Abstract
Purpose
Ultrasound (US) is a safer alternative to X-rays for bone imaging, and its popularity for orthopedic surgical navigation is growing. Routine use of intraoperative US for navigation requires fast, accurate and automatic alignment of tracked US to preoperative computed tomography (CT) patient models. Our group previously investigated image segmentation and registration to align untracked US to CT of only the partial pelvic anatomy. In this paper, we extend this to study the performance of these previously published techniques over the full pelvis in a tracked framework, to characterize their suitability in more realistic scenarios, along with an additional simplified segmentation method and similarity metric for registration.
Method
We evaluated phase symmetry segmentation, and Gaussian mixture model (GMM) and coherent point drift (CPD) registration methods on a pelvic phantom augmented with human soft tissue images. Additionally, we proposed and evaluated a simplified 3D bone segmentation algorithm we call Shadow–Peak (SP), which uses acoustic shadowing and peak intensities to detect bone surfaces. We paired this with a registration pipeline that optimizes the normalized cross-correlation (NCC) between distance maps of the segmented US–CT images.
Results
SP segmentation combined with the proposed NCC registration successfully aligned tracked US volumes to the preoperative CT model in all trials, in contrast to the other techniques. SP with NCC achieved a median target registration error (TRE) of 2.44 mm (maximum 4.06 mm), when imaging all three anterior pelvic structures, and a mean runtime of 27.3 s. SP segmentation with CPD registration was the next most accurate combination: median TRE of 3.19 mm (maximum 6.07 mm), though a much faster runtime of 4.2 s.
Conclusion
We demonstrate an accurate, automatic image processing pipeline for intraoperative alignment of US–CT over the full pelvis and compare its performance with the state-of-the-art methods. The proposed methods are amenable to clinical implementation due to their high accuracy on realistic data and acceptably low runtimes.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Pelvic fractures require effective surgical treatment in the form of accurate screw fixation. Existing methods for percutaneous screw insertion minimize the invasiveness of the surgery, but require intensive 2D fluoroscopic imaging to visualize the pelvic anatomy and screw trajectories, thus exposing patients and the surgical team to significant amounts of ionizing radiation. A recent study reported radiation exposure per procedure of up to 3 min [1]. Current measures used to protect the surgical team, e.g., in the form of lead aprons, remain cumbersome to use and their significant weight can cause long-term physical injury [2]. Furthermore, 2D X-ray images are difficult to interpret as they only depict projections of the bone structures, making surgical navigation a challenging mental task for the surgeon which can lead to significant errors and re-attempts in screw insertions.
Ultrasound (US) presents a non-ionizing, inexpensive, real-time modality that is readily capable of 3D imaging. In US images, bone is represented by high-intensity pixels at the incident osseous surface, coupled with deeper low intensities which represent acoustic shadows [3]. US-based bone detection for subsequent registration (alignment) to a preoperative patient model for the navigation of surgical tools has been explored for many years. For example, Tonetti et al. investigated the use of tracked US to guide screw insertion in pelvic surgeries as far back as 2001, but found that requiring surgeons to manually segment the bone in US images drastically increased operating times by approximately 25 min [4], which highlights the need for fast and automatic segmentation. However, given that US images are notorious for their poor signal-to-noise ratio and limited field of view of the desired anatomy, automatic segmentation and subsequent registration remained challenging.
A number of previous works aimed to address some of these challenges. In 2002, an intensity-based registration technique was proposed for surgical guidance of a lumbar spine procedure, but it required manual US segmentation [5]. US–CT intensity and surface-based registration techniques for the pelvis have been proposed [6, 7], but they required initialization through manual landmark identification for successful optimization, which slows or disrupts the surgical workflow. Statistical shape models (SSMs) were implemented for multimodal bone registration [8, 9]; however, these approaches are computationally expensive, requiring significant training data, and they are best suited to modeling healthy, non-fractured bones similar to those in the databases used in creating the SSMs. Our group previously proposed the phase symmetry bone segmentation technique, based on locally symmetric phase features to enable accurate US bone segmentation [10, 11] including an approach where model parameters were automatically optimized using data-driven approaches [12].
Phase features have also been used to optimize registration between US volumes and a CT model [13], but the authors report a runtime of approximately 6 min per volume which may be clinically infeasible. Point-based registration between phase symmetry-processed US and CT volumes has also been proposed [14,15,16], achieving submillimetric TRE and near real-time execution on restricted pelvic landmarks such as the iliac spine. Simulating US from preoperative CT intensity information was proposed to aid registration [17] and has been evaluated within a computer-assisted orthopedic surgery (CAOS) workflow on cadaver models of the femur, tibia and fibula, achieving a median target registration error of 3.7 mm, but a large maximum error of 22.7 mm [18]. Recently, machine learning methods have become popular and successful at automatic ultrasound bone segmentation. A patch-based random forest classifier and a U-net convolutional neural network (CNN) architecture were trained to detect bone in US images of lumbar vertebrae, achieving 88% recall and 94% precision with manual segmentation as the gold standard. Another CNN-based approach for bone segmentation and subsequent point surface registration was investigated and evaluated on femur, tibia and pelvis cadaver data, achieving a median TRE of 2.76 mm on the pelvis [19]. It remains to be explored how well the trained algorithms generalize to, for example, fractured bone images or data acquired with different imaging parameters [20]. Additionally, the learning process requires expensive computation on hundreds or thousands of training US images which may be difficult to acquire.
Despite the above advances, there remains an unmet need for automatic, fast and accurate registration of tracked US to a full-pelvis CT reference, to enable orthopedic surgical guidance. Our group previously explored the effectiveness of phase-based features for US bone segmentation [21] and point set registration [16, 22, 23] to align untracked US images to CT of partial pelvic models. In this paper, our goal is to assess the performance of these previously published methods on a full pelvic model with tracked US, to reflect a more clinically realistic setting. To this end, we also propose and evaluate a simplified 3D bone segmentation method based on our group’s and others’ previous work in bone shadow detection [24, 25], and pair this with a rigid US–CT volume registration framework to address the inherent challenges of ultrasound-based guidance. The proposed method is designed with the aim of practical integration into the clinical workflow where fast performance on standard hardware is typically necessary, without the need to learn on multiple annotated example images. Comparisons with current approaches suggest that our simplified methods, which are fully automated and do not require manual initialization, are capable of achieving clinically acceptable results over the complete pelvic anatomy.
Methods
Our approach to aligning tracked US to a preoperative segmented CT model is broken into two steps. We first segment bone surfaces from the spatially compounded US volume and then register this segmentation to the CT bone model. Segmentation is performed using the previously published phase symmetry segmentation technique and also our proposed Shadow–Peak method described below. Registration is performed using Gaussian mixture model and coherent point drift techniques. We also register using our proposed pipeline based on normalized cross-correlation between US and CT distance maps, described below.
Bone segmentation
Shadow–Peak segmentation
Here, we present a simplified 3D US bone segmentation technique that is guided by the physical characteristics of bone. Specifically, bone surfaces in US cast a deep shadow in the direction of the acoustic beam, and bone surfaces themselves appear brighter when the surface is more perpendicular to the acoustic beam [3]. Our segmentation technique extracts these features from US volumes.
We first normalize the intensities of the input B-mode US volume and then apply a 3D Gaussian filter (isotropic standard deviation of 1 voxel). We calculate, for each voxel, a measure of confidence that it represents a shadow:
where \(I_{x,y,z}\) represents the Gaussian filtered voxel intensity, x, y, z are the voxel coordinates (column, row, frame), and Y is the maximum y coordinate. The bone surface position in each scanline is then estimated by finding the peak intensity location of the shadow confidence map modulated by the original raw voxel intensities. The confidence of each estimate is given by the peak intensity value. Any confidence values that fall below half the standard deviation (SD) of the mean bone confidence are discarded, as they are likely to be false positives.
Connected-component analysis To reduce false positive bone surfaces after initial segmentation, we perform a 3D connected-component analysis to remove small isolated segmentations. Our assumptions in this process are that the locations of any bone fractures are remote from the US scanning site, and that non-displaced bone fractures are not imaged as discontinuities in the US image. We empirically set the minimum number of allowable connected voxels in each connected component to be 5% of the total number of connected voxels, smaller components are likely not to represent true bone surfaces. This relatively low threshold does not penalize large bone surfaces that may appear separate in the reconstructed volume, for instance, the left and right iliac spines. Figure 1 illustrates our complete segmentation pipeline. We call this algorithm Shadow–Peak (SP) segmentation.
Preoperative CT segmentation We presume that we have available a preoperatively acquired CT volume. We preprocess this volume by intensity thresholding and manually removing responses not corresponding to bone. We then segment the superficial CT bone surfaces by simple vertical tracing. The CT volume is also truncated 60 mm below the skin surface, as elements below this depth will not be visible in US due to the maximum US imaging depth.
Phase symmetry segmentation parameters
We compare Shadow–Peak with 3D phase symmetry (PS) for US bone segmentation techniques [21]. Following the notation in the original paper, we set the model parameters constant to \(m=1, \alpha =6, \lambda _\mathrm{min} = 20, \delta = 3, \sigma _\mathrm{alpha} = 0.45^{\circ }, \kappa = 3\) as recommended by the author. We enhance the signal-to-noise ratio of PS segmentation using intensity thresholding and bottom-up ray casting as described in [26].
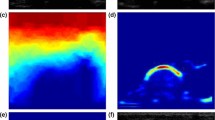
Qualitative comparison between Shadow–Peak (SP) and phase symmetry (PS) segmentation. a 2D slice from a 3D US volume of the human radius and ulna. b Shadow confidence map of a, given by Eq. 1. Higher intensities represent a higher confidence in shadow presence. c Final SP segmentation after peak detection, population thresholding and connected-component analysis. d PS analysis of a [21]. e Final PS segmentation after thresholding, ray casting and connected-component analysis [26]
Left: Distance map nonlinearity with \(0.01 \le \varvec{\Gamma } \le 0.05\) described in section “Normalized cross-correlation (NCC) registration of 3D distance maps” Right: Idealized contour segmentation in white and the effect of the inverse nonlinearity in magenta. Note the larger spread of intensities at the top of the image
Multimodal registration
Normalized cross-correlation (NCC) registration of 3D distance maps
Here, we present our volume registration pipeline. After segmentation is complete (either with SP or phase symmetry), the binary segmentations from US and CT are transformed into Euclidean distance maps through a 3D distance transform [27]. The distance maps are inverted through an exponential nonlinearity: \((I_\mathrm{dist}+\varvec{\Gamma })^{-2}\) where \(I_\mathrm{dist}\) represents the 3D Euclidean distance map. This nonlinearity assigns a maximum intensity at the segmented bone locations while providing a sharp drop-off in intensity away from the surface. The scaling parameter \(\varvec{\Gamma }\) is used to control the rate of intensity drop-off. It is varied linearly from 0.05 to 0.01 along the depth of the US and CT distance maps; see Fig. 2. We find that this significantly improves the registration accuracy, as it provides a larger weighting to more superficial bone surfaces, which are less likely to suffer from speed-of-sound distortions and hence their location is more certain than deeper bone structures.
The volumes are automatically aligned by their centers of gravity and along nominal anatomical directions prior to running the automatic registration algorithm. Rigid registration between the inverted distance maps of US and CT volumes is then performed by maximizing their normalized cross-correlation (NCC), defined as:
where \(\varvec{\mu }\) represents the parameters of the rigid transformation \({\varvec{T}}\). Stochastic gradient descent is used to efficiently converge to the NCC global maximum, and 1000 intensities are sampled at random coordinates over the CT domain to estimate the similarity metric at each iteration. The registration runs through five down-sampling and smoothing levels to iteratively obtain finer transformation estimates.
Gaussian mixture model (GMM) point set registration
We implement and evaluate GMM registration as described for US–CT registration of the pelvic anatomy [16]. We test this algorithm with 500 and 1000 particle-simulated points in both US and CT point clouds, as suggested in the original paper. We also evaluate the effect of curvature features on registration accuracy by inserting an additional 10% of points in areas of high curvature [16].
Coherent point drift (CPD) point set registration
CPD is another popular point set registration framework that has been used for multimodal alignment. We implement the original algorithm [22], also with 500 and 1000 points in both US (moving) and CT (fixed) point clouds. Points are randomly and uniformly sampled from the US and CT images of the pelvis, and we do not use particle simulation as with GMM registration.
a Pelvic phantom under construction. N.B. After construction, the pelvis was completely covered by the agar medium. b Example of bone surfaces scanned with tracked US (green) overlaid on the preoperative CT model (gray) when only the iliac spines were imaged, and c when the pubic symphysis was also imaged
Validation experiment
Phantom design To enable a controlled, quantitative evaluation of all combinations of the segmentation and registration techniques, we constructed a pelvis phantom consisting of a full pelvic bone model (Sawbones Inc, Vashon Island, WA, USA) embedded in a 5% (weight/weight) agar and water solution in an anatomically supine position. Agar was used to model the general acoustic response of soft tissue as it creates similar scattering and speckle patterns, and its acoustic velocity is similar to room temperature water [28]. A full pelvis was used, instead of a hemi-pelvis, as this more accurately captures the surgical scenario where US data have to be automatically registered to the complete pelvic CT model. This design choice allowed us to assess automatic registration performance when the intraoperative US bone surface is a fractionally small representation of the preoperative CT model. Forty-eight steel beads, of 2 mm diameter each, were embedded uniformly around the iliac spines and the pubic symphysis for subsequent evaluation of registration errors (Fig. 3).
Imaging We performed a CT scan of the phantom with a GE Healthcare CT750HD scanner (GE Healthcare, Chicago, IL, USA) to generate the reference CT volume, with a voxel size of 0.71\(\times \)0.71\(\times \)0.63 mm. This high resolution is also used clinically to assess pelvic fracture cases. For ultrasound imaging, we used a SonixTouch Q+ ultrasound machine with a 4DL14-5/38 linear transducer (BK Ultrasound, Peabody, MA, USA) imaging at 10 MHz, and tracked by an optical camera as described below. The transducer motor was fixed at a neutral 0\(^{\circ }\) and a maximum scanning depth of 60 mm. We collected repeated scans of the right and left iliac spines and the pubic symphysis of the pelvis, as they are accessible superficial bony structures (see Fig. 3). A total of 24 reconstructed US volumes were collected from the phantom, loosely aligned to anatomical axes. Twelve volumes contained scans from only the right and left iliac spines (“two-view” volumes) and the remaining 12 contained all three anatomical structures (both iliac spines and the pubic symphysis—“three-view” volumes). We collected these two groups of volumes to assess the robustness of the segmentation and registration techniques in cases where scanned data might be incomplete. Each US volume represented a region measuring approximately \(30\times 10\times 21\) cm (right/left, anterior/posterior, superior/inferior axes).
Soft tissue enhancement Because soft tissue structures were not directly incorporated in the agar phantom, we enhanced the phantom’s realism by re-sampling and overlaying soft tissue US images taken from the pelvic region of a human volunteer. Augmenting the phantom with human soft tissue images created a more realistic scenario to evaluate the segmentation and registration algorithms.
a Five examples of axial slices taken from the pelvis phantom with overlays of soft tissue images. The visible bony structures represent the right/left iliac spines and the pubic symphysis. b Application of Shadow–Peak segmentation and the inverse distance transform defined in section “Normalized cross-correlation (NCC) registration of 3D distance maps”
To do this, we collected three US volumes (using the imaging parameters described above) from anterior pelvic regions in a volunteer. These images were acquired from near the bony regions corresponding to the phantom’s design, but we ensured these images contained only soft tissue and no bony structures. The images were then automatically transformed to the phantom’s domain using nonrigid image registration. The nonrigid registration was performed between axial slices from the phantom and soft tissue images. From the phantom images, we extracted bone and shadow and overlaid the transformed soft tissue images using pixel-wise addition and intensity normalization. Figure 4 illustrates the original phantom data and the enhanced data. All segmentation and registration evaluations were performed on the soft-tissue-enhanced phantom data.
Tracking The US transducer and phantom were tracked with rigid passive markers using an infrared Polaris camera (Northern Digital Inc., Waterloo, ON, Canada; manufacturer reported accuracy of 0.35 mm RMS) at an acquisition rate of 50 Hz.
Implementation US images and optical tracking data were acquired, synchronized and spatially reconstructed using the open-source PLUS toolkit [29], 3D Slicer [30] and SlicerIGT [31]. Our segmentation technique was implemented in MATLAB 2017a (The MathWorks, Inc., Natick, MA, USA) and integrated into the open-source architecture using the “Matlab Bridge” Slicer module. Furthermore, we used the open-source registration framework “elastix” and the associated “SlicerElastix” extension to integrate our volume registration technique into the same architecture [32].
Before segmentation, the CT reference and US volumes were resampled to an isotropic pixel size of 0.8 mm, represented by 382\(\times \)214\(\times \)297 pixels for the CT model and by approximately 380\(\times \)110\(\times \)280 pixels for the US volumes. Each US volume was then segmented using either Shadow–Peak or phase symmetry—Fig. 5 illustrates a few segmented soft-tissue-enhanced images using Shadow–Peak. The CT volume was segmented as described in section “Shadow–Peak segmentation,” under the assumption that the relevant bone surfaces can be detected in the vertical direction, as the phantom has a single flat face that the US probe can access. With human specimens, this step would have to be modified to account for the additional directions through which the US probe could image the pelvis, which could be simply achieved by defining the normal to the skin surface in the preoperative CT model or transformation through a number of predefined angles. Each US–CT segmentation pair was tested with the three aforementioned registration techniques: GMM, CPD and NCC. A total of 14 segmentation and registration combinations were tested on each volume pair, when accounting for the variation in point cloud size and presence or absence of curvature features.
Evaluation The performance of Shadow–Peak and phase symmetry segmentation was characterized using the same recall and precision metrics proposed by Baka [33], which take into account the difficulty of comparing open contour segmentations (due to the thinness of the surfaces). Recall was measured as the percentage of manually segmented (considered as ground truth) pixels in a volume that were within a 1-mm vertical zone of the automatic segmentation. Precision was defined as the proportion of automatically segmented pixels that were within a 2-mm boundary of the manual segmentation. We also calculated the F-measure for each 3D segmentation, which is the harmonic mean between the recall and precision and is one method of evaluating the accuracy of a technique. The F-measure is defined as:
These metrics were analyzed with reference to the manual segmentations of 16 soft-tissue-enhanced US volumes taken from the phantom. The manual segmentations were performed in a repeatable manner by fixed intensity thresholding and removing false positives not corresponding to true bone surfaces.
To evaluate the registration performance, each US volume was first aligned to the CT model using steel fiducial markers (between 22 and 37 fiducials) to calculate the residual fiducial registration error (FRE), which can be taken to be an estimate of the lower bound on the target registration error. The performance of the automatic techniques was measured by the target registration error (TRE), defined as the root-mean-square error between the embedded fiducial points visible in US and CT. We also measured the surface registration error (SRE), defined as the root-mean-square error from the manually segmented US bone surface to the CT bone surface. The maximum nearest-neighbor surface discrepancy, i.e., the Hausdorff distance (HD), was also measured with the same directionality as for the SRE calculation. For all combinations of segmentation and registration techniques, a successful registration was defined as one where the resulting TRE was less than 5 mm, after subtracting the residual FRE—this threshold was based on the typical geometry of the tunnel which sacroiliac screws normally pass [34]. Note that the fiducial points and manually delineated surfaces were only used for evaluating errors and were not used in the automatic segmentation or registration techniques. The execution time of each segmentation and registration component was also measured.
Results
Segmentation accuracy Our analysis found that Shadow–Peak segmentation achieved a lower false positive rate, with a higher mean precision of 53.7% (SD 9.2%) compared to 41.3% (SD 5.7%) for phase symmetry. However, Shadow–Peak had a lower recall than phase symmetry: 76.7% (SD 8.4%) compared to 81.0% (SD 6.9%). Overall Shadow–Peak had a greater mean F-measure 62.6% (SD 7.3%) compared to phase symmetry which had a F-measure of 54.3% (SD 5.1%). These results are presented in Table 1.
Fiducial alignment for ground truth registration errors The mean FRE after fiducial-based manual alignment of the dataset was 1.54 mm (SD 0.09 mm), and the corresponding mean SRE was 1.62 mm (SD 0.22 mm). The mean HD was 7.93 mm (SD 0.87 mm), and the point of maximum distance was typically located on the deeper surfaces of the ilium. The initial preregistration TREs ranged between 27.03 and 88.52 mm.
Registration errors Figure 6 summarizes the total success rate for all segmentation and registration combinations on the dataset. Both CPD and NCC registrations had a success rate greater than 75% when SP segmentation was used, and their success rates with phase symmetry were lower. The combination of SP segmentation with NCC registration was the only one with a 100% success rate, even when only two anatomical views were imaged with US. Conversely, GMM registration did not surpass a 40% success rate in any scenario. Figure 7 plots the final TRE for successful segmentation and registration combinations. Overall, the mean TRE of SP and NCC was 3.22 mm (SD 1.03 mm). SP segmentation combined with 1000-point CPD registration achieved a mean TRE of 4.58 mm (SD 0.89 mm), and 3.89 mm (SD 1.27 mm) when only 500 points were used. In general, registrations preprocessed with phase symmetry segmentation resulted in higher registration errors—NCC 4.56 mm (SD 0.77 mm), 1000-CPD 4.39 mm (SD 0.75 mm), 500-CPD 4.15 mm (SD 1.07 mm). With only two views visible, SP with NCC had a mean TRE of 3.79 mm (SD 1.06 mm) compared to 2.65 mm (SD 0.62 mm) when both iliac spines and pubic symphysis were visible, which was the lowest mean error achieved by any combination of techniques.
SP segmentation with NCC registration achieved the lowest surface fit errors with a mean SRE of 1.35 mm (SD 0.17 mm), and SP combined with 500-point CPD and 1000-point CPD achieved mean SREs of 1.84 mm (SD 0.58 mm) and 1.99 mm (SD 0.44 mm), respectively. Phase symmetry segmentation with NCC registration achieved a mean SRE of 1.44 mm (SD 0.26 mm). The surface fit errors are summarized in Table 2.
Success rate (percentage) of all segmentation and registration combinations on 24 US–CT volume pairs, broken down by anatomical views visible in US. GMM—Gaussian mixture model registration with 500/1000 points; CPD—coherent point drift registration with 500/1000 points; NCC—normalized cross-correlation registration
Computational performance All experiments were conducted on an Intel Xeon E5-2630 v3 CPU @2.40GHz with 16GB of random access memory (RAM). SP segmentation took an average of 1.80s (SD 0.57s) to process an uncropped reconstructed US volume, and the proposed registration via NCC took an average of 25.98s (SD 2.48s). In contrast, phase symmetry segmentation took an average of 36.50s (SD 5.24s) to compute, and 500- and 1000-point CPD-based registrations took an average of 3.14s (SD 1.37s) and 10.62s (SD 3.73s), respectively. On average, GMM registration took longer than CPD to execute when combined with particle simulation techniques as in [16].
Discussion and conclusions
In this study, we evaluated the suitability of three previously published segmentation and registration techniques for aligning realistic intraoperative US to a full pelvic CT model. We also proposed and developed a simplified segmentation workflow based on SP segmentation and NCC registration. Overall, SP segmentation was more accurate than phase symmetry and resulted in more robust registration performance with the proposed NCC pipeline, compared to GMM or CPD registration.
The proposed SP and NCC combination resulted in a statistically lower mean TRE than any of the other successful combinations, with a significance level of p = 0.035 using a one-tailed, two-sample unequal variance t-test. On the other hand, CPD performs on average 15.8s quicker than NCC-based registration (p< 0.001) which makes it more attractive for clinical implementation. We also investigated the effect of increasing point cloud size beyond 1000 points in CPD and GMM registration, and found that this did not correlate with lower registration errors.
We were surprised to find that overall GMM registration was largely unsuccessful. Furthermore, we found that using curvature features on this dataset did not improve the registration accuracy as previously reported [16]. We further investigated the performance of GMM registration by manually reducing the CT field of view to more closely match the US field of view. On visual inspection, we found that the success of GMM registration depended on the fixed and moving point clouds sharing a very similar field of view and point cloud representation. This makes it less suitable for a clinical setting where there is no guarantee that the tracked US images would contain a similar surface representation to the CT preoperative model.
Our proposed segmentation and registration approach had similar target registration accuracy to other recently reported methods for US–CT registration. Wein et al. achieved a median TRE of 3.7 mm on cadaveric data [18], and Salehi et al. achieved a median TRE of 2.76 mm on cadaveric pelvis data when using a deep neural network to perform bone segmentation [19]. When considering only segmentation accuracy (i.e., without a corresponding registration step), random forest-based segmentation and a U-net deep neural architecture produced higher F-measures than SP or phase symmetry, with scores of 0.83 and 0.90, respectively, based on tests on 2D untracked lumbar vertebrae images reported in [33]. However, the generalizability of these learning algorithms on data collected with different imaging parameters remains to be tested. Moreover, SP segmentation runs in real-time without the need for high-performance hardware such as GPUs or multiple CPUs, unlike the learning-based methods. Similar to previous studies, we found that the integration of shadowing features in SP segmentation greatly improved noise rejection from surrounding soft tissue structures, leading to less noisy point cloud and distance map generation [24, 25]. However, our workflow simplified the shadow detection calculation, helping facilitate real-time operation on a standard CPU (approximately 150 frames/second for US volumes). Additionally, by performing a contour segmentation of the preoperative CT model, we optimized registration using the NCC similarity metric instead of more computationally expensive multimodal metrics like mutual information [35].
It is worth noting that a proportion of the TRE may be attributed to the relatively large errors in fiducial localization, limited tracking accuracy and speed-of-sound discrepancies between the phantom’s medium and what is expected by the US system. These errors contribute to the residual FRE when US and CT volumes are aligned using fiducials, and are also reflected in the relatively high SRE and HD after fiducial-based alignment. Although our NCC pipeline accounts for deeper bone surface localization uncertainty, by assigning higher weights to superficial surfaces using the inverse distance map, we expect further improvements in accuracy if speed-of-sound calibration is integrated into the workflow, e.g., as in [19]. Furthermore, since the pelvis represents a relatively large anatomical region, a more accurate tracking system with more consistent accuracy over that region may help further drive down the registration error.
Our future work will focus on validating our approaches on clinical ex vivo and in vivo data, as our investigation provided promising results in the presence of human soft tissue structures such as muscle, fat, vessels and nerves. We also plan to integrate speed-of-sound calibration. Finally, it is important to test the feasibility of applying our methods to more complex cases of displaced pelvic fractures.
References
Ecker T, Jost J, Cullmann J, Zech W, Djonov V, Keel M, Benneker L, Bastian J (2017) Percutaneous screw fixation of the iliosacral joint: a case-based preoperative planning approach reduces operating time and radiation exposure. Injury 48(8):1825–1830
Schueler BA (2010) Operator shielding: how and why. Tech Vasc Interv Radiol 13(3):167–171
Daanen V, Tonetti J, Troccaz J (2004) A fully automated method for the delineation of osseous interface in ultrasound images. In: Medical image computing and computer-assisted intervention—MICCAI 2004, 3216, pp 549–557
Tonetti J, Carrat L, Blendea S, Merloz P, Troccaz J, Lavalle S, Chirossel JP (2001) Clinical results of percutaneous pelvic surgery: computer assisted surgery using ultrasound compared to standard fluoroscopy. Comput Aided Surg 6(4):204–211
Brendel B, Rick SWA, Stockheim M, Ermert H (2002) Registration of 3D CT and ultrasound datasets of the spine using bone structures. Comput Aided Surg 7(3):146–155
Penney G, Barratt D, Chan C, Slomczykowski M, Carter T, Edwards P, Hawkes D (2006) Cadaver validation of intensity-based ultrasound to CT registration. Med Image Anal 10(3):385–395
Barratt D, Penney G, Chan C, Slomczykowski M, Carter T, Edwards P, Hawkes D (2006) Self-calibrating 3D-ultrasound-based bone registration for minimally invasive orthopedic surgery. IEEE Trans Med Imaging 25(3):312–323
Ghanavati S, Mousavi P, Fichtinger G, Foroughi P, Abolmaesumi P (2010) Multi-slice to volume registration of ultrasound data to a statistical atlas of human pelvis. In: Proceedings of SPIE, vol 7625, p 7625–7625–10
Ghanavati S, Mousavi P, Fichtinger G, Abolmaesumi P (2011) Phantom validation for ultrasound to statistical shape model registration of human pelvis. In: Proceedings of SPIE, vol 7964, pp 7964–7964–8
Hacihaliloglu I, Abugharbieh R, Hodgson AJ, Rohling RN (2009) Bone surface localization in ultrasound using image phase-based features. Ultrasound Med Biol 35(9):1475–1487
Hacihaliloglu I, Abugharbieh R, Hodgson AJ, Rohling RN, Guy P (2012) Automatic bone localization and fracture detection from volumetric ultrasound images using 3-D local phase features. Ultrasound Med Biol 38(1):128–144
Hacihaliloglu I, Abugharbieh R, Hodgson AJ, Rohling RN (2011) Automatic adaptive parameterization in local phase feature-based bone segmentation in ultrasound. Ultrasound Med Biol 37(10):1689–1703
Hacihaliloglu I, Wilson DR, Gilbart M, Hunt MA, Abolmaesumi P (2013) Non-iterative partial view 3D ultrasound to CT registration in ultrasound-guided computer-assisted orthopedic surgery. Int J Comput Assist Radiol Surg 8(2):157–168
Brounstein A, Hacihaliloglu I, Guy P, Hodgson A, Abugharbieh R (2011) Towards real-time 3D US to CT bone image registration using phase and curvature feature based GMM matching. In: Medical image computing and computer intervention—MICCAI 2011, vol 14, pp 235–242
Hacihaliloglu I, Brounstein A, Guy P, Hodgson A, Abugharbieh R (2012) 3D ultrasound-CT registration in orthopaedic trauma using GMM registration with optimized particle simulation-based data reduction. In: Medical image computing and computer-assisted intervention—MICCAI 2012, vol 15, pp 82–89
Brounstein A, Hacihaliloglu I, Guy P, Hodgson A, Abugharbieh R (2015) Fast and accurate data extraction for near real-time registration of 3-D ultrasound and computed tomography in orthopedic surgery. Ultrasound Med Biol 41(12):3194–3204
Wein W, Brunke S, Khamene A, Callstrom MR, Navab N (2008) Automatic CT-ultrasound registration for diagnostic imaging and image-guided intervention. Med Image Anal 12(5):577–585
Wein W, Karamalis A, Baumgartner A, Navab N (2015) Automatic bone detection and soft tissue aware ultrasound–CT registration for computer-aided orthopedic surgery. Int J Comput Assist Radiol Surg 10(6):971–979
Salehi M, Prevost R, Moctezuma JL, Navab N, Wein W (2017) Precise ultrasound bone registration with learning-based segmentation and speed of sound calibration. In: Descoteaux M, Maier-Hein L, Franz A, Jannin P, Collins DL, Duchesne S (eds) Medical Image Computing and Computer-assisted intervention—MICCAI 2017. Springer, Cham, pp 682–690
Brattain LJ, Telfer BA, Dhyani M, Grajo JR, Samir AE (2018) Machine learning for medical ultrasound: status, methods, and future opportunities. Abdom Radiol 43(4):786–799
Hacihaliloglu I, Abugharbieh R, Hodgson A, Rohling R (2008) Bone segmentation and fracture detection in ultrasound using 3d local phase features. In: Medical image computing and computer-assisted intervention—MICCAI 2008. Springer, Berlin, pp 287–295
Myronenko A, Song X (2010) Point set registration: coherent point drift. IEEE Trans Pattern Anal Mach Intell 32(12):2262–2275
Pandey P, Abugharbieh R, Hodgson AJ (2017) Trackerless 3d ultrasound stitching for computer-assisted orthopaedic surgery and pelvic fractures. In: CAOS 2017, EasyChair, EPiC Series in Health Sciences, vol 1, pp 318–321
Quader N, Hodgson A, Abugharbieh R (2014) Confidence weighted local phase features for robust bone surface segmentation in ultrasound. In: MICCAI workshop on clinical image-based procedures, pp 76–83
Karamalis A, Wein W, Klein T, Navab N (2012) Ultrasound confidence maps using random walks. Med Image Anal 16(6):1101–1112
Hacihaliloglu I, Guy P, Hodgson AJ, Abugharbieh R (2015) Automatic extraction of bone surfaces from 3D ultrasound images in orthopaedic trauma cases. Int J Comput Assist Radiol Surg 10(8):1279–1287
Maurer C, Qi Rensheng, Raghavan V (2003) A linear time algorithm for computing exact Euclidean distance transforms of binary images in arbitrary dimensions. IEEE Trans Pattern Anal Mach Intell 25(2):265–270
Madsen EL, Hobson MA, Shi H, Varghese T, Frank GR (2005) Tissue-mimicking agar/gelatin materials for use in heterogeneous elastography phantoms. Phys Med Biol 50(23):5597–5618
Lasso A, Heffter T, Rankin A, Pinter C, Ungi T, Fichtinger G (2014) PLUS: open-source toolkit for ultrasound-guided intervention systems. IEEE Trans Biomed Eng 61(10):2527–37
Kikinis R, Pieper SD, Vosburgh KG (2014) 3D slicer: a platform for subject-specific image analysis, visualization, and clinical support. Springer, New York, pp 277–289
Ungi T, Lasso A, Fichtinger G (2016) Open-source platforms for navigated image-guided interventions. Med Image Anal 33:181–186
Klein S, Staring M, Murphy K, Viergever MA, Pluim JPW (2010) Elastix: a toolbox for intensity-based medical image registration. IEEE Trans Med Imaging 29(1):196–205
Baka N, Leenstra S, van Walsum T (2017) Ultrasound aided vertebral level localization for lumbar surgery. IEEE Trans Med Imaging 36(10):1–1
Bates P, Starr A, Reinert C (2010) The percutaneous treatment of pelvic and acetabular fractures. Independent Publisher
Shams R, Sadeghi P, Kennedy R, Hartley R (2010) A survey of medical image registration on multicore and the GPU. IEEE Signal Process Mag 27(2):50–60
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was funded by the Natural Sciences and Engineering Research Council (Grant Number: CHRP 478466-15) and the Canadian Institutes of Health Research (Grant Number: CPG-140180).
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
This article does not contain patient data.
Rights and permissions
About this article
Cite this article
Pandey, P., Guy, P., Hodgson, A.J. et al. Fast and automatic bone segmentation and registration of 3D ultrasound to CT for the full pelvic anatomy: a comparative study. Int J CARS 13, 1515–1524 (2018). https://doi.org/10.1007/s11548-018-1788-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-018-1788-5