Abstract
Deep convolutional neural network (DCNN) models have been widely used to diagnose skin lesions, and some of them have achieved diagnostic results comparable to or even better than dermatologists. Most publicly available skin lesion datasets used to train DCNN were dermoscopic images. Expensive dermoscopic equipment is rarely available in rural clinics or small hospitals in remote areas. Therefore, it is of great significance to rely on clinical images for computer-aided diagnosis of skin lesions. This paper proposes an improved dual-branch fusion network called CR-Conformer. It integrates a DCNN branch that can effectively extract local features and a Transformer branch that can extract global features to capture more valuable features in clinical skin lesion images. In addition, we improved the DCNN branch to extract enhanced features in four directions through the convolutional rotation operation, further improving the classification performance of clinical skin lesion images. To verify the effectiveness of our proposed method, we conducted comprehensive tests on a private dataset named XJUSL, which contains ten types of clinical skin lesions. The test results indicate that our proposed method reduced the number of parameters by 11.17 M and improved the accuracy of clinical skin lesion image classification by 1.08%. It has the potential to realize automatic diagnosis of skin lesions in mobile devices.
Graphical Abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Skin cancer is one of the most common cancers in the world, among which melanoma has a very high fatality rate and poses a massive threat to people’s life. Its incidence has been increasing in recent years [1]. Typically, dermatologists rely on their own vision of a patient’s dermoscopic image or clinical skin biopsy to diagnose skin cancer. Dermoscopy imaging requires related equipment, and clinical skin biopsy requires patients to visit the corresponding dermatologist [2]. However, in some rural clinics or small hospitals in remote areas, due to a lack of expensive dermoscopic equipment and enough dermatologists, patients in the area cannot diagnose skin cancer in time, resulting in increased morbidity and mortality from melanoma [3]. Computer-aided diagnosis (CAD) technology based on machine learning and deep learning is a breakthrough in cancer detection [4]. Despite the lack of dermoscopic equipment and dermatologists in rural communities, the application of CAD to clinical skin lesion image classification enables local patients to self-detect the category of skin lesions and reduce the increased risk of death due to early undetected melanoma [5].
Early CAD techniques for dermoscopic image classification often relied on extracting hand-crafted features fed into traditional classifiers [6, 7]. In recent years, automatic skin cancer classification performance has been significantly improved using end-to-end training of deep convolutional neural networks (DCNN) [8,9,10,11,12]. Most of the existing DCNN-based methods use transfer learning methods [13]. For example, Kawahara et al. proposed a DCNN architecture using a pre-training model for skin lesion classification [14]. Esteva A et al. adopted the transfer learning method to fine-tune the network model based on Inception V3 and then trained the network model end-to-end to classify three skin lesions [11]. They only simply transfer DCNN models such as Deep Convolutional Network (VGGNet) [15] and Residual Network (ResNet) [16] to the task of skin lesion classification and has plenty of room for performance improvement.
In the ISIC skin lesions analysis challenge, the self-attention mechanism has been widely used in DCNN. Hu et al. [17] focused on channel relationships and proposed a new channel attention unit, called the squeeze and excitation module, which can accurately calibrate feature channels. Following this idea, Gessert et al. [8] exploited patch-based attention to aggregate contextual information. In contrast to most studies using channel attention, some studies use spatial attention to explore stochastic spatial dependencies. Zhang et al. proposed a spatial attention-based network, called Attention Residual Learning Convolutional Neural Network, for skin lesion classification [18]. Zenghui et al. proposed a dual-attention-based network for skin lesion classification with auxiliary learning [19], which utilized channel and spatial attention to improving network performance, combined with an auxiliary learning module to further focus on local features in skin lesions.
However, previous studies only used dermoscopic images to classify skin lesions. In rural clinics or small hospitals located in remote areas, dermoscopy equipment is often not available, making it important to consider computer-aided diagnosis of skin diseases using clinical images of lesions. To leverage the information contained in clinical images, most research has focused on the multi-modal domain. Yap et al. [20] utilized the ResNet-50 architecture to extract representations from both clinical and dermoscopic images, combining them with metadata representations for final classification. Kawahara et al. [21] attempted different combinations of clinical, dermoscopic, and metadata modalities using the Inception-V3 to find optimal performance. Bi et al. [22] proposed a hyper-connected convolutional neural network that connected representations from clinical and dermoscopic modalities for classification. Ge et al. [23] proposed a three-branch CNN architecture that extracted representations from clinical images, dermoscopic images, and their combination. Wang et al. [24] introduced an adversarial multi-modal fusion approach with attention mechanism (AMFAM), considering both the relevant and complementary information between clinical and dermoscopic modalities. Such multi-modal research requires the collection of more patient information, while clinical images of lesions can be easily obtained using digital cameras or smartphones.
In addition, Transformer was used early to solve problems in the natural language processing field. Recently, ViT was proposed to handle Transformer-based image recognition tasks [25]. It splits the image into several non-overlapping patches, then utilizes Transformer to calculate the global information between each token and adds an additional token for image recognition tasks. In addition, there are some variants of Transformer. For example, Swin Transformer uses shifted windows to compute local self-attention [26]. PVT combines a feature pyramid network with Transformer to capture features from multiple stages [27]. It has great potential as an alternative to the DCNN and has been validated by pre-training with a larger dataset with more than 12 million images [25, 26]. However, even without human annotation, the number of available skin lesion images is very limited in the skin lesion classification tasks. So sufficient skin lesion images cannot be collected to pre-train a robust initial model for skin lesion classification. Therefore, despite the great success of Transformer in general image classification, applying them to skin lesion classification still faces enormous difficulties.
In summary, both CNN and Transformer have certain limitations in terms of their global and local feature extraction capabilities. As a result, researchers have attempted to overcome these limitations in medical image analysis tasks. For instance, Yue et al. [28] improved segmentation performance in polyp segmentation tasks by integrating cross-level contextual information and utilizing edge information. To enable CNN to capture more comprehensive contextual features, they further proposed the Context Extraction Module (CEM) to retain local information and compress global information [29]. In the optic disc and optic cup segmentation task for glaucoma detection, Lei et al. [30] combined low-level and high-level features from edge prediction maps to capture similar morphological boundary information between the optic disc and optic cup. In skin lesion classification tasks, Wang et al. [31] introduced a global lesion localization module based on Class Activation Mapping (CAM) to guide CNN in learning consistent intra-class features and distinguishing inter-class features. Nakai et al. [32] proposed an enhanced deep bottleneck transformer model that can integrate any CNN model to enhance local interactions, achieving superior performance on two skin lesion datasets. Inspired by the aforementioned works, we aim to combine CNN and Transformer and improve the network specifically for clinical skin lesion classification tasks.
In this study, we propose a dual-branch network structure called CR-Conformer as a new method for clinical skin lesion image classification, which can combine the advantages of Transformer and DCNN to improve the classification accuracy of clinical skin lesion images. In CR-Conformer, the DCNN branch follows the design of ResNet [16]. It introduces convolutional rotation to extract diverse clinical skin lesion image features, and the Transformer branch follows the design of ViT [25]. These two branches can extract multi-scale local features in different directions and global representations and then fuse the extracted information with each other, further increasing the semantic information of skin lesion features obtained by each branch. Extensive experiments on two benchmark skin lesion datasets demonstrate that our proposed model can perform better than the baseline and advanced models in clinical skin lesion classification.
Overall, our main contributions are threefold:
-
1. Combining the advantages of DCNN and Transformer, a CR-Conformer dual-branch network model is proposed for clinical skin lesion image classification, which can capture local features and global representations of different scale features, respectively.
-
2. To improve the quality of the DCNN branch extracting clinical skin lesion image features, the convolutional rotation strategy is used to improve the branch to mine more semantic information.
-
3. Compared with the ISIC2018, a dermoscopic skin lesion image dataset, our proposed CR-Conformer model achieves better classification results on the XJUSL, a clinical skin lesion image dataset.
2 Materials and methods
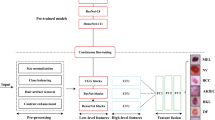
We propose CR-Conformer, a network model for clinical skin lesion classification. In the skin lesion classification task, CR-Conformer has a parallel dual-branch network structure similar to Conformer. Therefore, CR-Conformer shown in Fig. 1 includes (1) a DCNN branch that introduces convolutional rotation to extract finer local features, (2) a Transformer branch that leverages the fused local features to extract global features, and (3) a CR-FCU fusion module. These components are described in the various sections below. Notably, in the final stage, all features in the DCNN branch are pooled and fed into a classifier. The class token in the Transformer branch is extracted and provided to another classifier. In the training process, we use two cross-entropy losses with the same importance to supervise the two classifiers, respectively, and the calculation process of the loss function for predicting N samples containing K categories is shown in Eq. (1) and Eq. (2).
Where \(y\left(i,k\right)\) is the true label and \(p\left(i,k\right)\) is the probability that the i-th sample is predicted to be the k-th label. During inference, the prediction result is the sum of the outputs of the two classifiers.
2.1 DCNN branch
As shown in Fig. 1, the DCNN branch adopts a feature pyramid structure, similar to the Conformer, which contains 12 convolution blocks, and each convolution block contains several bottlenecks. Following the definition in ResNet, bottlenecks consist of 1 × 1 convolution (reduce the number of channels), 3 × 3 spatial convolution, 1 × 1 convolution (restore the number of channels), and a residual connection between the input and output. As shown in Fig. 2, we perform a convolutional rotation operation on the 1 × 1 convolution. The input feature map is divided into four parts on average according to the number of channels, which are rotated at 0°, 90°, 180°and 270°, respectively, and then fused as the input of the next convolution through a concatenation operation. The enhanced features of clinical skin lesion images extracted by the convolutional rotation are more diverse. The calculation process of the feature map Fi of the i-th layer is shown in Eq. (3).
Where Conv1 represents the convolution operation with the convolution kernel size of 1×1, and \({F}_{i-1}^{90^\circ }\) represents the rotation angle of the feature map of the i-1th layer is 90°. In this way, we utilize the DCNN branch for retaining the finer local features in the skin lesion images and continuously feed them to the Transformer branch.
2.2 Transformer branch
Similar to ViT, this branch contains 12 repeated Transformer blocks. As shown in Fig. 1, each Transformer block consists of a multi-head self-attention module and an MLP block. LayerNorm is applied before each layer, and residual connections exist in both the self-attention layer and the MLP block. First, the input image is converted to the image block sequence Z0. The calculation process is shown in Eq. (4).
Where \(\mathbf{E}\in {R}^{\left({P}^{2}\cdot C\right)\times D},{\mathbf{E}}_{pos}\in {R}^{\left(M+1\right)\times D}\), xp is the image block sequence after the image x is expanded. There are M image blocks in the sequence, P is the size of the image block, and C is the number of channels. Then it is embedded into the matrix E, performs a linear transformation on the flattened image block, and converts it into a D-dimension vector. The encoder uses the learnable vector xclass to explicitly predict the class, and the position encoding vector Epos is used to specify the spatial position information of the image block sequence. The total image block sequence Z0 is input into the encoder, and the forward calculation process is defined as Eq. (5) and Eq. (6).
Multi-head self-attention and LayerNorm iterate L times to get Z'l, MLP and LayerNorm iterate L times to get Zl. The fine local features provided by the DCNN branch are fused before the first LayerNorm, and the global features extracted after passing through the MLP block are provided to the DCNN branch.
2.3 CR-FCU fusion module
Since feature maps are used in the DCNN branch, and patch embeddings are used in the Transformer branch, eliminating the misalignment between them is an important issue. To address this issue, we propose CR-FCU that continuously integrates local features and global representations in an interactive manner.
As shown in Fig. 1, when a 64-channel feature map is fed to the Transformer branch, it first needs to align the patch embedding channel 384 through a 1 × 1 convolution. Then, a down-sampling module (as shown in Fig. 3) is used to complete the spatial dimension alignment. The down-sampling module uses the Average Pool method with a convolution kernel size of 4 and a stride of 4 to down-sample the feature map and then uses a reshaping operation to align spatial scale. Finally, the feature map is regularized by LayerNorm and is added to the patch embedding, as shown in Fig. 1. It is worth noting that the convolutional rotation operation has enhanced the feature map. When the patch embedding is fed back to the DCNN branch from the Transformer branch, it needs to be up-sampled (as shown in Fig. 3), which is similar to the down-sampling module, except that the up-sampling uses a linear interpolation method to align the spatial dimensions. Then, the channel dimension is aligned with the feature map enhanced by the convolutional rotation operation in the DCNN branch through 1 × 1 convolution. The BatchNorm method is used to regularize the features before fusion.
3 Results
3.1 Experimental setup
3.1.1 Datasets
To verify the classification performance of our method on dermoscopic and clinical skin lesion images, we use the public dermoscopic skin lesions dataset provided by ISIC 2018 challenge and the private clinical skin lesions dataset XJUSL provided by Xinjiang Urumqi People’s Hospital. The dataset details are as follows: (1) ISIC2018 contains seven types of skin lesions: melanocytic nevus, dermatofibroma, melanoma, actinic keratosis, benign keratosis, basal cell carcinoma, and vascular lesion. There are a total of 10,015 dermoscopic skin lesion images and labels, which we randomly divided into the training set (8019), validation set (497) and test set (1499) according to the ratio of 8:0.5:1.5. (2) XJUSL contains ten types of skin lesions: leucoderma (LEU), lichen planus (LIC), basal cell carcinoma (BAS), melanoma (MEL), solar keratosis (SOL), psoriasis (PSO), seborrheic keratosis (SEB), compound nevus (COM), junctional nevus (JUN), and intradermal nevus (INT). In order to remove the background noise in the clinical images, we crop them and keep only the skin area as the experimental dataset, and finally get 3131 clinical skin lesion images and their labels. Similarly, we randomly divide it into the training set (2511), validation set (156) and test set (464) in the ratio of 8:0.5:1.5. In all experiments, we resize and crop images from all datasets to 224 × 224 as the input to the model, and all pixel values in the images are normalized to 0–1. Since this work focuses on model innovation, to remove the confounding effects of data augmentation, we only use a simple geometric data augmentation strategy, namely random horizontal flipping, with random variables taken from uniform distributions.
3.1.2 Experimental configuration
All experiments are implemented on the mmcvContributors [33] framework, and we run all training and testing procedures on an NVIDIA RTX 3090 GPU with 24 GB of video memory. In order to make a fair comparison, we unified the parameter configuration of each comparison model as follows: (1) The AdamW optimizer is used uniformly, the learning rate is 0.0001, the weight decay is 0.01, the hyperparameter β1 is 0.9, and β2 is 0.999. The cosine annealing learning rate strategy is adopted to optimize the neural network. The learning rate curve is shown in Fig. 4. (2) Uniformly use the softmax function as the output layer and the cross entropy as the loss function to calculate the loss value. (3) Set the batch size to 32, train 100 epochs uniformly, and take the model with the highest validation accuracy as the test.
3.1.3 Evaluation metrics
We choose accuracy, precision, recall, and F1-score, which are widely used in image classification, as classification evaluation metrics of the skin lesion images. The definitions of these metrics are as follows:
where TP, TN, FP and FN are calculated based on the confusion matrix. They are the number of the True Positive (TP), True Negative (TN), False Positive (FP) and False Negative (FN) samples. Therefore, Accuracy is calculated to get the percentage of skin lesion samples correctly identified. Precision is used to calculate the precision rate, that is, how many skin lesion samples predicted to be True are actually True, to reflect the precision of the model prediction. Using Recall to calculate the recall rate, that is, how many skin lesion samples are found that are actually True, to reflect the comprehensiveness of the model prediction. F1-score takes into account both Precision and Recall to achieve the reconciliation of the two. For all these metrics, the larger values indicate better performance.
3.2 Experimental results and analysis
3.2.1 Ablation experiment
To evaluate the effectiveness of CR-Conformer for clinical skin lesion classification, we compare four different settings on two skin lesion datasets: (1) Only using ResNet of the DCNN branch to classify. (2) Only using ViT of the Transformer branch to classify. (3) Conformer that combines two branches. (4) CR-Conformer that uses convolutional rotation to enhance local features.
The results of the ablation experiments are shown in Table 1. It can be seen that the classification performance of our proposed CR-Conformer in clinical skin lesion images is better than that of Conformer, while the classification performance of dermoscopic skin lesion images is lower than that of Conformer, which shows the importance of using convolutional rotation for feature enhancement in the DCNN branch of the dual-branch network for clinical skin lesion images. Secondly, we can see that in the single-branch network, the classification accuracy of DCNN for clinical skin lesion images is 3.45% higher than that of Transformer, while the classification accuracy of dermoscopic skin lesion images is only 2.33% higher than that of Transformer, which indicates that the dermoscopic skin lesion images with higher image quality need to pay more attention to global features. However, the local features of clinical skin lesion images need more attention because they contain multiple lesions and more image noise. It also proves that we add convolutional rotation to the DCNN branch to extract finer local features, which can achieve better classification results on clinical skin lesion images.
To further analyze the effect of CR-Conformer on the classification of different types of clinical skin lesion images, we plotted the confusion matrix of our method and the baseline method using the prediction results of the test set, as shown in Fig. 5, where the numbers on the diagonal represent the number of correct predictions. It can be seen that compared with the baseline method, our proposed method has better predictive performance in the five skin lesion categories of LIC, MEL, BAS, SOL and PSO, especially the classification accuracy of melanoma with the least number of test cases increased from 16.7% to 50%, which reflects that our method is more meaningful for the auxiliary diagnosis of melanoma with greater harm. In other categories with slightly poorer classification performance, most lesions are very similar in appearance to black circles, such as various types of nevus, and lesions covering large areas of skin, such as leucoderma. It indicates that our convolutional rotation strategy has a poor feature enhancement effect for skin lesions with similar shapes and single colors. In contrast, it has a more obvious feature enhancement effect for skin lesions with more diverse appearance details.
3.2.2 Visualization
We further use gradient-weighted class activation maps (Grad-CAM) [34] to visually demonstrate the feature information capture of each clinical skin lesion image by the DCNN branch of Conformer and CR-Conformer. The Grad-CAM, in the form of a thermal map, highlights the important regions in an image when used to predict a given class, which we designate as the class predicted by the model.
To analyze which clinical skin lesion image classification our CR-Conformer method is more suitable for, we visualize one sample in each category, as shown in Fig. 6. The first row represents the original image of the selected sample, the second row represents the visualization of the DCNN branch in the baseline model Conformer, the third row represents the visualization of the DCNN branch in our model CR-Conformer, each column represents a different category. Figure 6(a) shows the categories whose classification accuracy of our model is higher than that of the baseline model. It can be seen that for some multiple lesions (LIC, SOL and PSO) or lesions with more obvious local features (MEL and BAS), the model can pay more attention to the lesion area after applying the convolution rotation strategy. Figure 6(b) shows the categories whose classification accuracy of our model is higher than that of the baseline model. It can be seen that for lesions covering a large area of skin (LEU and COM) or lesions with insignificant local features (INT, SEB and MAR), other features outside the lesion area will be paid attention to after the application of the convolution rotation strategy, such as folds in the normal skin area and the low-brightness part of the image.
3.2.3 Comparative experiment
Table 2 shows the test results of different classification models trained with the same parameter configuration to predict clinical skin lesion images. They include the classic DCNN-based model VGG and ResNet, the lightweight model MobileNet, and the advanced model ConvNeXt. Transformer-based classic model ViT, advanced model Swin Transformer, and the baseline model Conformer, which also uses a dual-branch network structure. It can be seen that CR-Conformer has better results than other models in the classification of clinical skin lesion images, and the three indicators have reached the optimum. CR-Conformer also achieves better classification results with fewer parameters than the baseline model Conformer. Compared with the two DCNN models with fewer parameters, the classification accuracy of our method is also higher by more than 10%, indicating that our network structure and the strategy of enhancing local features through convolution rotation are more suitable for clinical skin lesion images.
4 Discussion and conclusion
In this study, in order to achieve better classification results on clinical skin lesion images, we propose a dual-branch network structure CR-Conformer. Specifically, we use the DCNN branch to extract local features of clinical skin lesion images and the Transformer branch to extract global features. In order to obtain richer local features of clinical skin lesion images, we improve the DCNN branch of the Conformer, adopt the convolutional rotation strategy, and use the CR-FCU fusion module to interactively fuse it with the global features extracted by the Transformer branch. Comprehensive experiments are conducted on a public dermoscopic skin lesion image dataset ISIC2018 and a private clinical skin lesion image dataset XJUSL provided by the Dermatology Department of Xinjiang Urumqi People’s Hospital. The test results show that our method is more suitable for classifying clinical skin lesion images. In addition, by analyzing the classification of different types of clinical skin lesions by the model, we conclude that the convolutional rotation strategy is more suitable for skin lesions with multiple lesions or more obvious local features. Using only fewer parameters, CR-Conformer demonstrates its potential to automatically diagnose skin lesions in mobile devices.
In future work, we will explore the effect of different segmentation models applied to clinical skin lesion image segmentation and improve the network structure according to the features of clinical skin lesion images. In addition, we will cooperate with public hospitals to produce more standardized clinical skin lesion segmentation datasets to contribute to the auxiliary diagnosis of clinical skin lesions.
References
Yue G, Wei P, Zhou T et al (2022) Toward multicenter skin lesion classification using deep neural network with adaptively weighted balance loss. IEEE Trans Med Imaging 42(1):119–131
Vestergaard M, Macaskill P, Holt P, Menzies S (2008) Dermoscopy compared with naked eye examination for the diagnosis of primary melanoma: a meta-analysis of studies performed in a clinical setting. Br J Dermatol 159(3):669–676
Feng H, Berk-Krauss J, Feng PW, Stein JA (2018) Comparison of dermatologist density between urban and rural counties in the United States. JAMA Dermatol 154(11):1265–1271
Litjens G, Kooi T, Bejnordi BE, Setio AAA, Ciompi F, Ghafoorian M et al (2017) A survey on deep learning in medical image analysis. Med Image Anal 42:60–88
Adegun A, Viriri S (2021) Deep learning techniques for skin lesion analysis and melanoma cancer detection: a survey of state-of-the-art. Artif Intell Rev 54(2):811–841
Yang J, Sun X, Liang J, and Rosin P L (2018) Clinical skin lesion diagnosis using representations inspired by dermatologist criteria[C]//Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 1258–1266
Satheesha T, Satyanarayana F, Prasad MG, Dhruve KD (2017) Melanoma is skin deep: a 3D reconstruction technique for computerized dermoscopic skin lesion classification. IEEE J Transl Eng Health Med 5:1–17
Gessert N, Sentker T, Madesta F, Schmitz R, Kniep H, Baltruschat I et al (2019) Skin lesion classification using CNNs with patch-based attention and diagnosis-guided loss weighting. IEEE Trans Biomed Eng 67(2):495–503
Xie Y, Zhang J, Xia Y, Shen C (2020) A mutual bootstrapping model for automated skin lesion segmentation and classification. IEEE Trans Med Imaging 39(7):2482–2493
Yuan Y, Chao M, Lo YC (2017) Automatic skin lesion segmentation using deep fully convolutional networks with Jaccard distance. IEEE Trans Med Imaging 36(9):1876–1886
Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM et al (2017) Dermatologist-level classification of skin cancer with deep neural networks. Nature 542(7639):115–118
Liu Y, Jain A, Eng C, Way DH, Lee K, Bui P et al (2020) A deep learning system for differential diagnosis of skin diseases[J]. Nat Med 26(6):900–908
Pan SJ, Yang Q (2009) A survey on transfer learning[J]. IEEE Trans Knowl Data Eng 22(10):1345–1359
Kawahara J, Hamarneh G (2016) Multi-resolution-tract CNN with hybrid pretrained and skin-lesion trained layers[C]//International workshop on machine learning in medical imaging. Springer, Cham, pp 164–171
Simonyan K, and Zisserman A (2014) Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556
He K, Zhang X, Ren S, and Sun J (2016) Deep residual learning for image recognition[C]//Proceedings of the IEEE conference on computer vision and pattern recognition. 770–778
Hu J, Shen L, and Sun G (2018) Squeeze-and-excitation networks[C]//Proceedings of the IEEE conference on computer vision and pattern recognition. 7132–7141
Zhang J, Xie Y, Xia Y, Shen C (2019) Attention residual learning for skin lesion classification. IEEE Trans Med Imaging 38(9):2092–2103
Wei Z, Li Q, Song H (2022) Dual attention based network for skin lesion classification with auxiliary learning. Biomed Signal Process Control 74:103549
Yap J, Yolland W, Tschandl P (2018) Multimodal skin lesion classification using deep learning. Exp Dermatol 27(11):1261–1267
Kawahara J, Daneshvar S, Argenziano G et al (2018) Seven-point checklist and skin lesion classification using multitask multimodal neural nets. IEEE J Biomed Health Inform 23(2):538–546
Bi L, Feng DD, Fulham M et al (2020) Multi-label classification of multi-modality skin lesion via hyper-connected convolutional neural network. Pattern Recogn 107:107502
Ge Z,Demyanov S, Chakravorty R, et al. (2017) Skin disease recognition using deep saliency features and multimodal learning of dermoscopy and clinical images[C]//Medical Image Computing and Computer Assisted Intervention− MICCAI 2017: 20th International Conference, Quebec City, QC, Canada, September 11-13, 2017, Proceedings, Part III 20. Springer International Publishing 250-258
Wang Y, Feng Y, Zhang L et al (2022) Adversarial multimodal fusion with attention mechanism for skin lesion classification using clinical and dermoscopic images. Med Image Anal 81:102535
Dosovitskiy A, Beyer L, Kolesnikov A, Weissenborn D, Zhai X, Unterthiner T, et al. (2020) An image is worth 16x16 words: transformers for image recognition at scale. arXiv preprint arXiv:2010.11929
Liu Z, Lin Y, Cao Y, Hu H, Wei Y, Zhang Z, et al. (2021) Swin transformer: hierarchical vision transformer using shifted windows[C]//Proceedings of the IEEE/CVF International Conference on Computer Vision. 10012–10022
Wang W, Xie E, Li X, Fan D P, Song K, Liang D, et al. Pyramid vision transformer: a versatile backbone for dense prediction without convolutions[C]//Proceedings of the IEEE/CVF International Conference on Computer Vision. 2021: 568–578
Yue G, Han W, Jiang B et al (2022) Boundary constraint network with cross layer feature integration for polyp segmentation. IEEE J Biomed Health Inform 26(8):4090–4099
Yue G, Li S, Cong R et al (2023) Attention-guided pyramid context network for polyp segmentation in colonoscopy images. IEEE Trans Instrum Meas 72:1–13
Lei H, Liu W, Xie H et al (2021) Unsupervised domain adaptation based image synthesis and feature alignment for joint optic disc and cup segmentation. IEEE J Biomed Health Inform 26(1):90–102
Wang L, Zhang L, Shu X et al (2023) Intra-class consistency and inter-class discrimination feature learning for automatic skin lesion classification. Med Image Anal 85:102746
Nakai K, Chen YW, Han XH (2022) Enhanced deep bottleneck transformer model for skin lesion classification. Biomed Signal Process Control 78:103997
MMCV Contributors. MMCV: OpenMMLab computer vision foundation. https://github.com/open-mmlab/mmcv. Accessed Oct 14 2022
Selvaraju R R, Cogswell M, Das A, Vedantam R, Parikh D, and Batra D (2017) Grad-cam: visual explanations from deep networks via gradient-based localization[C]//Proceedings of the IEEE international conference on computer vision. 618–626
Funding
This work is partially supported by Xinjiang Uygur Autonomous Region Key R & D program under Grant 2021B03001-4 and National Natural Science Foundation of China 62362061.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Dezhi Zhang and Aolun Li have contributed equally to this work, should be regarded as co-first authorship.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, D., Li, A., Wu, W. et al. CR-Conformer: a fusion network for clinical skin lesion classification. Med Biol Eng Comput 62, 85–94 (2024). https://doi.org/10.1007/s11517-023-02904-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-023-02904-0