytic function of the Arabidopsis 26S proteasome is required for specifying leaf adaxial identity. Plant Cell 18:2479–2492
Hudson A (2000) Development of symmetry in plants. Annu Rev Plant Physiol Plant Mol Biol 51:349–370
Husbands A, Bell EM, Shuai B, Smith HMS, Springer PS (2007) LATERAL ORGAN BOUNDARIES defines a new family of DNA-binding transcription factors and can interact with specific bHLH proteins. Nucleic Acids Res 35:6663–6671
Jones JDG, Dunsmuir P, Bedbrook J (1985) High-level expression of introduced chimaeric genes in regenerated transformed plants. EMBO J 4:2411–2418
Kerstetter RA, Bollman K, Taylor RA, Bomblies K, Poethig RS (2001) KANADI regulates organ polarity in Arabidopsis. Nature 411:706–709
Kim J, Jung JH, Reyes JL, Kim YS, Kim SY, Chung KS, Kim JA, Lee M, Lee Y, Kim VN, Chua NH, Park CM (2005) MicroRNA cleavage of ATHB15 mRNA regulates vascular development in Arabidopsis inflorescence stems. Plant J 42:84–94
Lin WC, Shuai B, Springer PS (2003) The Arabidopsis LATERAL ORGAN BOUNDARIES-domain gene ASYMMETRIC LEAVES2 functions in the repression of KNOX gene expression and in adaxial-abaxial patterning. Plant Cell 15:2241–2252
Matsumura Y, Iwakawa H, Machida Y, Machida C (2009) Characterization of genes in the ASYMMETRIC LEAVES2/LATERAL ORGAN BOUNDARIES (AS2/LOB) family in Arabidopsis thaliana, and functional and molecular comparisons between AS2 and other family members. Plant J 58:525–537
McConnell JR, Barton MK (1998) Leaf polarity and meristem formation in Arabidopsis. Development 125:2935–2942
McConnell JR, Emery J, Eshed Y, Bao N, Bowman J, Barton MK (2001) Role of PHABULOSA and PHAVOLUTA in determining radial patterning in shoots. Nature 411:709–713
McHale NA, Koning RE (2004) PHANTASTICA regulates development of the adaxial mesophyll in Nicotiana leaves. Plant Cell 16:1251–1262
Meng LS (2015) Transcription coactivator Arabidopsis ANGUSTIFOLIA3 modulates anthocyanin accumulation and light-induced root elongation through transrepression of Constitutive Photomorphogenic1. Plant Cell Environ 38(4):838–851
Meng LS, Yao SQ (2015) Transcription co-activator Arabidopsis ANGUSTIFOLIA3 (AN3) regulates water-use efficiency and drought tolerance by modulating stomatal density and improving root architecture by the transrepression of YODA (YDA). Plant Biotechnol J. doi:10.1111/pbi.12324 (Epub ahead of print)
Meng LS, Wang YB, Yao SQ, Liu A (2015) Arabidopsis AINTEGUMENTA mediates salt tolerance by trans-repressing SCABP8. J Cell Sci 128:2919–2927
Meng LS, Ding WQ, Hu X, Wang CY (2009a) Transformation of PttKN1 gene to cockscomb. Acta Physiol Plant 31:683–691
Meng LS, Song JP, Sun SB, Wang CY (2009b) The ectopic expression of PttKN1 gene causes pleiotropic alternation of morphology in transgenic carnation (Dianthus caryophyllus L.). Acta Physiol Plant. 31:1155–1164
Napoli C, Lemieux C, Jorgensen R (1990) Introduction of a chimeric chalcone synthase gene into petunia results in reversible cosuppression of homologous genes in trans. Plant Cell 2:279–289
Oeller PW, Min-Wong L, Taylor LP, Pike DA, Theologis A (1991) Reversible inhibition of tomato fruit senescence by antisense RNA. Science 254:437–439
Peach C, Velten J (1991) Transgene expression variability (position effect) of CAT and GUS reporter genes driven by linked divergent T-DNA promoters. Plant Mol Biol 17:49–60
Prigge MJ, Otsuga D, Alonso JM, Ecker JR, Drews GN, Clark SE (2005) Class III homeodomain-leucine zipper gene family members have overlapping, antagonistic, and distinct roles in Arabidopsis development. Plant Cell 17:61–76
Rothstein SJ, DiMaio J, Strand M, Rice D (1987) Stable and heritable inhibition of the expression of nopaline synthase in tobacco expressing antisense RNA. Proc Natl Acad Sci USA 84:8439–8443
Sawa S, Watanabe K, Goto K, Kanaya E, Morita EH, Okada K (1999) FILAMENTOUS FLOWER, a meristem and organ identity gene of Arabidopsis, encodes a protein with a zinc finger and HMG-related domains. Genes Dev 13:1079–1088
Shuai B, Reynaga-Peña CG, Springer PS (2002) The lateral organ boundaries gene defines a novel, plant-specific gene family. Plant Physiol 129:747–761
Siegfried KR, Eshed Y, Baum SF, Otsuga D, Drews GN, Bowman JL (1999) Members of the YABBY gene family specify abaxial cell fate in Arabidopsis. Development 126:4117–4128
Smith CJS, Watson CF, Ray J, Blrd CR, Morris PC, Schuch W, Grierson D (1988) Antisense RNA inhibition of polygalacturonase gene expression in transgenic tomatoes. Nature 334:724–726
Sussex IM (1954) Experiments on the cause of dorsoventrality in leaves. Nature 174:351–352
Sussex IM (1955) Morphogenesis in Solanum tuberosum L: experiment investigation of leaf dorsoventrality and orientation in the juvenile shoot. Phytomorphology 5:286–300
Tang G, Reinhart BJ, Bartel DP, Zamore PD (2003) A biochemical framework for RNA silencing in plants. Genes Dev 17:49–63
Van der Krol AR, Lenting PE, Veenstra J, Van der Meer IM, Koes RE, Gerats AGM, Mol JNM, Stuitje AR (1988) An anti-sense chalcone synthase gene in transgenic plants inhibits flower pigmentation. Nature 333:866–869
van der Krol AR, Mur LA, Beld M, Mol JN, Stuitje AR (1990) Flavonoid genes in petunia: addition of a limited number of gene copies may lead to a suppression of gene expression. Plant Cell 2:291–299
Waites R, Hudson A (1995) Phantastica: a gene required for dorsiventrality of leaves in Antirrhinum majus. Development 121:2143–2154
Williams L, Grigg SP, Xie M, Christensen S, Fletcher JC (2005) Regulation of Arabidopsis shoot apical meristem and lateral organ formation by microRNA miR166 and its AtHD-ZIP target genes. Development 132:3657–3668
Xu L, Xu Y, Dong A, Sun Y, Pi L, Xu Y, Huang H (2003) Novel as1 and as2 defects in leaf adaxial-abaxial polarity reveal the requirement for ASYMMETRIC LEAVES1 and 2 and ERECTA functions in specifying adaxial identity. Development 130:4097–4107
Zhang H, Goodman HM, Jansson S (1997) Antisense inhibition of the photosystem I antenna protein Lhca4 in Arabidopsis thaliana. Plant Physiol 115:1525–1531
Acknowledgments
This research was supported by the Natural Science Foundation of China (Grant No.41161058) and by the Natural Science Foundation of GanSu Province (Grant No. 1107RJZE117). This research was also supported by ‘QingLan’ Talent Engineering Funds by Tianshui Normal University”.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J.-H Liu.
Y.-B. Wang and J.-P. Song contributed equally to this work.
Rights and permissions
About this article
Cite this article
Wang, YB., Song, JP. & Wang, ZB. ASYMMETRIC LEAVES2-LIKE38, one member of AS2/LOB gene family, involves in regulating ab-adaxial patterning in Arabidopsis lateral organs. Acta Physiol Plant 37, 185 (2015). https://doi.org/10.1007/s11738-015-1938-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-015-1938-0