Abstract
This study investigates the antibiotic resistance fate in the urban water cycle, evaluating the dynamics of antibiotic-resistant bacteria (ARB) and antibiotic-resistant genes (ARGs) in three different full-scale wastewater treatment plants (WWTPs) and two drinking water treatment plants (DWTPs) located in the same geographical area (North-West of Italy). ARB (tetracycline-, ampicillin-, and sulfonamide-resistant bacteria) were quantified by plate counting and the abundances of selected ARGs (i.e., tetA, blaTEM, and sulII) and intI1 gene were measured using quantitative real-time PCR (qPCR). Higher concentrations of ARB and ARGs were observed in the WWTPs with respect to the DWTPs identifying the WWTP as hotspot for the spread of antibiotic resistances. Although a significant reduction of ARB and ARGs was observed in WWTPs and DWTPs after the treatment, none of the detected ARB or ARGs was completely removed in drinking water. The stability of the antibiotic-resistant rates between inlet and outlet associated with the reduction of relative ARG abundances underlined that both the treatments (WWTs and DWTs) did not apply any selective pressure. The overall results highlighted the importance to investigate the antibiotic resistance dynamics in aquatic ecosystems involved in urban water cycle integrating the information obtained by culture-dependent method with the culture-independent one and the need to monitor the presence of ARB and ARGs mainly in drinking water that represents a potential route of transmission to human.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Antimicrobial resistance is of primary concern for the public health (WHO 2014). Globally, it is estimated that 700,000 people each year could die because of antimicrobial-resistant bacterial infections (Carvalho and Santos 2016). If effective interventions are not carried out to overcome infections attributable to microorganisms resistant to antimicrobials, there could be an increase in deaths estimated up to 10 million people in the world by 2050, each year (O’Neil 2016). The main driver for the spread and the persistence of antibiotic resistance is the overuse and misuse of antibiotics in human and animal medicine (Sanganyado and Gwenzi 2019). Antibiotic-resistant bacteria and resistance genes (ARB and ARGs) together with residues of antibiotics are released in sewer that constitutes a reservoir of ARB and ARGs (Rizzo et al. 2013).
In this context, waters, favoring dispersion and smoothing physical factors (e.g. temperature, UV radiation) reducing the survival rates of allochthonous bacteria in the environment, allow the spread of human- and animal-derived bacteria; therefore, it plays an important role in the release of antibiotic resistances into the environment. In particular, the urban water cycle that includes the collection, treatment, and discharge of treated wastewater into the environment as well as the abstraction, disinfection, and distribution of water for drinking purposes represents an interesting model for tracing the fate of antibiotic resistance in the environment and for assessing the risk of transmission to humans (Manaia et al. 2016; Almakki et al. 2019).
In the last two decades, several studies reported the presence of ARB and ARGs in different concentrations in aquatic environments all over the world (Baquero et al. 2008; Li et al. 2015; Yang et al. 2019). In particular, urban wastewater treatment plants (WWTPs) are among the most important receptors and hotspots for the release of antibiotic resistance into the environment (Rizzo et al. 2013). A recent overview on ARG occurrence in WWTPs highlights that the absolute abundance of the most frequently detected ARGs in influent worldwide ranged from 4.5 to 7 log copies/mL (Wang et al. 2020). Also antibiotic-resistant fecal indicators (Escherichia coli, coliforms) or ARB isolated from heterotrophic flora were frequently reported in WWTP influents, with a mean concentration of 4 and 6 log of colony-forming units (CFU)/mL detected in different studies for tetracycline- and sulfonamide-resistant bacteria, respectively (Munir et al. 2011; Gao et al. 2012).
Despite the wastewater treatments generally lead to a reduction of ARG and ARB abundance (~ 2–3 logs), some studies observed higher resistance rates in the effluents with respect to the influent, in relation to the different treatment processes investigated (Pazda et al. 2019; Wang et al. 2020; Stachurova et al. 2021).
Thus, it is important to underline that even a well-functioning WWTP equipped with secondary and even tertiary treatment will be able to release high concentrations of ARB and ARGs into the environment. Some studies reported that, in a final effluent of a WWTP, it is possible to detect about 109–1012 CFU of total bacteria per day per inhabitant equivalent; of these, at least 107–1010 showed some form of antibiotic resistance (Rizzo et al. 2013). Moreover, a recently pan-European survey on treated wastewater demonstrates that WWTPs are responsible for the discharge of considerable amounts of ARGs in the downstream water bodies (Cacace et al. 2019). These data underline the main role of WWTPs in the accumulation and spreading of ARB and ARGs into open waters.
Several studies evaluated the antibiotic resistance in WWTPs worldwide observing highly variable abundances of ARB and ARGs; these results suggested the influence of local environmental and anthropogenic factors (e.g., antimicrobial residue levels, bacterial taxonomic composition, local use of antimicrobials) and they highlighted the need to specifically evaluate antibiotic resistance in different geographical areas (Hendriksen et al. 2019).
The investigation of antibiotic resistance in the urban water cycle represents a relevant issue considering the risk that ARB and ARGs in the “unclean” phases of the cycle can reach the final consumer of the water (Manaia et al. 2016). Although the direct impact on human health by ARB and ARGs in drinking water is not well established, antibiotic resistance threats human health by two main different mechanisms: first, pathogenic and opportunistic ARB that can survive in drinking water may also enter the human microbiome following water consumption (Vaz-Moreira et al. 2014); second, ARGs in drinking water can occasionally be transferred by horizontal gene transfer (HGT) to human pathogenic bacteria (Manaia 2017). Therefore, drinking water can be considered as a potential transmission route of antibiotic resistance to humans (Chang et al. 2015; Manaia et al. 2016).
Although drinking water treatment drastically abates the overall bacterial numbers, including the number of ARB and ARGs, still a small but quantifiable number of cells and genes have been detected in several drinking water systems. Indeed, the standard drinking water treatment plants (DWTPs) are not specifically designed to reduce ARB and ARGs (Huang et al. 2021). Moreover, biofilm formation (Zhang et al. 2019), presence of chlorine residues (Bai et al. 2015), and heavy metals allow the persistence of antibiotic resistances in drinking water systems (Seiler and Berendonk 2012).
This study investigates the antibiotic resistance fate in the urban water cycle, evaluating the dynamics of ARB and ARGs in three different full-scale WWTPs and two DWTPs located in the North-West of Italy.
The combination of culture-dependent and culture-independent approach was used. In particular, tetracycline-, ampicillin-, and sulfonamide-resistant bacteria in the influent and in the final effluent of each WWTP and DWTP were quantified by plate counting. Moreover, the abundances of selected ARGs (i.e., tetA, blaTEM, and sulII; against tetracycline, β-lactams, and sulfonamides, respectively) as well as those of the integrase gene of the class 1 integrons (intI1), used as proxy of the anthropogenic pollution and of the antibiotic resistance in the environment (Gillings et al. 2015; Ma et al. 2017), were measured using quantitative real-time PCR (qPCR).
Materials and methods
Sampling
WWTPs
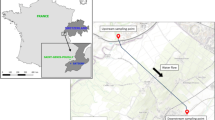
Influent and effluent samples were collected in three WWTPs (plants A, B, and C) located in North-West of Italy. The WWTP A (population equivalent of 3,800,000) employs preliminary treatment (screening and aerated grit removal), primary sedimentation, biological process (denitrification/oxidation/nitrification), secondary settling, then phosphorus removal and filtration steps (multilayer anthracite/sand beds). The WWTPs B and C (60,000 and 276,000 population equivalent, respectively) after screening and grit removal employ a denitrification step, the biological treatment, and the secondary settling. In addition, the WWTP B has a primary sedimentation step. Finally, a tertiary treatment is carried out with ultrafiltration in WWTP B and with a chlorination step in WWTP C.
Six wastewater samplings were performed during 1 year (March 2019–January 2020). One sample (2 L) was taken in each sampling point of the WWTPs. Samples collected in sterile plastic bottles were transported on ice to the laboratory and analyzed within 24 h.
DWTPs
Drinking water source (raw surface water from a large river) and finished water (drinking water) samples were collected in two DWTPs (plants D and E) in the same geographical area of WWTPs (~ 20 km2 in North-West of Italy). Plants D and E treat 130,000 m3/day and 86,400 m3/day of drinking water, respectively, provided by the same drinking water source.
The DWTP D involves pre-decantation, addition of powder-activated carbon (PAC), ozonation followed by clarification/flocculation, two granular-activated carbon (GAC) filtrations, and final disinfection using chlorine dioxide. The DWTP E employs chlorination, clarification, GAC filtration, and final disinfection using chlorine dioxide.
Six water samplings were performed during 1 year (March 2019–January 2020). One sample was taken in each sampling point of the DWTPs. The sampling volume was 3 L for raw surface water and 30 L for drinking water. Samples collected in sterile plastic bottles were transported on ice to the laboratory and analyzed within 24 h.
Sample processing and ARB quantification
ARB were isolated in a basic culture medium for the count of heterotrophic bacteria (R2Agar, VWR International, Milan) supplemented with ampicillin, tetracycline, and sulfamethoxazole at a concentration of 32 mg/L, 16 mg/L, and 50.4 mg/L, respectively.
The antibiotic concentrations tested were chosen considering the highest dose used in standard methods to establish resistance to antibiotics with clinical strains (breakpoint) (CLSI 2018) or the concentration reported in previous study (Gao et al. 2012). Total heterotrophic count (HPC) was determined on media without antibiotics.
Serial dilutions of wastewater samples (influent and effluent) were plated in duplicate on media with and without antibiotics. Serial dilution of surface water or different volumes of drinking water (0.5–1 L for HPC and 2 L for ARB) were filtered (0.22-µm pore nitrocellulose size filter membrane, Millipore) in duplicate and then the membranes were placed on R2Agar plates with and without antibiotics.
All plates were incubated at 30 °C for 7 days and the results are expressed as log CFU/mL. The antibiotic resistance rate for each antibiotic was calculated as the ratio between the CFU/mL of each ARB and the CFU/mL of HPC.
Sample processing and DNA extraction
Samples of wastewater influent (20 mL), wastewater effluent (250 mL), drinking water source (700 mL), and finished water (3 L) were filtered in triplicate on 0.22-µm pore size polycarbonate filter membrane. The filters were stored at − 20 °C until DNA extraction. Subsequently, the entire filters were processed for the DNA extraction using the DNeasy PowerWater kit (Qiagen) according to the manufacturer’s instructions. Concentration of the extracted DNA of each sample was quantified by spectrophotometry (NanoDrop® ND-1000, NanoDrop Technologies, Wilmington, DE).
Real-time qPCR of ARGs
The abundance of the selected ARGs (tetA, blaTEM, and sulII) of intI1 and of the 16S rRNA gene was measured by qPCR using a RT-thermocycler (CFX Connect, Bio-Rad). The protocol used in qPCR assays was previously described by Di Cesare et al. (2015). The qPCR program was 95 °C for 2 min, 35 cycles of 95 °C for 15 s, annealing temperature reported in Supplementary Table S1 for 30 s, and 72 °C for 15 s. Melt curve analysis was performed from 60 to 95 °C with increments of 0.5 °C/5 s. Standard calibration curves were carried out using the purified, quantified, and ten-fold diluted amplicon of each gene as described in Di Cesare et al. (2013). Each reaction was carried out in duplicate for each sample. The limits of quantification (LOQ) per each quantified gene were determined as described in Bustin et al. (2009). They were 1.55 × 103, 1.22 × 102, 3.57 × 101, 4.88 × 102, and 1.12 × 101 gene copy/µL for 16S rRNA, tetA, sulII, blaTEM, and intI1 genes respectively. The mean value ± standard deviation of the reaction efficiencies was 98.32 ± 8.23% and the R2 was always more than 0.97. The potential inhibition of the qPCRs due to the type of analyzed matrix was calculated by dilution method (Di Cesare et al. 2013) and no inhibition was obtained. The ARG and intI1 abundances were expressed as absolute abundance (log gene copies/mL) and relative abundance (gene copies/16S rRNA gene copy) and a mean value of the abundance for each gene was calculated. The interpretation of the results in case of abundance values lower than the LOQ and in case of discordance between the two replicates was made as previously reported (Di Cesare et al. 2015).
Statistical analyses
The statistical analysis of the data was carried out using the SPSS package (Version 25.0) for Windows. The bacterial concentrations and ARG abundances were converted to log10 (log CFU/mL or log copies/mL), whereas relative ARG abundances were transformed in the arcsine of their square root (arcsine of square root of gene copy/16 rRNA gene copy) (Crawley 2012). For the comparison of ARB concentration or antibiotic resistance rate or ARG abundance between influent vs effluent in the WWTPs or raw surface water vs drinking water in the DWTPs, the Student t-test was used. This test was also applied to evaluate the differences between DWTP D and DWTP E. The one-way ANOVA test, followed by Tukey’s post hoc analysis, was used to study the differences among the sampling and the WWTPs. The relationship among ARGs (relative abundance) and between ARGs (relative abundance) and ARB (antibiotic resistance rate) was analyzed with Pearson’s correlation test.
Results
Dynamics of ARB and ARGs in urban water cycle
The dynamics of antibiotic resistance in the urban water cycle that includes WWTPs and DWTPs highlighted a statistically significant difference (Table 1) of the ARB concentrations and ARGs (relative and absolute abundance) between WWTPs and DWTPs, with higher values in the WWTPs (Figs. 1, 2, 3, and 4). On the contrary, no difference was revealed for the antibiotic resistance rates (%) with respect to the plant (WWTPs vs DWTPs).
a Log-transformed concentrations of ampicillin-, tetracycline-, sulfonamide-resistant bacteria and HPC in influent and effluent of the WWTPs investigated. Box plots represent median and range values. b Antibiotic resistance rate of ampicillin-, tetracycline-, and sulfonamide-resistant bacteria in WWTPs. All values are normalized to HPC abundances. Box plots represent median and range values. WWTPs, wastewater treatment plants; HPC, total heterotrophic count; AmRB, ampicillin-resistant bacteria; TRB, tetracycline-resistant bacteria; SRB, sulfonamide-resistant bacteria
a Absolute abundance of tetA, sulII, blaTEM, and intI1 gene in influent and effluent of the WWTPs investigated. Box plots represent median and range values. b Relative abundance of tetA, blaTEM, intI1 (b1) and sulII (b2) gene in WWTPs. All values are normalized to 16S rRNA gene copy. Box plots represent median and range values. WWTPs, wastewater treatment plants
a Log-transformed concentrations of ampicillin-, tetracycline-, sulfonamide-resistant bacteria, and HPC in surface and drinking water of the DWTPs investigated. Box plots represent median and range values. b Antibiotic resistance rate of ampicillin-, tetracycline-, and sulfonamide-resistant bacteria in DWTPs. All values are normalized to HPC abundances. Box plots represent median and range values. DWTPs, drinking water treatment plants; HPC, total heterotrophic count; AmRB, ampicillin-resistant bacteria; TRB, tetracycline-resistant bacteria; SRB, sulfonamide-resistant bacteria
a Absolute abundance of tetA, sulII, blaTEM, and intI1 gene in surface and drinking water of the DWTPs investigated. Box plots represent median and range values. b Relative abundance of tetA, blaTEM, sulII, and intI1 gene in DWTPs. All values are normalized to 16S rRNA gene copy. Box plots represent median and range values. DWTPs, drinking water treatment plants
A generally higher ARB concentration and ARG relative and absolute abundance were observed in the inlet of the treatment plants (influent of WWTPs and surface raw water treated in the DWTPs) with respect to the outlet (effluent and drinking water), although no differences were detected when considering the antibiotic resistance rates (%).
The sampling period seemed to not affect the dynamics of antibiotic resistance.
Dynamics of ARB and ARGs in WWTPs
Total HPCs, ampicillin-resistant bacteria (AmRB), tetracycline-resistant bacteria (TRB), and sulfonamide-resistant bacteria (SRB) detected in the influents and in the effluents of the three WWTPs investigated are reported in Fig. 1. In the influents, the concentrations ranged from 5.7 to 6.4 for AmRB, from 4.5 to 6.1 for TRB, and from 5.4 to 7.0 log CFU/mL for SRB. The highest mean concentration reported was 6.4 log CFU/mL for SRB and the lowest 6.1 log CFU/mL for AmRB. The mean resistance rates of AmRB, TRB, and SRB were 9%, 1%, and 22%, respectively, and their highest ratio reached up to 21%, 4%, and 38%.
The concentrations in the effluents ranged from 2.1 to 4.5 for AmRB, from 1.3 to 3.9 for TRB, and from 2.1 to 4.9 log CFU/mL for SRB. Moreover, the highest mean concentration was 3.9 log CFU/mL for SRB and the lowest 2.8 log CFU/mL for TRB. As observed in the influent, the decreasing trend of the mean resistance rates of ARBs was SRB > AmRB > TRB (21% > 12% > 3%) and their highest ratio reached up to 35%, 33%, and 12%, respectively.
A statistically significant reduction of heterotrophic bacteria and ARB was observed in WWTPs after the treatment (Table 2, Fig. 1a). On the contrary, for all ARB monitored, the antibiotic resistance rate did not show a significant trend (influent vs effluent, Table 2), except for TRB (higher value in the effluent of WWTPs) (Fig. 1b). Moreover, no difference was observed among the different WWTPs both considering the results expressed as log CFU/mL and as antibiotic resistance rates. Also the sampling period seemed not to affect the abundances and the rates of TRB, SRB, and AmRB.
The absolute abundance of blaTEM, tetA, sulII, and intI1 in influents, when quantifiable, ranged from 8.0 to 9.0, 8.7 to 9.4, 8.8 to 9.8, and 8.4 to 9.5 log gene copies/mL, respectively (Fig. 2a). The relative abundances of blaTEM, tetA, sulII, and intI1 with respect to 16S rRNA gene, when quantifiable, ranged from 3.3 × 10−4 to 9.1 × 10−4, 1.1 × 10−3 to 3.7 × 10−3, 2.0 × 10−1 to 8.7 × 10−1, and 1.6 × 10−3 to 3.7 × 10−3 copies/16S rRNA gene copy, respectively. sulII showed for both absolute and relative abundances the highest mean values (mean: 9.4 log gene copies/mL and 3.8 × 10−1 copies/16S rRNA gene copy) (Fig. 2).
The absolute abundance of blaTEM, tetA, sulII, and intI1 when quantifiable ranged in the effluents from 7.3 to 8.2, 7.6 to 9.2, 8.8 to 10.3, and 8.4 to 9.6 log gene copies/mL, respectively (Fig. 2a). The relative abundances when quantifiable of blaTEM was 1.9 × 10−4 copies/16S rRNA gene copy, and the relative abundance of tetA, sulII, and intI1 ranged from 2.7 × 10−5 to 1.5 × 10−3, 1.3 × 10−1 to 9.1 × 10−1, and 5.6 × 10−4 to 3.1 × 10−3 copies/16S rRNA gene copy. As observed already in the influents, the highest mean value was obtained for sulII (mean: 9.5 log gene copies/mL and 3.6 × 10−1 copies/16S rRNA gene copy) (Fig. 2).
Although the absolute abundance of ARGs generally did not show differences between the influents and effluents of WWTPs, a significant reduction of the relative abundance of tetA, blaTEM, and intI1 was observed in the effluents.
Significant differences among the WWTPs were reported in effluents only for the relative abundance of sulII (WWTP2 > WWTP1 and WWTP3), tetA (WWTP1 > WWTP2), and intI1 (WWTP1 > WWTP2).
The absolute and relative abundances were not influenced by the sampling period.
A correlation between relative abundance of intI1 vs tetA (p < 0.001), intI1 vs blaTEM (p < 0.05), and blaTEM vs tetA (p < 0.001) was observed (p < 0.05). No relationship was reported for ARGs and ARB, except a correlation between TRB and tetA (p < 0.05) and TRB and intI1 (p < 0.05).
Dynamics of ARB and ARGs in DWTPs
ARB concentration and antibiotic resistance rate (%) detected in surface and drinking water of the two DWTPs are reported in Fig. 3.
ARB were observed in all analyzed surface water samples with concentrations ranging between 4.7 and 6.3 for AmRB, 3.3 and 6.2 for TRB, and 4.9 and 6.9 log CFU/mL for SRB (Fig. 3a). Moreover, the highest mean concentration was 6.1 log CFU/mL for SRB and the lowest 4.9 log CFU/mL for TRB. The highest ratio is 36% for SRB and the lowest 5% for TRB.
Considering the drinking water, SRB were observed in all analyzed samples with concentrations ranging from 1.0 to 2.1 log CFU/mL. On the contrary, TRB and AmRB were found in the 50% and 25% of the analyzed samples and the concentration when quantifiable ranged from 1.2 to 1.9 log CFU/mL for AmRB and from 0.4 to 1.5 log CFU/mL for TRB. Moreover, the highest mean concentration was 1.5 log CFU/mL for SRB and the lowest 0.4 log CFU/mL for AmRB. According to the ARB concentration, the highest resistance rate was observed for SRB (mean value: 27%).
A significant reduction (Table 3) was detected after the treatments for both heterotrophic bacteria and ARB (log CFU/mL) in DWTP (Fig. 3a), while no abatement was observed for antibiotic resistance rates (Fig. 3b). No difference was observed between the DWTPs and sampling period did not impact on abundances and rates of TRB, SRB, and AmRB.
In all the surface water sample, intI1, tetA, and sulII were observed, while blaTEM was never detected. The absolute abundance of tetA, sulII, and intI1 in these samples ranged from 8.2 to 9.6, 8.6 to 10.0, and 8.4 to 10.3 log gene copies/mL, respectively (Fig. 4a). The relative abundances of tetA, sulII, and intI1 ranged from 5.9 × 10−5 to 2.6 × 10−4, 5.8 × 10−5 to 9.1 × 10−4, and 1.8 × 10−4 to 1.5 × 10−3 copies/16S rRNA gene copy, respectively. intI1 showed the highest mean values for both absolute and relative abundances (mean: 9.6 log gene copies/mL and 7.5 × 10−4 copies/16S rRNA gene copy) (Fig. 4b).
blaTEM and tetA were not detected in drinking waters, except for three samples (1 for DWTP D, 2 for DWTP E) that presented only one replicate positive but not quantifiable for tetA, while all drinking water samples were positive for sulII, although this gene was not quantifiable. The absolute and relative abundance of intI1 was 7.5 log gene copies/mL and 1.3 × 10−3 copies/16S rRNA gene copy, respectively.
The relative and absolute abundance of tetA and sulII were reduced by each water treatment; moreover, for DWTP E also, the absolute abundance of intI1 was lower in the drinking water (Fig. 4a). The relative abundance of intI1 in the drinking water was the sole parameter evidencing a difference between the DWTPs (DWTP E > DWTP D; Fig. 4b). The absolute and relative ARG abundances were not influenced by the sampling period (Table 3).
Moreover, both relative and absolute abundances of tetA showed a correlation with the absolute abundance of sulII (p < 0.001). No relationship was detected between the values of ARGs and ARB, except for a correlation between SRB and sulII gene (p < 0.001).
Discussion
The urban water cycle that comprises both wastewater and drinking water treatment represents a possible route for spreading of ARB and ARGs into the environment with a subsequently potential return to humans with a direct impact on the human health (Manaia 2017). The treatment plants (WWTPs and DWTPs) investigated in our study, being in the same geographical area, are particularly interesting to track the fate of ARB and ARGs in the urban water cycle.
The overall ARB and ARG dynamics in the WWTPs and DWTPs sampled in this study highlighted, as expected, higher concentrations of antibiotic resistances in the WWTPs than in DWTPs. This is in agreement with several studies that identified the WWTP as hotspot for the spread of antibiotic resistances (Rizzo et al. 2013; Pazda et al. 2019). The concentrations of ARB in the influent of WWTPs observed in our study were similar or higher than those reported in other studies for TRB and SRB (Munir et al. 2011; Gao et al. 2012) and, generally, the monitored ARGs showed absolute abundances substantially higher than those reported by other authors (Narciso-da-Rocha et al. 2018; Fiorentino et al. 2019; Wang et al. 2020). The reduction of ARB concentrations between the influent and the effluent of the studied WWTPs was comparable with similar studies (Wang et al. 2020).
Moreover, the stability of the antibiotic-resistant rates in the influents and effluents (proportion of ARB in the HPC for each sample) underlined that the different treatments did not promote the selection of ARB. This was also confirmed by the relative abundance of the ARGs that showed a reduction for blaTEM, tetA, and intI1, with values significantly lower in the effluent. The results reported in literature that considered the effect of the treatment on the variation of the ARG relative abundances are discordant. Indeed, in two studies, performed in Canada and in China, no significant differences were observed in the relative abundance of ARGs between influent and effluent of two WWTPs (Mao et al. 2015; McConnell et al. 2018). On the contrary, Di Cesare et al. (2016) showed an increase of some ARGs (e.g., sulII) and of intI1 in the effluents of three Italian WWTPs.
Although the treatment process, the dimension, and the quality of the treated effluent varied in the investigated WWTPs, no significant difference in presence and abundance of ARB and ARGs was observed among the three WWTPs, highlighting that these factors had a limited impact on the fate of the resistances released by the plants. This is in agreement with the literature that hypothesizes a role for the biological process, present in all the investigated WWTPs, in the establishment of a bacterial community exerting specific ecological factors (e.g., enhanced competition, predation, cooperation) determining the presence of ARB and ARGs (Rizzo et al. 2013; Manaia et al. 2016).
As previously observed among ARB and ARGs investigated, SRB and sulII were the most abundant (Munir et al. 2011; Ferro et al. 2016; Ben et al. 2017).
In order to evaluate the dynamics of antibiotic resistance in the urban water cycle, it is also important to consider the spread of ARB and ARGs in surface waters utilized for drinking purposes that are influenced by the discharge of wastewater and by other anthropogenic activities. The role of surface water is confirmed in our study by the presence of all ARB and most of the quantified genes in all samples of raw surface water analyzed used as drinking water source, with high abundances of SRB and sulII, according to the results obtained in the three WWTPs that are in the same geographical area. The high abundance of antibiotic resistances against sulfonamides in surface water was observed also in other rivers impacted by human activities (Hu et al. 2019; Yang et al. 2020). The results obtained in the investigated surface water revealed a higher absolute abundance of ARGs and ARB (3–4 orders of magnitude) with respect to the data reported for other rivers in China (Hu et al. 2019; Yang et al. 2020) and Poland (Bondarczuk and Piotrowska-Seget 2019) influenced by anthropogenic activities (e.g., sewage discharges, agricultural runoff, swine farm) underlining its role in the spreading of antibiotic resistance. The presence of ARB and ARGs in surface water used as drinking water source highlights the importance to investigate all the phases composing the urban water cycle and to deepen the effect of the drinking water treatment.
None of the detected ARB or ARGs was completely removed in drinking water by the applied water treatments, according to Hu et al. (2019) and Siedlecka et al. (2021) who investigated ARGs in a DWTP in China and Poland. As previously reported in other studies, the presence and the total concentration of ARGs in drinking water significantly decreased in comparison to corresponding water source (Sanganyado and Gwenzi 2019; Stange et al. 2019). Moreover, the limited variations in antibiotic resistance rates between surface water and drinking water associated with the reduction of relative ARG abundances showed, as observed in WWTPs, that the treatment did not apply any selective pressure. These results highlight the key role played by the drinking water treatment in reducing ARGs and controlling ARB from the water source. As observed for WWTPs, the different drinking water treatments seemed to display a limited impact on the spread of SRB and ARGs, although DWTP D was equipped with an ozonation step that resulted the best treatment to reduce ARB and ARGs in some studies (Stange et al. 2019; Yang et al. 2020). It is important to highlight that the effectiveness of the disinfection step can be influenced by numerous factors such as the bacterial community composition in the raw water used as drinking water source, the effectiveness of other treatment steps, and the co-selection of antibiotic resistance enhanced by disinfection by-products (Sanganyado and Gwenzi, 2019).
Despite the observed reduction after drinking water treatment, the presence of ARB and ARGs in drinking water should be kept under observation, considering that ARB may also enter the human microbiome following water consumption (Vaz-Moreira et al. 2014) and ARGs can occasionally be transferred to human pathogenic bacteria (Manaia 2017). Moreover, it should be taken into account that the amount of ARB could increase in tap water, due to biofilm detachment in the distribution system (Zhang et al. 2018).
Considering that some genes can be utilized as proxy for the overall abundance of ARGs in waters (Su et al. 2018), Pearson’s correlation was carried out among the different measured ARGs in WWTPs and DWTPs to evaluate the suitability of the monitored target as indicator. The results highlighted that a positive correlation was observed only for blaTEM vs tetA in WWTPs and sulII vs tetA in DWTPs underlining that none of the selected genes could be used to predict the fate and the potential contamination by determinants of antibiotic resistance in the urban water cycle. A similar result was also observed for intI1 that was correlated only with blaTEM and tetA in WWTPs.
Another important topic is related to the meaning of the results obtained with the two approaches (cultivation dependent and independent) utilized in this study for characterizing the antibiotic resistance. Molecular results highlighted a trend (influent > effluent; raw surface water > drinking water) that was not shown with cultivation-dependent methods; moreover, no clear relationship between ARB and the corresponding ARGs was observed. This is probably related to the different information that the two approaches provide. The cultivation-dependent method offers a general overview of resistant bacteria but considers only a part of the viable bacteria because no information about VBNC (viable but not culturable bacteria) can be provided. Moreover, other drawbacks associated with this approach are present; for example, the results obtained can be influenced by the antibiotic tested and the concentration used (Manaia et al. 2016). On the contrary, the cultivation-independent method has high specificity and sensitivity and the detection is not influenced by the physiological status of bacteria; these characteristics make this method a useful tool to evaluate the potential of antibiotic resistance spreading in environment (Rizzo et al. 2013). However, as highlighted before, qPCR is a target-based quantification method that cannot provide a complete overview of the antibiotic resistome of a microbial community (Hendriksen et al. 2019).
Conclusion
The results obtained underline that to investigate the antibiotic resistance dynamics in the same geographical area is particularly interesting to track the fate of ARB and ARGs in the urban water cycle. More detailed description of the phenomenon was obtained in this study integrating the information obtained by culture-dependent method with those obtained by the culture-independent one. However, the results claim for the need to perform more studies by using untargeted based quantification approach, i.e., shotgun metagenomics, to characterize the overall antibiotic resistome (total content of ARGs) of a microbial community aiming to find the best targets as proxy of the antibiotic resistome in aquatic ecosystems.
Although the processes applied for treating wastewater and drinking water allowed to reduce the concentration of ARB and ARGs and did not apply selective pressures, the results highlighted the importance to monitor the presence of ARB and ARGs mainly in drinking water that represents a potential route of transmission to humans with a direct impact on human health.
Data availability
All data generated or analyzed during this study are included in this published article.
References
Almakki A, Jumas-Bilak E, Marchandin H, Licznar-Fajardo P (2019) Antibiotic resistance in urban runoff. Sci Tot Environ 667:64–76. https://doi.org/10.1016/j.scitotenv.2019.02.183
Bai X, Ma X, Xu F, Li J, Zhang H, Xiao X (2015) The drinking water treatment process as a potential source of affecting the bacterial antibiotic resistance. Sci Tot Environ 533:24–31. https://doi.org/10.1016/j.scitotenv.2015.06.082
Bondarczuk K, Piotrowska-Seget Z (2019) Microbial diversity and antibiotic resistance in a final effluent-receiving lake. Sci Total Environ 650:2951–2961. https://doi.org/10.1016/j.scitotenv.2018.10.050
Baquero F, Martınez JL, Canton R (2008) Antibiotics and antibiotic resistance in water environments. Curr Opin Biotechnol 19:260–265. https://doi.org/10.1016/j.copbio.2008.05.006
Ben W, Wang J, Cao R, Yang M, Zhang Y, Qiang Z (2017) Distribution of antibiotic resistance in the effluents of ten municipal wastewater treatment plants in China and the effect of treatment processes. Chemosphere 172:392–398. https://doi.org/10.1016/j.chemosphere.2017.01.041
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622. https://doi.org/10.1373/clinchem.2008.112797
Cacace D, Fatta-Kassinos D, Manaia CM, Cytryn E, Kreuzinger N, Rizzo L, Karaolia P, Schwartz T, Alexander J, Merlin C, Garelick H, Schmitt H, de Vries JD, Schwermer CU, Meric S, Ozkalt CB, Pons M, Kneis D, Berendonk TU (2019) Antibiotic resistance genes in treated wastewater and in the receiving water bodies: a pan-European survey of urban settings. Wat Res 162:320–330. https://doi.org/10.1016/j.watres.2019.06.039
Carvalho IT, Santos L (2016) Antibiotics in the aquatic environments: a review of the European scenario. Environ Int 94:736–757. https://doi.org/10.1016/j.envint.2016.06.025
Chang Q, Wang W, Regev-Yochay G, Lipsitch M, Hanage WP (2015) Antibiotics in agriculture and the risk to human health: how worried should we be? Evol Appl 8:240–2477. https://doi.org/10.1111/eva.12185
Clinical and Laboratory Standards Institute (CLSI) (2018) Performance standards for antimicrobial susceptibility testing. 28th ed. CLSI supplement M100 (ISBN 1–56238–838-X [Print]; ISBN 1–56238–839–8 [Electronic]). Clinical and Laboratory Standards Institute, 950 West Valley Road, Suite 2500, Wayne, Pennsylvania 19087 USA.
Crawley MJ (2012) The R book. John Wiley & Sons, Chichester
Di Cesare A, Luna GM, Vignaroli C, Pasquaroli S, Tota S et al (2013) Aquaculture can promote the presence and spread of antibiotic-resistant Enterococci in marine sediments. PLoS ONE 8:e62838. https://doi.org/10.1371/journal.pone.0062838
Di Cesare A, Eckert EM, D’Urso S, Bertoni R, Gillan DC, Wattiez R, Corno G (2016) Co-occurrence of integrase 1, antibiotic and heavy metal resistance genes in municipal wastewater treatment plants. Water Res 94:208–214. https://doi.org/10.1016/j.watres.2016.02.049
Di Cesare A, Eckert EM, Teruggi A, Fontaneto D, Bertoni R, Callieri C, Corno G (2015) Constitutive presence of antibiotic resistance genes within the bacterial community of a large subalpine lake. Mol Ecol 24:3888–3900. https://doi.org/10.1111/mec.13293
Ferro G, Guarino F, Castiglione S, Rizzo L (2016) Antibiotic resistance spread potential in urban wastewater effluents disinfected by UV/H2O2 process. Sci Total Environ 560–561:29–35. https://doi.org/10.1016/j.scitotenv.2016.04.047
Fiorentino A, Di Cesare A, Eckert EM, Rizzo L, Fontaneto D, Yang Y, Corno G (2019) Impact of industrial wastewater on the dynamics of antibiotic resistance genes in a full-scale urban wastewater treatment plant. Sci Total Environ 646:1204–1210. https://doi.org/10.1016/j.scitotenv.2018.07.370
Gao P, Munir M, Xagoraraki I (2012) Correlation of tetracycline and sulfonamide antibiotics with corresponding resistance genes and resistant bacteria in a conventional municipal wastewater treatment plant. Sci Total Environ 421–422:173–183. https://doi.org/10.1016/j.scitotenv.2012.01.061
Gillings MR, Gaze WH, Pruden A, Smalla K, Tiedje JM, Zhu Y (2015) Using the class 1 integron-integrase gene as a proxy for anthropogenic pollution. ISME J 9:1269e1279. https://doi.org/10.1038/ismej.2014.226
Hendriksen RS, Munk P, Njage P, van Bunnik B, McNally L, Lukjancenko O, Röder T, Nieuwenhuijse D, Karlsmose Pedersen S, Kjeldgaard J, Kaas RS, Clausen PTLC, Vogt JK, Leekitcharoenphon P, van de Schans MGM, Zuidema T, de Roda Husman AM, Rasmussen S, Petersen B, The Global Sewage Surveillance project consortium#, Amid C, Cochrane G, Sicheritz-Ponten T, Schmitt H, Alvarez JRM, Aidara-Kane A, Pamp SJ, Lund O, Hald T, Woolhouse M, Koopmans MP, Vigre H, Petersen TN, Aarestrup FM (2019) Global monitoring of antimicrobial resistance based on metagenomics analyses of urban sewage. Nat Commun 10:1124. https://doi.org/10.1038/s41467-019-08853-3
Hu Y, Jiang L, Zhang T, Jin L, Han Q, Zhang D, Lin K, Cui C (2018) Occurrence and removal of sulfonamide antibiotics and antibiotic resistance genes in conventional and advanced drinking water treatment processes. J Hazard Mater 360:364–372. https://doi.org/10.1016/j.jhazmat.2018.08.012
Hu Y, Zhang T, Jiang L, Luo Y, Yao S, Zhang D, Lin K, Cui C (2019) Occurrence and reduction of antibiotic resistance genes in conventional and advanced drinking water treatment processes. Sci Total Environ 669:777–784. https://doi.org/10.1016/j.scitotenv.2019.03.143
Huang F, Chen Q, Zhang X, Neilson R, Su J, Zhou S (2021) Dynamics of antibiotic resistance and its association with bacterial community in a drinking water treatment plant and the residential area. Environ Sci Pollut Res Int 28:55690–55699. https://doi.org/10.1007/s11356-021-14896-1
Li B, Yang Y, Ma L, Ju F, Guo F, Tiedje JM, Zhang T (2015) Metagenomic and network analysis reveal wide distribution and co-occurrence of environmental antibiotic resistance genes. ISME J 9:2490–2502. https://doi.org/10.1038/ismej.2015.59
Ma L, Li A, Yin X, Zhang T (2017) The prevalence of integrons as the carrier of antibiotic resistance genes in natural and man-made environments. Environ Sci Technol 51:5721–5728. https://doi.org/10.1021/acs.est.6b05887
Manaia CM (2017) Assessing the risk of antibiotic resistance transmission from the environment to humans: non-direct proportionality between abundance and risk. Trends Microbiol 25:3. https://doi.org/10.1016/j.tim.2016.11.014
Manaia CM, Macedo G, Fatta-Kassinos D, Nunes OC (2016) Antibiotic resistance in urban aquatic environments: can it be controller? Appl Microbiol Biotechnol 100:1543–1557. https://doi.org/10.1007/s00253-015-7202-0
Mao D, Yu S, Rysz M, Luo Y, Yang F, Li F, Hou J, Mu Q, Alvarez PJJ (2015) Prevalence and proliferation of antibiotic resistance genes in two municipal wastewater treatment plants. Water Res 85:458–466. https://doi.org/10.1016/j.watres.2015.09.010
McConnell MM, Hansen LT, Jamieson RC, Neudorf KD, Yost CK, Tong A (2018) Removal of antibiotic resistance genes in two tertiary level municipal wastewater treatment plants. Sci Total Environ 643:292–300. https://doi.org/10.1016/j.scitotenv.2018.06.212
Munir M, Wong K, Xagoraraki I (2011) Release of antibiotic resistant bacteria and genes in the effluent and biosolids of five wastewater utilities in Michigan. Water Res 45:681–693. https://doi.org/10.1016/j.watres.2010.08.033
Narciso-da-Rocha C, Rocha J, Vaz-Moreira I, Lira F, Tamames J, Henriques I, Martinez JL, Manaia CM (2018) Bacterial lineages putatively associated with the dissemination of antibiotic resistance genes in a full-scale urban wastewater treatment plant. Environ Inter 118:179–188. https://doi.org/10.1016/j.envint.2018.05.040
O’Neil J (2016) Review on Antimicrobial Resistance. Antimicrobial resistance: tackling drug-resistant infections globally: final report and recommendations.
Pazda M, Kumirska J, Stepnowski P, Mulkiewicz E (2019) Antibiotic resistance genes identified in wastewater treatment plant systems – a review. Sci Total Environ 697:134023. https://doi.org/10.1016/j.scitotenv.2019.134023
Rizzo L, Manaia C, Merlin C, Schwartz T, Dagot C, Ploy MC, Michael I, Fatta-Kassino D (2013) Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: a review. Sci Total Environ 447:345–360. https://doi.org/10.1016/j.scitotenv.2013.01.032
Sanganyado E, Gwenzi W (2019) Antibiotic resistance in drinking water systems: occurrence, removal, and human health risks. Sci Total Environ 669:785–797. https://doi.org/10.1016/j.scitotenv.2019.03.162
Seiler C, Berendonk TU (2012) Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front Microbiol 3:1–10. https://doi.org/10.3389/fmicb.2012.00399
Siedlecka A, Wolf-Baca M, Piekarska K (2021) Microbial communities of biofilms developed in a chlorinated drinking water distribution system: a field study of antibiotic resistance and biodiversity. Sci Total Environ 774:145113. https://doi.org/10.1016/j.scitotenv.2021.145113
Stachurova T, Pikovay A, Bartas M, Semerad J, Svobodova K, Malachova K (2021) Beta-lactam resistance development during the treatment processes of municipal wastewater treatment plants. Chemosphere 280:130749. https://doi.org/10.1016/j.chemosphere.2021.130749
Stange C, Yin D, Xu T, Guo X, Schäfer C, Tiehma A (2019) Distribution of clinically relevant antibiotic resistance genes in Lake Tai, China. Sci Total Environ 655:337–346. https://doi.org/10.1016/j.scitotenv.2018.11.211
Su HC, Liu YS, Pan CG, Chen J, He LY, Ying GG (2018) Persistence of antibiotic resistance genes and bacterial community changes in drinking water treatment system: from drinking water source to tap water. Sci Total Environ 616–617:453–461. https://doi.org/10.1016/j.scitotenv.2017.10.318
Vaz-Moreira I, Nunes OC, Manaia CM (2014) Bacterial diversity and antibiotic resistance in water habitats: searching the links with the human microbiome. FEMS Microbiol Lett 38:761–818. https://doi.org/10.1111/1574-6976.12062
Wang J, Chu L, Wojnárovits L, Takács E (2020) Occurrence and fate of antibiotics, antibiotic resistant genes (ARGs) and antibiotic resistant bacteria (ARB) in municipal wastewater treatment plant: an overview. Sci Total Environ 744:140997. https://doi.org/10.1016/j.scitotenv.2020.140997
WHO (2014) Antimicrobial resistance. Global Report on Surveillance. www.who.int/about/licensing/copyright_form/en/index.html
Yang J, Wang H, Roberts DJ, Du H, Yu X, Zhu N, Meng X (2020) Persistence of antibiotic resistance genes from river water to tap water in the Yangtze River Delta. Sci Total Environ 742:140592. https://doi.org/10.1016/j.scitotenv.2020.140592
Yang Y, Li Z, Song W, Due L, Ye C, Zhao B, Liu W, Deng D, Pan Y, Lin H, Cao X (2019) Metagenomic insights into the abundance and composition of resistance genes in aquatic environments: influence of stratification and geography. Environ Int 127:371–380. https://doi.org/10.1016/j.envint.2019.03.062
Zhang J, Li W, Chen J, Qi W, Wang F, Zhou Y (2018) Impact of biofilm formation and detachment on the transmission of bacterial antibiotic resistance in drinking water distribution systems. Chemosphere 203:368–380. https://doi.org/10.1016/j.chemosphere.2018.03.143
Zhang M, Wang L, Xu M, Zhou H, Wang S, Wang Y, Bai M, Zhang C (2019) Selective antibiotic resistance genes in multiphase samples during biofilm growth in a simulated drinking water distribution system: occurrence, correlation and low-pressure ultraviolet removal. Sci Total Environ 649:146–155. https://doi.org/10.1016/j.scitotenv.2018.08.297
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Experiments were performed by CP, MP, MM, and RS, while data collection and analysis was done by SiB, SaB, AD, and RS. The first draft of the manuscript was written by SiB, SaB, and AD and all authors commented on previous versions of the manuscript. GC and EC were in charge of supervision and greatly contributed to final review and editing of the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This is an original article that did not use other information that requires ethical approval.
Consent to participate
No consent of participation is to be claimed.
Consent for publication
All of the authors have read and approved the paper for publication. We confirmed that it has not been published previously nor is it being considered by any other peer-reviewed journal.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bonetta, S., Di Cesare, A., Pignata, C. et al. Occurrence of antibiotic-resistant bacteria and resistance genes in the urban water cycle. Environ Sci Pollut Res 30, 35294–35306 (2023). https://doi.org/10.1007/s11356-022-24650-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-24650-w