Abstract
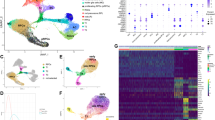
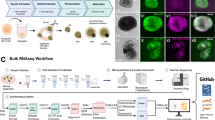
Human retina development involves multiple well-studied signaling pathways that promote the genesis of a wide arrange of different cell types in a complex architectural structure. Human embryonic stem cells (hESCs)-derived retinal organoids could recapitulate the human retinal development. We performed single-cell RNA-seq of retinal organoids from 5 time points (D36, D66, D96, D126, D186) and identified 9 distinct populations of cells. In addition, we analyzed the molecular characteristics of each main population and followed them from genesis to maturity by pseudotime analysis and characterized the cell-cell interactions between different cell types. Interestingly, we identified insulin receptor (INSR) as a specifically expressed receptor involved in the genesis of photoreceptors, and pleiothropin (PTN)-protein tyrosine phosphatase receptor type Z1 (PTPRZ1) as a mediator of a previously unknown interaction between Müller and retinal progenitor cells. Taken together, these findings provide a rich transcriptome-based lineage map for studying human retinal development and modeling developmental disorders in retinal organoids.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Berson, D. (2003). Strange vision: ganglion cells as circadian photoreceptors. Trends Neurosci 26, 314–320.
Butler, A., Hoffman, P., Smibert, P., Papalexi, E., and Satija, R. (2018). Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol 36, 411–420.
Butler, J.M., Sharif, U., Ali, M., McKibbin, M., Thompson, J.P., Gale, R., Yang, Y.C., Inglehearn, C., and Paraoan, L. (2015). A missense variant in CST3 exerts a recessive effect on susceptibility to age-related macular degeneration resembling its association with Alzheimer’s disease. Hum Genet 134, 705–715.
Camp, J.G., Wollny, D., and Treutlein, B. (2018). Single-cell genomics to guide human stem cell and tissue engineering. Nat Methods 15, 661–667.
Carlin, D.E., Demchak, B., Pratt, D., Sage, E., and Ideker, T. (2017). Network propagation in the cytoscape cyberinfrastructure. PLoS Comput Biol 13, e1005598.
Carter, D.A., Dick, A.D., and Mayer, E.J. (2009). CD133+ adult human retinal cells remain undifferentiated in Leukaemia Inhibitory Factor (LIF). BMC Ophthalmol 9, 1.
Chen, E.Y., Tan, C.M., Kou, Y., Duan, Q., Wang, Z., Meirelles, G., Clark, N.R., and Ma’ayan, A. (2013). Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinf 14, 128.
Collin, J., Queen, R., Zerti, D., Dorgau, B., Hussain, R., Coxhead, J., Cockell, S., and Lako, M. (2019). Deconstructing retinal organoids: single cell RNA-Seq reveals the cellular components of human pluripotent stem cell-derived retina. Stem Cells 37, 593–598.
da Cruz, L., Fynes, K., Georgiadis, O., Kerby, J., Luo, Y.H., Ahmado, A., Vernon, A., Daniels, J.T., Nommiste, B., Hasan, S.M., et al. (2018). Phase 1 clinical study ofan embryonic stem cell-derived retinal pigment epithelium patch in age-related macular degeneration. Nat Biotechnol 36, 328–337.
De Cat, B., Muyldermans, S.Y., Coomans, C., Degeest, G., Vanderschueren, B., Creemers, J., Biemar, F., Peers, B., and David, G. (2003). Processing by proprotein convertases is required for glypican-3 modulation of cell survival, Wnt signaling, and gastrulation movements. J Cell Biol 163, 625–635.
Dulken, B.W., Leeman, D.S., Boutet, S.C., Hebestreit, K., and Brunet, A. (2017). Single-cell transcriptomic analysis defines heterogeneity and transcriptional dynamics in the adult neural stem cell lineage. Cell Rep 18, 777–790.
Farrell, J.A., Wang, Y., Riesenfeld, S.J., Shekhar, K., Regev, A., and Schier, A.F. (2018). Single-cell reconstruction of developmental trajectories during zebrafish embryogenesis. Science 360, eaar3131.
Gonzalez-Cordero, A., Kruczek, K., Naeem, A., Fernando, M., Kloc, M., Ribeiro, J., Goh, D., Duran, Y., Blackford, S.J.I., Abelleira-Hervas, L., et al. (2017). Recapitulation of human retinal development from human pluripotent stem cells generates transplantable populations of cone photoreceptors. Stem Cell Rep 9, 820–837.
Han, X., Wang, R., Zhou, Y., Fei, L., Sun, H., Lai, S., Saadatpour, A., Zhou, Z., Chen, H., Ye, F., et al. (2018). Mapping the mouse cell atlas by microwell-seq. Cell 172, 1091–1107.e17.
Han, X., Zhou, Z., Fei, L., Sun, H., Wang, R., Chen, Y., Chen, H., Wang, J., Tang, H., Ge, W., et al. (2020). Construction of a human cell landscape at single-cell level. Nature 581, 303–309.
Hitchcock, P.F., Otteson, D.C., and Cirenza, P.F. (2001). Expression of the insulin receptor in the retina of the goldfish. Invest Ophthalmol Vis Sci 42, 2125–2129.
Hu, Y., Wang, X., Hu, B., Mao, Y., Chen, Y., Yan, L., Yong, J., Dong, J., Wei, Y., Wang, W., et al. (2019). Dissecting the transcriptome landscape of the human fetal neural retina and retinal pigment epithelium by single-cell RNA-seq analysis. PLoS Biol 17, e3000365.
Kaewkhaw, R., Kaya, K.D., Brooks, M., Homma, K., Zou, J., Chaitankar, V., Rao, M., and Swaroop, A. (2015). Transcriptome dynamics of developing photoreceptors in three-dimensional retina cultures recapitulates temporal sequence of human cone and rod differentiation revealing cell surface markers and gene networks. Stem Cells 33, 3504–3518.
Kim, S., Lowe, A., Dharmat, R., Lee, S., Owen, L.A., Wang, J., Shakoor, A., Li, Y., Morgan, D.J., Hejazi, A.A., et al. (2019). Generation, transcriptome profiling, and functional validation of cone-rich human retinal organoids. Proc Natl Acad Sci USA 116, 10824–10833.
Klein, A.M., Mazutis, L., Akartuna, I., Tallapragada, N., Veres, A., Li, V., Peshkin, L., Weitz, D.A., and Kirschner, M.W. (2015). Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201.
Krzyzanowski, P.M., and Andrade-Navarro, M.A. (2007). Identification of novel stem cell markers using gap analysis of gene expression data. Genome Biol 8, R193.
Kuwahara, A., Ozone, C., Nakano, T., Saito, K., Eiraku, M., and Sasai, Y. (2015). Generation of a ciliary margin-like stem cell niche from self-organizing human retinal tissue. Nat Commun 6, 6286.
La Manno, G., Gyllborg, D., Codeluppi, S., Nishimura, K., Salto, C., Zeisel, A., Borm, L.E., Stott, S.R.W., Toledo, E.M., Villaescusa, J.C., et al. (2016). Molecular diversity of midbrain development in mouse, human, and stem cells. Cell 167, 566–580.e19.
Laboissonniere, L.A., Goetz, J.J., Martin, G.M., Bi, R., Lund, T.J.S., Ellson, L., Lynch, M.R., Mooney, B., Wickham, H., Liu, P., et al. (2019). Molecular signatures of retinal ganglion cells revealed through single cell profiling. Sci Rep 9, 15778.
Lakowski, J., Han, Y.T., Pearson, R.A., Gonzalez-Cordero, A., West, E.L., Gualdoni, S., Barber, A.C., Hubank, M., Ali, R.R., and Sowden, J.C. (2011). Effective transplantation of photoreceptor precursor cells selected via cell surface antigen expression. Stem Cells 29, 1391–1404.
Lakowski, J., Gonzalez-Cordero, A., West, E.L., Han, Y.T., Welby, E., Naeem, A., Blackford, S.J.I., Bainbridge, J.W.B., Pearson, R.A., Ali, R. R., et al. (2015). Transplantation of photoreceptor precursors isolated via a cell surface biomarker panel from embryonic stem cell-derived self-forming retina. Stem Cells 33, 2469–2482.
Liu, Y., Fan, X., Wang, R., Lu, X., Dang, Y.L., Wang, H., Lin, H.Y., Zhu, C., Ge, H., Cross, J.C., et al. (2018). Single-cell RNA-seq reveals the diversity of trophoblast subtypes and patterns of differentiation in the human placenta. Cell Res 28, 819–832.
Macosko, E.Z., Basu, A., Satija, R., Nemesh, J., Shekhar, K., Goldman, M., Tirosh, I., Bialas, A.R., Kamitaki, N., Martersteck, E.M., et al. (2015). Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214.
Mao, X., An, Q., Xi, H., Yang, X.J., Zhang, X., Yuan, S., Wang, J., Hu, Y., Liu, Q., and Fan, G. (2019). Single-cell RNA sequencing of hESC-derived 3D retinal organoids reveals novel genes regulating RPC commitment in early human retinogenesis. Stem Cell Rep 13, 747–760.
Nakano, T., Ando, S., Takata, N., Kawada, M., Muguruma, K., Sekiguchi, K., Saito, K., Yonemura, S., Eiraku, M., and Sasai, Y. (2012). Self-formation of optic cups and storable stratified neural retina from human ESCs. Cell Stem Cell 10, 771–785.
Ochocinska, M.J., and Hitchcock, P.F. (2009). NeuroD regulates proliferation of photoreceptor progenitors in the retina of the zebrafish. Mech Dev 126, 128–141.
Otteson, D.C., Cirenza, P.F., and Hitchcock, P.F. (2002). Persistent neurogenesis in the teleost retina: evidence for regulation by the growth-hormone/insulin-like growth factor-I axis. Mech Dev 117, 137–149.
Paul, F., Arkin, Y., Giladi, A., Jaitin, D.A., Kenigsberg, E., Keren-Shaul, H., Winter, D., Lara-Astiaso, D., Gury, M., Weiner, A., et al. (2015). Transcriptional heterogeneity and lineage commitment in myeloid progenitors. Cell 163, 1663–1677.
Peng, Y.R., Shekhar, K., Yan, W., Herrmann, D., Sappington, A., Bryman, G.S., van Zyl, T., Do, M.T.H., Regev, A., and Sanes, J.R. (2019). Molecular classification and comparative taxonomics of foveal and peripheral cells in primate retina. Cell 176, 1222–1237.e22.
Phillips, M.J., Jiang, P., Howden, S., Barney, P., Min, J., York, N.W., Chu, L.F., Capowski, E.E., Cash, A., Jain, S., et al. (2018). A novel approach to single cell RNA-sequence analysis facilitates in silico gene reporting of human pluripotent stem cell-derived retinal cell types. Stem Cells 36, 313–324.
Qiu, X., Hill, A., Packer, J., Lin, D., Ma, Y.A., and Trapnell, C. (2017). Single-cell mRNA quantification and differential analysis with Census. Nat Methods 14, 309–315.
Rosenberg, A.B., Roco, C.M., Muscat, R.A., Kuchina, A., Sample, P., Yao, Z., Graybuck, L.T., Peeler, D.J., Mukherjee, S., Chen, W., et al. (2018). Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 360, 176–182.
Rozenblatt-Rosen, O., Stubbington, M.J.T., Regev, A., and Teichmann, S. A. (2017). The Human Cell Atlas: from vision to reality. Nature 550, 451–453.
Schwartz, S.D., Regillo, C.D., Lam, B.L., Eliott, D., Rosenfeld, P.J., Gregori, N.Z., Hubschman, J.P., Davis, J.L., Heilwell, G., Spirn, M., et al. (2015). Human embryonic stem cell-derived retinal pigment epithelium in patients with age-related macular degeneration and Stargardt’s macular dystrophy: follow-up of two open-label phase 1/2 studies. Lancet 385, 509–516.
Shekhar, K., Lapan, S.W., Whitney, I.E., Tran, N.M., Macosko, E.Z., Kowalczyk, M., Adiconis, X., Levin, J.Z., Nemesh, J., Goldman, M., et al. (2016). Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323.e30.
Shen, Y., Shen, H., Guo, D., Sun, X., Sun, Y., Hong, N., Wang, X., Xie, C., Zhao, Y., He, Q., et al. (2020). Recent developments in regenerative ophthalmology. Sci China Life Sci 63, 1450–1490.
Stern, J.H., Tian, Y., Funderburgh, J., Pellegrini, G., Zhang, K., Goldberg, J.L., Ali, R.R., Young, M., Xie, Y., and Temple, S. (2018). Regenerating eye tissues to preserve and restore vision. Cell Stem Cell 23, 453.
Sun, H., Zhou, Y., Fei, L., Chen, H., and Guo, G. (2019). scMCA: A tool to define mouse cell types based on single-cell digital expression. In: In: Yuan, G.C., ed. Computational Methods for Single-Cell Data Analysis. Methods in Molecular Biology. New York: Humana Press, 91–96.
Sun, Y., Kong, W., Falk, A., Hu, J., Zhou, L., Pollard, S., and Smith, A. (2009). CD133 (Prominin) negative human neural stem cells are clonogenic and tripotent. PLoS ONE 4, e5498.
Tasic, B., Menon, V., Nguyen, T.N., Kim, T.K., Jarsky, T., Yao, Z., Levi, B., Gray, L.T., Sorensen, S.A., Dolbeare, T., et al. (2016). Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat Neurosci 19, 335–346.
Treutlein, B., Brownfield, D.G., Wu, A.R., Neff, N.F., Mantalas, G.L., Espinoza, F.H., Desai, T.J., Krasnow, M.A., and Quake, S.R. (2014). Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq. Nature 509, 371–375.
Vento-Tormo, R., Efremova, M., Botting, R.A., Turco, M.Y., Vento-Tormo, M., Meyer, K.B., Park, J.E., Stephenson, E., Polanski, K., Goncalves, A., et al. (2018). Single-cell reconstruction of the early maternal-fetal interface in humans. Nature 563, 347–353.
Völkner, M., Zschätzsch, M., Rostovskaya, M., Overall, R.W., Busskamp, V., Anastassiadis, K., and Karl, M.O. (2016). Retinal organoids from pluripotent stem cells efficiently recapitulate retinogenesis. Stem Cell Rep 6, 525–538.
Wagner, D.E., Weinreb, C., Collins, Z.M., Briggs, J.A., Megason, S.G., and Klein, A.M. (2018). Single-cell mapping of gene expression landscapes and lineage in the zebrafish embryo. Science 360, 981–987.
Wang, S.J., Weng, C.H., Xu, H.W., Zhao, C.J., and Yin, Z.Q. (2014). Effect of optogenetic stimulus on the proliferation and cell cycle progression of neural stem cells. J Membr Biol 247, 493–500.
Wang, Y., Wang, R., Zhang, S., Song, S., Jiang, C., Han, G., Wang, M., Ajani, J., Futreal, A., and Wang, L. (2019). iTALK: an R package to characterize and illustrate intercellular communication. bioRxiv, 507871.
Wolf, F.A., Angerer, P., and Theis, F.J. (2018). SCANPY: large-scale single-cell gene expression data analysis. Genome Biol 19, 15.
Wu, B., An, C., Li, Y., Yin, Z., Gong, L., Li, Z., Liu, Y., Heng, B.C., Zhang, D., Ouyang, H., et al. (2017). Reconstructing lineage hierarchies of mouse uterus epithelial development using single-cell analysis. Stem Cell Rep 9, 381–396.
Yao, Z., Mich, J.K., Ku, S., Menon, V., Krostag, A.R., Martinez, R.A., Furchtgott, L., Mulholland, H., Bort, S., Fuqua, M.A., et al. (2017). A single-cell roadmap of lineage bifurcation in human ESC models of embryonic brain development. Cell Stem Cell 20, 120–134.
Zeisel, A., Munoz-Manchado, A.B., Codeluppi, S., Lonnerberg, P., La Manno, G., Jureus, A., Marques, S., Munguba, H., He, L., Betsholtz, C., et al. (2015). Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142.
Zeisel, A., Hochgerner, H., Lönnerberg, P., Johnsson, A., Memic, F., van der Zwan, J., Häring, M., Braun, E., Borm, L.E., La Manno, G., et al. (2018). Molecular architecture of the mouse nervous system. Cell 174, 999–1014.e22.
Zhong, S., Zhang, S., Fan, X., Wu, Q., Yan, L., Dong, J., Zhang, H., Li, L., Sun, L., Pan, N., et al. (2018). A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528.
Zhou, F., Li, X., Wang, W., Zhu, P., Zhou, J., He, W., Ding, M., Xiong, F., Zheng, X., Li, Z., et al. (2016). Tracing haematopoietic stem cell formation at single-cell resolution. Nature 533, 487–492.
Zhou, P.Y., Peng, G.H., Xu, H., and Yin, Z.Q. (2015). c-Kit+ cells isolated from human fetal retinas represent a new population of retinal progenitor cells. J Cell Sci 128, 2169–2178.
Zhu, J., Reynolds, J., Garcia, T., Cifuentes, H., Chew, S., Zeng, X., and Lamba, D.A. (2018). Generation of transplantable retinal photoreceptors from a current good manufacturing practice-manufactured human induced pluripotent stem cell Line. Stem Cells Transl Med 7, 210–219.
Zurdel, J., Finckh, U., Menzer, G., Nitsch, R.M., and Richard, G. (2002). CST3 genotype associated with exudative age related macular degeneration. British J Ophthalmol 86, 214–219.
Acknowledgements
This work was supported by the National Key Research and Development Program (2018YFA0107303), the National Natural Science Foundation of China (82070990), Initiation Fund of Distinguished Professor, Zhengzhou University (32310180), China Postdoctoral Research Fund (2017M613396).
Author information
Authors and Affiliations
Corresponding author
Additional information
Compliance and ethics
The author(s) declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Wang, S., Poli, S., Liang, X. et al. Longitudinal single-cell RNA-seq of hESCs-derived retinal organoids. Sci. China Life Sci. 64, 1661–1676 (2021). https://doi.org/10.1007/s11427-020-1836-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-020-1836-7