Abstract
Uc.40 is a long noncoding RNA that is highly conserved among different species, although its function is unknown. It is highly expressed in abnormal human embryonic heart. We previously reported that overexpression of uc.40 promoted apoptosis and inhibited proliferation of P19 cells, and downregulated PBX1, which was identified as a potential target gene of uc.40. The current study evaluated the effects of uc40-siRNA-44 (siRNA against uc.40) on the differentiation, proliferation, apoptosis, and mitochondrial function in P19 cells, and investigated the relationship between uc.40 and PBX1 in cardiomyocytes. The uc.40 silencing expression was confirmed by quantitative real-time polymerase chain reaction (RT-PCR). Observation of morphological changes in transfected P19 cells during different stages of differentiation revealed that uc40-siRNA-44 increased the number of cardiomyocyes. There was no significant difference in the morphology or time of differentiation between the uc40-siRNA-44 group and the control group. uc40-siRNA-44 significantly promoted proliferation of P19 cells and inhibited serum starvation-induced apoptosis. There was no significant difference in mitochondrial DNA copy number or cellular ATP level between the two groups, and ROS levels were significantly decreased in uc40-siRNA-44-transfected cells. The levels of PBX1 and myocardial markers of differentiation were examined in transfected P19 cells; uc40-siRNA-44 downregulated myocardial markers and upregulated PBX1 expression. These results suggest that uc.40 may play an important role during the differentiation of P19 cells by regulation of PBX1 to promote proliferation and inhibit apoptosis. These studies provide a foundation for further study of uc.40/PBX1 in cardiac development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Congenital heart disease (CHD), a common birth defect that includes cardiovascular malformations, is a main cause of infant death (Marino et al. 2012). Long noncoding RNA (lncRNA) is defined as noncoding RNA over 200 nt in length, and can participate in a variety of biological processes, including cancer and cardiovascular disease (Leung and Natarajan 2014; Han et al. 2015). The pathological mechanism of lncRNA action includes regulation of gene expression, cell differentiation, cell proliferation, and apoptosis (Chen and VanBuren 2014). In a chip study of lncRNAs expressed differently in the embryonic heart (Song et al. 2013), a group of lncRNAs were identified that contained ultraconserved elements attracted our attention. Recently, Li et al. (Li et al. 2017) found that overexpression of lnc-TUC40 (uc.40) inhibited differentiation of P19 cells into cardiomyocytes. NKX2.5 is associated with early heart development (Paffett-Lugassy et al. 2013), and SRF is involved in cyclization, differentiation, and maturation of heart cells. A correlation was found between uc.40 and heart development due to high expression of uc.40 in abnormal specimens, and inhibition of PBX1 genes associated with embryonic heart development (Qin et al. 2004; Stankunas et al. 2008). This evidence provides a molecular basis to understand the development of CHD, but it cannot completely prevent or treat CHD. Therefore, it is necessary to keep investigating the mechanism of embryonic heart development.
The P19 mouse embryonic carcinoma cell line is multipotent and can differentiate into cardiomyocytes with embryonic body formation in the presence of dimethyl sulfoxide (DMSO) (Yang et al. 2009). We utilized this cell line to investigate the effects of uc40-siRNA-44 (siRNA against uc.40) on differentiation, proliferation, apoptosis, and mitochondrial function. We hypothesized that silencing uc.40 using small interfering RNA (siRNA) would reduce abnormal cardiomyocytes differentiation.
Materials and Methods
Construction of the lncRNA-uc.40 (Lnc-TUC40) silencing plasmid
Vectors were constructed by Jima Biotechnology (GenePharma, Shanghai, China), and the recombinant vector was named pGPU6/GFP/Neo-Uc40-siRNA-44. The negative control vector (pGPU6/GFP/Neo-siNC), which expresses no homology to the target gene sequence, and the positive control vector (pGPU6/GFP/Neo-siGAPDH), which confirms the reliability of the method of RNA extraction and gene expression in the transfection experiment, were constructed simultaneously. The sequences of the three siRNAs are as follows: uc40-siRNA-44:5′-GGATTCTGCCAGGCGAA ATGA-3′; siNC:5′-GTTCTCCGAACGTGTCACGT-3′;siGAPDH:5′-GTATGACAACAGCCTCAAG-3′.
Cell culture, transfection, and differentiation
P19 cells were purchased from the American Type Culture Collection (Manassas, VA), and were cultured in modified Eagle medium (α-MEM; Gibco-BRL, Waltham, MA) containing 10% fetal bovine serum (Gibco-BRL), 100 U/mL penicillin, and100 μg/mL streptomycin (Gibco-BRL) in a 5% CO2 atmosphere at 37°C. The medium was replaced by complete medium without antibiotics 6 h before transfection. Uc40-siRNA-44, siNC, and siGAPDH vectors were transfected into P19 cells using the lipofectamine method (Lipofectamine 2000, Life Technology, Carlsbad, CA), and returned to the incubator for 36 h. Cell fluorescence was observed by fluorescence microscopy using a ZEISS AX10 (Zeiss, Oberkochen, Germany). The cells were also collected to confirm uc.40 silencing efficiency by real-time polymerase chain reaction (RT-PCR). Stable cell lines expressing the siRNAs were screened for 5 d, and clones were cultured with 0.5 mg/mL neomycin (Invitrogen, Carlsbad, CA) (Wu and Adamson 1993). During differentiation, the cells were maintained in suspension for the first 4 d in complete medium containing 1% DMSO (Sigma, Ronkonkoma, NY) in bacteriological dishes with a shallow high-pressured antiseptic agar, and were incubated in a 5% CO2 atmosphere at 37°C. On day 3, 70% of the medium was replaced with fresh complete medium containing 1% DMSO. On day 4, the cell aggregates were transferred to a new dish, and then cultured for 6 d (days 5–10) without DMSO. Culture medium was replaced every 2 d, and morphological changes of transfected P19 cells were observed and photographed using ZEISS AX10 (Zeiss). Cells at different stages of differentiation were collected for further use.
Cell cycle and proliferation analysis
The cell cycle was analyzed using the propidium iodide (PI) staining method by BD Accuri flow cytometer (FCM) (Shen et al. 2013). When the cells were approximately 70% confluent (70% of surface of flask covered by cell monolayer), they were serum starved for 24 h before replacing with complete medium and cell collection every 12 h. The cells were washed twice with phosphate-buffered saline (PBS), digested with trypsin, and resuspended in 1 mL cold PBS, before being pelleted and fixed with 1 mL cold 70% ethanol. The cells were then stained with 0.5 mL PI and incubated at 37°C in the dark for 30 min. The red fluorescence was detected at 488 nm using FCM, and cellular DNA content was analyzed using the Modfit software (Verity Software House, Topsham, ME).
Cell proliferation was analyzed using the cell counting kit 8 (CCK8) (Dojindo, Tokyo, Japan) according to the manufacturer’s instructions. In brief, 1 × 103 cells were inoculated into 96-well plates and incubated in 5% CO2 at 37°C for 24 h. Next, 10 μL of CCK8 solution was added to each well once a day for 5 d. The medium was changed every 2 d. Each set of 6 replicate wells was assessed by measuring absorbance at 450 nm after incubating in 5% CO2 at 37°C for an additional 2 h.
Apoptosis analysis
Cells with 70% confluent were serum starved for 18 h, and cell apoptosis was assessed using the Annexin V-PE/7-AAD method (BD Pharmingen, San Jose, CA). The cells were washed twice with PBS, trypsin digested, and washed twice again with cold PBS before being resuspended in cold 1× binding buffer to a final concentration of 1 × 106 cells/mL. An aliquot of cells (1 × 105 cells) was then transferred to a 5-mL culture tube, 5 μL of Annexin V-PE and 5 μL of 7-AAD were added, and the tubes were gently oscillated and incubated for 15 min at room temperature in the dark. Then, 1× binding buffer was added to each tube, and samples were analyzed by FCM within 1 h. The apoptosis ratio was calculated from the resulting plot, counting to the upper left quadrant as dead cells, the lower left quadrant as living cells, the right upper quadrant as late apoptotic cells, and the lower right quadrant as early apoptotic cells.
Caspase-3 activity was detected using a colorimetric assay kit (KeyGen, Nanjing, China) according to the manufacturer’s instructions. Cells were cultured in serum-free α-MEM for 24 h and collected by adding 100 μL lysis buffer and incubating on ice. Reaction buffer and caspase-3 substrate Ac-DEVD-pNA was added to cell lysates containing approximately 20 μg protein, before incubating at 37°C in the dark for 4 h. The absorbance value was measured at 405 nm in a microplate reader (BioTek, Winooski, VT).
Mitochondrial function analysis
The relative amounts of mitochondrial DNA (mtDNA) were determined using a DNA extraction kit (Omega, Norcross, GA) and by qPCR according to the manufacturer’s instructions. In brief, DNA from cells on the 10th day of differentiation was isolated and quantified by spectrophotometry. The ratio of mtDNA to nuclear DNA was assessed by amplifying a 110-bp mtDNA fragment containing the CytB gene and a 291-bp region of the nuclear 28 S internal reference gene by qPCR.
Cellular ATP production was assessed using a luciferase-based luminescence assay kit (Beyotime, Shanghai, China) according to the manufacturer’s instructions. In brief, the ATP concentration in differentiated P19 cells was determined by homogenizing the cells in cold ATP releasing buffer and determining ATP concentration with a Synergy H4 Hybrid multimode microplate reader (BioTek). The level of ATP was normalized to protein concentration (nmol/mg protein) while cellular protein content was detected using BCA method (Beyotime).
A DCFDA fluorescent probe (Beyotime) was used to determine intracellular reactive oxygen species (ROS) levels. The cells were differentiated for 10 d, and 5 μM of H2-DCFDA was added to each well and incubated for 30 min at 37°C. The cells were washed three times with prewarmed PBS, and the green fluorescence was observed and photographed using ZEISS AX10 (Zeiss). The cells were then digested with trypsin and centrifuged at 1500×g for 5 min at room temperature. Pellets were washed twice with PBS before being resuspended in PBS and analyzed by FCM to determine the relative levels of intracellular ROS.
Mitochondrial membrane potential (MMP) relative fluorescence intensity was estimated using a JC-1 probe from a MMP detection kit (Beyotime), according to the manufacturer’s instructions. Briefly, the cells were differentiated for 10 d, before being washed twice with PBS and incubated in 5-mL of JC-1 per dish for 30 min at 37°C. After washing three times with prewarmed PBS, green and red fluorescence was observed and photographed using ZEISS AX10 (Zeiss). Then, the cells were digested with trypsin and centrifuged at 1500×g for 5 min at room temperature. Pellets were resuspended in prewarmed PBS and analyzed by FCM to obtain the relative fluorescence intensity.
Quantitative RT-PCR and Western blot
qRT-PCR was performed to confirm the efficiency of uc.40 silencing, to determine the relative amount of mtDNA, and to analyze the expression of the potential target gene, PBX1, or cardiac markers, such as ANP and cTnI, during differentiation of uc40-siRNA-44-transfected P19 cells. Total RNA was extracted using TRIZOL reagent (Invitrogen), and cDNA was synthesized from 1 μg total RNA using a reverse transcriptase kit (Promega, Fitchburg, WI). The reaction protocol was as follows: 42°C for 60 min and 70°C for 10 min. RT-PCR was performed using the SYBR green select Master Mix (Life Technology) with strand-specific primers (shown in Table 1) in a LifeProFlex PCR detection system (Life Technology). The reaction process was as follows: 95°C for 10 min, followed by 40 cycles of 95°C for 15 s, 60°C for 60 s. The relative expression levels were quantified by the cycle threshold (Ct) value using the reference gene glyceraldehyde 3-phosphate dehydrogenase (GAPDH). Each sample was analyzed in triplicate in three independent experiments, and the expression levels were calculated using 2-ΔΔCt method in which ∆∆Ct is calculated by mean ∆Ct of transfected P19 cell minus mean ∆Ct of reference gene and here ∆Ct = Ctsample − CtGAPDH.
Statistical analysis
Data are presented as the mean ± standard deviation (SD) of three independent experiments performed in triplicate. All data were analyzed using the SPSS21.0 statistical software package (SPSS Inc., Chicago, IL). Student t test was used to determine the significance of differences between two datasets. The threshold of statistical significance was defined as p < 0.05.
Results
Confirmation of uc.40 silencing
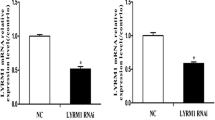
Vectors pGPU6/GFP/Neo-Uc40-siRNA-44 and pGPU6/GFP/Neo-siNC (Fig. 1A) were transiently transfected into P19 cells and expression of green fluorescent protein (GFP) was observed by fluorescence microscopy. Our results indicate that the transection efficiency was similar for all vectors from the images collected (Fig. 1B). The expression of uc.40 was normalized to the endogenous reference gene GAPDH by RT-PCR. As shown in Fig. 1C, P19 cells transfected with uc40-siRNA-44 express significantly lower levels of uc.40 RNA than those transfected with siNC (**p < 0.01). The steady-state expression of uc.40 RNA in uc40-siRNA-44-transfected P19 cells was only 33.3% of siNC-transfected P19 cells 36 h after transfection.
siRNA transfection and efficiency in P19 cells. (A) The structure of the vector. Uc40-siRNA-44, siNC, and siGAPDH were inserted into multiple cloning site (MCS). (B) The transfection efficiency after transfection for 12 and 36 h are observed by fluorescence microscope (×10). (C) Confirmation of uc.40 silencing in P19 cells by qRT-PCR. Uc.40 expression was inhibited by 66.7% in P19 cells transfected with uc40-siRNA-44 vector compared to cells transfected with siNC vector after 36 h. Values are means ± SD of three independent experiments (n = 6, **p < 0.01).
Uc40-siRNA-44 promotes the differentiation of P19 cells
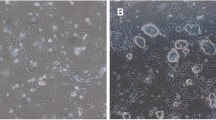
The morphological changes at different stages of differentiation of uc40-siRNA-44-transfected P19 cells were observed with an inverted microscope equipped with a digital camera. As shown in Fig. 2A, transfection with uc40-siRNA-44 promoted P19 cell differentiation into cardiomyocytes, increasing the size and amount of the embryonic-like bodies on the fifth day of differentiation (from the images collected). The cardiomyocytes aggregated on the fourth day of differentiation, and beating-like myocardial cells appeared after 8 to 9 d of differentiation. However, there were no significant changes to cell morphology or time to the appearance of beating-like myocardial cells between the two groups (Fig. 2b).
Effect of uc40-siRNA-44 on the differentiation of P19 cells. (A) Uc40-siRNA-44 can promote differentiation of P19 cells into cardiomyocytes and increased the size and amount of embryonic-like bodies from the images collected. (B) Morphological appearance of differentiated P19 cells. P19 cells transfected with uc40-siRNA-44 and siNC vectors were stimulated to differentiate for 10 d. Images of cells on days 0, 2, 4, 6, 8, and 10 were captured, but there were no obvious differences either in the morphology or time needed for the appearance of beating-like myocardial cells between the two groups.
Uc40-siRNA-44 promotes P19 cell proliferation and inhibits apoptosis
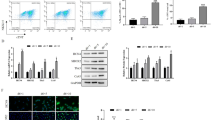
The cell cycle of uc40-siRNA-44- and siNC-transfected P19 cells was evaluated using PI staining, and cell proliferation was assessed using a CCK8 viability assay. The proportion of S phase cells in uc40-siRNA-44-transfected P19 cells was higher than in siNC-transfected P19 cells (Fig. 3A). Uc40-siRNA-44-transfected P19 cell had an increased growth rate compared to siNC-transfected P19 cells. The optical density values of uc40-siRNA-44-transfected P19 cell are significantly higher than siNC-transfected P19 cells from the fourth day of differentiation (Fig. 3B, *p < 0.05).
The effect of uc40-siRNA-44 on cell proliferation and apoptosis. (A) uc40-siRNA-44 promoted the proliferation of P19 cells through promoting more cells to enter the S phase. Cell cycle stages were assessed each 12 h. (B) Cell proliferation was measured over 5 consecutive days. Transfection with uc40-siRNA-44 promoted the proliferation of P19 cells. Cell apoptosis was detected using Annexin V-PE/7-AAD staining by FCM (C) or caspase-3 activity. The results from two assays indicated that uc40-siRNA-44 could inhibit serum deprivation-induced apoptosis.
The early stage of cell apoptosis was analyzed using the Annexin V-PE/7-AAD method (Hu et al. 2015). Uc40-siRNA-44 decreased the number of apoptotic cells in response to serum deprivation (Fig. 3C). These results indicate that uc40-siRNA-44 inhibited cell apoptosis after serum starvation.
Effects of uc40-siRNA on mtDNA copy number, cellular ATP production, intracellular ROS, and MMP in P19 cells
The relative mtDNA/nuclear DNA ratio was assessed by PCR; there was no significant difference between the two groups, which suggested that uc40-siRNA-44 has no significant effect on the numbers of copies of mtDNA, or that there are other compensatory mechanisms (Fig. 4A). ATP content in cells was detected by biological luminescence and was calibrated to protein concentration. There was no significant difference between the two groups (Fig. 4B).
(A) Effect of uc40-siRNA-44 on mitochondrial function of differentiated P19 cells. Effects of uc40-siRNA-44 on mtDNA copy number on the 10th day of differentiation; there was no significant difference between the two groups (n = 6, p > 0.05). (B) Effects of uc40-siRNA-44 on intracellular ATP level; there was no significant difference between the two groups (n = 6, p > 0.05). (C) Effects of uc40-siRNA-44 on intracellular ROS content in differentiated P19 cells; treatment with uc40-siRNA-44 significantly reduced intracellular ROS (n = 6, * p < 0.05). (D) The green fluorescence was observed and photographed using ZEISS AX10. (E) Effects of uc40-siRNA-44 on MMP in differentiated P19 cells; treatment with uc40-siRNA-44 significantly increases MMP (n = 6, *p < 0.05). The green and red fluorescence was observed and photographed using ZEISS AX10.
The relative levels of intracellular ROS in the uc40-siRNA-44 group were lower than that of the siNC group (Fig. 4C). Compared with the siNC group, the green fluorescence (JC-1 monomer) in the uc40-siRNA-44 group was weaker whereas the red one (JC-1 aggregates) was stronger from the images collected, which showed the cell membrane potential was higher (Fig. 4D). The results of FCM detection also showed that the relative fluorescence intensity (JC-1 aggregates) in the uc40-siRNA-44 group was higher than in the siNC group, suggesting that uc40-siRNA-44 causes and increase in MMP of myocardial-like cells (Fig. 4E).
Uc40-siRNA-44 can upregulate expression of the myocardial-specific gene PBX1 and downregulate markers of cardiomyocyte differentiation in P19 cells
The myocardial-specific gene PBX1 and myocardial-specific proteins, such as ANP and cTnI, are expressed in a developmentally controlled manner by embryonic stem cell-derived cardiomyocytes (Holmgren et al. 2005; Arrington et al. 2012; Suzuki et al. 2012). As shown in Fig. 5, the expression of all the markers investigated increased gradually during the process of differentiation. The expression of cTnI (day 8–day 10) and ANP (day 10) was significantly reduced, whereas the expression of PBX1 mRNA (day 8–day 10) was significantly increased in cells transfected with uc40-siRNA-44 (*p < 0.05, **p < 0.01).
Discussion
The latest research shows that the pathogeny of CHD is mainly related to genetic defects (Morlando et al. 2014). The heart is the first organ formed in the embryo, and any intrinsic or extrinsic factors that affect heart development may cause heart malformation. The growth and development of the fetal heart depends on the balance of proliferation and apoptosis of cardiomyocytes. Any imbalance of proliferation and apoptosis may lead to heart deformity or CHD (Ji et al. 2003; Bernard et al. 2010). Genes such as NKX2.5, TBX1, MEF2A, and MEF2C, and the Wnt/Notch signaling pathway, are involved in regulation of cardiac development, which provides some of the molecular basis for understanding the occurrence and development of CHD. However, there is still much that is unknown in the mechanism of embryonic heart development (Xu et al. 2004; Srivastava 2006; Prall et al. 2007).
The discovery of noncoding RNA greatly broadens the horizon for understanding the mechanism of disease occurrence and development (Bartel 2004). There are a large number of studies on microRNAs in cardiac development and cardiovascular diseases, but the studies on lncRNA are rare (Qin et al. 2013; Harada et al. 2014). The exploration of the mechanism of action of lncRNAs will promote understanding of the mechanisms of heart development greatly. LncRNA regulation is likely to be very complex, as it may be related to chromosome recombination or epigenetic factors, if they are mainly distributed in the nucleus (Gendrel and Heard 2014). Uc.40 is an lncRNA with an unknown role in embryonic heart malformations that is highly conserved in multiple species and is expressed in cardiomyocytes (Song et al. 2013). The high expression of uc.40 in abnormal samples, and its highly conserved sequence, suggests that it has an important function. Li et al. found that overexpression of uc.40 could inhibit differentiation of P19 cells into cardiomyocytes (Li et al. 2017). Furthermore, bioinformatic analyses indicated that PBX1 was a possible target gene of uc.40, providing us with an avenue of further investigation (Li et al. 2016). Uc40-siRNA-44-transfected P19 cells were less susceptible to apoptosis induced by serum starvation, with many cells gathering in the S phase, and the rate of proliferation in the uc40-siRNA-44 group was higher than that of the siNC group, suggesting that cell cycle shortening occurs in the uc40-siRNA-44 group, in addition to faster cell proliferation. This suggests that uc.40 plays a role in regulating cardiac cell proliferation and apoptosis, and may influence morphogenesis in the embryonic heart.
Mitochondria are important organelles that determine the survival and death of the cells, and changes in their function will affect the transmission and amplification of the apoptotic signal (Crow et al. 2004), leading to dynamic changes in the structure and function of mitochondria (Martinou and Youle 2011; Ugarte-Uribe and Garcia-Saez 2014). Therefore, we analyzed possible mechanisms by which uc40-siRNA-44 reduced apoptosis using mitochondrial function analysis. We found that uc40-siRNA-44 inhibited production of ROS in cardiomyocytes, which suggests that uc40-siRNA-44 may protect mitochondrial function. In addition, the size of MMP can directly reflect the function of mitochondria (Matsusaka and Ichikawa 1997). In the present study, we found that uc40-siRNA-44 could slightly increase the MMP of cardiomyocytes, suggesting that mitochondrial function was normal. Mitochondria are the energy sources of cardiomyocytes (Hatefi 1985); thus, cellular ATP levels can be used to assess the function of mitochondria. When mitochondrial function is damaged, the production of ATP is decreased, which leads to deficiency of the energy supply. In the uc40-siRNA-44 group, cellular ATP content was similar to that of the siNC group. These results indicate that the expression of uc40-siRNA-44, and reduction of the level of uc.40 in the cells, may be beneficial to the maintenance of mitochondrial function. Uc40-siRNA-44 expression may further exert its protective effects by inhibiting the initiation of apoptosis by reducing the number of ROS.
In addition, we analyzed the interaction between uc.40 and PBX1 to investigate the molecular mechanism of uc.40-induced cardiac malformations in embryos. In our study, uc40-siRNA-44 upregulated PBX1 gene expression, although the level by day 10 decreased rapidly. Therefore, we believe that uc.40 plays its role in the growth and development of myocardial cells through the regulation of its target gene PBX1. The expression of uc.40 is negatively correlated with the amount of its regulating gene PBX1, which means uc.40 plays a negative role at the pretranscription and /or transcriptional level. These data are consistent with a previous study suggesting that uc.40 overexpression reduced PBX1 expression and cardiomyocyte induction and differentiation and inhibited proliferation (Li et al. 2017). However, the expression of ANP and cTnl was deceased when the cells were transfected with uc40-siRNA-44. For a general case of paradoxical function, the different conditions of the microRNA can bring the threshold to binding to the different target molecules. The uc.40 downstream signal pathway should be further elucidated.
In summary, our study shows that inhibiting levels of uc.40 may promote the differentiation and proliferation of P19 cells, and inhibit the apoptosis of P19 cells, which may result in cardiac malformation. Uc.40 may also regulate the balance of proliferation and apoptosis through the regulation of PBX1. Although uc40-siRNA-44 can inhibit production of ROS, the MMP increased slightly and there was no significant difference in mtDNA and ATP between the two groups. However, the detailed molecular mechanisms and signal pathways that mediate the imbalance between proliferation and apoptosis in response to uc.40 silencing or overexpression require further exploration in future studies.
Conclusion
uc40-siRNA-44 may play an important role during the differentiation of P19 cells by regulation of PBX1 to promote proliferation and inhibit apoptosis, which may induce abnormal heart development. These studies provide a foundation for further study of uc.40/PBX1 in cardiac development.
References
Arrington CB, Dowse BR, Bleyl SB, Bowles NE (2012) Non-synonymous variants in pre-B cell leukemia homeobox (PBX) genes are associated with congenital heart defects. Eur J Med Genet 55:235–237
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Bernard D, Prasanth KV, Tripathi V, Colasse S, Nakamura T, Xuan Z, Zhang MQ, Sedel F, Jourdren L, Coulpier F, Triller A, Spector DL, Bessis A (2010) A long nuclear-retained non-coding RNA regulates synaptogenesis by modulating gene expression. EMBO J 29:3082–3093
Chen H, VanBuren V (2014) A provisional gene regulatory atlas for mouse heart development. PLoS One 9:e83364
Crow MT, Mani K, Nam YJ, Kitsis RN (2004) The mitochondrial death pathway and cardiac myocyte apoptosis. Circ Res 95:957–970
Gendrel AV, Heard E (2014) Noncoding RNAs and epigenetic mechanisms during X-chromosome inactivation. Annu Rev Cell Dev Biol 30:561–580
Han D, Wang M, Ma N, Xu Y, Jiang Y, Gao X (2015) Long noncoding RNAs: novel players in colorectal cancer. Cancer Lett 361:13–21
Harada M, Luo X, Murohara T, Yang B, Dobrev D, Nattel S (2014) MicroRNA regulation and cardiac calcium signaling: role in cardiac disease and therapeutic potential. Circ Res 114:689–705
Hatefi Y (1985) The mitochondrial electron transport and oxidative phosphorylation system. Annu Rev Biochem 54:1015–1069
Holmgren D, Westerlind A, Lundberg PA, Wahlander H (2005) Increased plasma levels of natriuretic peptide type B and A in children with congenital heart defects with left compared with right ventricular volume overload or pressure overload. Clin Physiol Funct Imaging 25:263–269
Hu Y, Lin X, Wang P, Xue YX, Li Z, Liu LB, Yu B, Feng TD, Liu YH (2015) CRM197 in combination with shRNA interference of VCAM-1 displays enhanced inhibitory effects on human glioblastoma cells. J Cell Physiol 230:1713–1728
Ji P, Diederichs S, Wang W, Boing S, Metzger R, Schneider PM, Tidow N, Brandt B, Buerger H, Bulk E, Thomas M, Berdel WE, Serve H, Muller-Tidow C (2003) MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 22:8031–8041
Leung A, Natarajan R (2014) Noncoding RNAs in vascular disease. Curr Opin Cardiol 29:199–206
Li HJ, Jiang L, Yu ZB, Han SP, Liu XH (2016) Bioinformatic and expression analysis of ventricular septal defect-associated long non-coding RNA TUC40. Zhongguo Yi Xue Ke Xue Yuan Xue Bao 38:1–8
Li H, Jiang L, Yu Z, Han S, Liu X, Li M, Zhu C, Qiao L, Huang L (2017) The role of a novel long noncoding RNA TUC40- in cardiomyocyte induction and maturation in P19 cells. Am J Med Sci 354:608–616
Marino BS, Lipkin PH, Newburger JW, Peacock G, Gerdes M, Gaynor JW, Mussatto KA, Uzark K, Goldberg CS, Johnson WH Jr, Li J, Smith SE, Bellinger DC, Mahle WT (2012) American Heart Association Congenital Heart Defects Committee CoCDitYCoCN and Stroke C. Neurodevelopmental outcomes in children with congenital heart disease: evaluation and management: a scientific statement from the American Heart Association. Circulation 126:1143–1172
Martinou JC, Youle RJ (2011) Mitochondria in apoptosis: Bcl-2 family members and mitochondrial dynamics. Dev Cell 21:92–101
Matsusaka T, Ichikawa I (1997) Biological functions of angiotensin and its receptors. Annu Rev Physiol 59:395–412
Morlando M, Ballarino M, Fatica A, Bozzoni I (2014) The role of long noncoding RNAs in the epigenetic control of gene expression. ChemMedChem 9:505–510
Paffett-Lugassy N, Singh R, Nevis KR, Guner-Ataman B, O'Loughlin E, Jahangiri L, Harvey RP, Burns CG, Burns CE (2013) Heart field origin of great vessel precursors relies on nkx2.5-mediated vasculogenesis. Nat Cell Biol 15:1362–1369
Prall OW, Menon MK, Solloway MJ, Watanabe Y, Zaffran S, Bajolle F, Biben C, McBride JJ, Robertson BR, Chaulet H, Stennard FA, Wise N, Schaft D, Wolstein O, Furtado MB, Shiratori H, Chien KR, Hamada H, Black BL, Saga Y, Robertson EJ, Buckingham ME, Harvey RP (2007) An Nkx2-5/Bmp2/Smad1 negative feedback loop controls heart progenitor specification and proliferation. Cell 128:947–959
Qin P, Haberbusch JM, Soprano KJ, Soprano DR (2004) Retinoic acid regulates the expression of PBX1, PBX2, and PBX3 in P19 cells both transcriptionally and post-translationally. J Cell Biochem 92:147–163
Qin DN, Qian L, Hu DL, Yu ZB, Han SP, Zhu C, Wang X, Hu X (2013) Effects of miR-19b overexpression on proliferation, differentiation, apoptosis and Wnt/beta-catenin signaling pathway in P19 cell model of cardiac differentiation in vitro. Cell Biochem Biophys 66:709–722
Shen Y, Song G, Liu Y, Zhou L, Liu H, Kong X, Sheng Y, Cao K, Qian L (2013) Silencing of FABP3 inhibits proliferation and promotes apoptosis in embryonic carcinoma cells. Cell Biochem Biophys 66:139–146
Song G, Shen Y, Zhu J, Liu H, Liu M, Shen YQ, Zhu S, Kong X, Yu Z, Qian L (2013) Integrated analysis of dysregulated lncRNA expression in fetal cardiac tissues with ventricular septal defect. PLoS One 8:e77492
Srivastava D (2006) Making or breaking the heart: from lineage determination to morphogenesis. Cell 126:1037–1048
Stankunas K, Shang C, Twu KY, Kao SC, Jenkins NA, Copeland NG, Sanyal M, Selleri L, Cleary ML, Chang CP (2008) Pbx/Meis deficiencies demonstrate multigenetic origins of congenital heart disease. Circ Res 103:702–709
Suzuki K, Uchida E, Schober KE, Niehaus A, Rings MD, Lakritz J (2012) Cardiac troponin I in calves with congenital heart disease. J Vet Intern Med 26:1056–1060
Ugarte-Uribe B, Garcia-Saez AJ (2014) Membranes in motion: mitochondrial dynamics and their role in apoptosis. Biol Chem 395:297–311
Wu JX, Adamson ED (1993) Inhibition of differentiation in P19 embryonal carcinoma cells by the expression of vectors encoding truncated or antisense EGF receptor. Dev Biol 159:208–222
Xu H, Morishima M, Wylie JN, Schwartz RJ, Bruneau BG, Lindsay EA, Baldini A (2004) Tbx1 has a dual role in the morphogenesis of the cardiac outflow tract. Development 131:3217–3227
Yang J, Ko SJ, Kim BS, Kim HS, Park S, Hong D, Hong SW, Choi JH, Park CY, Choi SC, Hong SJ, Lim DS (2009) Enhanced cardiomyogenic differentiation of P19 embryonal carcinoma stem cells. Korean Circ J 39:198–204
Acknowledgments
This study was supported by grants from the National Natural Science Foundation of China (grant no. 81470376), the National Natural Science Foundation of Jiangsu Province of China (grant no. BK20141077), Changzhou Municipal Science and Technology Bureau guidance project (grant no. 2016361), and Changzhou Science and Technique Development Foundation (grant no. 20170397).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Additional information
Editor: Tetsuji Okamoto
Rights and permissions
About this article
Cite this article
Wu, R., Xue, P., Wan, Y. et al. LncRNA-uc.40 silence promotes P19 embryonic cells differentiation to cardiomyocyte via the PBX1 gene. In Vitro Cell.Dev.Biol.-Animal 54, 600–609 (2018). https://doi.org/10.1007/s11626-018-0284-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11626-018-0284-0