Abstract
Purpose
To evaluate the feasibility of automated quantitative analysis with a three-dimensional (3D) computer-aided system (i.e., Gaussian histogram normalized correlation, GHNC) of computed tomography (CT) images from different scanners.
Materials and methods
Each institution’s review board approved the research protocol. Informed patient consent was not required. The participants in this multicenter prospective study were 80 patients (65 men, 15 women) with idiopathic pulmonary fibrosis. Their mean age was 70.6 years. Computed tomography (CT) images were obtained by four different scanners set at different exposures. We measured the extent of fibrosis using GHNC, and used Pearson’s correlation analysis, Bland–Altman plots, and kappa analysis to directly compare the GHNC results with manual scoring by radiologists. Multiple linear regression analysis was performed to determine the association between the CT data and forced vital capacity (FVC).
Results
For each scanner, the extent of fibrosis as determined by GHNC was significantly correlated with the radiologists’ score. In multivariate analysis, the extent of fibrosis as determined by GHNC was significantly correlated with FVC (p < 0.001). There was no significant difference between the results obtained using different CT scanners.
Conclusion
Gaussian histogram normalized correlation was feasible, irrespective of the type of CT scanner used.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Idiopathic pulmonary fibrosis (IPF) is a devastating lung disease that leads to breathlessness and ultimately respiratory failure and death [1]. Some new drugs (e.g., pirfenidone and nintedanib) have recently been developed that could significantly reduce the rate of disease progression [2–6]. Pirfenidone could improve progression-free survival [5]. The development of drugs for pulmonary fibrosis has increased the demand for biomarkers that can be used to evaluate disease progression and the effect of these drugs. In previous randomized trials [5, 6], a decline in the forced vital capacity (FVC) was used as one measure. However, the FVC is not decreased early in the disease, and the FVC is affected by emphysema. Pulmonary function tests (PFTs) are not easy to perform for patients with oxygen therapy and a history of pneumothorax. Thus, accurate surrogate markers are needed to manage patients with IPF.
Computed tomography (CT) is an essential imaging modality for detecting and diagnosing IPF [1, 7]. Many studies have demonstrated a high correlation between the severity of CT findings, pulmonary function, and prognosis [8–11]. Thus, if simple and reliable methods need to be established, quantitative analysis of CT is a potential candidate. Visual inspection and scoring are conventional methods for quantitatively evaluating lung abnormalities on CT [11]. Visual scoring, however, is a difficult task for less experienced radiologists. In addition, visual assessment by expert chest radiologists is associated with substantial intra- and inter-reader variability [12, 13].
To circumvent these shortfalls, automated and objective tools using computer-assisted diagnosis (CAD) have been proposed [10, 14–22]. From a technological standpoint, CAD for diffuse lung disease is commonly viewed as a texture analysis problem [23]. An artificial neural network has been applied to solve this problem [17, 19]. However, an artificial neural network requires a larger number of training datasets (e.g., more than one hundred datasets) [24]. This requirement is a disadvantage for relatively rare diseases such as IPF. Some systems based on artificial neural networks cannot be applied to different types of scanners from those used to train the network [22].
In this study, we examined the clinical utility of the Gaussian histogram normalized correlation (GHNC) system. Gaussian histogram normalized correlation requires a small number of datasets—approximately 20 datasets—for use as predesigned samples. The original two-dimensional (2D) version was reported by Asakura et al. [25], and the three-dimensional (3D) version was reported by Iwao et al. [26]. The feasibility and utility of the 2D version was demonstrated in a single-center study [27, 28]. We conducted a multicenter, multivendor prospective study to examine the 3D version of GHNC for analyzing images obtained by different scanners under different exposure conditions using a determined sample. We also compared the GHNC results with conventional radiologists’ scores and with FVC values. We evaluated the feasibility of using GHNC for different scanners.
Materials and methods
Patients
This research protocol was approved by each institution’s review board. Informed patient consent was not required. All IPF patients who underwent CT examination at each institution between January 2011 and April 2012 were potential candidates for this study. We enrolled 80 consecutive patients (20 patients each for four scanners) who underwent PFTs within 3 months of a CT examination. In this study, IPF was diagnosed based on the criteria developed by the American Thoracic Society [1]. Clinicians at each institution confirmed the diagnosis of IPF for each patient. The diagnosis was based on a review of the patient’s clinical history, occupational and environmental exposure, results of PFTs, thin-section CT images of the lungs, and, when available, transbronchial or surgical lung biopsy. The major exclusion criteria were patients with a history of thoracic surgery and patients with acute exacerbation, pneumothorax, or active respiratory infection. We also excluded patients who could not hold their breath. The inclusion and exclusion criteria were same regardless of the scanner used.
The PFTs were performed using three systems: CHESTAC-8800 (Chest M.I. Co., Tokyo, Japan), CHESTAC-33 (Chest M.I. Co.), and Fudac-77 (Fukuda Denshi, Tokyo, Japan). The forced vital capacity and forced expiratory volume in 1 s (FEV1) were obtained. Total lung capacity (TLC) and diffusing capacity were also measured in some patients using common standard measurement techniques [29]. The results were expressed as the percentage of predicted performance using standard values [29].
CT images
Thin-section CT images were obtained by four types of scanners during inspiration with the patient positioned supine in the scanner, and the routine exposure conditions of each institution were used. Table 1 lists the scanners and exposure conditions employed. The four types of scanners were: (CT-1) Light Speed, a 16-row multidetector CT (MDCT) (General Electric Medical Systems, Milwaukee, WI, USA); (CT-2) Brilliance iCT, a 128-row MDCT (Philips Healthcare, Best, the Netherlands); (CT-3) Aquilion-16, a 16-row MDCT (Toshiba, Tokyo, Japan); and (CT-4) Aquilion-64, a 64-row MDCT (Toshiba). The voltage was 120 kVp for all scanners. The current and range of the CT dose index (CTDI) of each scanner were as follows: variable milliamps × seconds (mAs) below 220 mAs and 10.97–14.8 mGy for CT-1; 200 mAs and 9.0 mGy for CT-2; 250 mAs and 16.4 mGy for CT-3, and variable mAs below 300 mAs (12.6–19.6 mGy) for CT-4. The slice thickness was 1.25 mm for CT-1, 1 mm for CT-2, and 0.5 mm for CT-3 and CT-4. For image reconstruction, filtered backprojection was used for the CT-1, CT-3, and CT-4 scanners, and iterative reconstruction was used for the CT-2 scanner. Each institution was requested to reconstruct images with the “soft” reconstruction kernel for GHNC analysis because in 2011 GHNC could not analyze the kernel for high-resolution CT (HRCT) (e.g., bone algorithm of the GE system) in which the distribution of pixel values within the local region changes [30].
CAD analysis
All CAD analyses were performed by one author. Before the segmentation process, we corrected the CT values by the mean attenuation value (−1000) in the tracheal gas [30]. The lung was extracted from the 3D CT dataset using the algorithm reported by Iwao et al. [26]. We extracted the lung using the optional threshold, adaptive density-based morphology, and minimal manual intervention. We segmented the bronchial tree and pulmonary vessels with the failure-recovery algorithm. After extracting the lung, each lesion was segmented using GHNC [25].
In brief, GHNC divides the pixels of the lung into five categories based on the predesigned samples using CT attenuation values and local histograms of them. The five categories were as follows: “normal,” “emphysema,” “ground glass opacity” (“GGO”), “consolidation,” and “fibrosis.” Fibrosis was also subdivided into “reticulation” and “honeycomb” (see Fig. 1 and the “Appendix”). For the analysis, we used 121 samples: 50 normal, 15 emphysema, 15 GGO, 5 consolidation, 21 reticulation, and 15 honeycomb samples. These samples were obtained from the CT datasets of 14 patients and seven normal individuals who were not patients included in this multicenter study. The volume of the diseased lung and the total CT lung volume (CTLV) were computed automatically by the GHNC system.
Scheme of the Gaussian histogram normalized correlation (GHNC) system. Samples of typical lesions are arranged to obtain histograms for the original images and the differential CT images (a differential image is an image with defined figure edges). We prepared Gaussian histograms of all pixels in the original images and the differential CT images. The normalized correlations between the Gaussian histogram of each pixel and the histograms of samples were analyzed. All pixels were divided into lesions and later computed into color images (i.e., GHNC images). The algorithm for the GHNC is summarized in the “Appendix”
Scoring of HRCT lesions by radiologists
The CT images were reviewed separately by two radiologists who were blinded to the clinical information and the GHNC results. Both radiologists were board-certified diagnostic radiologists who were majoring in the chest. Each radiologist had 15 years of experience.
The lungs were divided into eight zones (“upper,” “middle,” “lower,” and “bottom” on both sides). Each zone was evaluated separately. The upper lung zone was the area of the lung at the aortic arch; the middle lung zone was at the level of the tracheal carina; the lower lung zone was the area of the lung between the middle and bottom zones, and the bottom lung zone was the area of the lung 1 cm below the dome of the diaphragm.
The observers evaluated the extent of all radiological abnormalities of emphysema, GGO, consolidation, and fibrosis (i.e., reticulation and honeycombing). The definition of each lesion was based on the definitions provided in previous studies [9, 31]. When abnormal findings were present, the extent of lung involvement was evaluated visually and independently for each of the eight lung zones. The score was based on the percentage of the lung parenchyma that showed evidence of abnormality and was estimated to the nearest 5 % of the parenchyma. To calculate the score for the patient, we used the average of the scores for the eight zones. The radiologists evaluated the CT data once. We used the average of the two radiologists’ scores as the consensus result.
Statistical analysis
First, using Pearson’s correlation analysis, we compared the total extent of each lesion (based on the radiologists’ scores) and the volume (based on the GHNC results). Second, interobserver differences (i.e., between the two radiologists and between the radiologists and GHNC) were evaluated by weighted kappa analysis. The extent of each lesion was classified into 11 categories: from 0 to less than 5 %; 5 % or greater to less than 10 %; 10 % or greater to less than 20 %, and each subsequent 10 % in steps up to 100 %. Interobserver agreement was classified as slight (κ = 0.00–0.20), fair (κ = 0.21–0.40), moderate (κ = 0.41–0.60), substantial (κ = 0.61–0.80), or nearly perfect (κ = 0.81–1.00). Third, the limits of agreement between the radiologists’ scores and the GHNC results were also analyzed by Bland–Altman analysis. Fourth, we evaluated the relationship between the extent of fibrosis on CT and the FVC percentage. To investigate the influence of the type of CT scanner used, multiple linear regression analysis was performed. All statistical analyses were performed by SPSS v.20 software (SPSS Inc., Chicago, IL, USA). A p value of less than 0.05 was considered significant in all statistical analyses.
Results
Table 2 lists the patients’ characteristics. The patients comprised 65 men and 15 women with a mean age ± standard deviation of 70.6 ± 6.3 years and a median age of 71 years (range 48–83 years). There was no significant difference between the scanners with regard to patients’ age, sex, and PFTs; however, there was a significant difference in smoking history (p = 0.031).
The GHNC system successfully analyzed all CT images (Fig. 2). A period of 20–30 min was required to complete the analysis of a single patient: 10–15 min for the lung segmentation process and 10–15 min for the GHNC segmentation process. The processing time increased in accordance with the number of images. In addition, the lung segmentation process required longer when manual correction was needed.
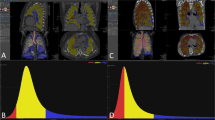
The Gaussian histogram normalized correlation (GHNC) results for the four CT scanners. a, d, g, j Coronal reconstruction images. b, e, h, k Coronal GHNC images. c, f, i, l Volume-rendered three-dimensional (3D) images. a, b, c The CT-1 scanner. d, e, f The CT-2 scanner. g, h, i The CT-3 scanner. j, k, l The CT-4 scanner. Pink normal lung tissue, dark blue emphysema, light green ground glass opacity, yellow and light blue fibrosis, green the bronchi, light orange the blood vessels
Table 3 shows the mean radiologists’ scores, the extent of each lesion based on GHNC, and the Pearson’s correlation coefficients between the mean radiologists’ scores and the GHNC results. There was no significant difference between the scanners in the extent of each lesion evaluated by the radiologists. The GHNC results were significantly correlated with the mean radiologists’ scores (p < 0.001) with a correlation coefficient of 0.895 for normal, 0.933 for emphysema, 0.751 for GGO, 0.388 for consolidation, and 0.884 for fibrosis in the 80 patients. Consolidation in CT-3 did not show any significant correlation.
Table 4 shows the results of a comparison of the scores of the two radiologists and a comparison of the consensus results from the radiologists versus the GHNC results, performed using weighted kappa analysis. Across all lesion types (except for consolidation), kappa ranged from 0.51 to 0.70 when the two radiologists’ scores were compared, and from 0.48 to 0.65 when the consensus results from the radiologists were compared to the results of GHNC. Consolidation was small (0.17 % of the mean) among the patients: 64 of 80 patients had no consolidation based on the radiologists’ consensus scores. Thus, the kappa values for consolidation when the consensus results from the radiologists were compared to the results of GHNC were not calculated because nearly all of the assessment results fell within the 0–5 % category.
Figure 3 show Bland–Altman plots of mean radiologists’ score versus the difference between the GHNC result and the mean radiologists’ score. The mean difference between the GHNC result and the mean radiologists’ score and the limits of agreement between them (in parentheses) were as follows: −2.3 % (−16.2 to −11.6 %) for normal; 0.1 % (−5.1 to 5.2 %) for emphysema; 2.6 % (−6.2 to 11.4 %) for GGO; 0.4 % (−1.4 to 2.2 %) for consolidation; and −0.8 % (−11.1 to 9.6 %) for fibrosis (see Table 5). There was no significant difference between the results obtained with different scanners.
Bland–Altman scatter plots showing the correlation between the GHNC analysis result and the mean radiologists’ score based on the images from 80 patients. The vertical axis shows the difference between the GHNC result and the mean radiologists’ score (i.e., the GHNC result minus the mean radiologists’ score). The solid line shows the mean of this difference. The dotted lines indicate the mean ± (2 × standard deviation). The scanners used were: CT-1 (unfilled circles), Light Speed Ultra 16 (General Electric Medical Systems); CT-2 (filled circles), Brilliance iCT (Philips Healthcare); CT-3 (unfilled triangles), Acquilion 16 (Toshiba); CT-4 (crosses), Acquilion 64 (Toshiba). GHNC Gaussian histogram normalized correlation
Figure 4 and Table 6 show the correlations of the percent of predicted FVC (FVCpred %) with the GHNC result and the FVCpred % with the mean radiologist’s score. The extent of each lesion (except for emphysema) and the FVCpred % showed a good correlation. The correlation coefficients were −0.524 for fibrosis based on the mean radiologists’ scores (p < 0.001) and −0.671 for fibrosis based on the GHNC results (p < 0.001).
Table 7 shows the results of using multiple linear regression analysis to explore the percent of FVC. We included CT scanner, patient age, sex, and smoking history in the analysis. Due to the multicollinearity problem, we also included the extent of fibrosis among the lesions based on either the mean radiologists’ score or the GHNC result. Data obtained using scanners CT-1, CT-2, and CT-3 were each treated as independent categorical data, and data obtained using scanner CT-4 were considered reference data. The extent of fibrosis (by percentage) based on the GHNC result was found to be a significant factor (p < 0.001). The extent of fibrosis based on the mean radiologists’ score was marginally significant (p = 0.050). The was no significant difference between the results obtained using different CT scanners.
Discussion
This multicenter study showed that the GHNC system successfully analyzed multivendor CT images that were obtained under different exposure conditions. Based on Pearson’s correlation analysis and kappa analysis, the GHNC result was strongly correlated with the mean radiologists’ score. The mean difference between these two measures of the extent of fibrosis was less than 3 % in the Bland–Altman plots. There was no significant difference between the results obtained using the different scanners, indicating that it is feasible to use GHNC with different scanners. Furthermore, multiple linear regression analysis showed that the extent of fibrosis on CT, based on the GHNC result, was significantly correlated with the FVC. Again, the choice of CT scanner did not significantly influence the results. The FVC is a popular biomarker that is used to measure the progression of IPF [5]. Thus, our results show the possibility of using GHNC results as a biomarker of disease severity and the effect of treatment in patients with IPF.
To the best of our knowledge, research into computer analysis of CT images of diffuse lung disease began in the 1990s [17]. The segmentation problem of diffuse lung disease is a common texture analysis problem. Previous studies have been characterized by viewpoints such as (i) 2D or 3D images, (ii) the type of regions of interest used for assessment, (iii) the texture features used for analysis, and (iv) the statistical methods used to identify these features [32]. In studies performed in the 1990s, the lungs were separated into small blocks, and each block was analyzed on limited 2D images [17]. Pixel-based analysis of 3D images such as GHNC has commonly been used in recent studies [16]. The texture features and statistical methods employed vary among the studies. First, an artificial neural network is applied to solve the segmentation problem. Uppaluri et al. [17] used the adaptive multiple feature method with 22 independent texture features to classify the tissue pattern. Rosas et al. [19] analyzed CT images with 25 texture features by using a smart vector machine. These artificial neural network methods involve relatively large computational costs. Thus, Zavaletta et al. [18] proposed a method that uses a histogram of CT attenuation values. They clustered the histogram into K clusters, and defined the histogram signature using the centroid and weight of the cluster. They established the canonical signature by using voxels of interest (VOIs), and expert radiologists selected VOIs that included 70 % or more typical lesions. They then calculated the similarity between the target voxels and the 809 sample VOIs using the Earth Mover’s Distance [18]. Maldonado et al. [10] analyzed the CT data of patients with IPF by using computer-aided lung informatics for pathology evaluation and rating (CALIPER), based on Zavaletta’s method; they found that the results of CALIPER correlated well with the PFT results and with the patient’s prognosis.
In the current study, we used GHNC, which is a histogram-based method. The differences between this method and Zavaletta’s method are that (i) GHNC estimates the similarity of the histogram using normalized correlation, and (ii) GHNC uses a histogram of the original images and the differential images. Usual interstitial pneumonia (UIP) pathologically appears as a patchwork pattern: severe fibrosis juxtaposed next to normal lung. Honeycomb lung includes air in the surrounding fibrosis [33]. Thus, UIP pattern fibrosis yields larger standard deviations, higher pixel values on differential images, and has a broader histogram than seen for other lesions (i.e., normal, emphysema, GGO) (Fig. 1). Thus, GHNC can easily detect the similarity, in terms of fibrosis, between the samples and target pixels. We believe that this is one reason for the smaller number of samples (121 in total) in our study than in Zavaletta’s (809 in total). The tested GHNC system requires a relatively long analysis time. We believe that this is primarily due to our computer’s performance (it was a built-to-order personal computer that utilized an Intel Core i7 CPU 950, 3.07 GHz, and Microsoft Windows 7 software, 64 bit. Further improvement of this system is necessary.
The present study has several limitations. First, we did not test all venders. The radiation exposure conditions varied depending on the scanner used, which may also be a limitation of our study. However, our results showed that GHNC could be used to analyze images obtained under various exposure conditions in a clinical setting. Vendor-independent feasibility is a clinically essential prerequisite for computer analysis.
Another limitation is that we used only “soft” reconstruction kernels in this study. In 2011, the feasibility of using GHNC for different kernels was not determined—especially for HRCT, in which the standard deviation of attenuation is larger than for an image with a soft kernel. Iwasawa et al. recently reported that GHNC can be used to analyze HRCT after altering images with an appropriate Gaussian filter (personal communication). Further study will be needed to test the feasibility of GHNC for multivendor HRCT.
In this study, we did not separate honeycombing and reticulation. A previous GHNC study [27] indicated that the kappa value between the radiologists’ scores and the GHNC results for honeycombing was poor. Watadani et al. [13] reported that the kappa value between expert radiologists’ evaluations of honeycombing was not high, and that recognition of honeycombing varied between expert radiologists. Thus, we grouped honeycombing and reticulation together as fibrosis, and then assessed it.
The GHNC results were highly correlated with the expert radiologists’ scores. The Bland–Altman plot showed that GHNC miscategorized 2.3 % of normal lungs. One reason for this is misregistration of the normal peripheral bronchi and vessels in the lung as lesions (especially as fibrosis). The false-positive rate of GHNC was reported to be 1.3 % in a previous study [34] which used only 10 normal individuals. The false-positive rate should be analyzed more precisely in a larger population.
The Bland–Altman plot showed that 2.6 % of the lungs with GGO according to CAD were miscategorized. The weighted kappa value was moderate. Artifacts with high attenuation values such as cardiac motion artifact were also misregistered as GGO and fibrosis [21]. In addition, GHNC mis-segmented the normal lung as GGO in some patients with advanced IPF. This was probably due to higher attenuation values of the normal lung caused by the redistribution of pulmonary blood flow from the severely affected portions to the portions that maintained relatively normal structures [35]. Another reason for the false GGO findings was an inherent flaw related to the inability of GHNC to account for traction bronchiectasis. Most radiologists classify GGO with traction bronchiectasis as fibrosis, whereas GHNC would consider such a region to have GGO, based on the attenuation values.
In this study, there was a poor correlation between the radiologists’ scores and the GHNC results in identifying consolidation. We could not calculate the kappa value for consolidation because the volume of consolidation was small in most patients. Consolidation is a very important finding suggesting acute exacerbation or complication of the infection in patients with IPF. Consolidation is common in secondary UIP, such as in vasculitis. Consolidation is an uncommon finding in people with stable IPF. Further study will be needed to evaluate the feasibility of evaluating consolidation by GHNC.
In conclusion, the GHNC system analyzed 3D CT images obtained by different scanners. The results were in moderate to good agreement with the radiologists’ scores. The volume of fibrosis measured by GHNC was highly correlated with the FVC, which is a representative marker of IPF.
A previous study showed that the 2D version of GHNC could detect the effect of pirfenidone [28]. We believe that the 3D version of GHNC can be feasibly employed in research (e.g., clinical trials that focus on pulmonary fibrosis) and in the clinical setting of IPF.
References
Raghu G, Collard HR, Egan JJ, Martinez FJ, Behr J, Brown KK, ATS/ERS/JRS/ALAT Committee on Idiopathic Pulmonary Fibrosis, et al. An official ATS/ERS/JRS/ALAT statement: idiopathic pulmonary fibrosis: evidence-based guidelines for diagnosis and management. Am J Respir Crit Care Med. 2011;183:788–824.
Azuma A, Nukiwa T, Tsuboi E, Suga M, Abe S, Nakata K, et al. Double-blind, placebo-controlled trial of pirfenidone in patients with idiopathic pulmonary fibrosis. Am J Respir Crit Care Med. 2005;171:1040–7.
Taniguchi H, Ebina M, Kondoh Y, Ogura T, Azuma A, Suga M, Pirfenidone Clinical Study Group in Japan, et al. Pirfenidone in idiopathic pulmonary fibrosis. Eur Respir J. 2010;35:821–9.
Richeldi L, du Bois RM, Raghu G, Azuma A, Brown KK, Costabe U, et al. INPULSIS Trial Investigators. Efficacy and safety of nintedanib in idiopathic pulmonary fibrosis. N Engl J Med. 2014;370:2071–82.
King TE Jr, Bradford WZ, Castro-Bernardini S, Fagan EA, Glaspole I, Glassberg MK, ASCEND Study Group, et al. A phase 3 trial of pirfenidone in patients with idiopathic pulmonary fibrosis. N Engl J Med. 2014;370:2083–92.
Noble PW, Albera C, Bradford WZ, Costabel U, Glassberg MK, Kardatzke D, CAPACITY Study Group, et al. Pirfenidone in patients with idiopathic pulmonary fibrosis (CAPACITY): two randomised trials. Lancet. 2011;377:1760–9.
Lynch DA, Travis WD, Müller NL, Galvin JR, Hansell DM, Grenier PA, et al. Idiopathic interstitial pneumonias: CT features. Radiology. 2005;236:10–21.
Fell CD, Martinez FJ, Liu LX, Murray S, Han MK, Kazerooni EA, et al. Clinical predictors of a diagnosis of idiopathic pulmonary fibrosis. Am J Respir Crit Care Med. 2010;181:832–7.
Sumikawa H, Johkoh T, Colby TV, Ichikado K, Suga M, Taniguchi H, et al. Computed tomography findings in pathological usual interstitial pneumonia: relationship to survival. Am J Respir Crit Care Med. 2008;177:433–9.
Maldonado F, Moua T, Rajagopalan S, Karwoski RA, Raghunath S, Decker PA, et al. Automated quantification of radiological patterns predicts survival in idiopathic pulmonary fibrosis. Eur Respir J. 2014;43:204–12.
Ley B, Elicker BM, Hartman TE, Ryerson CJ, Vittinghoff E, Ryu JH, et al. Idiopathic pulmonary fibrosis: CT and risk of death. Radiology. 2014;273:570–9.
Barr RG, Berkowitz EA, Bigazzi F, Bode F, Bon J, Bowler RP, COPDGene CT Workshop Group, et al. A combined pulmonary–radiology workshop for visual evaluation of COPD: study design, chest CT findings and concordance with quantitative evaluation. COPD. 2012;9:151–9.
Watadani T, Sakai F, Johkoh T, Noma S, Akira M, Fujimoto K, et al. Interobserver variability in the CT assessment of honeycombing in the lungs. Radiology. 2013;266:936–44.
Park SO, Seo JB, Kim N, Park SH, Lee YK, Park BW, et al. Feasibility of automated quantification of regional disease patterns depicted on high-resolution computed tomography in patients with various diffuse lung diseases. Korean J Radiol. 2009;10:455–63.
Hoffman EA, Reinhardt JM, Sonka M, Simon BA, Guo J, Saba O, et al. Characterization of the interstitial lung diseases via density-based and texture-based analysis of computed tomography images of lung structure and function. Acad Radiol. 2003;10:1104–18.
Korfiatis PD, Karahaliou AN, Kazantzi AD, Kalogeropoulou C, Costaridou LI. Texture-based identification and characterization of interstitial pneumonia patterns in lung multidetector CT. IEEE Trans Inf Technol Biomed. 2010;14:675–80.
Uppaluri R, Hoffman EA, Sonka M, Hunninghake GW, McLennan G. Interstitial lung disease: a quantitative study using the adaptive multiple feature method. Am J Respir Crit Care Med. 1999;159:519–25.
Zavaletta VA, Bartholmai BJ, Robb RA. High resolution multidetector CT-aided tissue analysis and quantification of lung fibrosis. Acad Radiol. 2007;14:772–87.
Rosas IO, Yao J, Avila NA, Chow CK, Gahl WA, Gochuico BR. Automated quantification of high-resolution CT scan findings in individuals at risk for pulmonary fibrosis. Chest. 2011;140:1590–7.
Wang J, Li F, Li Q. Automated segmentation of lungs with severe interstitial lung disease in CT. Med Phys. 2009;36:4592–9.
Kauczor HU, Heitmann K, Heussel CP, Marwede D, Uthmann T, Thelen M. Automatic detection and quantification of ground-glass opacities on high-resolution CT using multiple neural networks: comparison with a density mask. AJR Am J Roentgenol. 2000;175:1329–34.
Yoon RG, Seo JB, Kim N, Lee HJ, Lee SM, Lee YK, et al. Quantitative assessment of change in regional disease patterns on serial HRCT of fibrotic interstitial pneumonia with texture-based automated quantification system. Eur Radiol. 2013;23:692–701.
Sluimer IC, Prokop M, Hartmann I, van Ginneken B. Automated classification of hyperlucency, fibrosis, ground glass, solid, and focal lesions in high-resolution CT of the lung. Med Phys. 2006;33:2610–20.
Raghunath S, Rajagopalan S, Karwoski RA, Maldonado F, Peikert T, Moua T, et al. Quantitative stratification of diffuse parenchymal lung diseases. PLoS One. 2014;9:e93229.
Asakura A, Gotoh T, Iwasawa T, Saito K, Akasaka H. Classification system of the CT images with nonspecific interstitial pneumonia. J Inst Image Electron Eng Jpn. 2004;33:180–8.
Iwao Y, Gotoh T, Kagei S, Iwasawa T, Tsuzuki Mde S. Integrated lung field segmentation of injured region with anatomical structure analysis by failure–recovery algorithm from chest CT images. Biomed Signal Process Control. 2014;12:28–38.
Iwasawa T, Asakura A, Sakai F, Kanauchi T, Gotoh T, Ogura T, et al. Assessment of prognosis of patients with idiopathic pulmonary fibrosis by computer-aided analysis of CT images. J Thorac Imaging. 2009;24:216–22.
Iwasawa T, Ogura T, Sakai F, Kanauchi T, Komagata T, Baba T, et al. CT analysis of the effect of pirfenidone in patients with idiopathic pulmonary fibrosis. Eur J Radiol. 2014;83:32–8.
Japanese Respiratory Society. Guideline for pulmonary function test. 1st ed. Tokyo: Medical View; 2004.
Best AC, Lynch AM, Bozic CM, Miller D, Grunwald GK, Lynch DA. Quantitative CT indexes in idiopathic pulmonary fibrosis: relationship with physiologic impairment. Radiology. 2003;228:407–14.
Lynch DA, Godwin JD, Safrin S, Starko KM, Hormel P, Brown KK, et al. Idiopathic Pulmonary Fibrosis Study Group. High-resolution computed tomography in idiopathic pulmonary fibrosis: diagnosis and prognosis. Am J Respir Crit Care Med. 2005;172:488–93.
Sluimer I, Schilham A, Prokop M, van Ginneken B. Computer analysis of computed tomography scans of the lung: a survey. IEEE Trans Med Imaging. 2006;25:385–405.
Katzenstein AL. Idiopathic interstitial pneumonia. In: Katzenstein AL, editor. Katzenstein and Askin’s surgical pathology of non-neoplastic lung disease. 4th ed. Philadelphia: Saunders Elsevier; 2006. p. 51–84.
Iwasawa T, Iwao Y, Goto T, Asakura A, Komagata T, Ogura T, et al. Computer-aided segmentation system for 3D chest CT. Rinsho Hoshasen. 2012;57:41–7 (in Japanese).
Yamaguchi K, Mori M, Kawai A, Takasugi T, Oyamada Y, Koda E. Inhomogeneities of ventilation and the diffusing capacity to perfusion in various chronic lung diseases. Am J Respir Crit Care Med. 1997;156:86–93.
Acknowledgments
This study was approved by each institutional review board (IRB number, Kanagawa Cardiovascular & Respiratory Center 23-1, Saitama Cardiovascular and Respiratory Center 2011004).
This study was supported by the Kanagawa Cancer Research Fund, in part supported by a joint project from JSPS/CAPES under the Japan–Brazil Research Cooperative Program and Grants-in-Aid for Scientific Research (24500539), and in part supported by JSPS KAKENHI grant number 15K09901.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Appendix
Appendix
The Gaussian histogram normalized correlation (GHNC) method classifies pixels in a target image into different categories. In the GHNC method, a set of Gaussian histograms of CT attenuation values of the original image and the differential image are extracted from samples of each category in advance. The normalized correlations between the Gaussian histograms in the surrounding local 50-pixel area of a pixel and those of the categories are then compared for classification into the predefined categories. To avoid the effects of random noise, pixel attenuation is assumed to have random noise with a Gaussian distribution. Gaussian convolution filtering is applied to the extracted histograms from the local area and the samples in the categories.
The Gaussian histogram of the predesigned sample is obtained from the following formula:
in which α is the Hounsfield units of each pixel and N α is the number of pixels in the predesigned sample area D α. The variable σ is the standard deviation of the Gaussian random noise.
The Gaussian histogram of the target area D β is given by
in which β and N β also denote the Hounsfield units and the number of pixels in the target area D β, respectively.
The normalized correlation between α g(x) and β g(x) is given by the formula
In our method, the normalized correlations are calculated in the local area of each target pixel for the original and differential images. The product of both correlations is then used as the similarity of the target and the predesigned sample pattern in each category to classify the target pixels into the predesigned categories.
About this article
Cite this article
Iwasawa, T., Kanauchi, T., Hoshi, T. et al. Multicenter study of quantitative computed tomography analysis using a computer-aided three-dimensional system in patients with idiopathic pulmonary fibrosis. Jpn J Radiol 34, 16–27 (2016). https://doi.org/10.1007/s11604-015-0496-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11604-015-0496-0