Abstract
Untreated wastewater emanating from healthcare facilities are risk factors for the spread of antimicrobial resistance (AMR) at the human–environment interface. In this study, we investigated the determinants of resistance in three multidrug resistant strains of Proteus mirabilis isolated from untreated wastewater collected from three government owned hospitals in Ibadan, Nigeria. Despite showing low-level resistance to ciprofloxacin, whole genome sequencing revealed the transferable mechanism of quinolone resistance (TMQR) gene qnrD3 carried on Col3M plasmids in all the isolates. Core genome phylogenetic analysis showed the isolates are closely related differing from each other by ≤ 23 single nucleotide polymorphisms (SNP). Further, they shared the closest evolutionary relationship with isolates from China. Similarly, the Col3M plasmids is most closely related to p3M-2A found in P. vulgaris 3 M isolated from the intestine of shrimps in China. This to the best of our knowledge is the first report of Col3M plasmids carrying qnrD3 in environmental bacterial isolates. Our results indicate a possible silent spread of this important plasmid associated with the dissemination of qnrD3 in Nigeria, and further highlights the important role played by untreated wastewater from healthcare facilities in the spread of AMR in low- and middle-income countries.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The widespread contamination of the natural environment with clinically relevant antimicrobial resistance (AMR) has added a new dimension to the global challenge posed by AMR to human health. This challenge is more acute in low- and middle-income countries (LMIC) due to the pervasive lack of sanitation and hygiene facilities which makes the environment a receptacle for wastes from domestic, industrial, and agricultural sources. Thus, the United Nations Environment Programme (UNEP) recently placed environmental AMR at the top of six emerging issues of environmental concern currently facing mankind (UNEP 2017). Wastewater discharge from healthcare facilities is an important risk factor for environmental contamination with AMR in sub-Sharan Africa (Adelowo et al. 2008; 2018a). Residue of antimicrobials used in the management of infections in these facilities eventually find their way into natural ecosystems through the wastewater generated from these facilities. The risk of wastewater-associated AMR reaching the human population through wastewater is exacerbated by the acute lack of sanitation and hygiene facilities in many healthcare facilities in this region, which translates into a continuous release of untreated wastewater into the environment from these and other sources.
Fluoroquinolones, which were introduced into clinical practice in African countries in the early 2000s, have since become one of the most widely used antimicrobial drugs because of their broad-spectrum activity against enteric pathogens that commonly cause infections in this region (Orogade and Akuse 2004). Their widespread use has led to the emergence of fluoroquinolones resistance among bacteria in the African sub-region, most commonly associated with resistance to other antimicrobial agents (Lamikanra et al. 2011). Mutations in the quinolone resistance-determining regions on bacterial chromosomes were the first known mechanisms of resistance to fluoroquinolones. However, the first transferable mechanism of quinolone resistance (TMQR), qnrA1, was reported in a clinical strain of Klebsiella pneumoniae in 1998 and was followed quickly by reports of other TMQRs that confer resistance via efflux (qepA and oqxAB), antibiotic modification (aac(6′)-lb-cr), and target protection (qnrA, qnrB, qnrC, qnrD, qnrE, qnrS, and qnrVC) (Ruiz 2019).
qnrD, a member of the pentapeptide repeat protein family, was first described in 2009 on small (4.3 kb) non-conjugative plasmids (p2007057) found in clinical strains of Salmonella enterica serovars Bovismorbificans and Kentucky isolated in China (Cavaco et al. 2009). Subsequently, qnrD was reported on smaller sized plasmids (2.7 kb) in several members of the family Morganellaceae with pDIJ09-518a as the archetype (Guillard et al. 2014). To date, only three alleles of qnrD have been described (Ruiz 2019), frequently associated with members of the Morganellaceae and suggesting this family as a possible ancestor of the gene (Guillard et al. 2014). In this family, qnrD is frequently associated with two non-conjugative plasmids ranging in size from 2.7 to 4.2 kb, which are mobilizable by mobile insertion cassette elements (Guillard et al. 2014). Plasmids similar to pDIJ09-518a are disseminated worldwide among members of this family from human (Guillard et al. 2012; Chen et al. 2018; Bitar et al. 2020; Tchuinte et al. 2020) and animal sources (Jones-Dias et al. 2016; Rahman et al. 2020; Zhang et al. 2020). In addition to members of the family Morganellaceae, the plasmid has also been reported in Escherichia coli (Zhang et al. 2013), Citrobacter freundii from livestock (AbuOun et al. 2021), non-enteric Salmonella from swine (Elnekave et al. 2019), and Vibrio parahaemolyticus isolated from wild birds in China (Zheng et al. 2020).
Despite their widespread dissemination in human and animal, reports of qnrD-encoding plasmids from environmental sources have been limited to a few studies (Kraychete et al. 2019) with none of those studies originating from Nigeria. Moreover, sequence-based approaches, which enables a tracking of genes mediating resistance to antimicrobials and their mobile genetic support, is rarely deployed in studies investigating AMR in sub-Saharan Africa (Chattaway et al. 2016; Ikhimiukor et al. 2022) especially in bacteria isolated from environmental sources. In this study, we used whole genome sequencing approach to investigate determinants of low-level fluoroquinolone resistance in three species of Proteus mirabilis isolated from untreated wastewater collected from three government-owned hospitals in Ibadan, South West Nigeria.
Materials and methods
Bacterial species and antimicrobial susceptibility testing
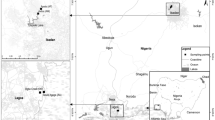
Untreated wastewater was collected in 50-mL sample bottles from three government-owned hospitals located in different areas of Ibadan, the capital city of Oyo State, southwestern Nigeria in May, June, and December 2019, and January 2020 as described (APHA 1998). The three hospitals were located in Yemetu in the Ibadan North Local Government area, Jericho in Ibadan North West, and Ring Road in Ibadan South West Local Government Areas of Oyo State. Samples were processed within 6 h of collection by plating on Eosin Methylene Blue (EMB) agar, McConkey Agar, Cetrimide agar, and Salmonella-Shigella Agar for the isolation of Gram-negative bacteria species. Purified colonies from the plates were stored frozen in 15% glycerol for further analysis. Preliminary identification of bacteria isolates was carried out through biochemical characterization tests as previously described (Bergey and Holt 2000, Cullimore 2019).
The susceptibility of the isolates to nine antibiotics was tested by disc diffusion as described by the Clinical and Laboratory Standards Institute (CLSI 2018). The antibiotics used were imipenem (IMP, 10 µg), cefotaxime (CTX, 30 µg), ceftazidime (CAZ, 30 µg), oxacilin (OX, 1 µg), cefoxitin (FOX, 30 µg), azithromycin (AZM, 15 µg), streptomycin (S, 10 µg), sulphamethoxazole (SXT, 25 µg), and ciprofloxacin (CIP, 5 µg). Discs containing the test antibiotics were layered on Mueller–Hinton agar plates inoculated with standardized saline suspensions (0.5 MacFarland standard) of the test bacteria. Plates were incubated overnight at 37 °C and zones of growth inhibition around each disc were measured and interpreted by zone diameter interpretive standard of the CLSI (CLSI 2018).
DNA extraction and whole genome sequencing
Total genomic DNA was extracted from the bacterial isolates using the DNeasy Blood and Tissue Kit (Qiagen) and the DNA quality checked using a Qubit Fluorometer (ThermoFisher Scientific). Sequencing libraries were prepared using the NEBNext® Ultra™ DNA Library Preparation Kit for Illumina® (New England Biolabs, Frankfurt, Germany) according to the manufacturer’s specifications. The libraries were sequenced on an Illumina MiSeq machine using v3 chemistry and pair-end approaches as described previously (Adelowo et al. 2018b). Raw sequences were subjected to adapter clipping and quality trimming using Trimmomatic (Bolger et al. 2014), reads were assembled with SPAdes v3.6.2 (Bankevich et al. 2012) and assembly quality of the genomes assessed with Quast v5.0.2 (Gurevich et al. 2013) while CheckM v1.0.4 (Parks et al. 2015) was used to check for contamination. Bacterial identity was verified using SpeciesFinder (https://cge.food.dtu.dk/services/SpeciesFinder/) which predicts bacteria identity to species level using the 16S ribosomal DNA sequence. Antibiotic resistance genes (ARG) were searched for using ResFinder 4.1 (Camacho et al. 2009; Bartolaia et al. 2020; Zankari et al. 2020) and the Comprehensive Antimicrobial Resistance Database (CARD) setting the parameter at perfect and strict hits only (Alcock et al. 2020). Plasmids and other mobile genetic elements were searched using PlasmidFinder (Carattoli et al. 2014; Camacho et al. 2009) and MobileElementFinder (MGE) (Johansson et al. 2021). The genomes were annotated using RASTKit v1.073 (Arkin et al. 2018) for further analysis.
Genetic relatedness of isolates
In addition to the three P. mirabilis isolates from our study, we searched for publicly available P. mirabilis genomes carrying qnrD from the National Centre for Biotechnology Information (NCBI) isolate Browser database, and included the database hits in our analysis to compare their genetic relatedness. Average nucleotide identity (ANI) of orthologous gene pairs within the isolates were defined using FastANI, and an ANI percentage > 95% was used to confirm the identities of the genomes to species level. The genomes were annotated using Prokka v.1.14.6 (Seemann 2014). Annotated genomes were used as input to carry out a pangenome analysis using Roary v3.13.0 (Page et al. 2015). Single nucleotide polymorphisms (SNP) were extracted from the resulting alignment of core genes (genes shared in > 95% of genomes) using SNP-sites (Page et al. 2016). The SNP alignment were used as input to infer a Maximum-Likelihood phylogeny using RAxML v8.2.12 using the GTR GAMMA model (Stamatakis 2014). The phylogenetic tree was visualized and annotated using figtree v.1.4.4 (http://tree.bio.ed.ac.uk/software/figtree/) and Interactive Tree of Life IToL (Letunic and Bork 2019). We used snp-dists v0.8.2 to determine pairwise SNP distance of core genome alignment (https://github.com/tseemann/snp-dists). The SNPs from the core genome alignment was used as input in building a minimum spanning tree using Grapetree (Zhou et al. 2018).
In silico plasmid analysis
The sequences of forty qnrD-bearing plasmids and plasmid p3M-2A were downloaded from GenBank (https://blast.ncbi.nlm.nih.gov) and aligned using MUSCLE (Madeira et al. 2022). The aligned sequences were used to plot a phylogenetic tree to show the evolutionary relationship of the plasmids (pNgM_1_qnrD3, pNgM_5_qnrD3 and pNgM_6_qnrD3) detected in the three Nigerian isolates with similar plasmids in other parts of the world.
Data availability
The whole genome sequences of the three isolates are deposited in the GenBank under BioProject Number PRJNA877695, BioSample Numbers SAMN30722190-SAMN30722192 and accession numbers JAODPC000000000- JAODPE000000000.
Results and discussion
Despite the current widespread distribution of TMQR genes which mediates reduced resistance to fluoroquinolones in clinical and environmental bacteria in different regions of the world, the prevalence of the qnrD variant has been low with reports to date limited to only three allelic variants (Ruiz 2019). While previous studies have reported TMQR genes (including qnrD) in bacteria from clinical and non-clinical sources in Nigeria (Ogbolu et al. 2011, 2016; Sumrall et al. 2014; Nsofor et al. 2021; Adekanmbi et al. 2022), none of these have reported the qnrD3 allelic variant either in clinical or environmental bacteria strains. Similarly, there is no detailed characterization of the plasmid backbone carrying this TMQR variant in Nigeria and their relationship to similar plasmid backbones from other regions of the world. Here we describe in details for the first time the presence of qnrD3 carried on Col3M plasmids in three P. mirabilis isolated from hospital wastewater in southwestern Nigeria.
Bacterial isolates and their susceptibility to antibiotics
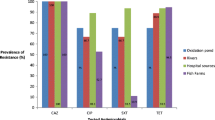
Three bacterial identified as P. mirabilis through biochemical characterization tests (Bergey and Holt 2000; Cullimore 2019) and analysis of the 16S rRNA gene ((https://cge.food.dtu.dk/services/SpeciesFinder/) were isolated from wastewater collected from three government-owned hospitals. The bacterial isolates were designated as strains M-1, M-5, and M-6. Strain M-1 was isolated in June 2019 from the wastewater collected at the State Government hospital located in Yemetu, while strains M-5 and M-6 were isolated in January 2020 and December 2019, respectively, from the State Government hospitals located in Jericho (M-5) and Ring Road (M-6), respectively. The three isolates were multidrug resistant showing resistance to five (M_1) and four (M_5 and M_6) different classes of antimicrobial drugs, respectively, including a common intermediate level resistance to CIP (Table 1). The three isolates shared phenotypic resistance to CAZ, CTX, FOX, OXA, and SXT. In addition, isolates M_1 and M_6 shared resistance to AZM, M_1 and M_5 shared resistance to streptomycin and isolate M_1 showed phenotypic resistance to IMP, the only carbapenem included in the antimicrobial susceptibility testing. Consistent with this observation, P. mirabilis, which usually show intrinsic resistance to many antibiotics including colistin, nitrofurans, tetracycline, and tigecycline and reduced or outright susceptibility to several others including imipenem and fluoroquinolones (Girlich et al. 2020) is currently emerging as an important reservoir of clinically relevant ARGs.
Whole genomes and phylogeny of genetic similarity of P. mirabilis
The isolates were whole genome sequenced to investigate their genetic relatedness and genotypic mechanisms of resistance to antimicrobials. The sizes of the three sequenced genomes from our study were 3.78 Mbp, 3.79 Mbp, and 3.78 Mbp for strains M_1, M_5, and M_6, respectively. The three genome assemblies have ≤ 51 contigs and a G + C content of 38.5% (Table 1). Based on ANI comparison using FastANI, the identities of the three Nigerian isolates was further confirmed as P. mirabilis (ANI identity of > 95%) (Fig. 1a). Alongside the three Nigeria isolates, thirty-seven publicly available high-quality genomes (originating from China (n = 32), USA (n = 1), Hong Kong (n = 1), and Germany (n = 1), and two isolates whose country of origin were unknown) having < 200 contigs, N50 > 40,000, and contamination < 5%, were included in pangenome construction (Supplementary Table 1).
Pangenome statistics identified 2430 core genes (genes present in ≥ 99% of the genomes), 336 soft core genes (genes present in 95 to < 99% of the genomes), 1542 shell genes (genes present in 15 to < 95% of the genomes) and 8197 cloud genes (genes present in < 15% of the genomes) in the genome collection. Pairwise SNPs distances of core gene alignment showed that the three P. mirabilis isolates from our study differed by ≤ 23 SNPs. The SNP distance between M5 and M6 = 1, M1 and M6 = 16, M1 and M5 = 23, thus showing that these strains are genetically highly similar (Fig. 1b, Supplementary Table 2). This suggests a possible silent dissemination of very closely related strains within the hospital source of the wastewater from where the bacteria species were isolated. Core genome phylogeny analysis revealed two large and distinct clades, one of which contained the three isolates from this study and another genome from China, sharing closest evolutionary relationship (Fig. 2). Core genome pairwise SNP distances between the genomes of the three isolates in the present study and the genome from China in this clade was 7629 (Supplementary Table 2).
The qnrD3-habouring P. mirabilis strains have exactly identical resistomes
To understand more about the genetic basis of the observed multidrug resistance among the isolates, we screened the genomes for ARGs through the ResFinder database. However, we observed there was no concordance between the phenotypic resistance and antibiotic resistance genotype of the three isolates. Despite showing phenotypic resistance to four and five classes of antibiotics, respectively, acquired ARG for beta-lactams, foliate inhibitors (SXT), macrolides (AZM) and aminoglycosides (S) were surprisingly not detected in the genomes of the three P. mirabilis isolates. ResFinder analysis however showed that the three isolates carried catA4 and qnrD3 conferring resistance to chloramphenicol and fluoroquinolones, respectively. Chloramphenicol was however not included in the antibiotics used for the antimicrobial susceptibility testing. The qnrD3 shared 100% identity with qnrD3 of E. coli EC68 (NG057448.1) isolated from bird feces in the United States while the catA4 shared 100% identity with catA of P. mirabilis strain S74-3–2 (CP073245.1) isolated from a tiger in China.
In addition to the aforementioned ARG, CARD analysis revealed strict hits corresponding to multidrug resistance efflux genes and genes that confer resistance by target alteration. Detected genes include three multi-antibiotic efflux pumps of the resistance nodulation-cell division (RND) family CRP, rsmA and adeF, two major facilitator superfamily (MFS) antibiotic efflux pumps kpnH and kpnF, and qacJ, an efflux pump of the small multidrug resistance (SMR) family. Between them, these efflux pumps confer resistance to penams, macrolides, fluoroquinolones, phenicols, diaminopyrimidine, tetracycline aminoglycosides, cephalosporins, peptide antibiotic, rifamycins, and carbapenems as well as disinfecting agents and antiseptics. Also detected in the genomes through CARD analysis are mutations in gyrB (S463A) and PBP3 (D350N) conferring resistance to fluoroquinolones and beta-lactams. It is quite possible that the presence of these genes and mutations in the genomes contributed to the multidrug resistance phenotype of the P. mirabilis isolates. This will however require further experimental proof beyond the scope of the present study.
Unlike the three Nigeria isolates, a large proportion of the 37 isolates included in the phylogenetic analysis however carried multiple acquired ARG in their genomes. This suggests that the evolution of the Nigerian isolates towards multidrug resistance through acquisition of ARG is still in the early stages.
qnrD3 was carried on Col3M plasmids
While no plasmids were detected in the genomes of the three isolates by PlasmidFinder, MGE analysis showed the presence of Col3M plasmids sharing 93% identity with Col3M plasmid p3M-2A (JX514065.1) found in P. vulgaris 3M isolated from shrimp in China (Zhang et al. 2020). p3M-2A (2656 bp) was found together with a different sized (5903 bp) but identical plasmid p3M-2B in P. vulgaris 3M. No additional plasmid similar to p3M-2B was detected in the three P. mirabilis isolates of the present study. Interestingly, no resistance gene was found on p3M-2A by Zhang et al (2020), but further analysis confirmed that p3M-2A has a positive regulatory effect on the qnrD carried by the sister plasmid p3M-2B leading to an increase in ciprofloxacin MIC in isolate 3M (Zhang et al. 2020). It thus appears that the plasmids in the present isolates may have evolved from p3M-2A but diversified through the recruitment of qnrD3 onto the plasmid backbone. Indeed, the current plasmids are only slightly bigger in size (by ~ 43 bp) than p3M-2A despite the incorporation of qnrD3 onto the p3M-2A backbone suggesting that a portion of the backbone was deleted to accommodate the qnrD3 in the present variant of the plasmid. Alignment of the three plasmid contigs with the sequence of p3M-2A indeed revealed deletions in p3M-2A and the plasmids detected in the Nigerian isolates.
Manual inspection of RASTKit annotated contigs corresponding to the Col3M plasmids in the genome of the Nigerian isolates showed that qnrD3 was split into two fragments with the insertion of two open reading frames (orf) in between the two halves. The first fragment of 519 bp was directly followed by an orf which on tblastn analysis did not share any significant similarity with any sequence in the GenBank database. This orf was directly followed by another orf which shared 82.4% identity (77% query coverage) with a portion coding for hypothetical protein on plasmid pHBNNC5-qnrD3 of P. mirabilis HBNNC5 (MT349900.1), one of two isolates carrying the qnrD3-bearing Col3M plasmids in the GenBank. The second orf was followed by another fragment (261 bp) in which the first 138 bp were a direct repeat of the last 138 bp of the first fragment. Similar to our observation on the first orf directly following the first fragment of qnrD3 on the present plasmid, orf1 of p3M-2A also did not show significant similarity with any sequence in the database but was shown to play a role in regulating the expression of qnrD3 on p3M-2B (Zhang et al. 2020). It however remains unknown whether this orf is performing similar regulatory function in the present plasmids. These observations indicate that much is still unknown about this plasmid lineage.
In fact, as at the time of this study, only two Col3M plasmids carrying qnrD3 are deposited in the GenBank database found in P. mirabilis strain HBNNC5 (MT349900.1) from dairy cow and P. mirabilis strain SCRJC7 (MT349902.1) from broiler chicken, both isolated in China, suggesting a recent acquisition of the qnrD3 onto the Col3M backbone. This to the best of our knowledge is the first report of qnrD3-bearing Col3M plasmids in environmental isolates of P. mirabilis and environmental isolates of the family Morganellaceae. Consistent with the characteristics of the two qnrD-bearing Col3M plasmids previously deposited in the GenBank (MT349900.1, MT349902.1), qnrD3 was the only ARG present on the Col3M plasmids found among the Nigerian isolates.
Comparison of the plasmids with others in the NCBI database showed that the three Nigerian plasmids shared the closest evolutionary relationship with Col3M plasmids found in S. enterica subsp. enterica serovar Heidelberg strain 69 (MK191843.1) isolated from swine in USA (Elnekave et al. 2019) and P. mirabilis strain CRE14IB (CP045539.1) isolated from a urine catheter in Italy (Bitar et al. 2020) (Fig. 3, Supplementary Table 3). However, while the allelic variant of the qnrD carried on the plasmids found in the isolates of Elnekave et al. (2019) was not specified, the plasmids found in the isolate of Bitar et al (2020) carried qnrD1. The plasmids in the Nigeria isolates are distantly related to p3M-2A and the qnrD3-bearing plasmids found in the two previously mentioned isolates (MT349900.1, MT349902.1) from China. However, in contrast to the single cluster formed by the three isolates in core genome phylogenetic analysis, the plasmids formed two closely related clusters. One cluster was formed by pNgM_1_qnrD3 and pNgM_5_qnrD3 while pNgM_6_qnrD3 formed a single cluster branching out from the M_1/M_5 cluster suggesting that the three plasmids in the Nigerian isolates are probably two closely related lineages.
Conclusion
We report the isolation of three strains of P. mirabilis carrying qnrD3 on Col3M plasmids from untreated wastewater collected from three government hospitals located in different areas of Ibadan, the third largest populated city in Nigeria. This is the first report of this important plasmid carrying qnrD3 in environmental isolates of the family Morganellaceae. The high degree of similarity between the isolates and the plasmids they carry may be an indication of a silent spread of the isolates/plasmids within the hospital origin of the wastewater from where the strains were isolated and possibly within the surrounding communities. Taking the present results into account, further genomic surveillance may be warranted to understand the ecology of this plasmids and their role in fluoroquinolone resistance within this setting.
Data availability
All relevant data are available in this manuscript and accompanying supplementary materials.
References
AbuOun M, Jones H, Stubberfield E et al (2021) Genomic epidemiological study shows that prevalence of microbial resistance in Enterobacterales is associated with the livestock host, as well as antimicrobial usage. Microb Genomics 7:000630. https://doi.org/10.1099/mgen.0.000630
Adelowo OO, Caucci S, Banjo OA et al (2018) Extended spectrum beta-lactamase (ESBL)-producing bacteria isolated from hospital wastewater, rivers and aquaculture sources in Nigeria. Environ Sci Pollut Res 25(3):2744–2755
Adelowo OO, Vollmers J, Mäusezahl I, Kaster AK, Müller JA (2018) Detection of the carbapenemase gene blaVIM-5 in members of the Pseudomonas putida group isolated from polluted Nigerian wetlands. Sci Rep 8:15116. https://doi.org/10.1038/s41598-018-33535-3
Adelowo OO, Fagade OE, Oke AJ (2008) Prevalence of co-resistance to disinfectants and clinically relevant antibiotics in bacterial isolates from three hospital laboratory wastewaters in southwestern Nigeria. World J Microbiol Biotechnol 24(9):1993–1997
Adekanmbi OA, Akinlabi OC, Olaposi AV (2022) High carriage of plasmid mediated quinolone resistance (PMQR) gene by cefotaxime-resistant Escherichia coli recovered from surface-leaking sanitary sewers. Arch Microbiol 204:131. https://doi.org/10.1007/s00203-021-02627-6
Alcock BP, Raphenya AR, Lau TTY et al (2020) CARD 2020: antibiotic resistome surveillance with the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res 48(D1):D517–D525. https://doi.org/10.1093/nar/gkz935
APHA (1998) Standard methods for the examination of water and wastewater. American Public Health Association, Washington DC
Arkin AP, Cottingham RW, Henry CS et al (2018) Kbase: The United States Department of Energy Systems Biology Knowledgebase. Nat Biotechnol 38:566–569
Bankevich A, Nurk S, Antipov D et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Bartolaia V, Kaas RF, Ruppe E et al (2020) ResFinder 4.0 for predictions of phenotypes from genotypes. J Antimicrob Chemother 75(12):3491–3500
Bergey DH, Holt JG (2000) Bergey’s manual of determinative bacteriology, 9th edn. Lippincott, Williams and Wilkins, Philadelphia
Bitar I, Marchetti VM, Mercato A et al (2020) Complete genome and plasmids sequences of a clinical Proteus mirabilis isolate producing plasmid mediated NDM-1 from Italy. Microorganisms 8:339. https://doi.org/10.3390/microorganisms8030339
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Carattoli A, Zankari E, Garcia-Fernandez A et al (2014) PlasmidFinder and pMLST: in silico detection and typing of plasmids. Antimicrob Agents Chemother 58(7):3895–3903
Camacho C, Coulouris G, Avagyan V et al (2009) BLAST+: architecture and applications. BMC Bioinformatics 10(1):421
Cavaco LM, Hasman H, Xia S, Aarestrup FM (2009) qnrD, a novel gene conferring transferable quinolone resistance in Salmonella enterica serovar Kentucky and Bovismorbificans strains of human origin. Antimicrob Agents Chemother 53:603–608. https://doi.org/10.1128/AAC.00997-08
Chattaway MA, Aboderin AO, Fashae K, Okoro CK, Opintan JA, Okeke IN (2016) Fluoroquinolone-resistant enteric bacteria in Sub-Saharan Africa: Clones, implications and research needs. Front Microbiol 7:558. https://doi.org/10.3389/fmicb.2016.00558
Chen L, Zhang Y, Du J, Zhang X, Li M, Chen H, Yu H, Sun Y, Zhou T (2018) Description and plasmid characterization of the qnrD determinant in Proteeae in Wenzhou, Southern China. J Microbiol Immunol Infect 51:115–122. https://doi.org/10.1016/j.jmii.2016.02.001
Clinical and Laboratory Standards Institute (CLSI) (2018) Performance standards for antimicrobial susceptibility testing. CLSI Supplement M100. 27th Edition. Wayne, USA
Cullimore RD (2019) Practical atlas for bacterial identification, 2nd edn. CRC Press, Florida
Elnekave E, Hong SL, Lim S, Hayer SS, Boxrud D, Taylor AJ, Lappi V, Noyes N, Johnson TJ, Rovira A, Davies P, Perez A, Alvarez J (2019) Circulation of plasmids harboring resistance genes to quinolones and/or extended spectrum cephalosporins in multiple Salmonella enterica serotypes from swine in the United States. Antimicrob Agents Chemother 63:e02602-e2618. https://doi.org/10.1128/AAC.02602-18
Girlich D, Bonnin RA, Dortet L, Naas T (2020) Genetics of acquired antibiotic resistance genes in Proteus spp. Front Microbiol 11:256. https://doi.org/10.3389/fmicb.2020.00256
Guillard T, Grillon A, de Champs C et al (2014) Mobile insertion cassette elements found in small non-transmissible plasmids in Proteeae may explain qnrD mobilization. PLoS One 9:e87801. https://doi.org/10.1371/journal.pone.0087801
Guillard T, Cambau E, Neuwirth C, Nenninger T, Mbadi A, Brasme L, Vernet-Garnier V, Bajolet O, de Champs C (2012) Description of a 2,683-base-pair plasmid containing qnrD in two Providencia rettgeri isolates. Antimicrob Agents Chemother 56:565–568. https://doi.org/10.1128/AAC.00081-11
Gurevich A, Saveliev V, Vyahhi N, Tesler G (2013) QUAST: quality assessment tool for genome assemblies. Bioinformatics 29(8):1072–1075
Ikhimiukor OO, Odih EE, Donado-Godoy P, Okeke IN (2022) A bottom-up view of antimicrobial resistance transmission in developing countries. Nat Microbiol 7:757–765. https://doi.org/10.1038/s41564-022-01124-w
Johansson MHK, Bortolaia V, Tansirichaiya S, Aarestrup FM, Roberts AP, Petersen TN (2021) Detection of mobile genetic elements associated with antibiotic resistance in Salmonella enterica using a newly developed web tool: MobileElementFinder. J Antimicrob Chemother 76:101–109. https://doi.org/10.1093/jac/dkaa390
Jones-Dias D, Clemente L, Moura IB, Sampaio DA, Albuquerque T, Vieira L, Manageiro V, Caniça M (2016) Draft genomic analysis of an avian multidrug resistant Morganella morganii isolate carrying qnrD1. Front Microbiol 7:1660. https://doi.org/10.3389/fmicb.2016.01660
Kraychete GB, Campana EH, Picão RC, Bonelli RR (2019) qnrD-harboring plasmids in Providencia spp. recovered from food and environmental Brazilian sources. Sci Total Environ 646:1290–1292. https://doi.org/10.1016/j.scitotenv.2018.07.378
Lamikanra A, Crowe JL, Lijek RS et al (2011) Rapid evolution of fluoroquinolone-resistant Escherichia coli in Nigeria is temporally associated with fluoroquinolone use. BMC Infect Dis 11:312. https://doi.org/10.1186/1471-2334-11-312
Letunic I, Bork P (2019) Interactive Tree of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47(W1):W256–W259. https://doi.org/10.1093/NAR/GKZ239
Madeira F, Pearce M, Tivey ARN, Basutkar P, Edbali O, Madhusoodanan N, Kolesnikov A, Lopez R (2022) Search and sequence analysis tools services from EMBL-EBI in 2022. Nucleic Acids Res 50(W1):W276–W279. https://doi.org/10.1093/nar/gkac240
Nsofor CM, Tattfeng MY, Nsofor CA (2021) High prevalence of qnrA and qnrB genes among fluoroquinolone-resistant Escherichia coli isolates from a tertiary hospital in southern Nigeria. Bullet National Res Centre 45:26. https://doi.org/10.1186/s42269-020-00475-w
Ogbolu DO, Daini OA, Ogunledun A, Alli AO, Webber MA (2011) High levels of multidrug resistance in clinical isolates of Gram negative pathogens from Nigeria. Int J Antimicrob Agents 37:62e6
Ogbolu DO, Alli AO, Anorue MC, Daini OA, Oluwadun A (2016) Distribution of plasmid-mediated quinolone resistance in Gram-negative bacteria from a tertiary hospital in Nigeria. Indian J Pathol Microbiol 59:322–326
Orogade AA, Akuse RM (2004) Changing patterns in sensitivity of causative organisms of septicaemia in children: the need for quinolones. Afr J Med Sci 33:69–72
Page AJ, Cummins CA, Hunt M, Wong VK, Reuter S, Holden MTG, Fookes M, Falush D, Keane JA, Parkhill J (2015) Roary: rapid large-scale prokaryote pangenome analysis. Bioinformatics 31(22):3691–3693
Page AJ, Taylor B, Delaney AJ, Soares J, Seemann T, Keane JA, Harris SR (2016) SNP-sites: rapid efficient extraction of SNPs from multi-FASTA alignments. Microb Genomics 2(4):e000056. https://doi.org/10.1099/MGEN.0.000056
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW (2015) CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25:1043–1055
Rahman A, Bhuiyan OF, Sadique A et al (2020) Whole genome sequencing provides genomic insights into three Morganella morganii strains isolated from bovine rectal swabs in Dhaka, Bangladesh. FEMS Microbiol Lett 367:fnaa043. https://doi.org/10.1093/femsle/fnaa043
Ruiz J (2019) Transferable mechanisms of quinolone resistance from 1998 onward. Clin Microbiol Rev 32:e00007-19. https://doi.org/10.1128/CMR.00007-19
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30(14):2068–2069. https://doi.org/10.1093/BIOINFORMATICS/BTU153
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313
Sumrall ET, Gallo EB, Aboderin AO, Lamikanra A, Okeke IN (2014) Dissemination of the transmissible quinolone-resistance gene qnrS1 by IncX Plasmids in Nigeria. PLoS One 9(10):e110279. https://doi.org/10.1371/journal.pone.0110279
Tchuinte PLS, Rabenandrasana MAN, Ramparany L, Ratsima E, Enouf V, Randrianirina F, Collard J-M (2020) Genome-based insight into the resistomes and mobilomes of two Providencia rettgeri strains isolated from wound infections in Madagascar. J Global Antimicrob Resist 20:178–182. https://doi.org/10.1016/j.jgar.2019.07.013
UNEP (2017) Frontiers 2017: emerging issues of environmental concern. United Nations Environment Programme, Nairobi
Zankari E, Allesøe R, Joensen KG, Cavaco LM, Lund O, Aarestrup FM (2020) PointFinder: a novel web tool for WGS-based detection of antimicrobial resistance associated with chromosomal point mutations in bacterial pathogens. J Antimicrob Chemother 72(10):2764–2768
Zhang S, Sun J, Liao X-P et al (2013) Prevalence and plasmid characterization of the qnrD determinant in Enterobacteriaceae isolated from animals, retail meat products, and humans. Microb Drug Resist 19:331–335. https://doi.org/10.1089/mdr.2012.0146
Zhang H, Chang M, Zhang X, Cai P, Dai Y, Song T, Wu Z, Xu H, Qiao M (2020) Functional identification and evolutionary analysis of two novel plasmids mediating quinolone resistance in Proteus vulgaris. Microorgnisms 8:1074. https://doi.org/10.3390/microorganisms8071074
Zheng L, Chen P, Guo JX, Zhu LW, Guan YJ, Wang Y, Jing J, Liang B, Ji X (2020) Virulence gene, antimicrobial resistance and phylogenetic characterization of Vibrio parahaemolyticus in migratory birds, Guangdong. PREPRINT (Version 1) available at Research Square. https://doi.org/10.21203/rs.3.rs-23919/v1. Accessed 25 July 2022
Zhou Z, Alikhan NF, Sergeant MJ et al (2018) GrapeTree: visualization of core genomic relationships among 100,000 bacterial pathogens. Genome Res 28(9):1395–404. https://genome.cshlp.org/content/28/9/1395.full
Funding
This study was supported by an Alexander von Humboldt (AvH) Foundation Renewed Research Stay Fellowship to OOA at the Institute of Biological Interfaces (IBG-5), Karlsruhe Institute of Technology, Germany, for the sequencing of the bacteria isolates.
Author information
Authors and Affiliations
Contributions
Adenike Omolola Ajayi-Odoko (AOA), Ayantade Dayo Victor Ayansina (ADVA), and Olawale Olufemi Adelowo (OOA) conceived and designed the study; AOA, OOA, Jochen A. Müller (JAM), and Odion O. Ikhimiukor performed experiment, collected, and analyzed data. OOA and JAM acquired funding. The first draft of the manuscript was written by OOA, all authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent to publish
This publication has been approved by all co-authors and the responsible authorities at the institutes where the work was carried out.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Diane Purchase
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
11356_2023_25618_MOESM1_ESM.csv
Supplementary file1 Genome assembly characteristics of the thirty-seven isolates included in the FastANI plot (CSV 4.13 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ajayi-Odoko, A.O., Ayansina, A.D.V., Ikhimiukor, O.O. et al. Proteus mirabilis isolated from untreated hospital wastewater, Ibadan, Southwestern Nigeria showed low-level resistance to fluoroquinolone and carried qnrD3 on Col3M plasmids. Environ Sci Pollut Res 30, 47158–47167 (2023). https://doi.org/10.1007/s11356-023-25618-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-25618-0