Abstract
Extra virgin olive oil is produced through either a cold press procedure or a centrifugation with no thermal and chemical treatments and it is considered as the best quality oil under the category of olive oils. The superior properties of olive oil due to its rich in phenolic and antioxidant content and its contribution to prevent several health problems has increased the demand for olive oil over the years. Consequently, it is nowadays sold at remarkably higher price than regular vegetable oils in the market. Unfortunately, extra virgin olive oil (EVOO) has been adulterated with other cheap oils due to potential high commercial profit. Even though, there are methods available to detect the adulteration in EVOO (such as chromatographic methods and PCR), alternative simpler and faster methods are being studied. In this study, performance of portable Raman spectroscopy to quantify soybean oil (SO) adulteration [up to 25 % (w/w)] in EVOO has been evaluated. Partial Least Square Regression (PLSR) calibration models were developed and both internally (using cross-validation, leave-one-out approach) and externally (using an independent sample set) validated. The model gave standard error of prediction (SEP) of 1.34 % (w/w) SO in EVOO and correlation coefficient of prediction (rPred) of 0.99. Additionally, the residual predictive deviation (RPD) value calculated for the model was found to be 5.71, indicating that the model was considered as “good” and could be used for routine analysis and quality control applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oil, produced from olive fruits (Olea europaea sativa Hoflm. Et Link), is a nutritious, healthy and tasty food product with a nice aroma and it is highly consumed throughout the world. Along with fresh consumption, olive oil is also utilized in other food applications such as cooking, seasonings and sauces as well as pharmaceutical applicaitons including its use as moisturizer [1] and protection against UV radiation [2, 3]. Olive oil is rich in phenolics and a good source of antioxidants and correspondingly, olive oil has been associated with prevention of several health problems such as diabetes, obesity, high blood pressure, heart disease, serum low density lipo-proteins (LDL), atherosclerosis and cancer [4–8].
In comparison with seed oils available, olive oil contains relatively lower level of saturated fatty acids and higher level of monounsaturated fatty acids [9]. Additionally, under the category of olive oil itself, products with varying qualities ranging from the best quality “extra virgin olive oil” to the lowest quality “pomace oil” are present [10]. In 2013, virgin olive oil production in the world was close to 3 million tones with the top five producer countries being Spain, Italy, Greece, Tunisia and Turkey in descending order [11]. EVOO is produced through either a cold press procedure or a centrifugation with no thermal and chemical treatments and its unsaturated fatty acid content is close to 90 % with 55–83 % oleic acid, 7.5–20 % palmitic acid and 3.5–21 % linoleic acid [10].
The unique properties of olive oil mentioned previously has increased the demand for it and therefore olive oil nowadays is sold at remarkably higher price than other regular vegetable oils. As a consequence, the adulteration of EVOO with other cheap oils is an issue due to temptation of high commercial profits by doing so. Adulteration of olive oil by fraudsters affects the consumers who purchase the product due to the health benefits. Very sensitive methods that can detect the minor components of the adulterants in EVOO such as high-performance liquid chromatography (HPLC) [12, 13], gas chromatography (GC) [13, 14], PCR analysis [15] and nuclear magnetic resonance (NMR) spectroscopy [16] have been abundant in the literature. However, these techniques require lengthy sample preparation, toxic and hazardous chemicals, sophisticated and costly instruments and skilled people to conduct the analyses. In recent years, simpler and faster methods have been becoming more and more popular in food analysis. Similarly, new approaches in detection of adulteration in olive oil including fluorescence spectroscopy [17, 18], laser-induced fluorescence [19], non-thermal plasma [20], UV spectrometry [21], mid-infrared spectroscopy [22], near-infrared spectroscopy, Raman spectroscopy, and voltammetric e-tongue [23] have been studied.
As part of the vibrational spectroscopy along with infrared, Raman spectroscopy is frequently used in food analysis and it is positioning as an attractive fingerprinting technique. Advances in spectroscopic instrumentation combined with multivariate data analysis have made this technology ideal for food analysis. This technique requires little or no sample preparation, no special accessories or sample preparations and allows measurements through transparent containers. Positive results from the past studies indicated the Raman spectroscopy as an ideal tool for oil adulteration [8, 9, 24–28].
The objective of this study was to develop a simple and rapid method to evaluate the performance of Raman spectroscopy to predict the % (w/w) level of SO adulteration in EVOO based on highly specific Raman signature profiles in combination with supervised pattern recognition techniques.
Materials and methods
Sample preparation
In this study, pure EVOO and SO were purchased from a local supermarket in Lincoln (Nebraska, USA). For each oil type, two different brands of oils were bought. One of each brands was used for preparing the samples to be included during calibration model development while the second brand was later used for the preparation of independent validation group. EVOO and SO were mixed based on weight at the ratios given in Table 1. For calibration model, a total of 26 samples were prepared. One of these samples was pure EVOO with no SO spiked (sample number one in Table 1). The next 25 samples in the calibration set was adulterated with SO from 1 % (w/w) to a maximum of 25 % (w/w) with increments of 1 % (w/w). Then, 13 EVOO samples adulterated with SO from 2 % (w/w) to a maximum of 25 % (w/w) with increments of 2 % (w/w) as shown in Table 1 were prepared as independent validation set samples. For the preparation of calibration and validation sets, different brands of oils were used. Each sample in this study was prepared as duplicates.
Raman spectra collection
Duplicate oil samples prepared previously were transferred to a short-form style glass vials with phenolic screw caps (internal diameter of 8 mm and the capacity of 2 mL) (VWR, Radnor, PA). Care was given to fill at least half of each vial with oil sample. Then, vials were sealed using the screw caps. Two vials were prepared for each sample and one Raman spectrum was collected from each vial (since there were two vials for each sample, two spectra in total were collected per sample). Raman spectra of the samples were collected using an Enwave Optronics EZRaman-M series field portable Raman spectrometer (Irvine, CA), which was connected to a laptop. All of the spectral data were displayed using the EZRaman Reader software provided with the Raman analyzer. For each oil sample, duplicate spectra were collected over a range of 3200–250 cm−1 by co-adding 64 scans at a resolution of 2 cm−1. The excitation wavelength of the laser was 785 nm, while the power of the laser was set at approximately 300–400 mW. Prior to each spectral collection, glass vials were carefully cleaned to avoid any fingerprints or other residues which may interfere with the measurements.
Multivariate analysis
The spectra collected using the portable Raman system were imported into the multivariate statistical program Pirouette 4.5 (Infometrix, Inc., Bothell, WA, USA). Quantitative model to determine the SO adulteration level in the EVOO samples were developed using PLSR. Duplicate spectra for each sample were averaged. Then, the averaged spectra were second derivative transformed (Savitzky-Golay second-order polynomial filter with a 25-point window) and smoothed (Savitzky-Golay second-order polynomial filter with a 35-point window).
PLSR is a multivariate regression analysis technique, which has been utilized to avoid overfitting during quantitative spectroscopic analysis [29]. PLSR is becoming a standard tool for modelling correlated relationships between multivariate measurements [30] and commonly used by both academia and industry. It compresses a large number of variables into a few orthogonal factors called ‘‘latent variables (PLS-factors)’’ representing the covariance of X (spectra) and Y (analyte’s concentration). Only these variables which are important to explain the variation in the model (usually <10) are used instead of thousands of wavenumbers [31].
In this study, calibration model developed was validated both internally (using leave-one-out approach of cross validation) and externally (using an independent sample set of 13 SO adulterated EVOOs as shown in Table 1). In chemometric analysis of our data, loading vectors, standard error of cross validation (SECV), standard error of prediction (SEP), correlation coefficient (r) and outlier diagnostics (Standard Residual of Sample vs. leverage) were used to evaluate the performance of the models. The SEP shows the magnitude of error expected when independent samples are introduced and predicted using the model. Number of factors included in developing the PLSR model was determined using the standard error of cross-validation (SECV) vs. number of PLS-factors without either underfitting or overfitting the data. Preparing a plot of the prediction residual error sum of squares vs. the number of factors was used to achieve that goal. Outliers were evaluated using X residuals and leverage and observations with large residuals or an unusual residual pattern considered as outliers. The leverage of a sample in the calibration model shows its potential contribution to the estimated calibration model. Samples containing abnormal standard residual (>2) and high leverage were reanalyzed and excluded if necessary, after which the calibration model was repeated.
Additionally, residual predictive deviation (RPD), which is a dimensionless statistic defined as the ratio of the standard deviation (SD) of the reference data in the validation set to the SEP, was used as an indicator of predictive ability of the models. RPD levels adopted from Williams [32] was used to further evaluate the performance of the model. According to the authors, there are six levels of RPD classification. An RPD values below 2.3 indicates very poor models and predictions, and this model would not be recommended to be used. Models with an RPD values between 2.3 and 3.0 could be used for rough screening, while RPD values between 3.1 and 4.9 shows that model can be used for screening. Higher RPD values (between 5.0 and 6.4) are considered as good and can be used for quality control applications. Lastly, RPD values of 6.5–8 and above 8.1 are considered as very good and excellent models, respectively.
Results and discussion
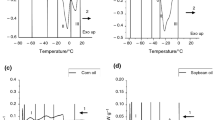
Samples used in calibration and validation sets are presented in Table 1. Calibration set contained 26 EVOO samples with SO spiked from 0 to 25 % (w/w) with 1 % (w/w) increments. Validation set included 13 EVOO samples with SO spiked from 2 to 25 % (w/w) with 2 % (w/w) increments. The data in Table 1 shows that calibration and validation set had similar characteristics considering their spiked SO % (w/w) range, mean and standard deviation values. Representative Raman spectra of pure EVOO and SO in the collected spectral region of 3200–250 cm−1 are displayed in Fig. 1a. Based on the figure, similar spectral patterns between the two types of oils used in this study were observed.
a Full raw Raman spectra of extra virgin olive oil and soybean oil in the region of between 3200 and 250 cm−1 with the major bands highlighted and b normalized Raman spectra of pure extra virgin olive oil, olive oil adulterated with soybean oil at 5, 10, 20 % and pure soybean oil in the region of between 1800 and 1000 cm−1 used for PLSR analysis
In order to build a PLSR model, initially the spectral region leading to better statistical performance was needed to be chosen. The spectral region selected and further used to build the PLSR model (1800–1000 cm−1) is highlighted in Fig. 1a and further demonstrated in Fig. 1b. According to normalized spectra of the selected samples in Fig. 1b, increasing intensity in the peaks centered at 1260 and 1665 cm−1 were observed as the SO concentration in the EVOO increased from 0 to 20 %. The band intensities at 1260 and 1665 cm−1 corresponding to the cis (=C–H) deformation and cis (C=C) stretching vibrations were previously shown to have a high correlation with increased mass percentage of SO by Zhang et al. [27]. However, the best performance statistics in our study were obtained for the model generated using the region between 1800 and 1000 cm−1 (Table 2). Additionally, selection of specific wavelengths improved the quality of the prediction compared to using the whole range in the spectrum by removing variables that may be irrelevant, noisy, or otherwise unreliable in the situation of interest [33]. The assignments of corresponding molecular vibrations occurred at the highlighted wavenumbers are shown in Table 2. In the selected region, the major vibrations were due to C–C, C=C and C=O stretchings and C=C and C–H bending vibrations. Detailed assignments for the major bands in the Raman spectra of the oils can be further listed in Table 2.
The performance statistics of the PLSR model developed in this study is shown in Table 3. Prior to PLSR analysis, all the spectra collected were initially second derivative transformed (Savitzky-Golay second-order polynomial filter with a 25-point window) and smoothed (Savitzky-Golay second-order polynomial filter with a 35-point window). Our PLSR model gave low standard error values [SEC, SECV and SEP values for the model were 1.24, 1.40 and 1.34 % (w/w)] and high correlation coefficients (rCal, rCV and rPred values ≥0.98). Lower standard error values indicate good prediction ability while higher correlation coefficients indicate better accuracy on the prediction, with correlation coefficient value above 0.80 considered good to obtain accurate predictions of the desirable variable [34]. Additionally, RPD, which is the ratio of standard deviation of spiked SO % (w/w) among the validation set samples to SEP value, was calculated as 5.71 for the model, indicating that model was good and it can be used in quality control applications. Figure 2 indicates the good correlation between the Raman estimated levels and spiked levels of SO for calibration and validation models obtained in this research. In the literature, Yang and Irudayaraj [9] evaluated near-infrared, mid-infrared and r Raman techniques to detect the lowest quality olive oil (olive pomace oil) adulteration in EVOO and compared the performance of the three techniques. The authors achieved the best performance using Raman instrument. They reported SEP of 1.72 % and rPred of 0.99. Our SEP value was slightly better than that of reported by Yang and Irudayaraj [9] (1.34 vs. 1.72 %). This could be due to higher range of SO level spiked in EVOO in their study (0–100 % SO vs. 0–25 % SO). Similarly, Mendes et al. [8] evaluated near infrared, mid-infrared and Raman spectroscopy to analyze soybean adulteration in EVOOs. They obtained the best results with PLSR model developed using Raman spectra giving SEP of 1.57 % and correlation coefficient of 0.99. Zhang et al. [27] focused on the band of 1265 cm−1 only for the evaluation of cheaper oil adulteration (soybean, corn and sunflower oil) in EVOO. They reported standard error of between 7.41 and 9.45 % for external standard method and between 4.55 and 6.96 % for support vector machine methods. The higher standard error values reported in this study could be due to the use of a single wavenumber for the model development as opposed to our study, where a defined region (1800–100 cm−1) abundant with the vibrations of functional groups related to fatty acids in the oil samples was used.
For PLSR model development, only three PLS-factors were found to be optimum (Fig. 3). The selection of the optimal number of PLS-factors was done by a cross-validation approach yielding the minimum root mean square error of cross validation (RMSECV). Typically, the quality of the prediction increases by increasing the number of orthogonal latent variables because they explain the relevant variance in the model; however, including too many latent variables can have an adverse effect in the prediction due to “overfitting” the data by incorporating random noise. PLSR loading plot in Fig. 4 showed the specific portions of the original spectra that were related with the highest variation in the calibration model. Regression coefficients of each wavenumber to spiked SO content (%, w/w) show the association between the highest variation in the calibration model and their corresponding bands of the spectrum, inferring the nature of the chemical components in the sample contributing to the regression. The PLS loading for the first PLS-factor indicated the bands at 1760, 1665, 1448, 1306, 1260 and 1075 cm−1 explained 99.7 % of the variation. These vibrations were relevant to the functional groups of fatty acids as can be interpreted from Table 2.
Conclusion
Overall, our results support the application of portable Raman spectroscopy and PLSR for the accurate prediction ability of SO adulteration in EVOO, showing high reliability of application in routine analysis and quality control. The technique used in this study have the potential to be used for fast, easy, low-cost and non-destructive detection and quantification of SO adulteration in EVOO with no sample preparation required.
References
M.N. Franco, T. Galeano-Díaz, Ó. J.G. López, Fernández-Bolaños, J. Sánchez, C. De Miguel, M.V. Gill, D. Martín-Vertedor, Food Chem. 163, 289–298 (2014)
S. D’Angelo, D. Ingrosso, V. Migliardi, A. Sorrentino, G. Donnarumma, A. Baroni, L. Masella, M.A. Tufano, M. Zappia, P. Galletti, Free Radic. Biol Med. 38(7), 908–919 (2005)
F. Soltanipoor, M. Delaram, S. Taavoni, H. Haghani, Complement Ther. Med. 20(5), 263–266 (2012)
M. Kratzb, P. Cullenc, Eur. J. Lipid Sci. Technol 104, 698–705 (2002)
J.L. Harwood, P. Yaqoob, Eur. J. Lipid Sci. Technol 104(9–10), 685–697 (2002)
S. Lopez, B. Bermudez, S. Montserrat-de la Paz, S. Jaramillo, L.M. Varela, A. Ortega-Gomez, F.J. Muriana, Biochim. Biophys. Acta Biomembr. 1838(6), 1638–1656 (2014)
S. Lamy, A. Ouanouki, R. Béliveau, R.R. Desrosiers, Exp. Cell Res 322(1), 89–98 (2014)
T.O. Mendes, R.A. da Rocha, B.L. Porto, M.A. de Oliveira, V.D.C. dos Anjos, M.J. Bell, Food Anal. Methods 8(9), 2339–2346 (2015)
H. Yang, J. Irudayaraj, JAOCS 78(9), 889–895 (2001)
IOOC (International Olive Oil Council). Trade standard applying to olive oil and olive–pomace oil. COI/T. 15/NC No 3/Rev. 7, May (2013)
FAOSTAT (Food and Agriculture Organization of the United Nations Statistics Division). http://faostat3.fao.org/browse/Q/QD/E. Accessed 13 June 2016
M. Lisa, M. Holcapek, M. Bohac, J. Agric. Food Chem. 57(15), 6888–6898 (2009)
H. Jabeur, A. Zribi, J. Makni, A. Rebai, R. Abdelhedi, M. Bouaziz, J. Agric. Food Chem. 62(21), 4893–4904 (2014)
H. Jabeur, A. Zribi, M. Bouaziz, Food Anal. Methods 9(3), 712–723 (2015)
M. Vietina, C. Agrimonti, N. Marmiroli, Food Chem. 141(4), 3820–3826 (2013)
G. Fragaki, A. Spyros, G. Siragakis, E. Salivaras, P. Dais, J. Agric. Food Chem. 53(8), 2810–2816 (2005)
K.I. Poulli, G.A. Mousdis, C.A. Georgiou, Anal. Bioanal. Chem. 386, 1571–1575 (2006)
Y. Xu, H. Li, Q. Chen, J. Zhao, Q. Ouyang, Int. J. Food Prop. 18(9), 2085–2098 (2015)
T. Mu, S. Chen, Y. Zhang, H. Chen, P. Guo, F. Meng, Food Anal. Methods 9(1), 275–279 (2016)
J. Van Durme, J. Vandamme, Food Chem. 208, 185–191 (2016)
L. Jiang, H. Zheng, H. Lu, J. Food Sci. Technol. 52(1), 479–485 (2015)
X. Sun, W.L.X. Lin, Q. Shen, H. Luo, Anal. Methods 7(9), 3939–3945 (2015)
I.M. Apetrei, C. Apetrei, Comput. Electron. Agric. 108, 148–154 (2014)
V. Baeten, M. Meurens, M.T. Morales, R. Aparicio, J. Agric. Food Chem. 44(8), 2225–2230 (1996)
M.Q. Zou, X.F. Zhang, X.H. Qi, H.L. Ma, Y. Dong, C.W. Liu, X. Guo, H. Wang, J. Agric. Food Chem. 57(14), 6001–6006 (2009)
X. Zhang, X. Qi, M. Zou, F. Liu, Anal. Lett. 44(12), 2209–2220 (2011a)
X.F. Zhang, M.Q. Zou, X.H. Qi, F. Liu, C. Zhang, F. Yin, J. Raman Spectrosc. 42(9), 1784–1788 (2011b)
W. Dong, Y. Zhang, B. Zhang, X. Wang, Anal. Methods 4(9), 2772–2777 (2012)
D.M. Haaland, E.V. Thomas, Anal. Chem. 60, 1193–1202 (1988)
S. Dejong, J. Chemom. 7, 551–557 (1993)
L. Moseholm, Environ. Pollut. 53, 313–331 (1988)
P.C. Williams, in Near-infrared technology in the agricultural and food industries, ed. by P.C. Williams, K.H. Norris (American Association of Cereal Chemistry, St. Paul, 2001), p. 169
R. Leardi, L. Nørgaard, J. Chemometr. 18(11), 486–497 (2004)
L. Zhang, M.A. Schultz, R. Cash, D.M. Barrett, M.J. McCarthy, Food Control 40(1), 214–223 (2014)
Acknowledgments
Funding from Higher Education Council (YÖK) as a visitor scientist by overseas scholarship (under Higher Education Law No. 2547 under Article 39) is acknowledged. The honorable Prof. Dr. Randy Wehling (from University of Nebraska-Lincoln, USA) is thanked for opening his lab and allowing the use of his equipment. Brandon Holder is further thanked for his friendship and help in the lab.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yildiz Tiryaki, G., Ayvaz, H. Quantification of soybean oil adulteration in extra virgin olive oil using portable raman spectroscopy. Food Measure 11, 523–529 (2017). https://doi.org/10.1007/s11694-016-9419-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11694-016-9419-8