Abstract
The mandarin fish Siniperca chuatsi is a freshwater fish that is endemic to East Asia. To study the different damages and molecular mechanisms caused by different salt (NaCl, Na2SO4, and NaHCO3) on Siniperca chuatsi, the fish were subjected to NaCl, Na2SO4, and NaHCO3 stresses with different concentration for 96 h for mortality assessment, moreover, the fish were exposed to these salt stresses with equal sodium ion concentration (Na+ = 210 mmol/L), then gill morphological changes were observed and gene expression was analyzed by high-throughput transcriptome sequencing and real-time quantitative PCR (qRT-PCR). The results showed that mandarin fish tolerated NaCl and Na2SO4 better than NaHCO3. NaHCO3 stress caused more damage to gill than NaCl and Na2SO4 stress. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses indicated that differentially expressed genes were enriched in damage and apoptosis upon NaHCO3 stress, whereas they were enriched in energy and immune-related pathways upon NaCl and Na2SO4 stress. Hub genes were different under all three stresses. MAPK pathway genes showed a trend in up-regulated expression under all salt stresses, but the expression patterns varied with time during salt exposure and freshwater recovery stage. Taken together, this study demonstrated the variation in the effects of NaCl, Na2SO4, and NaHCO3 stress on mandarin fish. The MAPK signaling pathway is important for regulating the response to salt stress.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Different types of saline-alkali soil and water are important potential agricultural and aquaculture resources. According to the Alegin water quality classification system, more than 10 types of saline alkali water have been discovered in China, which can be roughly divided into chloride, sulfate, and carbonate based on ion composition. The ionic components responsible for their differences are mainly Cl−, SO42−, and HCO3−. For example, 224 mmol/l (NaCl concentration) in Qinghai lake; 2450 mmol/l (NaCl concentration) in Beidachi lake; 11 mmol/l (Na2SO4 concentration) in Bosten lake, 966 mmol/l (MgSO4 concentration) in Ebinur lake; 4.6-10.1 mmol/L (HCO3- concentration) in Henan surface water; greater than 50 mmol/L (HCO3−/CO32− concentrations, pH 9.6) in Dali Nor lake (Chen et al. 2020b; Wang et al. 2021). Cl−, SO42−, and HCO3− can affect the survival of freshwater fishes (Schulz and Cañedo-Argüelles 2019). Water containing elevated Cl− can affect osmotic pressure regulation, endocrine homeostasis, and oxygen consumption in freshwater organisms (Breves et al. 2010). SO42− which is mainly derived from the dissolution of rocks, can cause osmotic stress or specific ion toxicity in aquatic organisms, as the gills do not have an active mechanism to absorb or regulate sulfate (Elphick et al. 2011; Karjalainen et al. 2023). HCO3− induces metabolic and respiratory toxicity in fish, and the increase of HCO3− in the water column leads to an increase in pH so that the ammonia nitrogen (NH3) metabolized by the organism is not efficiently converted to NH4+ for elimination (Wilkie and Wood 1996). In addition, HCO3− can cause surface ulceration, gill rot, and blindness in freshwater fish. NaCl and Na2SO4 are neutral salinities, whereas NaHCO3 and Na2CO3 are alkaline. Studies comparing the differential effects of different salts on organisms are more frequent in plants, such as wheat (Triticum aestivum) (Wang et al. 2022) and sorghum (Sorghum bicolor) (Niu et al. 2022). Then, plants can be selected for further breeding and cultivation based on the type of saline-alkali soil. In fish, acute stress experiments of juvenile Crucian carp (Carassius auratus) from Lake Dari showed that the toxic effect of NaHCO3 was greater than that of NaCl (Zhou et al. 2013). Sequencing analysis of hybrid tilapia revealed that the relevant pathways were biased in response to pathogen infection after alkalinity stress compared to salinity (Su et al. 2020). However, the physiological difference in the fish response to these salt stresses lacks in-depth investigation.
Different environmental stresses (dissolved oxygen, temperature, etc.) can have different effects on fish. The gills are in close and direct contact with the external environment, are sensitive to changes in water quality, and play important roles in response to external environmental changes, such as respiration, osmotic pressure regulation, and excretion (Evans et al. 2005). The gills exhibit defense mechanisms in response to external harmful substances, such as epithelial cell enlargement, hyperplasia, and fusion of gill lamellae. It has been shown that gill of extensively saline fish adapt to environmental stress in high salt and alkaline environments by regulating osmotically-related genes including chloride channel 2, sodium/potassium/chlorine transporter protein, carbonic anhydrase, and Na+/HCO3− cotransporter protein (Su et al. 2020). In recent years, transcriptomic analysis has been used to investigate salinity and alkalinity tolerance mechanisms in some fishes, such as Amur ide (Leuciscus waleckii) (Wang et al. 2021), Nile tilapia (Oreochromis niloticus) (Zhao et al. 2020), and Silver pomfret (Pampus argenteus) (Li et al. 2020). Although these results provide some candidate pathways or genes related to stress response and environmental adaptation, including signal transduction, ion regulation, osmoregulation, immune system, energy metabolism, and chemical detoxification, the molecular mechanism of fish response and adaptation to salt stress needs to be further studied.
Siniperca belongs to the order Perciformes. Siniperca is a freshwater fish endemic to Southeast Asia and is mainly distributed in China. Mandarin fish (Siniperca chuatsi), a fish of the genus Siniperca, is widely distributed in the Yangtze, Pearl, and Heilongjiang River basins of China, and is one of the 16 main freshwater aquaculture species in China. In 2020, the aquaculture production of mandarin fish reached 376,986 tons in China (Yue et al. 2023). Mandarin fish feeds on fish and other aquatic animals. It is adapted to temperatures 25 ~ 30°C, with dissolved oxygen 3 ~ 5 mg/L, and pH 6.5 ~ 7.5. No studies have been reported on the salt tolerance of mandarin fish.
To compare the different damages and molecular mechanisms caused by different salt on mandarin fish, we investigated the tolerance of mandarin fish to NaCl, Na2SO4, and NaHCO3 with 96 h acute exposure experiment respectively based on main types of saline-alkali water. We also detected the changes of gill morphology, transcriptome, and MAPK signaling pathway upon salt stress with an equal sodium ion concentration. The study contributes to develop reasonable domestication and screening scheme of mandarin fish according to different types of saline-alkali water.
Materials and methods
Ethical statement
This study was approved by the Animal Experimentation Ethics Committee of Shanghai Ocean University (Shanghai, China) (SHOU-DW-2016-004).
Fish and experimental conditions
The mandarin fish were reared in the Fish Germplasm Station, Shanghai Ocean University. Mandarin fish were fed with live prey fish (grass carp, Ctenopharyngodon idellus larvae) once a day. Before the stress treatment, all the fish were fasted for 24 h.
Yearling Fish (<1 year old, 1.76 ± 0.31 g) were transferred from freshwater to 6 concentrations of each salt stress, and the mortality of fish was calculated within 96 h. For NaCl stress, 102.56 mmol/L (6 g/L, pH = 7.4), 136.75 mmol/L (8 g/L, pH = 7.4), 170.94 mmol/L (10 g/L, pH = 7.5) , 205.13 mmol/L (12 g/L, pH = 7.6), 239.32 mmol/L (14 g/L, pH = 7.4) and 273.50 mmol/L (16 g/L, pH = 7.6) were prepared using tap water (pH = 7.5 ± 0.1) which were aerated for more than 48 h with NaCl (analytical reagent) by the conversion between the molar and mass. For Na2SO4 stress, 42.25 mmol/L (6 g/L, pH = 7.5), 56.33 mmol/L (8 g/L, pH = 7.5), 70.42 mmol/L (10 g/L, pH = 7.4), 84.50 mmol/L (12 g/L, pH = 7.5), 98.59 mmol/L (14 g/L, pH = 7.6) and 112.67 mmol/L (16 g/L, pH = 7.5) were prepared using tap water (pH = 7.5 ± 0.1) which were aerated for more than 48 h with Na2SO4 (analytical reagent). For NaHCO3 stress, 1.19 mmol/L (0.1 g/L, pH = 7.7) , 3.57 mmol/L (0.3 g/L, pH = 8.1), 5.95 mmol/L (0.5 g/L, pH = 8.1), 11.90 mmol/L (1 g/L, pH = 8.2), 17.86 mmol/L (1.5 g/L, pH = 8.2), and 23.81 mmol/L (2 g/L, pH = 8.3) were prepared using tap water (pH = 7.5 ± 0.1) which were aerated for more than 48 h with NaHCO3 (analytical reagent). pH was determined by a pH meter (Leici, Shanghai). The experiments were conducted in tanks (45 L) with 30 fish for each concentration of each salt stress and was repeated for three times (n=30×6×3=540, approximately half male and half female). The fish were not fed during the experiment. Approximately 90% of the solution in each tank was changed daily. Water temperature was maintained at 24.6 ± 0.5°C, dissolved oxygen was kept at about 6.7~7.3 mg/L. NH4+-N at <0.6 mg/L, and NO2-N at <0.04 mg/L. The number of dead fish was recorded frequently from the beginning of the stress experiment, and the death of fish is judged by no sign of gill activity and no response to external stimuli. The 96h semi-lethal concentrations (LC50) was determined using Probit regression with the SPSS 26.0 (SPSS, Chicago, Illinois, USA).
With equal cation concentrations (Na+ = 210 mmol/L), NaCl concentration of 210 mmol/L (pH = 7.6) (SW group), Na2SO4 concentration of 105 mmol/L (pH = 7.6) (SUW group), and NaHCO3 concentration of 210 mmol/L (pH = 8.5) (AW group) were set as stress treatments. The huge variation brings difficulties to the setting of the stress dose with equal cation concentration. If 96 h LC50 value of NaHCO3 stress is used, the NaCl stress is almost unaffected. To better understand the toxicity on fish, a higher stress dose, i.e., the 96 h LC50 value of NaCl is used as the stress dose, and the corresponding sampling time is in the early stages of stress. The experiment was conducted in a tank (45 L). Yearling fish (< 1 year old, 5.8 ± 1.3 g) were transferred directly from freshwater (FW group) into the SW, SUW, and AW groups. After the 24 h acute stress, the fish were transferred to fresh water for recovery. Each tank contained 30 fish and there were four replicates for each group (n=30×4×4=480, approximately half male and half female). The fish were not fed during the experiment. Gills were sampled from the SW and SUW groups at 0, 0.5, 4, 12, and 24 h after stress and 24 h after freshwater recovery. Gills were sampled from the AW group at 0 and 0.5 h after stress and 2 h after freshwater recovery (In the previous experiment all fish died within 1 h after 210 mmol/L NaHCO3 stress. Therefore, the time of NaHCO3 stress treatment was set at 0.5 h). Water temperature was maintained at 24.6 ± 0.5°C, dissolved oxygen was kept at about 6.7~7.3 mg/L. NH4+-N at < 0.6 mg/L, and NO2-N at < 0.04 mg/L. Gills from the FW, SW, and SUW groups at 24 h and the AW group at 0.5 h were taken for tissue observation and sequencing. A portion was washed with saline and put into Bouin's solution for tissue observation, while the other part was rapidly frozen in liquid nitrogen and stored at −80°C for sequencing (4 groups × 4 replicates = 16 samples). The rest of the samples were used for qRT-PCR. In order to detect the complete expression pattern and response mechanism of MAPK signaling pathway, we designed a freshwater recovery experiment. Gene expression of MAPK signaling pathway were determined during salt exposure and freshwater recovery stage.
Histology
Upon different salt stresses, gill tissues were sampled at the time point mentioned above and were washed with 0.9% saline and placed in Bouin's for 24 h. Tissues were dehydrated with 70% alcohol, and when the tissues became lighter in color, they were dehydrated using gradients of 80%, 90%, and 100% alcohol, with each gradient immersed for 40 min. The dehydrated tissues were immersed in a 1:1 mixture of xylene: alcohol for 15 min, and then immersed in pure xylene solution for 15 min until the tissues were transparently finished. Finally, the tissue was immersed using paraffin solution to complete the embedding.
Serial sectioning was performed using a Leica RM 2016 rotary microtome (Shanghai Langyi Medical Equipment Co., Shanghai, China) at a thickness of 5μm. The excised tissue sections were stained with hematoxylin-eosin (HE). After staining, the slides were sealed with neutral gum and observed under an inverted microscope (Olympus Industries Co., Shenzhen, China) in the laboratory for imaging. Gill tissue sections were randomly observed three times, and each observation was then combined into the final result.
Transcriptome library construction and sequencing
RNA extraction, RNA-Seq library preparation, and sequencing were performed by Biomarker Technologies (Beijing, China). RNA concentration and purity were measured using a NanoDrop 2000 (Thermo Fisher Scientific, Wilmington, Delaware, USA). RNA integrity was assessed using the RNA Nano6000 assay kit from Agilent Bioanalyzer 2100 systems (Agilent Technologies, Palo Alto, California, USA). Eukaryotic mRNA was enriched with magnetic beads with Oligo (dT). The mRNA was randomly interrupted by adding Fragmentation Buffer and the first cDNA strand was synthesized with six-base random primers (random hexamers) using mRNA as a template, followed by the addition of buffer, dNTPs, RNase H, and DNA polymerase I to synthesize the second cDNA strand. This was purified using AMPure XP beads (Beckman Coulter, Beverly, California, USA). The purified double-stranded cDNA was then end-repaired, A-tailed, and connected to sequencing junctions, followed by fragment size selection using AMPure XP beads. Finally, the cDNA library was enriched by PCR. After library construction, the effective library concentration (effective library concentration > 2 nM) was accurately quantified using qPCR which is a method of quantifying DNA based on PCR. After the libraries were qualified, different libraries were pooled according to the target downstream data volume and sequenced using the Illumina platform.
Reads containing splices and low-quality reads (including reads with a proportion of N greater than 10% and those with bases with a quality value Q ≤ 10 accounting for more than 50% of the whole read) were removed. High-quality clean reads were obtained after the above series of quality control.
Unigene expression was calculated using fragments per kilobase of transcript per million fragments mapped as a measure of transcript or gene expression level, and the obtained gene expression was used directly to compare gene expression differences between samples. In the process of differentially expressed gene detection, Fold Change ≥ 2 and false discovery rate (FDR) < 0.01 were used as screening criteria. Differentially expressed genes were subjected to Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis and the protein-protein interaction (PPI) network was determined with a corrected P < 0.05.
Real-time quantitative PCR (qRT-PCR) analysis
Using the same RNA samples, six differentially expressed genes were randomly selected to validate the RNA-Seq results. To elucidate the expression pattern of the MAPK signaling pathway in equal sodium ion concentration stress experiments, genes AREG (amphiregulin), MKNK2b (MAPK interacting serine/threonine kinase 2b), PDGFC (platelet derived growth factor c), and TAMALIN (trafficking regulator and scaffold protein tamalin) were selected. β-actin was used as an internal reference gene. See Supplementary Material 1 for primer information. The SYBR Green Premix Pro Taq HS qPCR Kit (Accurate Biotechnology Co., Ltd., Hunan, China) was used, and the PCR was performed according to the instructions. PCRs were conducted in a CFX96 Real-Time PCR system. PCR conditions were 50°C for 2 min, 95°C for 10 min, followed by 40 cycles of 95°C for 15 s and 60°C for 1 min. The 2-ΔΔCT method was used to calculate the relative gene expression.
Statistical analysis
SPSS 26.0 software was used for data analysis. The measured data are expressed as mean ± SD. Pairwise comparisons were performed using the least significant difference method. Comparison between groups was analyzed by one-way ANOVA. Data conforming to a normal distribution were compared by t-test. P < 0.05 was considered a significant difference.
Results
Mortality rate
The mortality of mandarin fish after salt stress were shown in Table 1. The regression equation of each salt was obtained by fitting the curve. The 96 h LC50 of mandarin fish under NaCl, Na2SO4, and NaHCO3 stresses were 210.85 mmol/L (95% confidence interval 196.35 ~ 227.43), 81.60 mmol/L (95% confidence interval 74.79 ~ 88.94), and 5.89 mmol/L (95% confidence interval 4.50~7.25), respectively.
Effects of NaCl, Na2SO4 and NaHCO3 stress on gills morphology of mandarin fish
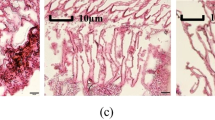
In the FW group, gill lamellae were evenly distributed on both sides of the gill filament, and pavement cells (PVCs) were covered with gill lamellae in a thin scale-like pattern, alternating with pillar cells (PCs) and blood cells (BCs) to form vascular channels. A thick epithelium was present between the gill lamellae and the gill lamellae, and this region contained mucous and chloride cells (Fig. 1a). After 24 h of NaCl stress, the gill filament vessels widened significantly and the stroma between the gill lamellae protruded slightly, while the number of chlorocytes in this region increased relative to the FW group. The gill lamellae showed shortening and thickening, and the PVCs covering the gill lamellae were severely broken and shed, and the hemocytes were larger in size (Fig. 1b). After 24 h of Na2SO4 stress, the stroma between the gill lamellae protruded and the gill lamellae showed different degrees of distortion or even breakage (Fig. 1c). After 0.5 h of NaHCO3 stress, the matrix was almost filled between the gill lamellae, and the number of chloride cells within the matrix increased. The gill lamellae appeared severely damaged, and the PVCs were shed in large numbers (Fig. 1d).
RNA-Seq data quality assessment and validation
A total of 103.75 Gb of clean data was obtained from the transcriptome analysis of 16 samples, and the clean data of each sample reached 5.71 Gb, with the percentage of Q30 bases at 92.96% and above. The clean reads of each sample were compared with the designated reference genome for sequence alignment, and the alignment efficiency ranged from 89.69% to 94.31% (Supplementary Material 3). Spearman's correlation coefficient r (r2) was used as an index for the assessment of biological duplicate correlation.
The r2 was closer to 1, indicating that the two duplicate samples are more correlated. The biological replicate correlation between each group in this experiment met the index (r2 ≥ 0.8) (Supplementary Material 2). The sequence data generated in this study have been stored in the NCBI SRA database (Accession number SUB11886438).
Six genes were randomly selected for detection by qRT-PCR. The results showed that the relative expression of genes CCL20b, GRS5, KCNE2, DAPK2a, IGλ, and HBB were down-regulated in the SW and SUW stress groups compared with the control group (FW), except for the up-regulated expression of CCL20b, and the down-regulated expression of GRS5, KCNE2, DAPK2a, IGλ, and HBB in the AW stress group. All genes were down regulated in the AW stress group. In addition, CCL20b did not change significantly in the AW group and DAPK2a did not change significantly in the SUW group. To ensure reproducible results, we used β-actin as an internal reference gene. The qRT-PCR analysis results were consistent with the RNA-Seq results of each stress group, indicating the reliability and accuracy of transcriptome analysis (Fig. 2).
Expression of six differential genes in different stress groups by qRT-PCR (left) and RNA-seq (right). The β-actin values were used to normalize the expression levels of different genes. Each column represents the mean ± standard deviation (n = 3), and the difference between the values of different letters (a-c) was statistically significant (P < 0.05), FW: control group; SW: NaCl stress; SUW: Na2SO4 stress; AW: NaHCO3 stress
Analysis of differential gene expression in mandarin fish upon NaCl, Na2SO4, and NaHCO3 stress
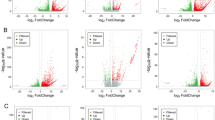
In the gill of mandarin fish, a total of 1218 genes were significantly differentially expressed in the FW vs SW group, of which 320 were up-regulated and 898 were down-regulated (P < 0.05); 466 genes were significantly differentially expressed in the FW vs SUW groups, of which 160 were up-regulated and 306 were down-regulated (Fold Change ≥ 2, FDR < 0.05); 313 genes were significantly differentially expressed in the FW vs AW groups (Supplementary Material 4). A total of 313 genes were significantly differentially expressed between the FW and AW groups, of which 155 were up-regulated and 158 were down-regulated (P < 0.05) (Supplementary Material 4). Compared with the FW group, the three stress groups shared 25 significantly differentially expressed genes, of which 23 were down-regulated (P < 0.05) (Fig. 3, Table 2).
There were 412 significantly differentially expressed genes in the SW vs SUW groups, of which 223 were up-regulated and 189 were down-regulated (P < 0.05); 2061 genes were significantly differentially expressed in the SW groups compared with AW group, of which 1354 were up-regulated and 707 were down-regulated (P < 0.05); 886 genes were significantly differentially expressed in the SUW group compared with the AW group, of which 644 were up regulated and 242 were down regulated (P < 0.05) (Supplementary Material 4).
Analysis of KEGG enrichment in mandarin fish upon NaCl, Na2SO4, and NaHCO3 stress
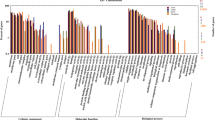
The number of KEGG pathways significantly enriched in the FW vs SW groups, FW vs SUW groups, and FW vs AW groups were 20, 4, and 8, respectively (P < 0.05). The KEGG pathways that were significantly enriched in the FW vs SW groups were cell adhesion molecules, cytokine-cytokine receptor interaction, carbon metabolism, biosynthesis of amino acids, ECM-receptor interaction, glycolysis/gluconeogenesis, and intestinal immune network for IgA production. KEGG pathways significantly enriched in the FW vs SUW groups were cell adhesion molecules, amino sugar and nucleotide sugar metabolism, biosynthesis of unsaturated fatty, and primary immunodeficiency. KEGG pathways significantly enriched in the FW vs AW groups were phagosome, Herpes simplex virus 1 infection, MAPK signaling pathway, cytokine-cytokine receptor interaction, intestinal immune network for IgA production, mucin-type O-glycan biosynthesis, steroid biosynthesis, and primary immunodeficiency (Fig. 4).
PPI analysis of differential genes
To identify the hub genes among the differentially expressed genes in each salinity or alkalinity group, a PPI network was constructed (Fig. 5). The larger the node the more edges associated with the protein, which also indicates that the protein interacts frequently with other proteins. In the SW group, the most frequent interaction between differential gene phases was Rho family GTPase 2 (RND2), which was up-regulated after stress. In the SUW group, the most frequent differential gene interaction was asporin (ASPN), which was down-regulated after stress. In the AW group, the most differential gene interaction was EPH receptor A4 (EPHA4a), which was up-regulated after stress.
Gene expression of MAPK signaling pathway during salt exposure and freshwater recovery stage
The expression of TAMALIN and PDGFC was down-regulated at 0.5 h in the SW group at the beginning of stress. The expression of MAPK signaling pathway genes was up-regulated in all three salts at 0.5 h. With increasing duration of stress, AREG and MKNK2b in the SW group peaked at 24 h of stress and PDGFC at 12 h of stress. AREG in the SUW group peaked at 24 h, TAMALIN at 0.5 h, and MKNK2b and PDGFC both at 4 h of stress (Fig. 6).
During the freshwater recovery stage, the expression level of MAPK signaling pathway genes in the AW group was up-regulated after 2 h freshwater recovery, on the contrary, these genes recovered to or below initial levels in both SW and SUW group, except for the SUW group where AREG expression did not drop to the initial stage after 24 h freshwater recovery (Fig. 6).
Discussion
Generally, in this study the 96 h LC50 value of NaHCO3 stress (5.89 mmol/L) was much lower than that of NaCl (210.85 mmol/L) and Na2SO4 (81.60 mmol/L) stress in mandarin fish. All fish died after 1 h exposure to 210 mmol/L NaHCO3 stress whereas some fish still survive after 96 h exposure to 210 mmol/L NaCl stress. The damage to gills, differentially expressed genes, KEGG signaling pathway enrichment, and hub genes among the three stresses with equal sodium ion concentration were different. These results strongly suggested that there were variation effects of the salt stresses (NaCl, Na2SO4, and NaHCO3) on mandarin fish. Detailed analysis will be conducted separately in the following text. It is assumed that the variation in salt stress effect is related to the difference in ionic composition, whereas, the direct cause of the variation remain unclear and further research is needed, because differences in ionic composition can lead to changes in the parameters of the water, such as pH changes.
The LC50 directly reflects the difference in tolerance of fish to different salts. The NaCl stress tolerance of mandarin fish (weight 5.8 ± 1.3 g , 96 h LC50 = 210.85 mmol/L) was weaker than that of Nile tilapia (weight 6.0 ~ 7.0 g, 96 h LC50 = 372.82 mmol/L) (Liang et al. 2015), Chalcalburnus chalcoides aralensis (weight 2.6 ~ 4.6 g, 96 h LC50 = 363.07 mmol/L) (Lin et al. 2015) and przewalski's naked carp (Gymnocypris przewalskii) (weight 12.5 ± 0.3 g, 96 h LC50 = 311.11 mmol/L) (Wu et al. 2017a), but stronger than that of grass carp (juvenile, 96 h LC50 = 182.73 mmol/L) and silver carp (Hypophthalmichthys molitrix) (juvenile, 96 h LC50 = 160.34 mmol/L) (Yang et al. 2014). The NaHCO3 stress tolerance of mandarin fish (96 h LC50 = 5.89 mmol/L) was weaker than that of przewalski’s naked carp (96 h alkalinity LC50 = 150.18 mmol/L) (Wu et al. 2017a), Chalcalburnus chalcoides aralensis (96 h alkalinity LC50 = 112.23 mmol/L) (Lin et al. 2015), Nile tilapia (96 h alkalinity LC50 = 101.30 mmol/L) (Liang et al. 2015), grass carp (96 h alkalinity LC50 = 92.94 mmol/L) and silver carp (96 h alkalinity LC50 = 86.25 mmol/L) (Weight is the same as above) (Yang et al. 2014).
Differences in morphological and phenotypic changes upon different salts were found by observing the gill. Gill is direct contact with water and it is important organs for respiration and ion exchange for fish. Ion regulation is accomplished in large part by 'branchial ionocytes', i.e., specialized cells in the gill epithelium in all fishes. In this experiment, the blood vessels of mandarin fish gill filaments were significantly dilated, and the volume of BCs became larger under NaCl stress. It was hypothesized that NaCl stress caused many ions to enter the plasma of fish, increasing the osmotic pressure of blood. To balance the osmotic pressure balance inside and outside the cell membrane, many ions enter the cell causing the blood cell volume to increase. Under Na2SO4 stress, which is also a neutral salt, it did not show the phenomenon of blood cell volume enlargement. It was manifested by the distortion and deformation of gill filaments. In contrast, the gill of the NaHCO3 stress group showed more damage than that of NaCl and Na2SO4 stress group, and the bulge at the base of the gill lamellae was severe or even completely wrapped around the gill lamellae after stress. This change may delay the entry of harmful substances into the interior of the organism but seriously affected the role of gill in respiration. Therefore, it is a possible reason for the high mortality under NaHCO3 stress in mandarin fish. In addition, the pH of water with different NaCl and Na2SO4 concentration stress was almost the same, whereas the pH increased with increasing NaHCO3 concentration. The increased pH in water can also cause the intolerance of mandarin fish to bicarbonate, since that high pH can damage fish gills and skin, reduce oxygen absorption by gills, disrupt the body's acid-base balance and cause blood alkalosis (Wright and Wood 1985; Zhao et al. 2023).
Differential gene expression and pathway analysis further reveal molecular differences in response to different salt stresses. High salinity affects fish plasma osmolality, antioxidant capacity, protease hydrolysis, and the energy metabolism (Sardella and Brauner 2008; Wu et al. 2017b; Liu et al. 2022). Alkalinity is another factor affecting freshwater fishes. Excessive alkalinity causes inflammation and immune regulation (Shang et al. 2021). The immune response is an important measure of environmental stress in fish and cell adhesion molecules, cell cytokine-cytokine receptor interactions, and ECM-receptor interactions play key roles in inflammation and immune responses (Zhong et al. 2018; Wu et al. 2020; Olbei et al. 2021). Carbon metabolism, biosynthesis of amino acids, and glycolysis/gluconeogenesis pathways are associated with energy supply (Yang et al. 2018; Tsogtbaatar et al. 2020). In the present study, the enrichment of these pathways and the down-regulation of pathway genes suggested that mandarin fish had reduced energy supply and immune response capacity upon NaCl stress. Under Na2SO4 stress in the mandarin fish, differential gene expressions were associated with cell adhesion molecules, amino and nucleotide sugar metabolism, and the biosynthesis of unsaturated fatty and primary immunodeficiency. These pathways are associated with immune response, glucose utilization, and regulation of osmotic pressure in fish (Chen et al. 2020a). It was suggested that Na2SO4 stress caused a decrease in immune capacity and glucose utilization in mandarin fish. In this experiment, NaHCO3 stress caused the enrichment of pathways such as phagosomes, herpes simplex virus type 1 infection, MAPK signaling, and cytokine-cytokine receptor interactions. Phagosomes can maintain cellular homeostasis by removing dead cells or foreign object. And it is a fundamental defense mechanism that activates immune and inflammatory responses against invading pathogens (Uribe-Querol and Rosales 2017). The enrichment of these pathways was associated with tissue damage under NaHCO3 stress in the gills of mandarin fish, which was consistent with the histology results.
PPI networks can highlight the modularity of cellular processes (Fionda 2019). In this experiment, RND2 were up-regulated and RND2 was the core gene that had the most connections in the network under NaCl stress in the gills of mandarin fish. RND2 target the p38 MAPK signalling pathway which were highly expressed after alkali stress in tilapia (Xu et al. 2020; Zhao et al. 2020). In the Na2SO4 stress group, ASPN was the core gene. ASPN is an ECM protein and A interacts with the insulin growth factor receptor (IGF1r) and is essential for activating IGFr-mediated intracellular signaling pathways (Simkova et al. 2016). EPHA4a was the core gene in the NaHCO3 stress group. EPHA4a signaling is located downstream of the mechanical barrier and plays an important role in maintaining cell separation on the surface by regulating actin to separate cells into compartments (Calzolari et al. 2014). The significant association and variability of these hub genes in different salt stress suggest that these genes and their associated genes are involved in salt stress response and that there are differences in the response mechanisms induced by different salt stresses.
The MAPK signaling pathway is an important bridge in the switch from extracellular signals to intracellular responses, including stress to the environment and inflammatory responses (Braicu et al. 2019). In previous studies in Nile tilapia (Zhou et al. 2020; Zhao et al. 2020), target genes of miRNAs associated with alkaline water stress were enriched in the MAPK signaling pathway. In this experiment, differential expressed genes were also enriched in the MAPK signaling pathway under NaHCO3 by transcriptome analysis. Furthermore, qRT-PCR experiments showed that MAPK pathway genes had up-regulated expression trends upon all three salt stresses, but the expression patterns varied with the duration of stress. The results suggest that the MAPK signaling pathway is an important regulatory mechanism in response to salt stress.
Notably, gene expression of MAPK signaling pathway significant increased at freshwater recovery stage upon NaHCO3 stress, while they decreased upon NaCl, Na2SO4 stress, indicating that responding to NaHCO3 stress is stronger than NaCl and Na2SO4 stress with 210 mmol/L Na+.
While gene expression differs upon different salt stresses, there are also some similarities. There were 25 common differentially expressed genes upon NaCl, Na2SO4, and NaHCO3 stress compared to the FW group, of which 23 were consistently down-regulated. Among these down-regulated differential genes, C2H2 and FOXQ1a are transcription factors, GFRA1b and RGS5a are involved in signal transduction; HBB, LOC122868549, and LOC122873121 are involved in oxygen transport; KCNE2 and KCNAB3 are potassium ion voltage-gated channels, DAPK2a and SCEL are associated with apoptosis, and LOC122869198, LOC122869137, NewGene_1818, LOC122869171, and CCL20b are involved in the immune response. The immune response is an important stress response, and the activation of this response in fish during environmental stress and virus infestation is crucial for organismal adaptation (Li et al. 2020; Zhao et al. 2020; Yin et al. 2022). In our study, down-regulation of immune-related genes was common upon all salt stress. It is suspected that high-stress concentrations suppressed immune responses and reduced physiological activity.
In this study, the toxicity and molecular response mechanism of different salts (NaCl, Na2SO4 and NaHCO3) on mandarin fish were reported for the first time, but there were also some shortcomings:Concentrations of the salts (210 mmol/L Na+) are a bit high in acute exposure experiments, especially for NaHCO3 stress. However, it reflects the fact that it causes serious damage upon NaHCO3. In addition, the tolerance of different size of fish to the salt stress also varies. In future studies, low-concentration stress and different of size fish should be chosen to evaluate toxicity of salt stress on mandarin fish comprehensively. To determine the median lethal time (LT50) is also a good way for evaluating toxicity, considering the short survival time for fish exposed to NaHCO3. Besides, the 96 h LC50 was calculated by the stress of artificially configured solution using single chemical agent in these experiments. There were huge differences between the make-up salt solution and the natural saline-alkaline water. Thus, such make-up salt water will low evaluate the tolerance of mandarin fish to these ions in real saline-alkaline water.
Conclusion
In conclusion, there was variation in effects of the three studied salts on mandarin fish. The tolerance of mandarin fish to NaHCO3 stress was weaker than that of NaCl and Na2SO4. The damage to gill tissues by NaHCO3 stress was more severe than that of NaCl and Na2SO4 at equal sodium ion concentrations (210 mmol/L Na+). The MAPK signaling pathway is an important regulatory mechanism in response to salt stress.
Data availability
Not applicable.
References
Braicu C, Buse M, Busuioc C, Drula R, Gulei D, Raduly L, Rusu A, Irimie A, Atanasov AG, Slaby O, Ionescu C, Berindan-Neagoe I (2019) A Comprehensive Review on MAPK: A Promising Therapeutic Target in Cancer. Cancers (Basel) 11:1618. https://doi.org/10.3390/cancers11101618
Breves JP, Hasegawa S, Yoshioka M, Fox BK, Davis LK, Lerner DT, Takei Y, Hirano T, Grau EG (2010) Acute salinity challenges in Mozambique and Nile tilapia: differential responses of plasma prolactin, growth hormone and branchial expression of ion transporters. Gen Comp Endocrinol 167:135–142. https://doi.org/10.1016/j.ygcen.2010.01.022
Calzolari S, Terriente J, Pujades C (2014) Cell segregation in the vertebrate hindbrain relies on actomyosin cables located at the interhombomeric boundaries. EMBO J 33:686–701. https://doi.org/10.1002/embj.201386003
Chen XZ, Lai QF, Yao ZL, Gao HY, Han ZX (2020a) Saline-alkali aquaculture technology model. China. Fisheries 538:61–63 https://kns.cnki.net/kcms2 (in Chinese)
Chen Y, Le Mauff F, Wang Y, Lu RY, Sheppard DC, Lu L, Zhang SZ (2020b) The transcription factor SomA synchronously regulates biofilm formation and cell wall homeostasis in Aspergillus fumigatus. mBio 11:e02329–e02320. https://doi.org/10.1128/mBio.02329-20
Elphick JR, Davies M, Gilron G, Canaria EC, Lo B, Bailey HC (2011) An aquatic toxicological evaluation of sulfate: The case for considering hardness as a modifying factor in setting water quality guidelines. Environ Toxicol Chem 30:247–253. https://doi.org/10.1002/etc.363
Evans DH, Piermarini PM, Choe KP (2005) The multifunctional fish gill: dominant site of gas exchange, osmoregulation, acid-base regulation, and excretion of nitrogenous waste. Physiol Rev 85:97–177. https://doi.org/10.1152/physrev.00050.2003
Fionda V (2019) Networks in Biology. In: Ranganathan S, Gribskov M, Nakai K, Schönbach C (eds) Encyclopedia of Bioinformatics and Computational Biology. Academic Press, Oxford, pp 915–921
Karjalainen J, Hu X, Mäkinen M, Karjalainen A, Järvistö J, Järvenpää K, Sepponen M, Leppänen MT (2023) Sulfate sensitivity of aquatic organism in soft freshwaters explored by toxicity tests and species sensitivity distribution. Ecotoxicol Environ Saf 258:114984. https://doi.org/10.1016/j.ecoenv.2023.114984
Li J, Xue LY, Cao MY, Zhang Y, Wang YJ, Xu SL, Zheng BX, Lou ZJ (2020) Gill transcriptomes reveal expression changes of genes related with immune and ion transport under salinity stress in silvery pomfret (Pampus argenteus). Fish Physiol Biochem 4:1255–1277. https://doi.org/10.1007/s10695-020-00786-9
Liang CF, Jia JH, Zhang YH, Ren BC, Wu JW, Wang Y, Wang Y, Zhao JL (2015) Evaluation on salinity-alkalinity tolerance and growth of the first selection generation of Oreochromis niloticus. Guangdong Agric Sci 42:115–119. https://doi.org/10.16768/j.issn.1004-874x.2015.09.029 (in Chinese)
Lin YH, Wang XH, Ding CL (2015) Research Progress of Chalcalburnus chalcoides Aralensis. J Jilin Agric Univ 37:512–519. https://doi.org/10.13327/j.jjlau.2015.2472 (in Chinese)
Liu YJ, Yao MZ, Li S, Wei XF, Ding L, Han SP, Wang P, Lv BC, Chen ZX, Sun YC (2022) Integrated application of multi-omics approach and biochemical assays provides insights into physiological responses to saline-alkaline stress in the gills of crucian carp (Carassius auratus). Sci Total Environ 822:153622. https://doi.org/10.1016/j.scitotenv.2022.153622
Niu J, Li Z, Zhu J, Wu R, Kong L, Niu T, Li X, Cheng X, Li J, Dai L (2022) Genome-wide identification and characterization of the C2 domain family in Sorghum bicolor (L.) and expression profiles in response to saline-alkali stress. Physiol Mol Biol Plants 28:1695–1711. https://doi.org/10.1007/s12298-022-01222-3
Olbei ML, Thomas J, Hautefort I, Treveil A, Bohar B, Madgwick M, Potari-Gul L, Csabai L, Modos D, Korcsmaros T (2021) CytokineLink: a cytokine communication map to analyse immune responses in inflammatory and infectious diseases. bioRxiv 455874. https://doi.org/10.1101/2021.08.10.455874
Sardella BA, Brauner CJ (2008) The effect of elevated salinity on ‘California’ Mozambique tilapia (Oreochromis mossambicus x O. urolepis hornorum) metabolism. Comp Biochem Physiol Part D Genomics Proteomics 148:430–436. https://doi.org/10.1016/j.cbpc.2008.05.006
Schulz C-J, Cañedo-Argüelles M (2019) Lost in translation: the German literature on freshwater salinization. Philos Trans R Soc Lond Ser B Biol Sci 374:20180007. https://doi.org/10.1098/rstb.2018.0007
Shang X, Geng L, Yang J, Zhang Y, Xu W (2021) Transcriptome analysis reveals the mechanism of alkalinity exposure on spleen oxidative stress, inflammation and immune function of Luciobarbus capito. Ecotoxicol Environ Saf 225:112748. https://doi.org/10.1016/j.ecoenv.2021.112748
Simkova D, Kharaishvili G, Slabakova E, Murray PG, Bouchal J (2016) Glycoprotein asporin as a novel player in tumour microenvironment and cancer progression. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub 160:467–473. https://doi.org/10.5507/bp.2016.037
Su HH, Ma DM, Zhu HP, Liu ZG, Gao FY (2020) Transcriptomic response to three osmotic stresses in gills of hybrid tilapia (Oreochromis mossambicus female× O. urolepis hornorum male). BMC Genomics 21:1–20. https://doi.org/10.1186/s12864-020-6512-5
Tsogtbaatar E, Landin C, Minter-Dykhouse K, Folmes CD (2020) Energy metabolism regulates stem cell pluripotency. Front Cell Dev Biol 8:87. https://doi.org/10.3389/fcell.2020.00087
Uribe-Querol E, Rosales C (2017) Control of phagocytosis by microbial pathogens. Front Immunol 8:1368. https://doi.org/10.3389/fimmu.2017.01368
Wang S, Kuang Y, Liang L, Sun B, Zhao X, Zhang L, Chang Y (2021) Resequencing and SNP discovery of Amur ide (Leuciscus waleckii) provides insights into local adaptations to extreme environments. Sci Rep 11:1–14. https://doi.org/10.1038/s41598-021-84652-5
Wang W, Zhao Y, Wang W, Xu K, Niu L, Yu L, Zou JW, Wang Z, Zhang YJ, Wang F, Zhang S, Yang X (2022) Evaluation of Salt Tolerance Mechanism and Study on Salt Tolerance Relationship of Different Salt-Tolerant Wheat Varieties. Comput Intell Neurosci 2022:2395568. https://doi.org/10.1155/2022/2395568
Wilkie MP, Wood CM (1996) The adaptations of fish to extremely alkaline environments. Comp Biochem Physiol B Biochem Mol Biol 113:665–673. https://doi.org/10.1016/0305-0491(95)02092-6
Wright P, Wood C (1985) An analysis of branchial ammonia excretion in the freshwater rainbow trout: effects of environmental pH change and sodium uptake blockade. J Exp Biol 114:329–353. https://doi.org/10.1242/jeb.114.1.329
Wu HF, Liu JL, Lu Z, Xu LL, Ji CL, Wang Q, Zhao JM (2017b) Metabolite and gene expression responses in juvenile flounder Paralichthys olivaceus exposed to reduced salinities. Fish Shellfish Immunol 63:417–423. https://doi.org/10.1016/j.fsi.2017.02.042
Wu PF, Geng LW, Jiang HF, Tong GX, Li CY, Xu W (2017a) Tolerance of three Cobitidae fish species to high salinity and alkalinity. J Fish Sci China 24:248–257. https://doi.org/10.3724/SP.J.1118.2017.16151 (in Chinese)
Wu RX, Ma C, Liang YX, Chen FM, Liu XH (2020) ECM-mimicking nanofibrous matrix coaxes macrophages toward an anti-inflammatory phenotype: Cellular behaviors and transcriptome analysis. Appl Mater Today 18:100508. https://doi.org/10.1016/j.apmt.2019.100508
Xu Y, Sun Q, Yuan FE, Dong HM, Zhang HK, Geng RX, Qi YZ, Xiong XX, Chen QX, Liu BH (2020) RND2 attenuates apoptosis and autophagy in glioblastoma cells by targeting the p38 MAPK signalling pathway. J Exp Clin Cancer Res 39:1–17. https://doi.org/10.1186/s13046-020-01671-2
Yang J, Xu W, Geng LW, Guan HH, Dang YF, Jiang HF (2014) Effects of salinity on survival, gill and kidney tissue in juveniles of 5 species. Freshwater Fisheries 44:7-12. https://doi.org/10.13721/j.cnki.dsyy.2014.04.002 (in Chinese)
Yang SP, Li X, Liu XF, Ding XB, Xin XB, Jin CF, Zhang S, Li GP, Guo H (2018) Parallel comparative proteomics and phosphoproteomics reveal that cattle myostatin regulates phosphorylation of key enzymes in glycogen metabolism and glycolysis pathway. Oncotarget 9:11352. https://doi.org/10.18632/oncotarget.24250
Yin XX, Bai H, Mu LL, Chen N, Qi WW, Huang Y, Xu HY, Jian JC, Wang AL, Ye JM (2022) Expression and functional characterization of the mannose receptor (MR) from Nile tilapia (Oreochromis niloticus) in response to bacterial infection. Dev Comp Immunol 126:104257. https://doi.org/10.1016/j.dci.2021.104257
Yue GH, Tay YX, Wong J, Shen Y, Xia J (2023) Aquaculture species diversification in China. Aquaculture and Fisheries. https://doi.org/10.1016/j.aaf.2022.12.001
Zhao Y, Wang Y, Zhang CS, Zhou HT, Song LY, Tu HQ, Zhao JL (2023) Variation in pH, HCO3−, carbonic anhydrases, and HCO3− transporters in Nile tilapia during carbonate alkalinity stress. Hydrobiologia 850:2447–2459. https://doi.org/10.1007/s10750-022-05020-6
Zhao Y, Zhang CS, Zhou HT, Song LY, Wang J, Zhao JL (2020) Transcriptome changes for Nile tilapia (Oreochromis niloticus) in response to alkalinity stress. Comp Biochem Physiol Part D Genomics Proteomics 33:100651. https://doi.org/10.1016/j.cbd.2019.100651
Zhong L, Simard MJ, Huot J (2018) Endothelial microRNAs regulating the NF-κB pathway and cell adhesion molecules during inflammation. FASEB J 32:4070–4084. https://doi.org/10.1096/fj.201701536R
Zhou HT, Zhao Y, Zhang CS, Wang YL, Zhao JL (2020) Analysis and Identification of Micro RNA Transcriptome by Deep Sequencing of Nile Tilapia (Oreochromis niloticus) upon Alkalinity Stress. Genom Appl Biol 39:4964–4974. https://doi.org/10.13417/j.gab.039.004964 (in Chinese)
Zhou WJ, Chang YM, Liang LQ, Tang R, Li SG, Ren B, Zhang LM, Dou XJ, Tao R, Han GC, Gao YK (2013) Toxic effects of salinity (NaCl) and alkalinity (NaHCO3) on crucian carp Carassius auratus Linnaeus in Dali Lake. Dalian Ocean Univ 28:340–346. https://doi.org/10.16535/j.cnki.dlhyxb.2013.04.009 (in Chinese)
Acknowledgments
We thank International Science Editing (http://www.internationalscienceediting.com ) for editing the English text of a draft of this manuscript.
Funding
This research was supported by the earmarked fund for CARS-46 (Grant No. CARS-46), the National Key R&D Program of China (2020YFD0900400).
Author information
Authors and Affiliations
Contributions
Yan Zhao contributed to the study design. Material preparation, data collection and analysis were performed by Shuaishuai Li, Shoujie Tang, Yanling Wang, Xiaoli Yao, and Jinyang Xie. The first draft of the manuscript was written by Yan Zhao and Shuaishuai Li. Jinliang Zhao provided funding for the project and reviewed the manuscript. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This study was approved by the Animal Experimentation Ethics Committee of Shanghai Ocean University (Shanghai, China) (SHOU-DW-2016-004).
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Wei Liu
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 397 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, Y., Li, S., Tang, S. et al. Effects of chloride, sulfate, and bicarbonate stress on mortality rate, gill tissue morphology, and gene expression in mandarin fish (Siniperca chuatsi). Environ Sci Pollut Res 30, 99440–99453 (2023). https://doi.org/10.1007/s11356-023-29411-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-29411-x