Abstract
Phaeosphaeriaceae is a species-rich family in the order Pleosporales, encompassing species with diverse lifestyles viz., endophytic, epiphytic, lichenicolous, phytopathogenic, saprobic and even human pathogenic. In a survey on biodiversity of fungal species associated with leaf spot diseases of herbaceous plants in Iran, a coelomycetous fungus was recovered from symptomatic leaves of Fallopia convolvulus. Morphologically, the fungal isolates resembled species in the genus Parastagonospora. Although the phylogenetic analysis based on combined LSU and ITS sequence data placed these isolates within the family Phaeosphaeriaceae, they clustered distinct from presently known genera in the family. The monotypic genus Parastagonosporella (Phaeosphaeriaceae) is therefore introduced, with Parastagonosporella fallopiae as type species. A detailed description is provided, with notes discussing allied genera in the family.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Phaeosphaeriaceae, which is characterized by coelomycetous asexual morphs, constitutes a large family in the order Pleosporales, which is the largest and most diverse order in the class Dothideomycetes (Phookamsak et al. 2014). The family was introduced by Barr (1979) and typified by Phaeosphaeria with Phaeosphaeria oryzae (Miyake 1909) as the type species.
The family Phaeosphaeriaceae has a cosmopolitan distribution, and many species in this family are important plant pathogens, infecting several major crops (Carson 2005; Stukenbrock et al. 2006; Quaedvlieg et al. 2013), while others may be saprobes (Shoemaker 1984; Schoch et al. 2006; Zhang et al. 2012; Hyde et al. 2013; Quaedvlieg et al. 2013), endophytes (Wang et al. 2005; Sánchez Márquez et al. 2007) or even lichenicolous (Lawrey et al. 2012). Furthermore, some species have been reported to cause human infections (Ahmed et al. 2017).
The taxonomy of the family Phaeosphaeriaceae has been subject of several changes in recent years. Barr (1979) introduced the family with 15 genera, but during the past 10 years, various phylogenetic studies have revealed the Phaeosphaeriaceae to be heterogeneous, and recent studies have introduced several new genera or transferred some known genera to other families (Zhang et al. 2012; Hyde et al. 2013; Quaedvlieg et al. 2013; Phookamsak et al. 2014; Trakunyingcharoen et al. 2014; Crous et al. 2015; Ertz et al. 2015). In this regard, Phookamsak et al. (2014) revised the family and published a monograph of Phaeosphaeriaceae based on morphology and phylogeny and accepted 30 genera. More recently, further novel genera were placed in the Phaeosphaeriaceae (Phukhamsakda et al. 2015; Senanayake et al. 2015; Tennakoon et al. 2016; Tibpromma et al. 2016, b; Ahmed et al. 2017; Wanasinghe et al. 2018) based on morphological characteristics and phylogenetic analyses. Presently, more than 50 sexual and asexual genera are accepted in the family. These genera include the following: Acericola, Allophaeosphaeria, Amarenomyces, Ampelomyces, Bhatiellae, Camarosporioides, Chaetosphaeronema, Dactylidina, Dematiopleospora, Didymocyrtis, Embarria, Equiseticola, Galiicola, Hawksworthiana, Italica, Juncaceicola, Leptosphaeria, Leptospora, Loratospora, Melnikia, Muriphaeosphaeria, Neosetophoma, Neostagonospora, Neosulcatispora, Nodulosphaeria, Ophiobolopsis, Ophiobolus, Ophiosimulans, Ophiosphaerella, Paraophiobolus, Paraphoma, Parastagonospora, Phaeopoacea, Phaeosphaeria, Phaeosphaeriopsis, Poaceicola, Populocrescentia, Pseudoophiobolus, Pseudophaeosphaeria, Sclerostagonospora, Scolicosporium, Septoriella, Setomelanomma, Setophoma, Sulcispora, Tintelnotia, Vagicola, Vrystaatia, Wojnowicia, Wojnowiciella, Xenoseptoria and Yunnanensis (Quaedvlieg et al. 2013; Wijayawardene et al. 2014, 2016; Phookamsak et al. 2014, 2017; Ariyawansa et al. 2015, b; Ertz et al. 2015; Li et al. 2015; Phukhamsakda et al. 2015; Senanayake et al. 2015; Tennakoon et al. 2016; Tibpromma et al. 2016, b; Ahmed et al. 2017; Wanasinghe et al. 2018).
During a recent survey exploring the fungal species associated with leaf spot diseases of herbaceous plants in Iran, a coelomycetous fungus was recovered from Black Bindweed, Fallopia convolvulus. A subsequent phylogenetic study based on different gene regions revealed this fungus to represent an undescribed genus in Phaeosphaeriaceae. The aim of this study was thus to resolve the taxonomy of this genus and elucidate the phylogenetic relationship to allied genera in Phaeosphaeriaceae.
Materials and methods
Fungal isolates
During a field excursion in the Jowkandan region, Talesh country, Guilan province, Iran, symptomatic Black Bindweed (Fallopia convolvulus) leaves were collected and returned to the laboratory. Leaves were examined directly under a Nikon SMZ 1500 dissecting microscope to observe sporulation. Single conidial isolates were obtained in pure culture by direct transfer of spores onto plates containing 2% malt extract agar (MEA; Fluka, Hamburg, Germany) using a procedure previously described by Bakhshi et al. (2011).
Dried specimens were preserved in the Fungarium of the Iranian Research Institute of Plant Protection, Tehran, Iran (IRAN). Representative cultures were deposited in the culture collection of the Westerdijk Fungal Biodiversity Institute (CBS), Utrecht, The Netherlands, and the culture collection of Tabriz University (CCTU), Tabriz, Iran.
DNA extraction, amplification and sequencing
Fungal genomic DNA was extracted from fresh mycelium harvested from colonies grown on MEA for 10 days at 25 °C in the dark, according to the protocol described by Möller et al. (1992). Seven genomic loci were targeted for PCR amplification and sequencing, namely the 28S nrRNA gene (LSU), internal transcribed spacer regions and intervening 5.8S nrRNA gene (ITS) of the nrDNA operon, actin (actA), histone H3 (his3), translation elongation factor 1-α (tef1), calmodulin (cmdA) and DNA-directed RNA polymerase II second largest subunit (rpb2). PCR amplifications were performed in a total volume of 12.5 μL solutions on a GeneAmp PCR System 9700 (Applied Biosystems, Foster City, California, USA). The primers, protocols and conditions for standard amplification and subsequent sequencing of the loci were according to Bakhshi et al. (2015) for LSU, ITS and his3 loci, and Quaedvlieg et al. (2013) for the remaining loci (actA, tef1, cmdA and rpb2).
Sequence alignment and phylogenetic analyses
The reference nucleotide sequences (Table 1) of representative genera in the Phaeosphaeriaceae were retrieved from GenBank and recently published alignments (Quaedvlieg et al. 2013; Ertz et al. 2015; Jayasiri et al. 2015; Phukhamsakda et al. 2015; Tennakoon et al. 2016; Tibpromma et al. 2016; Karunarathna et al. 2017; Phookamsak et al. 2017; Wanasinghe et al. 2018) (Table 1). The obtained sequences from GenBank, together with the generated sequences in this study, were aligned with the MAFFT v. 7 online interface using default settings (http://mafft.cbrc.jp/alignment/server/) (Katoh and Standley 2013) for each gene and improved manually where necessary using MEGA v. 6.06 (Molecular Evolutionary Genetics Analysis) (Tamura et al. 2013). The alignments were concatenated with Mesquite v. 3.10 (Maddison and Maddison 2015). The best nucleotide substitution model for each data partition was determined by MrModeltest v. 2.3 (Nylander 2004), and a Bayesian phylogenetic reconstruction was performed with MrBayes v. 3.2.2 (Ronquist et al. 2012). The heating parameter was set at 0.15, and the Markov Chain Monte Carlo (MCMC) analysis of four chains was started in parallel from a random tree topology and lasted until the average standard deviation of split frequencies reached a value of 0.01. Burn-in was set to 25%, and trees were saved each 1000 generations. The resulting phylogenetic tree was printed with Geneious v. 8.1.8 (Kearse et al. 2012). All new sequences generated in this study were deposited in GenBank (www.ncbi.nlm.nih.gov).
Taxonomy
Morphological descriptions were based on isolates sporulating in vitro and in planta. In this regard, colonies were sub-cultured onto synthetic nutrient-poor agar plates (SNA; Crous et al. 2009) containing sterile Urtica dioica (stinging nettle) stems (Quaedvlieg et al. 2013). Cultures were incubated at 25 °C under continuous near-ultraviolet light for 14–30 days to promote sporulation. Freehand sections of fungal conidiomata were prepared, and fungal structures were mounted in clear lactic acid. For the morphological study in planta, hand sections were made from infected leaves and mounted in lactic acid. Observations were made with a Nikon Eclipse 80i compound microscope with differential inference contrast (DIC) illumination at 1000× magnification and a mounted Nikon digital sight DS-f1 high-definition colour camera. Thirty measurements were made of all relevant morphological features, and the 95% percentiles are presented, with extremes given between brackets. Photographic plates were edited and combined using Adobe Photoshop CS5. Growth rates and culture characters were noted on MEA and Oatmeal Agar (OA; Crous et al. 2009) after 20 days in the dark at 25 °C. Colony colour was rated according to the mycological colour charts of Rayner (1970).
Results
Phylogenetic analyses
The final concatenated alignment contained 92 ingroup taxa within the family Phaeosphaeriaceae with 1427 characters including gaps (gene boundaries of LSU, 1–790; ITS, 792–1427). These characters contained 586 unique site patterns (195 and 391 for LSU and ITS, respectively). One taxon of Coniothyriaceae (Coniothyrium carteri, CBS 105.91) was used as outgroup.
The results of MrModeltest recommended a SYM + I + G for ITS and GTR + I + G for LSU. All partitions had Dirichlet base frequencies. The Bayesian analysis lasted 4,405,000 generations and saved a total of 8812 trees. After discarding the first 25% of sampled trees, the consensus trees and posterior probabilities were calculated from the remaining 6610 trees (Fig. 1).
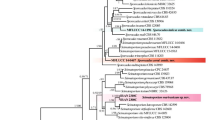
Consensus phylogram (50% majority rule) of 8812 trees resulting from a Bayesian inference analysis of the combined two-loci (LSU and ITS) sequence alignment showing the phylogenetic relationship of the new genus Parastagonosporella among the other Phaeosphaeriaceae genera. Type species are indicated with boldface type. The scale bar indicates 0.02 expected changes per site. The tree was rooted to Coniothyrium carteri (CBS 105.91)
Taxonomy
In the multi-locus phylogeny inferred from the combined dataset shown in Fig. 1, the two isolates occurring on Fallopia convolvulus clustered in a separate clade, distinct from other genera in the family Phaeosphaeriaceae, suggesting that they represent a novel genus in this family. Therefore, a monotypic genus Parastagonosporella, typified by Parastagonosporella fallopiae, is introduced in the family Phaeosphaeriaceae.
Parastagonosporella M. Bakhshi, Arzanlou & Crous, gen. nov.
MycoBank: MB 826900.
Diagnosis: Morphologically distinct from the genus Parastagonospora by having conidiomata with more or less papillate neck, and walls of 4–8 layers of brown textura angularis.
Type species: Parastagonosporella fallopiae M. Bakhshi, Arzanlou & Crous, sp. nov.
Etymology: Morphologically resembling to the genus Parastagonospora, but distinct.
Parastagonosporella fallopiae M. Bakhshi, Arzanlou & Crous, sp. nov. Fig. 2
MycoBank: MB 826901.
Type: IRAN, Guilan Province, Talesh, Jowkandan, on Fallopia convolvulus, Jul. 2012, M. Bakhshi (holotype IRAN 17010 F, culture ex-type CBS 135981). GenBank accessions for sequences obtained from ex-type culture: LSU = MH460545; ITS = MH460543; actA = MH460537; his3 = MH460541; tef1 = MH460549; cmdA = MH460539; rpb2 = MH460547.
Etymology: Named after the host genus from which it was isolated, Fallopia.
Description in planta: Leaf spots numerous, small, 2–3 mm in diameter, circular to angular, and often merging to form irregular patterns, amphigenous, brown in centre, surrounded by raised dark brown margin, diffuse outward to form a halo. Conidiomata pycnidial, dark brown, subepidermal, amphigenous, several in each leaf spot, subglobose, immersed, up to 200 μm in diameter, releasing conidia in creamy to white cirrhi; wall of 4–8 layers of brown textura angularis; ostiolum central, circular, with papillate neck, 15–35 μm wide. Conidiophores reduced to conidiogenous cells. Conidiogenous cells hyaline, smooth, lining the inner cavity, broadly to narrowly ampulliform to subcylindrical, occasionally phialidic, with prominent periclinal thickening or annellidic, proliferating percurrently with more or less distinct annelations, 7–13 × 2.5–6 μm. Conidia hyaline, smooth, thin-walled, scolecosporous, subcylindrical, granular to multi-guttulate, with obtuse apex and truncate to subtruncate base, 3–10-septate, (18–)35–40(−50) × (3–)4–6 μm.
Description in vitro: On sterile Urtica dioica stems on SNA. Conidiomata as in planta, pycnidial, dark brown, erumpent, up to 250 μm in diameter, exuding pale orange to creamy conidial mass. Conidiophores reduced to conidiogenous cells. Conidiogenous cells as in planta, 7–15 × 3–6 μm. Conidia similar in shape as in planta, 5–12-septate, (24–)32–45(−55) × (3–)4.5–6 μm.
Culture characteristics: Colonies on MEA after 20 days in the dark at 25 °C, up to 40 mm in diameter, flat, with even margin and white aerial mycelium, surface olivaceous, reverse iron-grey; on OA surface flat, smooth, entire edge, with aerial mycelium, ochraceous white in centre, olivaceous grey in outer part, reaching 35 mm in diameter after 20 days at 25 °C.
Additional material examined: IRAN, Guilan Province, Talesh, Jowkandan, on Fallopia convolvulus, Jul. 2012, M. Bakhshi (CCTU 1151.1). GenBank accessions: LSU = MH460546; ITS = MH460544; actA = MH460538; his3 = MH460542; tef1 = MH460550; cmdA = MH460540; rpb2 = MH460548.
Discussion
A coelomycetous fungus associated with leaf spot disease of Black Bindweed was subjected to phylogenetic study and morphological analyses. By combining LSU and ITS sequence data as well as detailed morphological data, we were able to delimit a new genus Parastagonosporella, among the coelomycetous genera in the family Phaeosphaeriaceae within the order Pleosporales.
Morphological characters traditionally used to delineate genera in coelomycetes include conidiomatal structure, structure of the conidiophores, conidiogenous cells and conidial features such as septation, pigmentation, and conidial appendages (Sutton 1980; Nag Raj 1993). Recent molecular studies have shown that these features are not always appropriate to delineate genera as natural units, and they may vary even between sibling species (Crous et al. 2012; Quaedvlieg et al. 2013; Phookamsak et al. 2014).
As is the case with many other coelomycetous genera, the lack of useful morphological characters combined with the high level of variation therein and the need for high levels of expertise in morphology-based identification makes it difficult to distinguish individual genera within the Phaeosphaeriaceae based solely on their morphological characters (Quaedvlieg et al. 2013). Hence, genera and species recognition using the molecular phylogeny of several unlinked DNA loci has already resulted in the natural and reliable delimitation of genera within this family as well as in other fungal families of Dothideomycetes, such as the Botryosphaeriaceae (Phillips et al. 2013), Mycosphaerellaceae (Videira et al. 2017) and Pleosporaceae (Ariyawansa et al. 2015). By using molecular techniques, several novel taxa have been described in the family Phaeosphaeriaceae in recent years (Wijayawardene et al. 2014, 2016; Phookamsak et al. 2014, 2017; Ariyawansa et al. 2015; Ertz et al. 2015; Phukhamsakda et al. 2015; Senanayake et al. 2015; Tennakoon et al. 2016; Tibpromma et al. 2016, b; Ahmed et al. 2017; Moslemi et al. 2018; Wanasinghe et al. 2018). The loci used in these and similar recent studies typically include LSU and ITS data as these loci can distinguish most of the presently known genera within the Phaeosphaeriaceae. In the combined (LSU/ITS) phylogenetic tree, these data were sufficient to clearly separate the novel genus Parastagonosporella from other known genera within the Phaeosphaeriaceae. Furthermore, data for the additional loci generated in this study (actA, his3, tef1, rpb2 and cmdA) were deposited in GenBank, as this would aid future studies on the family.
Phylogenetic analyses of combined LSU and ITS sequence data (Fig. 1) indicated that Parastagonosporella is a distinct genus in Phaeosphaeriaceae, which is closely related to the genera Paraphoma, Pseudophaeosphaeria, Setomelanomma and Xenoseptoria. Paraphoma is distinctly different from Parastagonosporella in having ellipsoid and aseptate conidia (Quaedvlieg et al. 2013). Pseudophaeosphaeria (Hyde et al. 2016) and Setomelanomma (Wu et al. 2014) also accommodate species that reproduce sexually. Xenoseptoria differs from Parastagonosporella in having (1–)3-septate conidia tapering to subobtuse apex and obtuse base (Quaedvlieg et al. 2013). Phylogenetically, these genera are also clearly distinct (Fig. 1). Parastagonosporella is morphologically similar to the genus Parastagonospora by having pycnidial conidiomata, hyaline, smooth, ampulliform to subcylindrical conidiogenous cells, with euseptate, hyaline, granular to multi-guttulate conidia with truncate bases, but distinct in having conidiomata with more or less papillate necks and walls of 4–8 layers of brown textura angularis, versus 2–3 layers in Parastagonospora and phialidic or annellidic conidiogenous cells, versus phialidic in Parastagonospora (Quaedvlieg et al. 2013). Phylogenetically, these genera are also clearly distinguishable from each other (Fig. 1). Based on these clear morphological and phylogenetic data, Parastagonosporella is introduced as a new genus.
Quaedvlieg et al. (2013) comprehensively studied the phylogeny of the genus Septoria and other morphologically similar genera such as Stagonospora, Sphaerulina and Phaeosphaeria. Their results surprisingly revealed that “Stagonospora” nodorum (causal agent of nodorum blotch of cereals), clustered in a distinct genus, unrelated to Stagonospora s. str. within the family Massarinaceae. Consequently, they introduced the genus Parastagonospora (with P. nodorum as the type species) in the family Phaeosphaeriaceae, based on multi-locus molecular data to accommodate several cereal pathogens that could not be placed in Stagonospora or Phaeosphaeria. Current literature further indicates that the sole morphology-based classification of coelomycete families as well as their associated genera and species can be misleading. Here we introduce the novel genus Parastagonosporella to accommodate the isolates occurring on Black Bindweed which are parastagonospora-like in morphology, but cluster apart from Parastagonospora by forming a well-supported separate clade with high Bayesian posterior probability.
Based on literature, several coelomycetous fungi have been reported to be present on the host genus Fallopia within the family Polygonaceae, including Discosia sp. (Amphisphaeriaceae, Amphisphaeriales), Phyllosticta fallopiae, Phyllosticta polygonorum (Phyllostictaceae, Botryosphaeriales), Pilidium lythri (Chaetomellaceae, Chaetomellales) and Septoria polygonorum (Mycosphaerellaceae, Capnodiales) (Farr and Rossman 2018). To our knowledge, the new species Parastagonosporella fallopiae is the first association of a fungus belonging to the family Phaeosphaeriaceae on the plant genus Fallopia.
The present study adds a new genus to the Phaeosphaeriaceae, which is a family that has been intensively studied in recent years due to its economic importance (Quaedvlieg et al. 2013; Wijayawardene et al. 2014, 2016; Phookamsak et al. 2014, 2017; Ariyawansa et al. 2015; Ertz et al. 2015; Phukhamsakda et al. 2015; Senanayake et al. 2015; Tennakoon et al. 2016; Tibpromma et al. 2016, b; Ahmed et al. 2017; Wanasinghe et al. 2018). Here, we further demonstrate that the delimitation of taxa in this family based solely on morphological features is not feasible and emphasize the necessity of using DNA sequence data along with morphology and ecology to facilitate the accurate identification in the Phaeosphaeriaceae.
References
Abd-Elsalam KA, Tibpromma S, Wanasinghe DN, Camporesi E, Hyde KD (2016) Equiseticola gen. nov. (Phaeosphaeriaceae), from Equisetum sp. in Italy. Phytotaxa 284:169–180
Ahmed SA, Hofmüller W, Seibold M, de Hoog GS, Harak H, Tammer I, Van Diepeningen AD, Behrens-Baumann W (2017) Tintelnotia, a new genus in Phaeosphaeriaceae harbouring agents of cornea and nail infections in humans. Mycoses 60:244–253
Ariyawansa HA, Hyde KD, Jayasiri SC, Buyck B, Chethana KT, Dai DQ, Dai YC, Daranagama DA, Jayawardena RS, Lücking R, Ghobad-Nejhad M, Niskanen T, Thambugala KM, Voigt K, Zhao RL, Li GJ, Doilom M, Boonmee S, Yang ZL, Cai Q, Cui YY, Bahkali AH, Chen J, Cui BK, Chen JJ, Dayarathne MC, Dissanayake AJ, Ekanayaka AH, Hashimoto A, Hongsanan H, Jones EBG, Larsson E, Li WJ, Li QR, Liu JK, Luo ZL, Maharachchikumbura SSN, Mapook A, McKenzie EHC, Norphanphoun C, Konta S, Pang KL, Perera RH, Phookamsak R, Phukhamsakda C, Pinruan U, Randrianjohany E, Singtripop C, Tanaka K, Tian CM, Tibpromma S, Wahab MA, Wanasinghe DN, Wijayawardene NN, Zhang JF, Zhang H, Abdel-Aziz FA, Wedin M, Westberg M, Ammirati JF, Bulgakov TS, Lima DX, Callaghan TM, Callac P, Chang CH, Coca LF, Dal-Forno M, Dollhofer V, Fliegerová K, Greiner K, Griffith GW, Ho HM, Hofstetter V, Jeewon R, Kang JC, Wen TC, Kirk PM, Kytövuori I, Lawrey JD, Xing J, Li H, Liu ZY, Liu XZ, Liimatainen K, Lumbsch HT, Matsumura M, Moncada B, Nuankaew S, Parnmen S, de Azevedo SALCM, Sommai S, Song Y, de Souza CAF, de Souza-Motta CM, Su HY, Suetrong S, Wang Y, Wei SF, Wen TC, Yuan HS, Zhou LW, Réblová M, Fournier J, Camporesi E, Luangsa-ard J, Tasanathai K, Khonsanit A, Thanakitpipattana D, Somrithipol S, Diederich P, Millanes AM, Common RS, Stadler M, Yan JY, Li XH, Lee HW, Nguyen TT, Lee HB, Battistin E, Marsico O, Vizzini A, Vila J, Ercole E, Eberhardt U, Simonini G, Wen HA, Chen XH, Miettinen O, Spirin V (2015) Fungal diversity notes 111–252—taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers 75:27–274
Ariyawansa HA, Thambugala KM, Manamgoda DS, Jayawardena R, Camporesi E, Boonmee S, Wanasinghe DN, Phookamsak R, Hongsanan S, Singtripop C (2015) Towards a natural classification and backbone tree for Pleosporaceae. Fungal Divers 71:85–139
Bakhshi M, Arzanlou M, Babai-Ahari A (2011) Uneven distribution of mating type alleles in Iranian populations of Cercospora beticola, the causal agent of Cercospora leaf spot disease of sugar beet. Phytopathol Mediterr 50:101–109
Bakhshi M, Arzanlou M, Babai-Ahari A, Groenewald JZ, Crous PW (2015) Is morphology in Cercospora a reliable reflection of generic affinity? Phytotaxa 213:22–34
Barr ME (1979) A classification of Loculoascomycetes. Mycologia 71:935–957
Carson ML (2005) Yield loss potential of Phaeosphaeria leaf spot of maize caused by Phaeosphaeria maydis in the United States. Plant Dis 89:986–988
Chen Q, Jiang JR, Zhang GZ, Cai L, Crous PW (2015) Resolving the Phoma enigma. Stud Mycol 82:137–217
Crous PW, Carris LM, Giraldo A, Groenewald JZ, Hawksworth DL, Hernández-Restrepo M, Jaklitsch WM, Lebrun M-H, Schumacher RK, Stielow JB, Van der Linde EJ, Vilcane J, Voglmayr H, Wood AR (2015) The genera of fungi-fixing the application of the type species of generic names–G 2: Allantophomopsis, Latorua, Macrodiplodiopsis, Macrohilum, Milospium, Protostegia, Pyricularia, Robillarda, Rotula, Septoriella, Torula, and Wojnowicia. IMA Fungus 6:163–198
Crous PW, Groenewald JZ, Lombard L, Wingfield MJ (2012) Homortomyces gen. nov., a new dothidealean pycnidial fungus from the Cradle of Humankind. IMA Fungus 3:109–115
Crous PW, Shivas RG, Quaedvlieg W, Van der Bank M, Zhang Y, Summerell BA, Guarro J, Wingfield MJ, Wood AR, Alfenas AC, Braun U, Cano-Lira JF, García D, Marin-Felix Y, Alvarado P, Andrade JP, Armengol J, Assefa A, den Breeÿen A, Camele I, Cheewangkoon R, De Souza JT, Duong TA, Esteve-Raventós F, Fournier J, Frisullo S, García-Jiménez J, Gardiennet A, Gené J, Hernández-Restrepo M, Hirooka Y, Hospenthal DR, King A, Lechat C, Lombard L, Mang SM, Marbach PAS, Marincowitz S, Marin-Felix Y, Montaño-Mata NJ, Moreno G, Perez CA, Pérez Sierra AM, Robertson JL, Roux J, Rubio E, Schumacher RK, Stchige AM, Sutton DA, Tan YP, Thompson EH, Van der Linde E, Walker AK, Walker DM, Wickes BL, Wong PTW, Groenewald JZ (2014) Fungal planet description sheets: 214–280. Persoonia 32:184–306
Crous PW, Verkley GJM, Christensen M, Castañeda-Ruiz RF, Groenewald JZ (2012) How important are conidial appendages? Persoonia 28:126–137
Crous PW, Verkley GJM, Groenewald JZ (2006) Eucalyptus microfungi known from culture. 1. Cladoriella and Fulvoflamma genera nova, with notes on some other poorly known taxa. Stud Mycol 55:53–63
Crous PW, Verkley GJM, Groenewald JZ, Samson RA (2009) Fungal biodiversity. CBS laboratory manual series 1. Centraalbureau voor Schimmelcultures, Utrecht
Crous PW, Wingfield MJ, Burgess TI, Hardy GSJ, Barber PA, Alvarado P, Barnes CW, Buchanan PK, Heykoop M, Moreno G, Thangavel R, Van der Spuy S, Barili A, Barrett S, Cacciola SO, Cano-Lira JF, Crane C, Decock C, Gibertoni TB, Guarro J, Guevara-Suarez M, Hubka V, Kolařík M, Lira CRS, Ordoñez ME, Padamsee M, Ryvarden L, Soares AM, Stchigel AM, Sutton DA, Vizzini A, Weir BS, Acharya K, Aloi F, Baseia IG, Blanchette RA, Bordallo JJ, Bratek ZT, Butler T, Cano-Canals J, Carlavilla JR, Chander J, Cheewangkoon R, RHSF C, da Silva M, Dutta AK, Ercole E, Escobio V, Esteve-Raventós F, Flores JA, Gené J, Góis JS, Haines L, Held BW, Horta Jung M, Hosaka K, Jung T, Jurjević Z, Kautman V, Kautmanova I, Kiyashko AA, Kozanek M, Kubátová A, Lafourcade M, La Spada F, KPD L, Madrid H, Malysheva EF, Manimohan P, Manjón JL, Martín MP, Mata M, Merényi Z, Morte A, Nagy I, Normand AC, Paloi S, Pattison N, Pawłowska J, Pereira OL, Petterson ME, Picillo B, KNA R, Roberts A, Rodríguez A, Rodríguez-Campo FJ, Romański M, Ruszkiewicz-Michalska M, Scanu B, Schena L, Semelbauer M, Sharma R, Shouche YS, Silva V, Staniaszek-Kik M, Stielow JB, Tapia C, Taylor PWJ, Toome-Heller M, Vabeikhokhei JMC, Van Diepeningen AD, Van Hoa N, Van Tri M, Wiederhold NP, Wrzosek M, Zothanzama J, Groenewald JZ (2017) Fungal planet description sheets: 558–624. Persoonia 38:240–384
Crous PW, Wingfield MJ, Guarro J, Hernández-Restrepo M, Sutton DA, Acharya K, Barber PA, Boekhout T, Dimitrov RA, Dueñas M, Dutta AK, Gené J, Gouliamova DE, Groenewald M, Lombard L, Morozova OV, Sarkar J, Smith MT, Stchigel AM, Wiederhold NP, Alexandrova AV, Antelmi I, Armengol J, Barnes I, Cano-Lira JF, Castañeda Ruiz RF, Contu M, Courtecuisse PR, da Silveira AL, Decock CA, de Goes A, Edathodu J, Ercole E, Firmino AC, Fourie A, Fournier J, Furtado EL, Geering ADW, Gershenzon J, Giraldo A, Gramaje D, Hammerbacher A, He XL, Haryadi D, Khemmuk W, Kovalenko AE, Krawczynski R, Laich F, Lechat C, Lopes UP, Madrid H, Malysheva EF, Marín-Felix Y, Martín MP, Mostert L, Nigro F, Pereira OL, Picillo B, Pinho DB, Popov ES, Rodas Peláez CA, Rooney-Latham S, Sandoval-Denis M, Shivas RG, Silva V, Stoilova-Disheva NM, Telleria MT, Ullah C, Unsicker SB, Van der Merwe NA, Vizzini A, Wagner HG, Wong RTW, Wood AR, Groenewald JZ (2015) Fungal planet description sheets: 320–370. Persoonia 34:167–266
Crous PW, Wingfield MJ, Le Roux JJ, Richardson DM, Strasberg D, Shivas RG, Alvarado P, Edwards J, Moreno G, Sharma R, Sonawane MS, Tan YP, Altés A, Barasubiye T, Barnes CW, Blanchette RA, Boertmann D, Bogo A, Carlavilla JR, Cheewangkoon R, Daniel R, de Beer ZW, de Jesús Yáñez-Morales M, Duong TA, Fernández-Vicente J, Geering ADW, Guest DI, Held BW, Heykoop M, Hubka V, Ismail AM, Kajale SC, Khemmuk W, Kolařík M, Kurli R, Lebeuf R, Lévesque CA, Lombard L, Magista D, Manjón JL, Marincowitz S, Mohedano JM, Nováková A, Oberlies NH, Otto EC, Paguigan ND, Pascoe IG, Pérez-Butrón JL, Perrone G, Rahi P, Raja HA, Rintoul T, Sanhueza RMV, Scarlett K, Shouche YS, Shuttleworth LA, Taylor PWJ, Thorn RG, Vawdrey LL, Solano-Vidal R, Voitk A, Wong PTW, Wood AR, Zamora JC, Groenewald JZ (2015) Fungal planet description sheets: 371–399. Persoonia 35:264–327
De Gruyter J, Aveskamp MM, Woudenberg JH, Verkley GJM, Groenewald JZ, Crous PW (2009) Molecular phylogeny of Phoma and allied anamorph genera: towards a reclassification of the Phoma complex. Mycol Res 113:508–519
Ertz D, Diederich P, Lawrey JD, Berger F, Freebury CE, Coppins B, Gardiennet A, Hafellner J (2015) Phylogenetic insights resolve Dacampiaceae (Pleosporales) as polyphyletic: Didymocyrtis (Pleosporales, Phaeosphaeriaceae) with Phoma-like anamorphs resurrected and segregated from Polycoccum (Trypetheliales, Polycoccaceae fam. nov.). Fungal Divers 74:53–89
Farr DF, Rossman AY (2018) Fungal databases, U.S. national fungus collections, ARS, USDA. https://nt.ars-grin.gov/fungaldatabases/
Hyde KD, Hongsanan S, Jeewon R, Bhat DJ, McKenzie EHC, Jones EBG, Phookamsak R, Ariyawansa HA, Boonmee S, Zhao Q, Abdel-Aziz FA, Abdel-Wahab MA, Banmai S, Chomnunti P, Cui BK, Daranagama DA, Das K, Dayarathne MC, de Silva NI, Dissanayake AJ, Doilom M, Ekanayaka AH, Gibertoni TB, Go’es-Neto A, Huang SK, Jayasiri SC, Jayawardena RS, Konta S, Lee HB, Li WJ, Lin CG, Liu JK, Lu YZ, Luo ZL, Manawasinghe IS, Manimohan P, Mapook A, Niskanen T, Norphanphoun C, Papizadeh M, Perera RH, Phukhamsakda C, Richter C, Santiago ALCM, Drechsler-Santos ER, Senanayake IC, Tanaka K (2016) Fungal diversity notes 367–490: taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers 80:1–270
Hyde KD, Jones EG, Liu J-K, Ariyawansa H, Boehm E, Boonmee S, Braun U, Chomnunti P, Crous PW, Dai D-Q (2013) Families of dothideomycetes. Fungal Divers 63:1–313
Jayasiri SC, Wanasinghe DN, Ariyawansa HA, Jones EBG, Kang JC, Promputtha I, Bahkali AH, Bhat J, Camporesi E, Hyde KD (2015) Two novel species of Vagicola (Phaeosphaeriaceae) from Italy. Mycosphere 6:716–728
Karunarathna A, Papizadeh M, Senanayake IC, Jeewon R, Phookamsak R, Goonasekara ID, Wanasinghe DN, Wijayawardene NN, Amoozegar MA, Shahzadeh Fazeli SA (2017) Novel fungal species of Phaeosphaeriaceae with an asexual/sexual morph connection. Mycosphere 8:1818–1834
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Lawrey JD, Diederich P, Nelsen MP, Freebury C, Van den Broeck D, Sikaroodi M, Ertz D (2012) Phylogenetic placement of lichenicolous Phoma species in the Phaeosphaeriaceae (Pleosporales, Dothideomycetes). Fungal Divers 55:195–213
Li WJ, Bhat DJ, Camporesi E, Tian Q, Wijayawardene NN, Dai DQ, Phookamsak R, Chomnunti P, Bahkali AH, Hyde KD (2015) New asexual morph taxa in Phaeosphaeriaceae. Mycosphere 6:681–708
Liu JK, Hyde KD, Jones EG, Ariyawansa HA, Bhat DJ, Boonmee S, Maharachchikumbura SS, McKenzie EHC, Phookamsak R, Phukhamsakda C, Shenoy BD, Abdel-Wahab MA, Buyck B, Chen J, Chethana KWT, Singtripop C, Dai DQ, Dai YC, Daranagama DA, Dissanayake AJ, Doliom M, D’souza MJ, Fan XL, Goonasekara ID, Hirayama K, Hongsanan S, Jayasiri SC, Jayawardena RS, Karunarathna SC, Li WJ, Mapook A, Norphanphoun C, Pang KL, Perera RH, Peršoh D, Pinruan U, Senanayake IC, Somrithipol S, Suetrong S, Tanaka K, Thambugala KM, Tian Q, Tibpromma S, Udayanga D, Wijayawardene NN, Wanasinghe DN, Wisitrassameewong K, Zeng XY, Abdel-Aziz FA, Adamčík S, Bahkali AH, Boonyuen N, Bulgakov T, Callac P, Chomnunti P, Greiner K, Hashimoto A, Hofstetter V, Kang JC, Lewis DA, Li XH, Liu XX, Liu ZY, Matsumura M, Mortimer PE, Rambold G, Randrianjohany E, Sato G, Indrasutdhi VS, Tian CM, Verbeken A, von Brackel W, Wang Y, Wen TC, Xu JC, Yan JY, Zhao RL, Camporesi E (2015) Fungal diversity notes 1–110: taxonomic and phylogenetic contributions to fungal species. Fungal Divers 72:1–197
Maddison WP, Maddison DR (2015) Mesquite: a modular system for evolutionary analysis. Version 3.10. http://mesquiteproject.org
Mapook A, Boonmee S, Ariyawansa HA, Tibpromma S, Campesori E, Jones EG, Bahkali AH, Hyde K (2016) Taxonomic and phylogenetic placement of Nodulosphaeria. Mycol Prog 15:34
Miyake I (1909) Studies on the parasitic fungi of rice in Japan. Bot Mag Tokyo 23:85–97
Möller E, Bahnweg G, Sandermann H, Geiger HH (1992) A simple and efficient protocol for isolation of high molecular weight DNA from filamentous fungi, fruit bodies, and infected plant tissues. Nucleic Acids Res 20:6115–6116
Moslemi A, Ades PK, Crous PW, Groom T, Scott JB, Nicolas ME, Taylor PW (2018) Paraphoma chlamydocopiosa sp. nov. and Paraphoma pye sp. nov., two new species associated with leaf and crown infection of pyrethrum. Plant Pathol 67:124–135
Nag Raj T (1993) Coelomycetous anamorphs with appendage-bearing conidia. Mycologue Publications, Waterloo
Nylander JAA (2004) MrModeltest v2.0. Program distributed by the author. Evolutionary Biology Centre, Uppsala University, Uppsala
Phillips AJL, Alves A, Abdollahzadeh J, Slippers B, Wingfield MJ, Groenewald JZ, Crous PW (2013) The Botryosphaeriaceae: genera and species known from culture. Stud Mycol 76:51–167
Phookamsak R, Liu J-K, McKenzie EHC, Manamgoda DS, Ariyawansa H, Thambugala KM, Dai D-Q, Camporesi E, Chukeatirote E, Wijayawardene NN (2014) Revision of Phaeosphaeriaceae. Fungal Divers 68:159–238
Phookamsak R, Wanasinghe DN, Hongsanan S, Phukhamsakda C, Huang S-K, Tennakoon DS, Norphanphoun C, Camporesi E, Bulgakov TS, Promputtha I, Mortimer PE, Xu JC, Hyde KD (2017) Towards a natural classification of Ophiobolus and ophiobolus-like taxa; introducing three novel genera Ophiobolopsis, Paraophiobolus and Pseudoophiobolus in Phaeosphaeriaceae (Pleosporales). Fungal Divers 87:299–339
Phukhamsakda C, Ariyawansa HA, Phookamsak R, Chomnunti P, Bulgakov TS, Yange JB, Bhat DJ, Bahkalih AH, Hyde KD (2015) Muriphaeosphaeria galatellae gen. et sp. nov. in Phaeosphaeriaceae (Pleosporales). Phytotaxa 227:55–65
Quaedvlieg W, Verkley GJM, Shin H-D, Barreto RW, Alfenas AC, Swart WJ, Groenewald JZ, Crous PW (2013) Sizing up Septoria. Stud Mycol 75:307–390
Rayner RW (1970) A mycological colour chart. CMI and British Mycological Society, Kew
Ronquist F, Teslenko M, Van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Sánchez Márquez S, Bills GF, Zabalgogeazcoa I (2007) The endophytic mycobiota of the grass Dactylis glomerata. Fungal Divers 27:171–195
Schoch CL, Crous PW, Groenewald JZ, Boehm E, Burgess TI, De Gruyter J, De Hoog GS, Dixon LJ, Grube M, Gueidan C (2009) A class-wide phylogenetic assessment of Dothideomycetes. Stud Mycol 64:1–15
Schoch CL, Shoemaker RA, Seifert KA, Hambleton S, Spatafora JW, Crous PW (2006) A multigene phylogeny of the Dothideomycetes using four nuclear loci. Mycologia 98:1041–1052
Senanayake IC, Maharachchikumbura SS, Hyde KD, Bhat JD, Jones EG, McKenzie EH, Dai DQ, Daranagama DA, Dayarathne MC, Goonasekara ID, Konta S, Li WJ, Shang QJ, Stadler M, Wijayawardene NN, Xiao YP, Norphanphoun C, Li Q, Liu XZ, Bahkali AH, Kang JC, Wang Y, Wen TC, Wendt L, Xu JC, Camporesi E (2015) Towards unraveling relationships in Xylariomycetidae (Sordariomycetes). Fungal Divers 73:73–144
Shoemaker RA (1984) Canadian and some extralimital Nodulosphaeria and Entodesmium species. Can J Bot 62:2730–2753
Slippers B, Boissin E, Phillips A, Groenewald JZ, Lombard L, Wingfield MJ, Postma A, Burgess T, Crous PW (2013) Phylogenetic lineages in the Botryosphaeriales: a systematic and evolutionary framework. Stud Mycol 76:31–49
Stukenbrock EH, Banke S, McDonald BA (2006) Global migration patterns in the fungal wheat pathogen Phaeosphaeria nodorum. Mol Ecol 15:2895–2904
Sutton BC (1980) The Coelomycetes: fungi imperfecti with pycnidia, acervuli and stromata. Commonwealth Mycological Institute, Kew
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA 6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tennakoon DS, Hyde KD, Phookamsak R, Wanasinghe DN, Camporesi E, Promputtha I (2016) Taxonomy and phylogeny of Juncaceicola gen. nov. (Phaeosphaeriaceae, Pleosporinae, Pleosporales). Cryptogam Mycol 37:135–156
Thambugala KM, Wanasinghe DN, Phillips AJL, Camporesi E, Bulgakov TS, Phukhamsakda C, Ariyawansa HA, Goonasekara ID, Phookamsak R, Dissanayake A, Tennakoon DS, Tibpromma S, Chen YY, Liu ZY, Hyde KD (2017) Mycosphere notes 1–50: grass (Poaceae) inhabiting Dothideomycetes. Mycosphere 8:697–796
Tibpromma S, Liu J-K, Promputtha I, Camporesi E, Bhakali AH, Hyde KD, Boonmee S (2016) Ophiosimulans tanaceti gen. et sp. nov. (Phaeosphaeriaceae) on Tanacetum sp. (Asteraceae) from Italy. Mycol Prog 15:1–11
Tibpromma S, Promputtha I, Phookamsak R, Boonmee S, Camporesi E, Yang J-B, Bhakali AH, McKenzie EH, Hyde KD (2015) Phylogeny and morphology of Premilcurensis gen. nov. (Pleosporales) from stems of Senecio in Italy. Phytotaxa 236:40–52
Tibpromma S, Wijayawardene NN, Manamgoda DS, Boonmee S, Wanasinghe DN, Camporesi E, Yang J-B, Hyde KD (2016) Camarosporium arezzoensis on Cytisus sp., an addition to sexual state of Camarosporium sensu stricto. Saudi J Biol Sci 23:1–8
Trakunyingcharoen T, Lombard L, Groenewald JZ, Cheewangkoon R, To-anun C, Alfenas AC, Crous PW (2014) Mycoparasitic species of Sphaerellopsis, and allied lichenicolous and other genera. IMA Fungus 5:391–414
Videira SIR, Groenewald JZ, Nakashima C, Braun U, Barreto RW, de Wit PJGM, Crous PW (2017) Mycosphaerellaceae—chaos or clarity? Stud Mycol 87:257–421
Wanasinghe DN, Jones EG, Camporesi E, Boonmee S, Karunarathna SC, Thines M, Mortimer PE, Xu J, Hyde KD (2014) Dematiopleospora mariae gen. sp. nov., from Ononis spinosa in Italy. Cryptogam Mycol 35:105–117
Wanasinghe DN, Phukhamsakda C, Hyde KD, Jeewon R, Lee HB, Jones EG, Tibpromma S, Tennakoon DS, Dissanayake AJ, Jayasiri SC, Gafforov Y, Camporesi E, Bulgakov TS, Ekanayake AH, Perera RH, Samarakoon MC, Goonasekara ID, Mapook A, Li WJ, Senanayake IC, Li J, Norphanphoun C, Doilom M, Bahkali AH, Xu J, Mortimer PE, Tibell L, Tibell S, Karunarathna SC (2018) Fungal diversity notes 709–839: taxonomic and phylogenetic contributions to fungal taxa with an emphasis on fungi on Rosaceae. Fungal Divers 89:1–236
Wang Y, Guo L-D, Hyde KD (2005) Taxonomic placement of sterile morphotypes of endophytic fungi from Pinus tabulaeformis (Pinaceae) in northeast China based on rDNA sequences. Fungal Divers 20:235–260
Wijayawardene NN, Camporesi E, Song Y, Dai D-Q, Bhat DJ, Mckenzie EH, Chukeatirote E, Mel'Nik VA, Wang Y, Hyde KD (2013) Multi-gene analyses reveal taxonomic placement of Scolicosporium minkeviciusii in Phaeosphaeriaceae (Pleosporales). Cryptogam Mycol 34:357–366
Wijayawardene N, Crous PW, Kirk PM, Hawksworth DL, Boonmee S, Braun U, Dai DQ, D’souza MJ, Diederich P, Dissanayake A, Doilom M, Hongsanan S, Jones EBG, Groenewald JZ, Jayawardena R, Lawrey JD, Liu JK, Lücking R, Madrid H, Manamgoda DS, Muggia L, Nelsen MP, Phookamsak R, Suetrong S, Tanaka K, Thambugala KM, Wanasinghe DN, Wikee S, Zhang Y, Aptroot A, Ariyawansa HA, Bahkali AH, Bhat DJ, Gueidan C, Chomnunti P, Hoog GSD, Knudsen K, Li WJ, McKenzie EHC, Miller AN, Phillips AJL, Piątek M, Raja HA, Shivas RS, Slippers B, Taylor JE, Tian Q, Wang Y, Woudenberg JHC, Cai L, Jaklitsch WM, Hyde KD (2014) Naming and outline of Dothideomycetes—2014 including proposals for the protection or suppression of generic names. Fungal Divers 69:1–55
Wijayawardene NN, Hyde KD, Wanasinghe DN, Papizadeh M, Goonasekara ID, Camporesi E, Bhat DJ, McKenzie EH, Phillips AJ, Diederich P, Tanaka K, Li WJ, Tangthirasunun N, Phookamsak R, Dai DQ, Dissanayake AJ, Weerakoon G, Maharachchikumbura SSN, Hashimoto A, Matsumura M, Bahkali AH, Wang Y (2016) Taxonomy and phylogeny of dematiaceous coelomycetes. Fungal Divers 77:1–316
Wu ZQ, Fan XL, Yang T, Tian CM, Liang YM, Yan-Fang Ma YF, Zhang SL (2014) New record of Setomelanomma holmii on Picea crassifolia in China based on morphological and molecular data. Mycotaxon 128:105–111
Yang JW, Yeh YH, Kirschner R (2016) A new endophytic species of Neostagonospora (Pleosporales) from the coastal grass Spinifex littoreus in Taiwan. Botany 94:593–598
Zhang Y, Crous PW, Schoch CL, Hyde KD (2012) Pleosporales. Fungal Divers 53:1–221
Acknowledgements
We acknowledge the Iran National Science Foundation (INSF), the Research Deputy of the Iranian Research Institute of Plant Protection, Agricultural Research, Education and Extension Organization (AREEO) and the Westerdijk Fungal Biodiversity Institute for financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Section Editor: Marc Stadler and Hans-Josef Schroers
This article is part of the “Special Issue on Hyphomycete Taxonomy and Diversity in honour of Walter Gams who passed away in April 2017”.
Rights and permissions
About this article
Cite this article
Bakhshi, M., Arzanlou, M., Groenewald, J.Z. et al. Parastagonosporella fallopiae gen. et sp. nov. (Phaeosphaeriaceae) on Fallopia convolvulus from Iran. Mycol Progress 18, 203–214 (2019). https://doi.org/10.1007/s11557-018-1428-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-018-1428-z