Abstract
Interest in the identification of thermostable lipases for industrial applications is growing. Herein, the novel thermostable lipase TDL2 was identified from the thermophilic fungus Thermomyces dupontii ATCC 16461 and its gene was cloned and overexpressed in Pichia pastoris. The recombinant TDL2 was purified and biochemically characterized. The optimal conditions for hydrolytic activity were identified as 50 °C and pH 8.5. Interestingly, the enzyme was stable at 50 °C and stable over the pH range of 6.5–9.0. Additionally, the activity analysis for various p-nitrophenyl acyl esters indicated that the lipase TDL2 showed higher specific activity for medium- and long-chain substrates, as well as the highest activity toward p-nitrophenyl laurate. Furthermore, the lipase TDL2 exhibited excellent enantioselectivity (E > 200) in the kinetic resolution of 2-carboxyethyl-3-cyano-5-methylhexanoic acid ethyl ester. Given the excellent enantioselectivity, TDL2 is an interesting enzyme for industrial applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lipases (EC 3.1.1.3) are versatile enzymes that catalyze a great variety of reactions, including ester hydrolysis and synthesis, transesterification, acidolysis, alcoholysis, and aminolysis (Jaeger and Reetz 1998; Gupta et al. 2004; Kapoor and Gupta 2012). Given their unique catalytic properties, such as stability in organic solvents; broad substrate spectrum; high chemo-, region-, and enantioselectivity, lipases are arguably the most popular biocatalysts for various biochemical processes in the detergent, food, medical, paper, and pharmaceutical industries (Gupta et al. 2004; Sarmah et al. 2018; Zheng et al. 2019). Demand for the discovery and production of high-performing lipases is increasing because of their continuously expanding utility as biocatalysts in various industrially relevant processes.
Lipases are widespread in higher eukaryotes and microbial organisms. The most widely utilized microbial lipases have been mainly derived from bacteria, yeasts, and filamentous fungi. However, in many industrial processes, enzymes are subjected to extreme physicochemical conditions that are very drastically different from their cellular environment (Littlechild 2015). Thermostable enzymes are more resistant to organic solvents besides high thermal stability compared with general enzymes and which is more conducive to industrial storage (Li et al. 2005). The ubiquitous thermophilic fungus Thermomyces lanuginosus, also known under the obsolete synonym Humicola lanuginosa, produces several thermostable enzymes for industrial applications (de Oliveira et al. 2015; Ronald et al. 2014). Notably, TLL, the lipase from T. lanuginosus, has been used for the efficient kinetic resolution of 2-carboxyethyl-3-cyano-5-methylhexanoic acid ethyl ester (CNDE) to produce the desired (S)-mono acid enantiomer; this process is the most challenging step in the chemoenzymatic synthesis of pregabalin (Martinez et al. 2008; Zheng et al. 2016).
In addition to TLL, the two other thermostable lipases, namely Lip (Li et al. 2014a; Li et al. 2014b) and TTL (Ding et al. 2018a; Ding et al. 2018b), had also been used for the chemoenzymatic route to pregabalin. Interestingly, TLL, Lip, and TTL show high amino acid sequence similarity. TLL and Lip are both derived from the thermophilic fungus T. lanuginosus, while TTL comes from the thermophilic fungus Thermomyces dupontii, which is also known under the obsolete synonym Talaromyces thermophilus (de Oliveira et al. 2015; Ronald et al. 2014). The optimal growth temperature and maximum temperature for this thermophilic fungus are 45–50 °C and 60 °C, respectively (de Oliveira et al. 2015). In recent years, a number of enzymes, many of which have been proven to be highly thermostable (Wang et al. 2019), have been isolated and characterized from T. dupontii. These enzymes include the α-amylase TdAmyA (Wang et al. 2019), D-aspartate oxidase (Takahashi et al. 2019), β-galactosidase (Nakkharat and Haltrich 2006), β-glucosidase (Mallek-Fakhfakh and Belghith 2016; Mallek-Fakhfakh et al. 2017), lipase (Romdhane et al. 2012), xylanase (Mallek-Fakhfakh et al. 2017), β-xylosidase (Guerfali et al. 2009), and an α-arabinofuranosidase (Guerfali et al. 2011). Hence, T. dupontii is also an excellent producer of hydrolases. However, these reported enzymes are often obtained via the fermentation of wild-type T. dupontii rather than heterologous expression, owing to a lack of knowledge of the corresponding genes. The application of these thermostable enzymes is limited by the cost of enzyme production and the amounts that can be obtained from wild-type hosts. Only a few enzyme-coding genes including the lipase TTL (Romdhane et al. 2012), the D-aspartate oxidase TdDDO (Takahashi et al. 2019), and the α-amylase TdAmyA (Wang et al. 2019) have been cloned from T. dupontii. Thus, this useful source organism of stable enzymes has not yet been characterized to its fullest, and cloning novel thermostable enzymes from T. dupontii has received considerable interest.
In our previous study, the engineered lipase Lip from T. lanuginosus DSM 10635 was demonstrated to be a useful biocatalyst for the kinetic resolution of CNDE to produce (3S)-2-carboxyethyl-3-cyano-5-methylhexanoic acid (Li et al. 2014a; Li et al. 2014b). Furthermore, the lipases TLL (Martinez et al. 2008; Zheng et al. 2016) and TTL (Ding et al. 2018a; Ding et al. 2018b) attracted our attention due to their excellent performance in the kinetic resolution of CNDE. Interestingly, amino acid sequence alignment revealed a high sequence identity of 78–90% among the three lipases. Although TTL was derived from the thermophilic fungus T. dupontii, the homology between TLL and TTL (90%) was higher than that between TLL and Lip (78%). Moreover, TLL and TTL shared the same signal peptide. Furthermore, the phylogenetic tree analysis of the three lipases indicated that TTL clustered together with TLL, whereas Lip clustered in an independent clade. Considering the microbial origin and sequence similarity of these three lipases, we hypothesized that, (1) T. dupontii might possess another undiscovered lipase with high identity with Lip, and (2) this undiscovered lipase might show high enantioselectivity toward CNDE.
In the present study, the gene encoding the novel thermostable lipase TDL2 was successfully cloned from the thermophilic fungus T. dupontii ATCC 16461. After successful heterologous expression in Pichia pastoris, the biochemical properties of purified TDL2 were studied. Finally, the biocatalytic application potential of TDL2 was investigated by using the kinetic resolution of CNDE as a model reaction.
Materials and methods
Materials
PrimeSTAR HS DNA polymerase, LA Taq DNA polymerase, and 3´-rapid amplification of cDNA ends (RACE) kits were purchased from TaKaRa (Dalian, China). ZeocinTM was acquired from Thermo Scientific (Shanghai, China) and p-nitrophenyl acyl esters with different chain lengths were procured from J&K Scientific Ltd. (Beijing, China). Primers were synthesized by Sangon Biotech Co., Ltd. (Shanghai, China). Unless otherwise specified, all other chemicals used in this work were of analytical grade and were sourced from local suppliers.
Strains, plasmids, and culture conditions
Thermomyces dupontii ATCC 16461 (syn. Talaromyces thermophilus) was grown in YPD medium at 40 °C. Escherichia coli JM109, which was used as the host strain for the cloning of lipase-encoding genes, was routinely cultured at 37 °C in LB medium. Pichia pastoris X33, which was used for lipase expression, was grown in YPD medium at 30 °C. The vectors pGEM-T (Promega, Beijing, China) and pPICZαA (Thermo Scientific, Shanghai, China) were used for cloning and expression, respectively.
Methods
DNA manipulation and cloning
The total RNA extracted from T. dupontii ATCC 16461 was used as a template for cDNA synthesis. The DNA sequence encoding T. dupontii lipase (TDL2) was obtained via PCR amplification with cDNA as the template, purified from the gel, ligated into pGEM-T, and used to transform E. coli JM109. The correct clones were confirmed by sequencing. In accordance with the bioinformatic analyses of the nucleotide and amino acid sequences of the lipases TLL, Lip and TTL, the primers P1 and P2 (Table 1) were designed on the basis of the signal peptide of lipase Lip and the sequences that were highly conserved among the three lipases. These primers were used to amplify the 5´-flanking region of TDL2 by using LA Taq DNA polymerase. The 3´-flanking region of the cDNAs was amplified by using a 3´-RACE kit in accordance with the manufacturer’s recommendations. The specific primers for RACE-PCR (P3 and P4) were designed on the basis of the previously obtained fragment sequences. The products of 3´-RACE were further cloned into the pGEM-T vector and sequenced to construct the specific primers P5 and P6 (Table 1), which were used to amplify the full-length sequence of TDL2 by using PrimeSTAR HS DNA polymerase.
Amino acid sequence alignment, phylogenetic analysis, and homology-modeling of TDL2
The web tool ORF Finder on NCBI was used to identify the open reading frame. The DNA sequences were analyzed and translated by utilizing Vector NTI Suite 6 software package (Invitrogen, USA). Sequences with high similarity to TDL2 were searched by using BLAST. The sequence alignment was visualized with ESPript3.0 (Robert and Gouet 2014). Clustal X1.83 (Larkin et al. 2007) and MEGA 5.0 (Kumar et al. 2018) were applied to align TDL2 sequences with related sequences and to construct the phylogenetic tree, respectively. The evolutionary tree was constructed on the basis of the manually adjusted alignment by using the neighbor-joining method with 1000 bootstrap iterations. The putative signal peptide was identified with the SignalP-5.0 server (http://www.cbs.dtu.dk/services/SignalP/) (Armenteros et al. 2019). A three-dimensional model for TDL2 was constructed using the SWISS-MODEL protein structure homology-modeling server (Waterhouse et al. 2018; Bienert et al. 2017), with the reported 3D structure of Thermomyces lanuginosus lipase (1EIN) (Brzozowski 2000), which shows 81% sequence identity with TDL2, as the template.
Overexpression of TDL2 in P. pastoris
TDL2 was expressed heterologously in P. pastoris strain X33 by using the vector pPICZαA. The lipase was expressed under the transcriptional control of the strong inducible AOX1 promoter, which allows the methanol-inducible, high-level expression of the genes of interest in P. pastoris. The gene fragment coding for the mature protein without the signal peptide was amplified by PCR with the primers P7 and P8 (Table 1). The resulting PCR product cloned between the EcoRI and XbaI sites of pPICZαA was used to construct the P. pastoris secretory expression plasmid pPICZαA-TDL2. The resulting plasmid was confirmed by sequencing, linearized by using SacI, and used to transform P. pastoris X33 by electroporation with the Bio-Rad gene Pulser in accordance with the operating manual. The linearized empty pPICZαA without an insert was used as the negative control. Transformants were selected on YPDS medium with 200 mg/L ZeocinTM and confirmed by PCR.
A colony of P. pastoris expressing TDL2 was used to inoculate buffered glycerol complex medium (BMGY: 1.0% yeast extract, 2.0% peptone, 1.34% yeast nitrogen base with ammonium sulfate without amino acids, 4 × 10−5% biotin, 100 mM potassium phosphate, pH 6.0; with 1.0% glycerol) and grown at 30 °C and 220 rpm. After 24 h, the cells were harvested via centrifugation and resuspended in buffered methanol complex medium (BMMY: same as BMGY but with 1% methanol instead of glycerol). The culture was fed with 0.5% (final vol/vol) methanol every 12 h to ensure continuous induction.
Purification of TDL2
The culture supernatant of the recombinant P. pastoris containing TDL2 was collected by centrifugation at 12,000 g for 15 min and concentrated using by an Amplicon Ultra-15 ultrafiltration membrane with a 10 kDa molecular weight cutoff (Millipore, USA). After concentrating the protein to approximately 30 mg/mL, the protein was loaded onto a 10-mL Ni-NTA column that was pre-equilibrated with binding buffer, and washed with washing buffer (50 mM Tris-HCl, 50 mM imidazole, 500 mM NaCl, pH 8.0). The hexahistidine-tagged TDL2 was released by using buffer (50 mM Tris-HCl, 300 mM imidazole, 500 mM NaCl, pH 8.0). The collected fractions containing the enzyme were concentrated with the same 10 kDa ultrafiltration membrane, lyophilized, and stored at − 20 °C until further experiments. The purity of the enzyme was confirmed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) on a 12% acrylamide gel. The concentration of lipase was measured through the Coomassie brilliant blue method (Bradford 1976) with bovine serum albumin as the standard.
Enzyme activity assays
Lipase activity was measured by using p-nitrophenyl acyl esters as substrates. Enzyme reaction solution was composed of 0.1 ml lipase solution, 2.8 ml Tris-HCl buffer (50 mM, pH 8.0) and 0.1 ml p-nitrophenyl esters (30 mM, dissolved in acetonitrile) in the test tube, which reacted for 5 min in a water bath at 40 °C and halted the reaction by adding ethanol (using inactivated enzyme solution as the blank control), then the absorbance at 410 nm (OD410) was measured. One unit of lipase activity was defined as the amount of enzyme that released 1 μmol of p-nitrophenol per minute. Under the assay conditions (50 mM Tris-HCl buffer, pH 8.0 at 40 °C), the molar absorptivity is 1.46 × 105 L mol-1 cm-1. Unless stated otherwise, all enzymatic activity assays were performed in triplicate. The data were expressed as the mean value ± standard deviation of the sample.
The activity analysis for various p-nitrophenyl acyl esters
The activity of TDL2 for p-nitrophenyl acyl esters with fatty chains with lengths ranging from C2 to C16 was determined as described previously (Li et al. 2014c).
Effects of temperature and pH
The optimal pH of TDL2 was determined by measuring the enzyme’s activity in 50 mM buffers with pH values ranging from 6.0 to 10.0 (pH 6.0–7.0 was citric acid/sodium phosphate buffer, pH 7.5–8.5 was Tris-HCl buffer, and pH 9.0–10.0 was Gly-NaOH buffer) at 40 °C. The optimal temperature was determined by measuring TDL2 activity at the optimal pH at temperatures between 25 and 70 °C. The enzyme solution was incubated with a microplate reader (Molecular Devices, Spectramax M5) at a specific temperature for 5 min, the absorbance at 410 nm was measured every 10 S. The lipase activity was obtained by determining the initial velocities of reaction.
The pH and temperature stability of TDL2 were analyzed with p-nitrophenyl laurate as the substrate. TDL2 was pre-incubated in Tris-HCl buffer (pH 8.0) at 50, 55, and 60 °C to evaluate thermal stability, and residual activity was measured under the optimal conditions (50 °C and pH 8.5) at fixed time intervals. The enzyme’s pH stability was estimated by incubating the enzyme in the above-mentioned buffers at pH 6.0–10.0 for 12 h at 40 °C. The residual lipase activity was then measured under the optimal conditions (50 °C and pH 8.5) described above, and all enzymatic activity assays were performed in triplicate.
Effects of metal ions
The effects of metal ions (Na+, K+, Mg2+, Ba2+, Co2+, Ni2+, Fe2+, Cu2+, Zn2+, Ca2+, Mn2+, and Al3+) on enzyme activity were measured as described previously (Li et al. 2014c), and the concentrations of metal ions used were set in accordance with the standard described by Kamarudin (Kamarudin et al. 2014). The residual lipase activity was estimated according to the standard activity assay described above. The activity of TDL2 incubated in the absence of metal ions was defined as 100%, and all enzymatic activity assays were performed in triplicate.
Enantioselective hydrolysis of CNDE
TDL2 was used in the kinetic resolution of CNDE to the corresponding acid. The indicated amount of TDL2 was mixed with Tris-HCl buffer (100 mM, pH 8.0) and 5 mM calcium acetate. Enantioselective hydrolysis was initiated by adding 100 mM CNDE. The reactions were carried out in a shaking incubator at 40 °C and 150 rpm, and samples were periodically withdrawn from the reaction mixture to determine the conversion ratio and enantiomeric excess (e.e.) by chiral-GC analysis as described previously (Li et al. 2014a; Li et al. 2014b; Li et al. 2014c). The conversion (C) and enantiomeric ratio (E) were calculated based on eeS and eeP according to the method as described previously (Zhong et al., 2020).
Results and discussion
Cloning of TDL2
The sequences of the three homologous lipases TLL, Lip, and TTL were analyzed, and the degenerate primers P1 and P2 were designed on the basis of the signal peptide of Lip and the sequences that were highly conserved among the three homologous lipases, respectively. PCR with the primers P1 and P2 resulted in a 596 bp amplicon showing high sequence identity with the 5´-flanking regions of TLL, Lip, and TTL. The resulting 5´-flanking fragment sequence was used for 3´-RACE-PCR to unveil the sequence of the complete ORF. The 3´-RACE with the primers P3 and P4 resulted in a 650 bp amplicon of the 3´-flanking region of the cDNA (Fig. S1). On the basis of these sequencing results, the complete ORF was revealed and obtained through PCR with the primers P5 and P6. Nucleotide sequence analysis revealed that the newly cloned 876-bp ORF coded for a putative lipase that we named TDL2 and deposited in the GenBank database under accession number MN782511.
Analysis of TDL2 sequence
The analysis of the deduced amino acid sequence of TDL2 (291 amino acids) revealed that it shares the highest identity (86.9%) with the lipase Lip from T. lanuginosus (AGH70111.1), as well as 83.5 and 81.7% identity with the lipases from T. thermophilus (TTL, AEE61324.1) and T. lanuginosus (TLL, O59952.1), respectively. An alignment of the amino acid sequence of TDL2 with the amino acid sequences of TLL, Lip, and TTL is shown in Fig. 1. TDL2 was also found to share 46.9–59.1% identities with several lipases and putative proteins from Aspergillus and Penicillium.
Multiple sequence alignment of the amino acid sequence of lipases TDL2, TLL, Lip, and TTL. The amino acid residues of the putative catalytic triad Ser-His-Asp and the oxyanion hole are indicated by triangles and circles, respectively. The signal peptide, propeptide, and lid are indicated by a blue arrow and lines, respectively
The putative signal peptide with a cleavage site located between residues 17 and 18 (ALA-RP) was predicted by using the SignalP-5.0 server. Residues 18 to 22 (RPVRR) were predicted to act as a propeptide (Fig. 1). The mature form of the protein (residues 23 to 291, 269 amino acid residues) had a predicted molecular mass of 29.0 kDa and a theoretical pI value of 4.49. The sequence analysis of mature TDL2 revealed the presence of the conserved motif GHSLG, which was a part of a characteristic nucleophilic elbow that included a catalytic serine residue (Ser146, counted from 1 after removing the signal peptide and propeptide). A histidine and an aspartic acid, at positions 258 and 201, respectively, were predicted as the two additional amino acids of the catalytic triad of TDL2. Moreover, Ser83 and Leu147 were predicted to be involved in the putative oxyanion hole. Similar to the other lipases of the Rhizomucor miehei lipase-like family (abH23.01), TDL2 also had a lid. The lid of TDL2 was predicted to comprise an α-helix fragment (86IENWIGN92) plus eight amino acids (82GSRS85 and 93LIVG96) that acted as two hinges around the lid. In this lipase family, lid movement can be described as a hinge bending motion of a helical lid (Svendsen 2000).
In order to obtain structural insights into TDL2, a 3D structural model with open-lid conformation was generated using the SWISS-MODEL protein structure homology-modeling server (Waterhouse et al. 2018; Bienert et al. 2017), with the reported X-ray structure of Thermomyces lanuginosus lipase (PDB 1EIN (Brzozowski et al. 2000) as the template. The 3D model with an estimated precision of 100% covers the residues 1 to 269. The GMQE value of the TDL2 model was 0.97, which indicates that the reliability of the model is very high. The Ramachandran plot of the final TDL2, generated using the PROCHECK program, showed that 98.7% of residues were either in the most favored or in the additional allowed regions (Fig. S2). The overall 3D homology model and detail showing the catalytic pocket of TDL2 are shown in Fig. 2. Similar to basic features of most microbial lipases, the 3D structure model of TDL2 has a globular shape with a typical α/β hydrolase fold (Fig. 2A and B). TDL2 consists of 8 α-helices and 10 β-strands (Fig. 2A). The residues of the catalytic triad (Ser146-His258-Asp201) are present at the bottom of a catalytic groove and are not accessible at the surface of the molecule (Fig. 2B and C). The amino acid residues that constitute the oxyanion hole (Ser83 and Leu147) are located near the catalytic Ser146 residue. In the orientation shown in Figure 2, the lid of TDL2 is directly to the left of the active site. TDL2 has three putative disulfide bonds in the structure, Cys22-Cys268, Cys36-Cys41 and Cys104-Cys107, which might play a role in stabilizing the overall conformation of the enzyme.
Homology model of the overall structure of lipase TDL2. A Cartoon representation of TDL2 showing 8 α-helices and 10 β-strands. The amino acid residues of the catalytic triad (Ser146-His258-Asp201) and the oxyanion hole (Ser83 and Leu147) are shown as sticks, and the lid of TDL2 is shown in magenta. B Surface representation of TDL2. The catalytic residue Ser146 located at the bottom of a catalytic groove is shown in yellow. C Detail showing the catalytic pocket of TDL2. The amino acid residues of the catalytic triad and the oxyanion hole (Ser83 and Leu147) were labeled and represented as sticks, respectively. The lid is shown in magenta
Phylogenetic position of TDL2
On the basis of sequence homology analysis in the Lipase Engineering Database (Fischer and Pleiss 2003), TDL2 was assigned to the GX class in accordance with the oxyanion hole structure and sequence. TDL2 could be further classified into the Rhizomucor miehei lipase-like family (abH23.01), which in turn belongs to the superfamily of filamentous fungi lipases (abH23). For phylogenetic analysis, 21 lipases and putative proteins displaying high sequence identity (46.9–86.9%) were selected. A neighbor-joining tree was constructed to analyze the relationship between TDL2 and other lipases. TDL2 clustered together with the lipases from T. lanuginosus and T. thermophilus, whereas those from Aspergillus and Penicillium constituted different branches on the phylogenetic tree (Fig.3). TDL2 was grouped into a clade that was different from that of TTL, and clustered together with Lip, indicating that TDL2 had a closer evolutionary relationship with Lip than with TTL. This result was notable because TDL2 and TTL are derived from T. dupontii, whereas TLL and Lip are derived from T. lanuginosus.
Molecular phylogenetic analysis of lipase TDL2. Phylogenetic analyses were conducted by MEGA software using the neighbor-joining method. The percentage of trees in which the associated sequences clustered together in the bootstrap test (1000 iterations) is shown next to the branches. The tree is drawn to scale, and the scale bar indicates the number of substitutions per site. The GenBank accession numbers of the sequences are shown in brackets
Expression and purification of TDL2
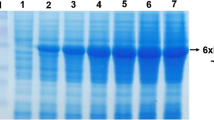
To analyze the enzymatic characteristics of TDL2, the lipase was heterologously expressed in P. pastoris X33. The supernatant of the induced recombinant P. pastoris cells was analyzed by using SDS-PAGE to confirm the successful secretion of TDL2 (Fig. 4). The SDS-PAGE gel showed a band that corresponded to a molecular mass of approximately 35 kDa, which was similar to the theoretical value of TDL2. The secreted TDL2 from the supernatant of the recombinant P. pastoris cells was purified by affinity chromatography on a Ni-NTA column, and the protein sample was concentrated by using an ultrafiltration membrane with a 10 kDa molecular weight cutoff. The purified lipase presented as a single band in SDS-PAGE (Fig. 4). This result indicated that the enzyme was recovered as an electrophoretically homogeneous preparation. The described purification procedure resulted in a 17-fold concentration of enzymatic activity, with an overall yield of 34%.
Activity of TDL2 for various p-nitrophenyl esters
Lipolytic enzymes, which catalyze the hydrolysis of triacylglycerols into glycerol and fatty acids, include lipases (EC 3.1.1.3) and esterases (EC 3.1.1.1), which differ in terms of their specific activities for the acyl moiety with long- or short-chain fatty acids. Lipases display high specific activity for long-chain acylglycerols and other long-chain acyl esters, whereas esterases usually only hydrolyze short-chain acylglycerols (Bornscheuer 2002). The activity of purified TDL2 against different p-nitrophenyl ester substrates with chain-lengths ranging from C2 to C16 is shown in Fig. 5. The specific activities of enzyme were higher for the esters of moderate- to long-chain fatty acids (C > 10). Under 40 °C and pH 8.5, TDL2 displayed the highest activity (285 U/mg) toward p-nitrophenyl laurate (C12). Conversely, the esters of short-chain fatty acids were poor substrates for TDL2. These results indicated that TDL2 was indeed a lipase rather than an esterase.
Activity of TDL2 against p-nitrophenyl esters with different chain lengths (C2–C16). The enzymatic activity assays were performed in triplicate, and the error bars represent standard deviations of the sample. The activity (285 U/mg) toward p-nitrophenyl laurate (C12) was defined as 100% and used to normalize the results for the other substrates
Effects of metal ions on lipase activity
Various metal ions were added at two different concentrations (1 and 5 mM) to test the influence of metals, and the activity of TDL2 was measured with p-nitrophenyl laurate as the substrate (Table 2). At the low concentration of 1 mM, the 12 tested metal ions did not show any significant effects on TDL2 activity. At the high concentration of 5 mM, the activity of TDL2 was slightly elevated in the presence of Co2+ and Al3+. However, Ni2+, K+, and Ca2+ had a strong inhibitory effect on the activity of TDL2 at the high concentration of 5 mM, with reductions of in relative activity of 34.4%, 29.1%, and 26.3%, respectively. Other metal ions, such as Cu2+, Mg2+, Fe2+, and Zn2+, had no significant effect on the lipase even at the high concentrations.
Effects of pH and temperature
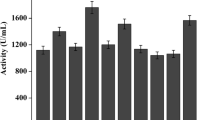
The effects of pH and temperature on the activity and stability of the lipase TDL2 were analyzed by using p-nitrophenyl laurate as the substrate (Fig. 6). The pH and temperature optima for the lipase activity of TDL2 were 8.5 and 50 °C (Fig. 6A and C), respectively. Under the optimal conditions (pH 8.5 and 50 °C), TDL2 displayed maximum enzyme activity (840 U/mg) toward p-nitrophenyl laurate. TDL2 displayed high activity at pH values between 8.0 and 9.0. However, its relative activity decreased significantly above pH 9.0. In addition, although TDL2 was stable over the pH range of 6.5–9.0 (Fig. 6B), its relative activity significantly decreased above pH 9.5. Notably, this lipase had a temperature optimum of 50 °C and retained 78% of its maximum activity at 55 °C (Fig. 6C). However, relative activity decreased significantly at temperatures over 60 °C. TDL2 was stable below 55 °C (Fig. 6D). It retained 91% and 68% of its initial activity after 3 h of incubation at 50 °C and 55 °C, respectively, and even maintained 50% of its initial activity after incubation at 60 °C for 2 h. These results indicated that TDL2 was a moderately thermophilic lipase.
Effects of pH and temperature on the activity (A and B) and stability (C and D) of TDL2. The temperature in Figures 6A and 6C is 40 °C, the pH in Figures 6B and 6D is pH 8.5, respectively. The enzymatic activity assays were performed in triplicate, and the error bars represent standard deviations of the sample
Kinetic resolution of CNDE
The kinetic resolution of CNDE was performed as a model reaction to evaluate the enantioselectivity and biocatalytic potential of lipase TDL2. The reaction profile is shown in Fig. 7. A conversion of 49.1% with an enantiomeric excess of the product (e.e.P) of 98.1% was achieved after 12 h. Most importantly, TDL2 showed a high preference for (S)-CNDE. The e.e. of the hydrolysis product (3S)-2-carboxyethyl-3-cyano-5-methylhexanoic acid remained high (e.e.P >98%) throughout the reaction. The enantiomeric ratio (E) of the reaction was 384. This result (E >200) indicated that TDL2 had high enantioselectivity in the conversion of CNDE. Therefore, the lipase TDL2 is a potential biocatalyst for the kinetic resolution of CNDE.
In addition to TDL2, several biocatalysts, including lipases (Martinez et al. 2008; Zheng et al. 2016; Li et al. 2014a; Li et al. 2014b; Ding et al. 2018a; Ding et al. 2018b), esterases (Martinez et al. 2008; Li et al. 2014c; Xu et al. 2015), and microbial cells (Zheng et al. 2013), have been used for the kinetic resolution of CNDE with high enantioselectivity (E>50) in Table 3. Most of these lipases and esterases displayed S-selectivity with CNDE as the substrate. Compared with lipases, esterases generally show lower enantioselectivity in the hydrolysis of CNDE. Notably, similar to TLL, Lip, TTL, and Rhizopus delemar lipase, TDL2 could hydrolyze CNDE with the highest enantioselectivity (E>200). If enantioselectivity is the only concern, TDL2 is certainly a new potential biocatalyst for the enantioselective hydrolysis of CNDE. However, the activity and substrate load of wild-type TDL2 (12 h, 100 mM) was clearly inferior to that of TLL (24 h, 3 M CNDE), engineered Lip (24 h, 3 M CNDE), and especially engineered TTL (1.5 h, 1 M CNDE). Further studies on the engineering of TDL2 to match the industrial applications are in progress. The findings of these studies will be reported in a future publication.
Conclusions
In the present study, the novel lipase TDL2 was successfully cloned from Thermomyces dupontii ATCC 16461. The deduced mature TDL2 was heterologously expressed, purified, and characterized. As a moderately thermophilic lipase, the lipase showed an optimum pH and temperature of 8.5 and 50 °C, respectively. Notably, TDL2 was able to hydrolyze CNDE with the highest enantioselectivity (E > 200). The characteristics, especially the excellent enantioselectivity, make TDL2 an interesting enzyme for industrial applications.
References
Armenteros JJA, Tsirigos KD, Sønderby CK, Petersen TN, Winther O, Brunak S, von Heijne G, Nielsen H (2019) Signal P 5.0 improves signal peptide predictions using deep neural networks. Nat Biotechnol 37:420–423. https://doi.org/10.1038/s41587-019-0036-z
Bienert S, Waterhouse A, Beer TAPd, Tauriello G, Studer G, Bordoli L, Schwede T (2017) The SWISS-MODEL repository-new features and functionality. Nucleic Acids Res 45(D1):313–319. https://doi.org/10.1093/nar/gkw1132
Bornscheuer UT (2002) Microbial carboxyl esterases: classification, properties and application in biocatalysis. FEMS Microbiol Rev 26:73–81. https://doi.org/10.1111/j.1574-6976.2002.tb00599.x
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1006/abio.1976.9999
Brzozowski AM, Savage H, Verma CS, Turkenburg JP, Lawson DM, Svendsen A, Patkar S (2000) Structural origins of the interfacial activation in Thermomyces (Humicola) lanuginosa lipase. Biochemistry 39(49):15071–15082. https://doi.org/10.1021/bi0013905
de Oliveira TD, Gomes E, Rodrigues A (2015) Thermophilic fungi in the new age of fungal taxonomy. Extremophiles 19:31–37. https://doi.org/10.1007/s00792-014-0707-0
Ding X, Tang XL, Zheng RC, Zheng YG (2018) Efficient biocatalytic synthesis of chiral intermediate of pregabalin using immobilized Talaromyces thermophilus lipase. BioMed Res Int 12:1–7. https://doi.org/10.1155/2018/6192059
Ding X, Zheng RC, Tang XL, Zheng YG (2018) Engineering of Talaromyces thermophilus lipase by altering its crevice-like binding site for highly efficient biocatalytic synthesis of chiral intermediate of Pregablin. Bioorg Chem 77:330–338. https://doi.org/10.1016/j.bioorg.2018.01.018
Fischer M, Pleiss J (2003) The lipase engineering database: a navigation and analysis tool for protein families. Nucleic Acids Res 31:319–321. https://doi.org/10.1093/nar/gkg015
Guerfali M, Maalej I, Gargouri A, Belghith H (2009) Catalytic properties of the immobilized Talaromyces thermophilus β-xylosidase and its use for xylose and xylooligosaccharides production. J Mol Catal B-Enzym 57:242–249. https://doi.org/10.1016/j.molcatb.2008.09.011
Guerfali M, Gargouri A, Belghith H (2011) Catalytic properties of Talaromyces thermophilus α-l-arabinofuranosidase and its synergistic action with immobilized endo-β-1,4-xylanase. J Mol Catal B-Enzym 68:192–199. https://doi.org/10.1016/j.molcatb.2010.11.003
Gupta R, Gupta N, Rathi P (2004) Bacterial lipases: an overview of production, purification and biochemical properties. Appl Microbiol Biotechnol 64:763–781. https://doi.org/10.1007/s00253-004-1568-8
Jaeger KE, Reetz MT (1998) Microbial lipases form versatile tools for biotechnology. Trends Biotechnol 16:396–403. https://doi.org/10.1016/s0167-7799(98)01195-0
Kamarudin NH, Rahman RN, Ali MS, Leow TC, Basri M, Salleh AB (2014) A new cold-adapted, organic solvent stable lipase from mesophilic Staphylococcus epidermidis AT2. Protein J 33:296–307. https://doi.org/10.1007/s10930-014-9560-3
Kapoor M, Gupta MN (2012) Lipase promiscuity and its biochemical applications. Process Biochem 47:555–569. https://doi.org/10.1016/j.procbio.2012.01.011
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Li H, Zhang X (2005) Characterization of thermostable lipase from thermophilic Geobacillus sp. TW1. Protein Expression & Purif 42:153–159. https://doi.org/10.1016/j.pep.2005.03.011
Li XJ, Zheng RC, Ma HY, Huang JF, Zheng YG (2014) Key residues responsible for enhancement of catalytic efficiency of Thermomyces lanuginosus lipase Lip revealed by complementary protein engineering strategy. J Biotechnol 188:29–35. https://doi.org/10.1016/j.jbiotec.2014.08.004
Li XJ, Zheng RC, Ma HY, Zheng YG (2014) Engineering of Thermomyces lanuginosus lipase Lip: creation of novel biocatalyst for efficient biosynthesis of chiral intermediate of Pregabalin. Appl Microbiol Biotechnol 98:2473–2483. https://doi.org/10.1007/s00253-013-5136-y
Li XJ, Zheng RC, Wu ZM, Ding X, Zheng YG (2014) Thermophilic esterase from Thermomyces lanuginosus: molecular cloning, functional expression and biochemical characterization. Protein Expre Purif 101:1–7. https://doi.org/10.1016/j.pep.2014.05.006
Littlechild JA (2015) Enzymes from extreme environments and their industrial applications. Front Bioeng Biotech 3:1–9. https://doi.org/10.3389/fbioe.2015.00161
Mallek-Fakhfakh H, Belghith H (2016) Physicochemical properties of thermotolerant extracellular β-glucosidase from Talaromyces thermophilus and enzymatic synthesis of cello-oligosaccharides. Carbohyd Res 419:41–50. https://doi.org/10.1016/j.carres.2015.10.014
Mallek-Fakhfakh H, Fakhfakh J, Walha K, Hassairi H, Gargouri A, Belghith H (2017) Enzymatic hydrolysis of pretreated Alfa fibers (Stipa tenacissima) using β-d-glucosidase and xylanase of Talaromyces thermophilus from solid-state fermentation. Int J Biol Macromol 103:543–553. https://doi.org/10.1016/j.ijbiomac.2017.05.078
Martinez CA, Hu S, Dumond Y, Tao J, Kelleher P, Tully L (2008) Development of a chemoenzymatic manufacturing process for Pregabalin. Org Process Res Dev 12:392–398. https://doi.org/10.1021/op7002248
Nakkharat P, Haltrich D (2006) Lactose hydrolysis and formation of galactooligosaccharides by a novel immobilized β-galactosidase from the thermophilic fungus Talaromyces thermophilus. Appl Biochem Biotech 129:215–225. https://doi.org/10.1385/abab:129:1:215
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42:320–324. https://doi.org/10.1093/nar/gku316
Romdhane IB, Frikha F, Maalej-Achouri I, Gargouri A, Belghith H (2012) Gene cloning and molecular characterization of the Talaromyces thermophilus lipase catalyzed efficient hydrolysis and synthesis of esters. Gene 494:110–118. https://doi.org/10.1016/j.gene.2011.11.059
Ronald JH, Vries RD, Samson RA (2014) Modern taxonomy of biotechnologically important Aspergillus and Penicillium species. Adv Appl Microbiol 86:199–249. https://doi.org/10.1016/B978-0-12-800262-9.00004-4
Sarmah N, Revathi D, Sheelu G, Yamuna Rani K, Sridhar S, Mehtab V, Sumana C (2018) Recent advances on sources and industrial applications of lipases. Biotechnol Progr 34:5–28. https://doi.org/10.1002/btpr.2581
Svendsen A (2000) Lipase protein engineering. BBA-Protein Struct Mol Enzymol 1543:223–238. https://doi.org/10.1016/s0167-4838(00)00239-9
Takahashi S, Osugi K, Shimekake Y, Shinbo A, Abe K, Kera Y (2019) Characterization and improvement of substrate-binding affinity of d-aspartate oxidase of the thermophilic fungus Thermomyces dupontii. Appl Microbiol Biotechnol 103:4053–4064. https://doi.org/10.1007/s00253-019-09787-y
Wang YC, Zhao N, Ma JW, Liu J, Yan QJ, Jiang ZQ (2019) High-level expression of a novel α-amylase from Thermomyces dupontii in Pichia pastoris and its application in maltose syrup production. Int J Biol Macromol 127:683–692. https://doi.org/10.1016/j.ijbiomac.2019.01.162
Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer FT, de Beer TAP, Rempfer C, Bordoli L, Lepore R, Schwede T (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46(2):296–303. https://doi.org/10.1093/nar/gky427
Xu F, Chen S, Xu G, Wu J, Yang L (2015) Discovery and expression of a Pseudomonas sp. esterase as a novel biocatalyst for the efficient biosynthesis of a chiral intermediate of pregabalin. Biotechnol Bioproc Eng 20:473–487. https://doi.org/10.1007/s12257-015-0069-1
Zheng RC, Wang TZ, Fu DJ, Li AP, Li XJ, Zheng YG (2013) Biocatalytic synthesis of chiral intermediate of pregabalin with high substrate loading by a newly isolated Morgarella morganii ZJB-09203. Appl Microbiol Biotechnol 97:4839–4847. https://doi.org/10.1007/s00253-013-4810-4
Zheng RC, Ruan LT, Ma HY, Tang XL, Zheng YG (2016) Enhanced activity of Thermomyces lanuginosus lipase by site-saturation mutagenesis for efficient biosynthesis of chiral intermediate of pregabalin. Biochem Eng J 113:12–18. https://doi.org/10.1016/j.bej.2016.05.007
Zheng JY, Lan X, Li XJ, Huang LJ, Zhang YJ, Wang Z (2019) High-level expression and characterization of a stereoselective lipase from Aspergillus oryzae in Pichia pastoris. Protein Expre Purif 155:1–7. https://doi.org/10.1016/j.pep.2018.10.012
Zhong W, Zhang M, Li X, Zhang Y, Wang Z, Zheng J (2020) Enantioselective resolution of (R, S)-2-phenoxy-propionic acid methyl ester by covalent immobilized lipase from Aspergillus oryzae. Appl Biochem Biotechnol 190:1049–1059. https://doi.org/10.1007/s12010-019-03145-4
Acknowledgements
The authors greatly appreciate the help of reviewers for their kindness in editing this manuscript. This research was financially supported by the National Natural Science Foundation of China (Grant Nos. 31660247 and 31600639) and Science and Technology Project Founded by the Education Department of Jiangxi Province (Grant No. GJJ202305).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declared that they have no conflict of interest to this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, XJ., Li, Q., Zhan, XX. et al. Expression and characterization of a thermostable lipase from Thermomyces dupontii. Chem. Pap. 76, 2811–2821 (2022). https://doi.org/10.1007/s11696-022-02068-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11696-022-02068-5