Abstract
A survey of freshwater ascomycetes conducted in Thailand yielded a number of aquatic hyphomycetes. In this study, fresh collections of three chaetosphaeriaceous species from submerged wood in freshwater are characterized based on morphology and molecular phylogeny. Dictyochaeta siamensis sp. nov. and Tainosphaeria siamensis sp. nov. are introduced based on morphological and molecular data. A detailed description of Menisporopsis theobromae from the new collection is presented and the first molecular data for this genus are provided. Phylogenetic analysis of combined LSU and ITS sequence data was carried out to determine the phylogenetic placement of these species within the family Chaetosphaeriaceae. Menisporopsis theobromae showed a close phylogenetic relationship with Rattania, but has significantly different morphology; D. siamensis clustered together with Codinaeopsis, Menispora, and Zignoëlla; while T. siamensis clustered together with T. crassiparies but presented as a distinct clade. The phylogenetic relationship between the new taxa and their relatives are compared and discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hyphomycetes are asexual fungi producing conidia and conidiogenous cells directly on the mycelium (Seifert et al. 2011). Currently, 1800 genera comprising 9000 species have been described worldwide (Kirk et al. 2008). Despite the large number of hyphomycetes documented, little is known connecting them to their sexual forms and, hence, their systematic placement. However, it is possible to link asexual morph taxa to their sexual morphs using molecular phylogenetic analysis.
Aquatic hyphomycetes are a dominant and diverse group of asexual fungi which are involved in litter degradation in freshwater ecosystems (Ho et al. 2001; Cai et al. 2003; Tsui and Hyde 2004; Krauss et al. 2011; Hyde et al. 2016). Taxa in this habitat are generally saprobic; however, some may also be pathogens or symbionts (Wong et al. 1998). The present study deals with three saprobic hyphomycetes in Chaetosphaeriaceae, a family which was established for Chaetosphaeria and its allies (Réblová et al. 1999). The type genus Chaetosphaeria was introduced by Tulasne and Tulasne (1863) based on the type species, C. innumera. Species of Chaetosphaeria produce both perithecial ascomata and conidia, while some species apparently produce only conidia. Among those species for which both sexual and asexual morphs are known, the species differences are more readily seen in the asexual morphs than in the sexual morphs. Therefore, the diagnostic taxonomic value of individual morphological characters of the asexual morphs, particularly those of conidia and conidiogenous cells, has been discussed several times (Gams and Holubová-Jechová 1976; Kendrick 1980; Arambarri and Cabello 1989; Réblová 2000; Li et al. 2012). However, there are no review papers addressing the characters and phylogeny of asexual morphs. Réblová (2000) suggested that ontogenetic characters, such as conidiogenous cells and conidia, are useful to show the phylogenetic relationships and classify the asexual morph genus Chaetosphaeria. Lumbsch and Huhndorf (2010) listed ten genera (mostly sexual genera) in the family Chaetosphaeriaceae and Maharachchikumbura et al. (2015) listed 35 genera, which included most of the sexual and asexual genera, based on the literature.
In this study, we collected freshwater chaetosphaeriaceous hyphomycetes in Thailand and our studies yielded two novel species and one collection of Menisporopsis theobromae (type of the genus). The new species Dictyochaeta siamensis and Tainosphaeria siamensis are introduced, along with the first sequence data for M. theobromae. For those genera in Chaetosphaeriaceae that have been sequenced, phylogenetic analyses are presented to provide further evidence for the uniqueness of these taxa, and the new taxa are described and compared with related taxa.
Materials and methods
Isolation and morphology
Specimens of submerged decaying wood were collected from Chiang Rai and Prachuap Khiri Khan Provinces, Thailand during November to December 2014 and returned to the laboratory in plastic bags. The samples were incubated in plastic boxes lined with moistened tissue paper at room temperature for one week. The samples were processed and examined following the methods described by Taylor and Hyde (2003). Morphological observations were made using a Motic SMZ168 Series stereomicroscope and photographed by a Nikon E80i microscope camera system. Measurements were made with Tarosoft (R) Image Frame Work (Liu et al. 2010).
Isolations were made from single spores as described by Liu et al. (2010). Type material is deposited at the herbarium of Guizhou Academy of Agriculture Sciences (GZAAS), Guiyang, China and Mae Fah Luang University (MFLU), Chiang Rai, Thailand. Fungi isolated in our study were deposited at Mae Fah Luang University Culture Collection (MFLUCC) and Guizhou Culture Collection, China (GZCC). Facesoffungi numbers and Index Fungorum numbers are provided as outlined by Jayasiri et al. (2015) and in Index Fungorum (2016).
DNA extraction, PCR amplification, and sequencing
Fungal isolates were grown on PDA for 21 days at 28 °C in the dark. Genomic DNA was extracted from the fresh mycelium using the Biospin Fungus Genomic DNA Extraction Kit (BioFlux®), following the manufacturer’s protocol (Hangzhou, P.R. China).
DNA amplification was performed by polymerase chain reaction (PCR). Two partial gene portions were used in this study: the internal transcribed spacers (ITS) and the large subunits of the nuclear ribosomal RNA genes (LSU). The primers used were ITS5 and ITS4 (White et al. 1990) for ITS, and LROR and LR5 (Vilgalys and Hester 1990) for LSU. The PCR thermal cycle program for ITS and LSU amplification was as follows: initially denaturing step of 94 °C for 3 min, followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 55 °C for 50 s, elongation at 72 °C for 1 min, and a final extension at 72 °C for 10 min.
PCR products were purified using minicolumns, purification resin, and buffer according to the manufacturer’s protocols (Amersham product code: 27-9602-01). Sequence analysis was carried out by Shanghai Sangon Biological Engineering Technology & Services Co., Ltd. (Shanghai, P.R. China).
Phylogenetic analysis
Sequences generated from different primers were analyzed with other sequences obtained from GenBank. The related sequences were determined by using a BLAST search to reveal the closest matches with taxa in Chaetosphaeriaceae and recent relevant publications (Lumbsch and Huhndorf 2010; Crous et al. 2012; Hashimoto et al. 2015a, b). Sequences were aligned using BioEdit 7.2.5 (Hall 1999) and ClustalX v.1.83 (Thompson et al. 1997). The alignments were checked visually and improved manually where necessary. Phylogenetic analyses were performed by using PAUP v.4.0b10 (Swofford 2002) for maximum parsimony (MP) and MrBayes v.3.0b4 (Huelsenbeck and Ronquist 2001) for Bayesian analyses.
A maximum likelihood (ML) analysis was performed at the CIPRES web portal (Miller et al. 2010) using RAxML v.7.2.8 as part of the “RAxML-HPC2 on TG” tool (Stamatakis 2006). A general time-reversible model (GTR) was applied with a discrete gamma distribution and four rate classes. Fifty thorough ML tree searches were done in RAxML v.7.2.7 under the same model. One thousand non-parametric bootstrap iterations were run with the GTR model and a discrete gamma distribution. The resulting replicates were plotted onto the best scoring tree obtained previously.
MP analyses were performed using the heuristic search option with 1000 random taxa addition and tree bisection and reconnection (TBR) as the branch-swapping algorithm. All characters were unordered and of equal weight, and gaps were treated as missing data. Maxtrees were unlimited, branches of zero length were collapsed and all multiple, equally parsimonious trees were saved. Clade stability was assessed using a bootstrap (BT) analysis with 1000 replicates, each with 10 replicates of random stepwise addition of taxa (Hillis and Bull 1993). The model of evolution was estimated by using MrModeltest 2.2 (Nylander 2004). Posterior probabilities (PP) (Rannala and Yang 1996; Zhaxybayeva and Gogarten 2002) were determined by Markov Chain Monte Carlo sampling (BMCMC) in MrBayes v.3.0b4 (Huelsenbeck and Ronquist 2001). Six simultaneous Markov chains were run for 3,000,000 generations and trees were sampled every 1000th generation a tree was sampled. The MCMC heated chain was set with a “temperature” value of 0.15. All sampled topologies beneath the asymptote (20 %) were discarded as part of a burn-in procedure; the remaining trees were used for calculating posterior probabilities (PP) in the majority rule consensus tree.
Phylogenetic trees were drawn using TreeView (Page 1996) and MEGA5 (Tamura et al. 2011). The sequences derived in this study are deposited in GenBank (Table 1).
Results
Phylogenetic analysis
Three isolates of hyphomycetes obtained from the incubated specimens of submerged wood were identified in the family Chaetosphaeriaceae. ITS and LSU sequence data and morphological characters were used to assign the species or genus and to describe novel taxa with a comparison with similar taxa.
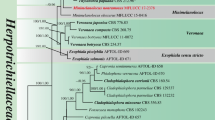
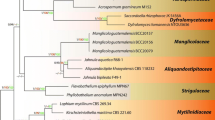
The combined LSU and ITS dataset comprised 45 taxa, with Lasiosphaeria ovina (SMH 4605) as the outgroup taxon. The dataset comprises 1648 characters after alignment: 1112 characters were constant and 367 characters were parsimony informative, while 169 variable characters were parsimony-uninformative. RAxML, MP, and Bayesian analysis of the combined dataset resulted in phylogenetic reconstructions with largely similar topologies, and the Bayesian tree is shown in Fig. 1.
Consensus phylogram (50 %) majority rule resulting from a Bayesian analysis of a combined LSU and ITS sequence alignment of Chaetosphaeriaceae. Bayesian posterior probabilities (PP) above 0.95 and ML bootstrap proportion (BP) greater than 50 % are presented at the nodes as BP/PP. Branches with more than 75 % bootstrap (ML) are in bold. The original isolate numbers are noted after the species names, the new isolates are in red, and ex-type isolates are in bold. The scale bar shows 0.05 changes and the tree is rooted to Lasiosphaeria ovina (SMH 4605)
Representatives of the sequenced genera (with molecular data) of Chaetosphaeriaceae (Réblová and Winka 2000; Lumbsch and Huhndorf 2010; Crous et al. 2012; Hashimoto et al. 2015a, b; Liu et al. 2015; Maharachchikumbura et al. 2015) are included in our phylogenetic analysis (Fig. 1). Twenty-six genera are represented by at least one species in Chaetosphaeriaceae. Some were introduced or studied recently, such as Neopseudolachnella and Pseudodinemasporium (Hashimoto et al. 2015b), Brunneodinemasporium, Dendrophoma, and Dinemasporium (Crous et al. 2012), Infundibulomyces (Somrithipol et al. 2008), Tainosphaeria (Fernández and Huhndorf 2005), and Pyrigemmula (Magyar et al. 2011). Most of them are asexual taxa. Our three isolates were also included in the analysis of combined LSU and ITS sequence data. One isolate was identified as Menisporopsis theobromae and clustered with Rattania with high statistical support, but is quite different morphologically. Dictyochaeta siamensis formed a sister clade to the Codinaeopsis, Menispora, and Zignoëlla clade in Chaetosphaeriales. The third isolate clustered with Tainosphaeria crassiparies in a well-supported clade. It was, however, phylogenetically and morphologically distinct and is introduced as T. siamensis sp. nov. in this paper.
Taxonomy
Dictyochaeta siamensis J. Yang, K.D. Hyde & J.K. Liu, sp. nov. Fig. 2
Dictyochaeta siamensis (MFLU 15-1149, holotype). a Substrate. b Colonies. c Conidiophore and conidiogenous cell. d, e Setae. f–h Conidiophores and conidiogenous cell. i–n Conidia. o Germinating conidium. p–q Culture. Scale bars: b = 200 μm; c, d = 50 μm; e, f, h, o = 30 μm; g, l = 20 μm; j–n = 15 μm
Index Fungorum number: IF552464; Facesoffungi number: FoF 02188
Holotype: MFLU 15-1149
Etymology Named after the country from where this fungus was collected, Thailand.
Saprobic on decaying plant twigs in freshwater. Colonies effuse, brown, with long hairy mycelium, with white glistening conidial mass. Mycelium partly immersed, partly superficial, consisting of branched, septate, smooth, thin-walled, brown hyphae. Asexual morph: Setae erect, straight, dark brown at the base, fading towards the apex, acerose to subacerose, septate, unbranched, smooth, fertile, and tapering at distal ends, 165–365 × 3–6 μm. Conidiophores mononematous, macronematous, brown at the base, fading to pale brown towards the apex, 4–6 septate, unbranched, cylindrical, straight or slightly curved, erect, smooth, arising singly or in groups from the mycelial knots from the bases of setae, 60–100 × 2–5 μm. Conidiogenous cells mostly monophialidic or rarely polyphialidic, sometimes percurrently proliferating, integrated, terminal, and determinate, with conspicuous collarettes. Conidia 15.5–21 × 2.5–4 μm (x̄ = 17.5 × 3 μm, n = 20), aggregated in slimy mass at the apex of the conidiophore, acrogenous, enteroblastic, hyaline, aseptate, cylindrical or long fusiform, curved, lunar, with 7–12-μm-long hair-like appendages at both ends, smooth, rounded at apex. Sexual morph: Not observed
Material examined
THAILAND, Prachuap Khiri Khan Province, Hua Hin, Kaeng Krachan, on plant twigs in stream running from the national park, 25 December 2014, Jaap van Strien Site 4 15-2 (MFLU 15-1149, holotype); ex-type living culture MFLUCC 15-0614; GZCC 15-0060.
Notes
The combined LSU and ITS phylogenetic analysis showed that Dictyochaeta siamensis clustered together with Menispora, Zignoëlla, and Codinaeopsis, and, although they are phylogenetically related, they are morphologically distinct genera. Codinaeopsis has discrete conidiogenous cells produced in verticels along an erect seta; Menispora also has mainly discrete conidiogenous cells on conidiophores, the upper part of which is usually sterile. The new taxon D. siamensis has five or six conidiophores, each grouped around a single seta, and these morphological characters suggest that the taxon is a Dictyochaeta species. A key to Dictyochaeta species was provided by Whitton et al. (2000); however, D. siamensis cannot be assigned to any species. In addition, D. siamensis differs from the recently described species D. aciculata by its long fusiform, aseptate conidia, while the latter species has 3-septate, acicular conidia (Silva and Gusmão 2013). The molecular phylogeny of Dictyochaeta species is unresolved and there are few reliable sequences available in GenBank, although many species have been described in the genus. In our phylogenetic analysis, D. simplex (CBS 966.69) was included to represent Dictyochaeta and the result showed that the new taxon is not related; however, this is not an ex-type species. New collections and molecular data are required for Dictyochaeta species to determine their phylogenetic placement in Chaetosphaeriaceae, especially the type species. Menispora and Zignoëlla were reported as related genera (Fernández et al. 2006; Somrithipol et al. 2008); however, the relationship between these two genera requires further data for confirmation.
Tainosphaeria siamensis J. Yang, K.D. Hyde & J.K. Liu, sp. nov. Fig. 3
Index Fungorum number: IF552113; Facesoffungi number: FoF 02189
Holotype: MFLU 15-1142
Etymology Named after the country from where this fungus was collected, Thailand.
Colonies on submerged wood, effuse, aggregate, brown, as a hyaline, glistening conidial mass, hairy short. Asexual morph: Conidiophores mononematous, macronematous, brown at the base fading to pale brown towards the apex, 40.5–86 μm long, 2.5–4 μm wide, 2–4 septate, unbranched, cylindrical, straight or slightly curved, erect, solitary, smooth. Conidiogenous cells monophialidic, integrated, terminal, determinate, with conspicuous collarettes. Conidia 13.5–19 × 2–3.5 μm (x̄ = 16 × 2.5 μm, n = 20), aggregated in slimy mass at the apex of the conidiophore, acrogenous, enteroblastic, phialidic, hyaline, aseptate, cylindrical or long fusiform, with 3.6–10-μm-long hair-like appendages at both ends, smooth, apex rounded and slightly. Sexual morph: Not observed.
Material examined
THAILAND, Prachuap Khiri Khan Province, Hua Hin, Kaeng Krachan, on submerged wood, 25 December 2014, Jaap van Strien, Site 4 15-2 (MFLU 15-1142, holotype); ex-type living culture MFLUCC 15-0607; GZCC 15-0056.
Notes
Tainosphaeria was introduced by Fernández and Huhndorf (2005) as a monotypic genus with T. crassiparies as the type species; both the asexual and sexual morphs were found from the substrate and described. Tainosphaeria siamensis was found on submerged wood with only the asexual morph; the morphology matches the genus Tainosphaeria. In a MegaBLAST search of GenBank, the closest hits for the LSU sequence data are Tainosphaeria crassiparies strain SMH 1934 [GenBank AF466089, identities 811/826 (98 %)] and Pseudolachnea fraxini strain CBS 113701 [GenBank JQ889301, identities 800/829 (97 %)]; for ITS, the closest hit is Dictyochaeta simplex strain ICMP 14613 [GenBank EF029193, identities 456/515 (89 %)]; the latter BLAST hits are probably because of the lack of ITS sequences for genera closely related to Tainosphaeria. The combined LSU and ITS sequence analysis showed that these two species of Tainosphaeria are phylogenetically distinct and form a well-supported (100 % ML, 1.00 PP) clade in the family Chaetosphaeriaceae. This also confirms Tainosphaeria as a well-supported genus of Chaetosphaeriaceae. Morphologically, T. siamensis differs from T. crassiparies by its larger conidia (13.5–19 × 2–3.5 μm versus 10.5–14.8 × 2–3 μm in T. crassiparies) (Fernández and Huhndorf 2005).
Menisporopsis theobromae S. Hughes., Mycol Pap 48: 59 (1952) Fig. 4
Colonies on submerged wood, superficial, scattered, effuse, white to pale brown. Mycelium partly immersed, composed of brown hyphae. Asexual morph: Setae central, 223 × 5.5 μm, solitary, erect, brown, septate, thick-walled, lower part encased tightly by compact conidiophores and obviously wider than each conidiophore. Conidiophores synnematous, macronematous, brown, smooth, thin to thick-walled, septate, unbranched, cylindrical, lower part narrow, upper part wider, erect, straight or slightly flexuous, up to 112.5 μm long. Conidiogenous cells terminal, monophialidic, integrated, pale brown, with collarettes. Conidia 14–19 × 2–3 μm (x̄ = 17 × 2.5 μm, n = 20), hyaline, aseptate, thin-walled, smooth, acrogenous, aggregated in slimy masses at the apex of the synnemata, fusiform, gently curved or straight, with a single and unbranched, flexuous, hyaline, hair-like appendages at each end, 6–7.5 μm long. Sexual morph: Not observed.
Material examined
THAILAND, Chiang Rai Province, stream flowing near Tham Luang Nang Non Cave, on submerged wood, 25 November 2014, Yang Jing (MFLU 15-1168, reference specimen designated here), living culture MFLUCC 15-0055; GUCC 16-0003.
Notes
The production of a synnematous group of conidiophores around a conspicuous polar setulum places our new collection in Menisporopsis. Menisporopsis was introduced with M. theobromae as the type and was found on decaying leaves of Theobroma cacao in Ghana (Hughes 1952). Most Menisporopsis species were originally described as occurring on decaying leaves or wood with a pantropical distribution (Seifert et al. 2011), and M. theobromae is widespread. There are nine species accepted in the genus and keys were provided by Tsui et al. (1999) and Castañeda Ruiz et al. (2001). The newly collected fungus from submerged wood in Thailand is morphologically similar to M. kobensis and M. theobromae. However, its smaller conidia identifies this species as M. theobromae, while the conidia of M. kobensis are 16–32 × 3–5 μm and 14–19 × 2–3 μm in M. theobromae, which is in agreement with the original description by Hughes (1952). We, therefore, designate it as a reference species (sensu Ariyawansa et al. 2014).
The phylogenetic analysis placed the Menisporopsis theobromae isolate close to Rattania, which is a monotypic genus. However, it differs from Rattania by its long synnematous conidiophores and the single seta associated with each synnema, while Rattania has very short conidiophores arranged in a sporodochium with multiple setae. Menisporopsis theobromae also shares similar conidia characters with Dinemasporium and Pseudolachnella, but the latter genera are coelomycetous. The isolate of M. theobromae formed a distinct clade in Fig. 1, and based on both the morphology and molecular phylogeny, this genus is distinct.
References
Arambarri AM, Cabello MN (1989) A numerical taxonomic study of some phialidic genera of hyphomycetes: cluster analysis. Mycotaxon 34:679–696
Ariyawansa HA, Hawksworth DL, Hyde KD, Jones EBG, Maharachchikumbura SSN, Manamgoda DS, Thambugala KM, Udayanga D, Camporesi E, Daranagama A, Jayawardena R, Liu JK, McKenzie EHC, Phookamsak R, Senanayake IC, Shivas RG, Tian Q, Xu JC (2014) Epitypification and neotypification: guidelines with appropriate and inappropriate examples. Fungal Divers 69:57–91
Cai L, Zhang KQ, McKenzie EH, Hyde KD (2003) Freshwater fungi from bamboo and wood submerged in the Liput River in the Philippines. Fungal Divers 13:1–12
Castañeda Ruiz RF, Iturriaga T, Saikawa M, Cano J, Guarro J (2001) The genus Menisporopsis in Venezuela with the addition of M. anisospora anam. sp. nov. from a palm tree. Cryptogamie Mycol 22:259–263
Crous PW, Verkley GJ, Christensen M, Castañeda-Ruiz RF, Groenewald JZ (2012) How important are conidial appendages? Persoonia 28:126–137
Fernández FA, Huhndorf SM (2005) New species of Chaetosphaeria, Melanopsammella and Tainosphaeria gen. nov. from the Americas. Fungal Divers 18:15–57
Fernández FA, Miller AN, Huhndorf SM, Lutzoni FM, Zoller S (2006) Systematics of the genus Chaetosphaeria and its allied genera: morphological and phylogenetic diversity in north temperate and neotropical taxa. Mycologia 98:121–130
Gams W, Holubová-Jechová V (1976) Chloridium and some other dematiaceous hyphomycetes growing on decaying wood. Stud Mycol 13:1–99
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser (Oxf) 41:95–98
Hashimoto A, Sato G, Matsuda T, Hirayama K, Hatakeyama S, Harada Y, Shirouzu T, Tanaka K (2015a) Molecular taxonomy of Dinemasporium and its allied genera. Mycoscience 56:86–101
Hashimoto A, Sato G, Matsuda T, Matsumura M, Hatakeyama S, Harada Y, Ikeda H, Tanaka K (2015b) Taxonomic revision of Pseudolachnea and Pseudolachnella and establishment of Neopseudolachnella and Pseudodinemasporium gen. nov. Mycologia 107:383–408
Hillis DM, Bull JJ (1993) An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Syst Biol 42(2):182–192
Ho WH, Hyde KD, Hodgkiss IJ, Yanna (2001) Fungal communities on submerged wood from streams in Brunei, Hong Kong, and Malaysia. Mycol Res 105:1492–1501
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17(8):754–755
Hughes SJ (1952) Fungi from the Gold Coast I. Mycol Pap 48:1–91
Hyde KD, Fryar S, Tian Q, Bahkali AH, Xu JC (2016) Lignicolous freshwater fungi along a north–south latitudinal gradient in the Asian/Australian region; can we predict the impact of global warming on biodiversity and function? Fungal Ecol 19:190–200
Index Fungorum (2016) Home page at: http://www.indexfungorum.org/names/names.asp
Jayasiri SC, Hyde KD, Ariyawansa HA, Bhat J, Buyck B, Cai L, Dai YC, Abd-Elsalam KA, Ertz D, Hidayat I, Jeewon R, Jones EBG, Bahkali AH, Karunarathna SC, Liu JK, Luangsa-ard JJ, Lumbsch HT, Maharachchikumbura SSN, McKenzie EHC, Moncalvo JM, Ghobad-Nejhad M, Nilsson H, Pang KL, Pereira OL, Phillips AJL, Raspé O, Rollins AW, Romero AI, Etayo J, Selçuk F, Stephenson SL, Suetrong S, Taylor JE, Tsui CKM, Vizzini A, Abdel-Wahab MA, Wen TC, Boonmee S, Dai DQ, Daranagama DA, Dissanayake AJ, Ekanayaka AH, Fryar SC, Hongsanan S, Jayawardena RS, Li WJ, Perera RH, Phookamsak R, Silva NI, Thambugala KM, Tian Q, Wijayawardene NN, Zhao RL, Zhao Q, Kang JC, Promputtha I (2015) The Faces of Fungi database: fungal names linked with morphology, phylogeny and human impacts. Fungal Divers 74:3–18
Kendrick B (1980) The generic concept in Hyphomycetes—a reappraisal. Mycotaxon 11:339–364
Kirk PM, Cannon PF, Minter DW, Stalpers JA (2008) Dictionary of the fungi, 10th edn. CABI Publishers, Wallingford, UK, 771 pp
Krauss GJ, Solé M, Krauss G, Schlosser D, Wesenberg D, Bärlocher F (2011) Fungi in freshwaters: ecology, physiology and biochemical potential. FEMS Microbiol Rev 35(4):620–651
Li DW, Kendrick B, Chen J (2012) Two new hyphomycetes: Codinaea sinensis sp. nov. and Parapleurotheciopsis quercicola sp. nov., and two new records from Quercus phillyraeoides leaf litter. Mycol Prog 11:899–905
Liu JK, Chomnunti P, Cai L, Phookamsak R, Chukeatirote E, Jones EBG, Moslem M, Hyde KD (2010) Phylogeny and morphology of Neodeightonia palmicola sp. nov. from palms. Sydowia 62:261–276
Liu JK, Hyde KD, Jones EBG, Ariyawansa HA, Bhat DJ, Boonmee S, Maharachchikumbura SS, McKenzie EHC, Phookamsak R, Phukhamsakda C, Shenoy BD, Abdel-Wahab MA, Buyck B, Chen J, Chethana KWT, Singtripop C, Dai DQ, Dai YC, Daranagama DA, Dissanayake AJ, Doliom M, Fan LX, Goonasekara D, Hirayama K, Hongsanan S, Jayasiri SC, Jayawardena RS, Karunarathna SC, Li WJ, Mapook A, Norphanphoun C, Pang KL, Perera RH, Peršoh D, Pinruan U, Senanayake IC, Somrithipol S, Satinee S, Tanaka K, Thambugala KM, Tian Q, Tibpromma S, Udayanga D, Wijayawardene NN, Wanasinghe D, Abdel-Aziz FA, Adamcík S, Bahkali AH, Boonyuen N, Bulgakov T, Callac P, Chomnunti P, Greiner K, Hashimoto A, Hofstetter V, Kang JC, Li XH, Liu ZY, Matumura M, Mortimer PE, Rambold R, Randrianjohany E, Sato G, Indrasutdhi VS, Verbeken A, Brackel W, Wang Y, Wen TC, Xu JC, Yan JY, Zhao RL, Camporesi E (2015) Fungal diversity notes 1–110: taxonomic and phylogenetic contributions to fungal species. Fungal Divers 72:1–197
Lumbsch HT, Huhndorf SM (2010) Outline of Ascomycota—2009. Myconet 14:1–64
Magyar D, Shoemaker RA, Bobvos J, Crous PW, Groenewald JZ (2011) Pyrigemmula, a novel hyphomycete genus on grapevine and tree bark. Mycol Prog 10:307–314
Maharachchikumbura SSN, Hyde KD, Jones EG, McKenzie EH, Huang SK, Abdel-Wahab MA, Daranagama DA, Dayarathne M, D’souza MJ, Goonasekar ID, Hongsanan S, Jayawardena RS, Kirk PM, Konta S, Liu JK, Liu ZY, Norphanphoun C, Pang KL, Perera RH, Senanayake IC, Shang QJ, Shenoy BD, Xiao YP, Bahkali AH, Kang JC, Somrothipol S, Suetrong S, Wen TC, Xu JC (2015) Towards a natural classification and backbone tree for Sordariomycetes. Fungal Divers 72:199–301
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In: Proceedings of the Gateway Computing Environments Workshop 2010 (GCE), New Orleans, Louisiana, November 2010, pp 1–8
Nylander JAA (2004) MrModeltest 2.0. Program distributed by the author. Evolutionary Biology Centre, Uppsala University
Page RDM (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Rannala B, Yang Z (1996) Probability distribution of molecular evolutionary trees: a new method of phylogenetic inference. J Mol Evol 43:304–311
Réblová M (2000) The genus Chaetosphaeria and its anamorphs. Stud Mycol 45:149–168
Réblová M, Barr ME, Samuels GJ (1999) Chaetosphaeriaceae, a new family for Chaetosphaeria and its relatives. Sydowia 51:49–70
Réblová M, Winka K (2000) Phylogeny of Chaetosphaeria and its anamorphs based on morphological and molecular data. Mycologia 22:939–954
Seifert KA, Morgan-Jones G, Gams W, Kendrick B (2011) The genera of hyphomycetes. CBS-KNAW Fungal Biodiversity Centre, Utrecht
Silva SS, Gusmão LFP (2013) Conidial fungi from the semi-arid Caatinga Biome of Brazil. A new species of Dictyochaeta. Mycosphere 4(4):701–705
Somrithipol S, Sakayaroj J, Rungjindamai N, Plaingam N, Jones EBG (2008) Phylogenetic relationship of the coelomycete genus Infundibulomyces based on nuclear rDNA data. Mycologia 100:735–741
Stamatakis A (2006) RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Swofford DL (2002) PAUP*. Phylogenetic analysis using parsimony (*and other methods). Version 4. Sinauer Associates, Sunderland, MA
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
Taylor JE, Hyde KD (2003) Microfungi of tropical and temperate palms. Fungal Diversity Press, Hong Kong
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882
Tsui CKM, Goh TK, Hyde KD, Hodgkiss IJ (1999) Reflections on Menisporopsis, with the addition of M. multisetulata sp. nov. from submerged wood in Hong Kong. Mycol Res 103:148–152
Tsui CKM, Hyde KD (2004) Biodiversity of fungi on submerged wood in a stream and its estuary in the Tai Ho Bay, Hong Kong. Fungal Divers 15:205–220
Tulasne LR, Tulasne C (1863) Selecta Fungorum Carpo l (Arambarri and Cabello 1989; Kendrick 1980) ogia: Xylariei- Valsei-Spaeriei. 2
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246
White TJ, Bruns T, Lee SJ, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols: Guide Methods Appl 18:315–322
Whitton SR, McKenzie EHC, Hyde KD (2000) Dictyochaeta and Dictyochaetopsis species from the Pandanaceae. Fungal Divers 4:133–158
Wong MK, Goh TK, Hodgkiss IJ, Hyde KD, Ranghoo VM, Tsui CKM, Ho WH, Wong WS, Yuen TK (1998) Role of fungi in freshwater ecosystems. Biodivers Conserv 7:1187–1206
Zhaxybayeva O, Gogarten JP (2002) Bootstrap, Bayesian probability and maximum likelihood mapping: exploring new tools for comparative genome analyses. BMC Genomics 3(1):4
Acknowledgments
The authors would like to thank the Featured microbial resources and diversity investigation in Southwest Karst area (2014FY120100). J.-K Liu would like to thank the Phylogeny and Biodiversity of Botryosphaeriaceae in Southwest China (QianKeHe LH 2015-7061).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Section editors: Roland Kirschner and Pedro W. Crous
This article is part of the Special Issue “Biodiversity of Hyphomycetes - Special Issue in honor of Dr. Subramanian”.
Rights and permissions
About this article
Cite this article
Liu, JK., Yang, J., Maharachchikumbura, S.S.N. et al. Novel chaetosphaeriaceous hyphomycetes from aquatic habitats. Mycol Progress 15, 1157–1167 (2016). https://doi.org/10.1007/s11557-016-1237-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-016-1237-1