Abstract
Pancreatic cancer (PC) is one of the most common malignant tumors in the world with a poor prognosis. There were limited studies investigating the genetic signatures associated with inflammatory responses, tumor microenvironment (TME), and tumor drug sensitivity prediction. In the Cancer Genome Atlas (TCGA) dataset, we constructed an inflammatory response–related genes prognostic signature for PC, and predictive ability of the model was assessed via the International Cancer Genome Consortium (ICGC) database. Then, we explored the differences of TME, immune checkpoint genes and drug resistance genes, and the cancer cell sensitivity to chemotherapy drugs between different risk score group. Based on the TCGA and ICGC databases, we constructed and validated a prognostic model, which consisted of 5 genes (including AHR, F3, GNA15, IL18, and INHBA). Moreover, the prognostic model was independent prognostic factors affecting overall survival (OS). The low-risk score group had better OS, and lower stromal score, compared with patients in the high-risk score group. The difference of antigen-presenting cells, T cell regulation, and drug resistance genes between different risk score groups was found. In addition, the immune checkpoint genes were positively correlation to risk score. The expression levels of AHR, GNA15, IL18, and INHBA were related to the sensitivity of anti-tumor chemotherapy drugs. Gene set enrichment analysis (GSEA) showed significant pathway such as calcium signaling pathway and p53 signaling pathway. We successfully constructed a 5-inflammatory response–related gene signature to predict survival, TME, and cancer cell sensitivity to chemotherapy drugs in PC patients. Furthermore, substantiation was warranted to verify the role of these genes in tumorigenesis.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
PC is the 7th leading cause of cancer-related death globally, with 496,000 new cases and 466,000 deaths in the past year (Sung et al. 2021), and a dismal 5-year survival rate of roughly 5% (Li et al. 2019). Despite its low incidence, PC is projected to surpass breast cancer as the third leading cause of cancer death by 2025, according to a survey conducted in 28 European countries (Ferlay et al. 2016). Surgical resection is the mainstay curative treatment modality (Okasha et al. 2017), significantly improving the 5-year survival rate by 20 to 30% (Okasha et al. 2017). However, because early PC symptoms are insidious, 20–-80% of patients have locally advanced or distant metastases at diagnosis (Ansari et al. 2016); hence, only around 9% of the patients are eligible for resection (Kamisawa et al. 2016). Due to the high degree of malignancy, rapid progression, atypical clinical manifestations, and lack of effective screening methods, the prognosis of PC is poor (Mizrahi et al. 2020).
Treatment options for patients who cannot undergo radical surgery include radiation therapy, chemotherapy, immunotherapy (Houot et al. 2015; Lutz et al. 2014), stem cell therapy (Kure et al. 2012; Raj et al. 2015), and metabolism targeting therapy (Blum and Kloog 2014; Son et al. 2013). Although these methods have demonstrated therapeutic efficacy, it is debatable whether these treatments could ultimately benefit patients since side effects such as nausea, vomiting, fatigue, hypertension, renal impairment, and decreased immunity are quite common. Therefore, a model for predicting patient prognosis is essential for individualized treatment.
Public databases and next-generation gene sequencing facilitate further in-depth tumor studies, and prognostic gene signatures at the mRNA level can predict OS in PC (including ductal and lobular neoplasms, adenomas, and adenocarcinomas). Cystic, mucinous, and serous neoplasms have too few samples to participate in model construction. It should be noted that pancreatic neuroendocrine tumor, especially nonfunctional neuroendocrine tumor of the pancreas, was a rare case, and we may only learn about this disease from published case report (Mogoanta et al. 2015).
Studies have confirmed that inflammatory response (chronic inflammation, infection, autoimmunity, and tumor elicited inflammation) is related to the occurrence and proliferation of tumors (Dominguez et al. 2017; Greten and Grivennikov 2019; Mantovani et al. 2008; Panigrahy et al. 2019; Takahashi et al. 2010).
Inflammation, inflammatory secretion, and infiltration of immune cells have played a vital role in the launch of cancer and promoting and developing into malignant metastasis (Shadhu and Xi 2019).
In addition, high systemic inflammatory response indexes, such as the neutrophil-albumin ratio (Tingle et al. 2018), platelet–lymphocyte ratio (Lee et al. 2018), C-reactive protein–lymphocyte ratio (Fan et al. 2020), and monocyte–lymphocyte ratio (Sierzega et al. 2017), have been categorically demonstrated to contribute to poor prognosis in PC. A previous study constructed an inflammatory response–related gene signature which could reveal distinct survival outcome in PC (Xie et al. 2022); however, this study did not explored the relationship of the gene signature and immune checkpoint genes and drug-resistance genes.
To the best of our knowledge, genetic signatures associated with inflammatory responses, TME, and tumor drug sensitivity prediction have not been investigated to this date. Herein, an inflammatory response–related gene signature was constructed, and the efficacy, as well as the sensitivity of this signature, was evaluated.
Methods
Data collection
Gene expression information and corresponding clinical information of 182 and 262 PC patients were downloaded from the TCGA database (https://portal.gdc.cancer.gov/) and ICGC database (ICGC-CA) (https://docs.icgc.org/https://docs.icgc.org/), respectively. The clinical information (including OS, survival state, age, gender, and AJCC stage) of PC patients in TCGA and ICGC was summarized in Table 1. These data are publicly available. The detailed information, such as when these patients been recruited, and how many years were they followed, was not found. 200 inflammatory response–related genes were queried in the molecular signature database (Supplementary Table 1). Samples in datasets that meet the following inclusion criteria were included in this study: (1) the sample with both mRNA sequencing data and clinical information; (2) the sample with prognosis information. The preprocessing of TCGA RNA-seq was as follows: (1) PC samples without clinical information were removed; (2) data on normal PC tissue samples were removed; (3) the genes in which the “count” was zero in more than half of the PC samples were excluded. The ICGC data were preprocessed as follows: (1) data on normal PC tissue samples were removed, whereas data on primary tumor were retained; (2) the OS period was converted from day/month to year; (3) and only the expression profile of immune-related genes was included.
Constructing and validating a prognostic inflammatory response–related gene signature
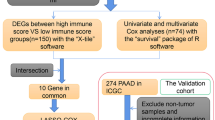
In the TCGA databases, differentially expressed genes (DEGs) in tumor and normal tissues were screened using the “limma” package in R, with the criteria |log2 fold change (FC)|> 1 and adjusted P < 0.05(Lin et al. 2021). Univariate Cox regression analysis was used to screen prognostic-related inflammatory response genes, and the P value was adjusted by Benjamini and Hochberg (BH) correction method. To prevent overfitting, LASSO regression analysis was utilized to construct the model (Simon et al. 2011; Tibshirani 1997). The LASSO algorithm was used to select and shrink variables with the “glmnet” R package, so that some regression coefficients were strictly equal to 0, thereby obtaining an interpretable model. The normalized expression matrix of candidate prognostic DEGs was the independent variable in regression, and the dependent variable was the OS and status of patients in the TCGA cohort. The optimal penalty parameter λ related to the smallest tenfold cross-validation in the TCGA dataset was confirmed.
Next, according to the constructed gene signature, the risk score of each patient was quantified. The calculation formula used is as follows: the expression level of each gene * the corresponding regression coefficient (Huo et al. 2021; Mo et al. 2020). Patients were then stratified into high-risk and low-risk groups based on the median risk score of all patients. Principal component analysis (PCA) analysis and t-distributed stochastic neighbor embedding (t-SNE) analysis using the “Rtsne” and “ggplot2” packages in R were used to investigate the distribution of high and low risk patients.
To explore the predictive power of the prognostic model, survival analysis was conducted using the “survminer” package, and by drawing receiver operating characteristic (ROC) curve using “timeROC” in R. Subsequently, univariate and multivariate Cox analyses were performed using the “survival” package.
Functional enrichment analysis
To explore potential molecular mechanisms, GSEA was conducted in the high- and low-risk groups with GSEA software 4.1.0 (Canzler and Hackermuller 2020; Croken et al. 2014). The enrichment pathway with P < 0.05 represented significant pathway enrichment in the high- and low-risk groups. The infiltration scores of 16 immune cells and the activities of 13 immune-related pathways in different risk score groups were calculated using the single-sample GSEA computed by the “GSVA” R package (Hanzelmann et al. 2013).
TME and chemotherapy sensitivity analysis
We estimated the stromal, immune, and estimate scores in the TME using the ESTIMATE algorithm (Charoentong et al. 2017). The immune and stromal scores were used to evaluate the infiltration of immune cells and stromal cells in tumor tissue (Yoshihara et al. 2013). The enrichment scores of different immune cells, related functions, and pathways were quantified using the ssGSEA.
Based on the NCI-60 database (https://discover.nci.nih.gov/cellminer), Pearson correlation analysis between prognostic gene expression and the sensitivity of anti-tumor drugs was conducted (including 216 drugs approved by FDA; see Supplementary Table 2). The specific method was as follows: (1) download the RNA-seq file and compound activity: DTP NCI-60 file from the NCI-60 database, (2) extract 216 drugs approved by FDA from the DTP NCI-60 file, (3) extract the information of prognostic gene expression, and (4) Pearson correlation analysis was performed to explore the relevance.
Statistical analysis
Wilcoxon test was used to select DEGs. Survival analysis between high-and low-risk score groups was compared using the Kaplan–Meier analysis (log-rank test). The univariate Cox analysis was used to select variables related to OS. The multivariate Cox analyses were used to determine independent risk factors affecting patient prognosis. The Spearman or Pearson correlation analysis was used to compare the relevance between risk score and immune checkpoint genes (including PD-1, PD-L1, PD-L2, and CTLA-4) and drug resistance genes (including MRP1, MRP2, MRP3, MRP4, MRP5, and MRP6, MRP7, MRP8, and MRP9). The ssGSEA scores of immune cells or immune pathways in different risk groups were compared by the Wilcoxon test. The Wilcoxon test was used to compare the expression level of immune checkpoint genes and drug resistance genes. Wilcoxon test was used to compare the risk score in different clinical information (including age, gender, stage, and grade), TME scores (including immune score, stromal score, and estimate score), and immune components (including C1 (wound healing), C2 (INF-g dominant), C3 (inflammatory), and C6 (TGF-b dominant)). Correlation analysis between prognostic gene expression and the sensitivity of anti-tumor drugs was conducted by the Spearman or Pearson test. The R software (Version 3.6.3) with packages venn, igraph, ggplot2, pheatmap, ggpubr, corrplot, and survminer was used to create plots. In all statistical results, a two-tailed P value < 0.05 indicated statistical significance.
Result
Identification of prognostic inflammatory response–related DEGs in the TCGA cohort
A differential analysis using the “limma” package was performed with the selection criteria | log2 FC|> 1 and P < 0.05 in the TCGA database. Moreover, a total of 27 DEGs were screened (Supplementary Table 3). Next, using the univariate Cox analysis, 78 inflammatory response–related genes were analyzed based on their prognostic value (Fig. 1A). Finally, 14 overlapping genes (including AHR, AXL, CD55, F3, GNA15, HBEGF, IL18, INHBA, MMP14, PDPN, PLAUR, SLC7A2, TNFAIP6, and TPBG) between DEGs and inflammatory response–related genes were selected (Fig. 1B, C, D).
Identification of the candidate inflammatory response–related genes based on TCGA cohort. A Forest map of 78 genes with prognostic value. B Venn diagram to identify 14 overlapping genes between DEGs and inflammatory response–related genes with prognostic value. C Forest map of the 14 overlapping genes. D The heat map of 14 overlapping genes expression in PC tissues and adjacent normal tissue samples. PC, pancreatic cancer
Construction of a prognostic model in the TCGA cohort
A LASSO analysis was further conducted to construct a prognostic model, and a 5-inflammatory response–related gene signature was established (Fig. 2A–B). The risk score was calculated as follows: Risk score = AHR × 0.02623 + F3 × 0.01848 + GNA15 × 0.12838 + IL18 × 0.19575 + INHBA × 0.01221. According to the median value of risk scores, samples were stratified into the high- and low-risk score groups. Survival analysis (log-rank test) revealed that patients in the low-risk score group had a better prognosis than those in the high-risk group (Fig. 2C). In addition, a relatively robust area under the curve (AUC) value that was obtained for 1-, 2-, and 3-year OS was 0.679, 0.656, and 0.702, respectively) (Fig. 2D). The survival status map showed that the number of mortalities increased as the risk score increased (Fig. 2E–F). Furthermore, the PCA analysis and T-SNE analysis maps could clearly distinguish patients in the high- or low-risk score groups (Fig. 2G–H).
Construction of a 5-inflammatory response–related gene signature using the TCGA cohort. A Cross-validation for tuning parameter selection in the LASSO model. B LASSO coefficient profiles of 6 prognostic inflammatory response–related genes. C Kaplan–Meier curves of OS of PC patients in the training cohort using the risk score. D ROC plot. E–F The distribution of the risk score, and the survival status of patients. G The PCA plot. H The t-SNE plot. OS, overall survival; PC, pancreatic cancer; ROC, receiver operating characteristic; PCA, principal component analysis; t-SNE, t-distributed stochastic neighbor embedding
External validation of the 5-inflammatory response–related gene signature in the ICGC cohort
External cohort verification (186 HCC patients from the ICGC cohort) was used to confirm the stability of the above model. The formula to calculate the risk score of the ICGC cohort was the same as the TCGA cohort. Based on the median value mentioned previously, we stratified the risk score of the ICGC cohort into a high- or low-risk group.
The survival curve of the low-risk score group was significantly better than the high-risk score group (Fig. 3A). The AUCs for 1, 2, and 3-year OS were 0.753, 0.722, and 0.684, respectively (Fig. 3B). The survival status map showed that the number of mortalities increased as the risk score increased (Fig. 3C–D). The PCA analysis and T-SNE analysis maps could clearly distinguish patients with different risk score groups (Fig. 3E–F), consistent with the results of the TCGA cohort.
Validation of the 5-inflammatory response–related gene signature in the ICGC cohort. A Kaplan–Meier curves of OS of PC patients in the test cohort using the risk score. B ROC curves. C–D The distribution of the risk score, and the survival status of patients. E The PCA plot. F The t-SNE plot. OS, overall survival; PC, pancreatic cancer; ROC, receiver operating characteristic; PCA, principal component analysis; t-SNE, t-distributed stochastic neighbor embedding
Analysis of independent prognostic factors affecting OS based on the TCGA cohort and the ICGC cohort
We included the risk score and other clinical factors (age, gender, grade, and AJCC stage) in univariate and multivariate Cox regression analyses based on the TCGA cohort (clinical factors include age, gender, grade, and AJCC TNM stage) and the ICGC cohort (clinical factors include age, gender, and AJCC TNM stage.) to explore independent prognostic factors affecting OS. Interestingly, we obtained a similar result, with the risk score being an independent factor affecting patient prognosis (Figure 4A, B, C, D).
Prognostic model risk score and clinical features
In addition, a Wilcoxon test between risk scores and clinical features was conducted. The results revealed that a higher risk score was associated with poorer tumor differentiation (P < 0.05) (Fig. 5A, B, C, D).
Immune status and TME analysis
ssGSEA was utilized to explore the relationship between the risk score and immune status, related functions and pathways. The results showed that the fractions of CD8 + T cells, mast cells, and NK cells in the high-risk score group are significantly lower than those in the low-risk score group in the TCGA cohort (P < 0.05) (Fig. 6A). In the ICGC cohort, the fractions of B cells and TIL in the high-risk score group are significantly lower compared to the low-risk score group (P < 0.05) (Fig. 6C). Furthermore, the cytolytic activity and type-II IFN response scores in the TCGA cohort were lower in the high-risk score group than in the low-risk score group. In contrast, the type-I IFN response score was higher in the high-risk score group compared to the low-risk score group (Fig. 6B). In the ICGC cohort, the HLA, MHC class I, T cell co-stimulation, and type-II IFN response scores were lower in the high-risk score group than in the low-risk score group (Fig. 6D). This showed the difference in antigen-presenting cells and T cell regulation between different risk score groups. Next, we explored the potential contribution of the risk score to the TME. The result showed that stromal score experienced a significantly higher trend in the risk-low group (Fig. 6E).
Immune status and TME analysis in different risk-score groups based on TCGA cohort (A, B) and ICGC cohort (C, D). A, C The boxplots of the scores of 16 immune cells. B, D The boxplots of the 13 immune-related functions. E Comparison between the risk score and stromal score, immune score, and estimate score. F Comparison of the risk score in different immune infiltration subtypes. TME, tumor microenvironment
In addition, the correlation between risk scores and immune components was further studied. There are six types of immunogenic permeates in human tumors, corresponding to tumor-promoting and tumor-suppressing functions (Tamborero et al. 2018), namely, C1 (wound healing), C2 (INF-g dominant), C3 (inflammatory), C4 (lymphocyte depleted), C5 (immunologically quiet), and C6 (TGF-b dominant). The results showed that a high-risk score was significantly related to C1, C2, and C6, while a low-risk score was significantly related to C3 (Fig. 6F).
Relationship between risk score and immune checkpoint genes and drug resistance genes
To explore the relationship between risk scores, immune checkpoint genes, and drug resistance genes, the Spearman or Pearson correlation analysis test was performed. Although the results showed that the expression levels of cytotoxic T lymphocyte antigen 4 (CTLA-4), programmed cell death 1 (PD-1), programmed cell death-ligand 1 (PD-L1), programmed cell death-ligand 2 (PD-L2) had no significant difference in high and low-risk score groups (P > 0.05), we found that the expression levels of these genes differed in different risk score groups (Supplementary Fig. 1A-1H). In addition, the expression level of PD-L1 was positively correlated with risk scores. This showed that the prognostic score could provide a theoretical basis for immunotherapy.
The expression levels of MRP1, MRP3, and MRP8 in the high-risk score group were significantly higher than that in the low-risk score group (Supplementary Fig. 2A, 2E, and 2I). The expression levels of MRP1, MRP3, and MRP8 were positively correlated with risk scores (Supplementary Fig. 2B, 2F, and 2J). In contrast, the expression levels of MRP2 and MRP4 in the low-risk score group were significantly higher compared to in the high-risk score group (Supplementary Fig. 2C and 2G). The expression levels of MRP2 and MRP4 were negatively correlated with the risk score (Supplementary Fig. 2D and 2H). However, there was no significant difference in the expression levels of MRP5, MRP6, MRP7, and MRP9 in the high- and low-risk score groups (Supplementary Fig. 3A, 3C, 3E and 3G). The expression levels of MRP6 and MRP9 were negatively correlated with the risk score (Supplementary Fig. 3D, 3H). In comparison, the expression levels of MRP5 and MRP7 were not significantly correlated with the risk score (Supplementary Fig. 3B, 3F).
Prognostic gene expression and cancer cell sensitivity to chemotherapy
Figure 7 shows the relationship between prognostic gene expression and drug sensitivity. The higher the expression level of GNA15, the more drug-sensitive cancer cells were to Nelarabine, Fludarabine, Asparaginase, Cladribine, and Hydroxyurea. The higher the expression level of AHR, the less drug-resistant cancer cells were to Oxaliplatin, Arsenic trioxide, and Lomustine ifosfamide. The higher the expression level of INHBA, the more drug-sensitive cancer cells were to Zoledronate and Dasatinib. The higher the expression, the more drug-sensitive cancer cells were to Tyrothricin. The higher the expression level of IL18, the less drug-resistant cancer cells were to Pipamperone, Bortezomib, Actinomycin D, and Estramustine.
Biological function and pathway analyses
To explore potential mechanisms, GSEA enrichment analysis was conducted, and the results showed that calcium signaling pathway, serine, and threonine metabolism, maturity-onset diabetes of the young, neuroactive ligand-receptor interaction, p53 signaling pathway, pathogenic Escherichia coli infection, pentose phosphate pathway, proteasome, thyroid cancer, and tryptophan metabolism were significantly enriched. Collectively, our results provide new ideas for further targeted gene therapy (Fig. 8).
Validation of the six mRNA expressions
In the TCGA database, all five genes were highly expressed compared to adjacent non-tumor liver tissue. Next, as showed in Fig. 9, we further performed expression verification of the 5 genes from HPA database (The Human Protein Atlas) and GEPIA database (http://gepia.cancer-pku.cn/). Compared to normal pancreatic tissues, AHR, F3, GNA15, IL18, and INHBA were significantly overexpressed in PC tissues. Using clinical samples from the Human Protein Atlas, we analyzed the expression of the proteins encoded by the 5 genes to determine their clinical relevance. Compared to normal pancreatic tissues, IL18 was strongly expressed, while AHR, F3, and GNA15 were moderately expressed. However, the website did not list INHBA.
Differences in protein expression induced by five genes were verified in human tissue samples. A–D The mRNA expression levels in TCGA pancreatic cancer tumor tissue and matching normal tissue from data of TCGA and the genotype-tissue expression (GTEx) database. E–I The representative protein expression of the five genes in pancreatic cancer tumor tissue and normal tissue. Data was obtained from the human protein atlas (https://www.proteinatlas.org/)
Discussion
Public databases greatly facilitate cancer research as well as treatment development and are extensively used nowadays. Inflammation-related genes are receiving more and more attention. To our knowledge, we are the first to investigate the role of inflammation-related genes in PC. Herein, we constructed a prognostic model related to inflammatory response genes using the TCGA database. Notably, we achieved a good AUC for our prognostic model. Additionally, the ICGC database was utilized to validate the prognostic model’s prediction capabilities. We achieved good AUCs for the TCGA and ICGC datasets, and patients with a high-risk score had a worse OS, with the risk score being an independent predictive factor for patients. Therefore, further investigation of genes associated with the 5-inflammatory response can shed light on the mechanism underlying our prognostic model.
AHR is a ligand-activated transcription factor that promotes cancer cell growth and malignancy (Puga et al. 2009; Safe et al. 2013). AHR activation results in immune-tolerant cells such as a dendritic cell (DCS) and Tregs, which co-cultivates the TME, which is defective in identifying and eliminating cancer cells (Cheong and Sun 2018). AHR can activate KYN to further increase the growth and invasiveness of PC cells (Wang et al. 2020). The reallocation of methylated cysteines in the GNA15 gene is associated with the presence of Gα15 in pancreatic ductal adenocarcinoma (PDAC), starting with precursor lesions and continuing through all stages of PDAC (Innamorati et al. 2021). Studies have confirmed that IL-18 plays an essential role in the pathogenesis of chronic and eosinophilic pancreatitis (Manohar et al. 2018, 2017). In addition, macrophage-induced IL-18-mediated inflammatory responses play a crucial function in malignant pancreatic tumors initiation and progression (Kandikattu et al. 2021). Studies have confirmed that INHBA plays a vital role in different tumors (Chen et al. 2019; Lyu et al. 2018). High levels of INHBA expression were positively associated with lower 5-year survival rates in colorectal and gastric cancer patients (Okano et al. 2013; Zhao et al. 2021). The role of F3 in the occurrence and development of PC was still unclear, and further research was needed.
This study conducted a correlation analysis between the immune status and the TME to show the differences in antigen-presenting cells and T cell regulation in different risk groups. We found that type-II IFN response scores in the high-risk group were significantly lower than the low-risk score group. Notably, Type II interferon plays a key role in the killing of tumor cells (Mitra-Kaushik et al. 2004; Street et al. 2002; Wang et al. 2015). We also found that the stromal score of patients in the low-risk group were higher than in the high-risk group. Furthermore, the relationship between the risk score and immune components was investigated. The results revealed that the high-risk scores were associated with C1, C2, and C6, while low-risk scores were associated with C3. These results suggest that C1 could promote tumor initiation and progression, while C3 could have a protective factor, consistent with the findings of a previous study (Tamborero et al. 2018). GSEA analysis revealed an enrichment of oncogenic pathways in the high-risk group. We hypothesized that changes in inflammatory response–related gene expression levels could enhance tumor incidence and growth by altering the TME. The Wilcoxon test of risk score and clinical characteristics revealed that the greater the risk score, the less differentiated the tumor. Together, our findings provided more contexts for the poor outcome of patients in the high-risk group.
To explore the clinical significance of the model, a correlation analysis between prognostic models and the expression levels of immune checkpoint genes and drug resistance genes was carried out. Although the expression levels of CTLA-4, PD-1, PD-L1, and PD-L2 had no significant difference in the high-and low-risk score groups (P > 0.05), we found that the expression levels of immune checkpoint genes were different in different risk-score groups. In addition, the results showed that PD-L1 expression levels were positively correlated with risk scores. This showed that the prognostic score could provide a theoretical basis for immunotherapy.
MRP1 to MRP9 was the primary drug resistance gene against the anti-tumor drug, and the major mechanism is to discharge the anti-tumor chemotherapy drug outside the cells, resulting in tumor drug resistance (Sodani et al. 2012). So, the relationship between risk score and drug resistance genes, including MRP1, MRP2, MRP3, MRP4, and MRP8, indicated that targeting tumor resistance genes appears to have a therapeutic potential for high-risk patients.
Using the NCI-60 database, we uncovered that some genes, such as GNA15 and INHBA, were upregulated. Importantly, these genes were negatively associated with increased drug resistance to Nelarabine, Fludarabine, Asparaginase, Cladribine, Hydroxyurea, Zoledronate, and Dasatinib in cancer cells. However, increased expression of AHR, INHBA, and IL18 was associated with increased drug resistance to Oxaliplatin, Tyrothricin, Pipamperone, Bortezomib, Arsenic trioxide, Lomustine, Actinomycin D, and Estramustine in vivo. Collectively, the above findings might be an important foundation for tumor treatment.
The risk factors for pancreatic cancer include alcohol consumption, smoking, a diet rich in saturated fats, and viral infections such as chronic infection with hepatitis B and C viruses (Gheorghe et al. 2022). The symptomatology remains nonspecific until its advanced stages, and the retroperitoneal location of the pancreas makes imaging diagnosis more difficult. With the establishment of next-generation sequencing technology and the beginning of the era of precision medicine, various treatments for PC have been developed. Large-scale gene sequencing is another heavy economic burden for PC patients. We successfully constructed and validated a 5-inflammatory response–related prognostic model. According to the relationship between prognostic model and immune checkpoint genes or drug resistance genes, the corresponding immunotherapy or chemotherapy may be applied to patients in different risk groups.
Also, the 5-inflammatory response–related gene signature (including AHR, F3, GNA15, IL18, and INHBA) could not be used for the diagnosis of pancreatic cancer in early stages. Clinical manifestations, endoscopic ultrasound, endoscopic retrograde cholangiopancreatography (ERCP), CT, MRI, and PET, and methylation of ADAMTS1 and BNC1 showed a certain sensitivity and specificity for the early diagnosis of PC (Gheorghe et al. 2020).
We have not performed further experiments to explore the mechanisms of these inflammatory response–related genes, but we will address this in the future work. Moreover, as for the PC patients who were mostly diagnosed by imaging modalities and treated with nonsurgical methods, the value of this model may be limited, because our model need to quantify the expression levels of five specific genes in tumor tissues. Therefore, it is necessary to continue performing multicenter prospective research on this subject.
Conclusion
In conclusion, we successfully constructed a 5-inflammatory response–related gene signature based on the TCGA database and verified the stability of the model using the ICGC database. Nevertheless, further substantiation is warranted to verify the role of these genes in tumorigenesis. Taken together, our findings might shed new light on immune responses, the TME, and drug resistance to promote the development of personalized cancer therapies.
Data availability
The datasets analyzed for this study were obtained from The Cancer Genome Atlas (TCGA) (https://portal.gdc.cancer.gov/) and International Cancer Genome Consortium (ICGC) (https://icgc.org/).
References
Ansari D, Tingstedt B, Andersson B, Holmquist F, Sturesson C, Williamsson C, Sasor A, Borg D, Bauden M, Andersson R (2016) Pancreatic cancer: yesterday, today and tomorrow. Future Oncol 12:1929–1946
Blum R, Kloog Y (2014) Metabolism addiction in pancreatic cancer. Cell Death Dis 5:e1065
Canzler S, Hackermuller J (2020) multiGSEA: a GSEA-based pathway enrichment analysis for multi-omics data. BMC Bioinformatics 21:561
Charoentong P, Finotello F, Angelova M, Mayer C, Efremova M, Rieder D, Hackl H, Trajanoski Z (2017) Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep 18:248–262
Chen ZL, Qin L, Peng XB, Hu Y, Liu B (2019) INHBA gene silencing inhibits gastric cancer cell migration and invasion by impeding activation of the TGF-beta signaling pathway. J Cell Physiol 234:18065–18074
Cheong JE, Sun L (2018) Targeting the IDO1/TDO2-KYN-AhR pathway for cancer immunotherapy — challenges and opportunities. Trends Pharmacol Sci 39:307–325
Croken MM, Qiu W, White MW, Kim K (2014) Gene set enrichment analysis (GSEA) of Toxoplasma gondii expression datasets links cell cycle progression and the bradyzoite developmental program. BMC Genomics 15:515
Dominguez C, David JM, Palena C (2017) Epithelial-mesenchymal transition and inflammation at the site of the primary tumor. Semin Cancer Biol 47:177–184
Fan Z, Luo G, Gong Y, Xu H, Qian Y, Deng S, Huang Q, Yang C, Cheng H, Jin K et al (2020) Prognostic value of the C-reactive protein/lymphocyte ratio in pancreatic cancer. Ann Surg Oncol 27:4017–4025
Ferlay J, Partensky C, Bray F (2016) More deaths from pancreatic cancer than breast cancer in the EU by 2017. Acta Oncol 55:1158–1160
Gheorghe G, Bungau S, Ilie M, Behl T, Vesa CM, Brisc C, Bacalbasa N, Turi V, Costache RS, Diaconu CC (2020) Early diagnosis of pancreatic cancer: the key for survival. Diagnostics (Basel) 10(11):869
Gheorghe G, Diaconu CC, Ionescu V, Constantinescu G, Bacalbasa N, Bungau S, Gaman MA, Stan-Ilie M (2022) Risk factors for pancreatic cancer: emerging role of viral hepatitis. J Pers Med 12(1):83
Greten FR, Grivennikov SI (2019) Inflammation and cancer: triggers, mechanisms, and consequences. Immunity 51:27–41
Hanzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14:7
Houot R, Schultz LM, Marabelle A, Kohrt H (2015) T-cell-based immunotherapy: adoptive cell transfer and checkpoint inhibition. Cancer Immunol Res 3:1115–1122
Huo J, Wu L, Zang Y (2021) Development and validation of a novel metabolic-related signature predicting overall survival for pancreatic cancer. Front Genet 12:561254
Innamorati G, Wilkie TM, Malpeli G, Paiella S, Grasso S, Rusev B, Leone BE, Valenti MT, Carbonare LD, Cheri S et al (2021) Galpha15 in early onset of pancreatic ductal adenocarcinoma. Sci Rep 11:14922
Kamisawa T, Wood LD, Itoi T, Takaori K (2016) Pancreatic cancer. Lancet 388:73–85
Kandikattu HK, Manohar M, Verma AK, Kumar S, Yadavalli CS, Upparahalli Venkateshaiah S, Mishra A (2021) Macrophages-induced IL-18-mediated eosinophilia promotes characteristics of pancreatic malignancy. Life Sci Alliance 4(8):e202000979
Kure S, Matsuda Y, Hagio M, Ueda J, Naito Z, Ishiwata T (2012) Expression of cancer stem cell markers in pancreatic intraepithelial neoplasias and pancreatic ductal adenocarcinomas. Int J Oncol 41:1314–1324
Lee BM, Chung SY, Chang JS, Lee KJ, Seong J (2018) The neutrophil-lymphocyte ratio and platelet-lymphocyte ratio are prognostic factors in patients with locally advanced pancreatic cancer treated with chemoradiotherapy. Gut Liver 12:342–352
Li S, Xu HX, Wu CT, Wang WQ, Jin W, Gao HL, Li H, Zhang SR, Xu JZ, Qi ZH et al (2019) Angiogenesis in pancreatic cancer: current research status and clinical implications. Angiogenesis 22:15–36
Lin Z, Xu Q, Miao D, Yu F (2021) An inflammatory response-related gene signature can impact the immune status and predict the prognosis of hepatocellular carcinoma. Front Oncol 11:644416
Lutz ER, Kinkead H, Jaffee EM, Zheng L (2014) Priming the pancreatic cancer tumor microenvironment for checkpoint-inhibitor immunotherapy. Oncoimmunology 3:e962401
Lyu S, Jiang C, Xu R, Huang Y, Yan S (2018) INHBA upregulation correlates with poorer prognosis in patients with esophageal squamous cell carcinoma. Cancer Manag Res 10:1585–1596
Manohar M, Verma AK, Venkateshaiah SU, Mishra A (2018) Role of eosinophils in the initiation and progression of pancreatitis pathogenesis. Am J Physiol Gastrointest Liver Physiol 314:G211–G222
Manohar M, Verma AK, Venkateshaiah SU, Sanders NL, Mishra A (2017) Pathogenic mechanisms of pancreatitis. World J Gastrointest Pharmacol Ther 8:10–25
Mantovani A, Allavena P, Sica A, Balkwill F (2008) Cancer-related inflammation. Nature 454:436–444
Mitra-Kaushik S, Harding J, Hess J, Schreiber R, Ratner L (2004) Enhanced tumorigenesis in HTLV-1 tax-transgenic mice deficient in interferon-gamma. Blood 104:3305–3311
Mizrahi JD, Surana R, Valle JW, Shroff RT (2020) Pancreatic cancer. Lancet 395:2008–2020
Mo Z, Yu L, Cao Z, Hu H, Luo S, Zhang S (2020) Identification of a hypoxia-associated signature for lung adenocarcinoma. Front Genet 11:647
Mogoanta SS, Costache A, Mutiu G, Bungau SG, Ghilusi M, Grosu F, Vasile M, Vilcea ID, Gherghinescu MC, Mogoanta L et al (2015) A nonfunctional neuroendocrine tumor of the pancreas — a case report. Rom J Morphol Embryol 56:511–519
Okano M, Yamamoto H, Ohkuma H, Kano Y, Kim H, Nishikawa S, Konno M, Kawamoto K, Haraguchi N, Takemasa I et al (2013) Significance of INHBA expression in human colorectal cancer. Oncol Rep 30:2903–2908
Okasha H, Elkholy S, El-Sayed R, Wifi MN, El-Nady M, El-Nabawi W, El-Dayem WA, Radwan MI, Farag A, El-Sherif Y et al (2017) Real time endoscopic ultrasound elastography and strain ratio in the diagnosis of solid pancreatic lesions. World J Gastroenterol 23:5962–5968
Panigrahy D, Gartung A, Yang J, Yang H, Gilligan MM, Sulciner ML, Bhasin SS, Bielenberg DR, Chang J, Schmidt BA et al (2019) Preoperative stimulation of resolution and inflammation blockade eradicates micrometastases. J Clin Invest 129:2964–2979
Puga A, Ma C, Marlowe JL (2009) The aryl hydrocarbon receptor cross-talks with multiple signal transduction pathways. Biochem Pharmacol 77:713–722
Raj D, Aicher A, Heeschen C (2015) Concise review: stem cells in pancreatic cancer: from concept to translation. Stem Cells 33:2893–2902
Safe S, Lee SO, Jin UH (2013) Role of the aryl hydrocarbon receptor in carcinogenesis and potential as a drug target. Toxicol Sci 135:1–16
Shadhu K, Xi C (2019) Inflammation and pancreatic cancer: an updated review. Saudi J Gastroenterol 25:3–13
Sierzega M, Lenart M, Rutkowska M, Surman M, Mytar B, Matyja A, Siedlar M, Kulig J (2017) Preoperative neutrophil-lymphocyte and lymphocyte-monocyte ratios reflect immune cell population rearrangement in resectable pancreatic cancer. Ann Surg Oncol 24:808–815
Simon N, Friedman J, Hastie T, Tibshirani R (2011) Regularization paths for Cox’s proportional hazards model via coordinate descent. J Stat Softw 39:1–13
Sodani K, Patel A, Kathawala RJ, Chen ZS (2012) Multidrug resistance associated proteins in multidrug resistance. Chin J Cancer 31:58–72
Son J, Lyssiotis CA, Ying H, Wang X, Hua S, Ligorio M, Perera RM, Ferrone CR, Mullarky E, Shyh-Chang N et al (2013) Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496:101–105
Street SE, Trapani JA, MacGregor D, Smyth MJ (2002) Suppression of lymphoma and epithelial malignancies effected by interferon gamma. J Exp Med 196:129–134
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global Cancer Statistics 2020: GLOBOCAN Estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249
Takahashi H, Ogata H, Nishigaki R, Broide DH, Karin M (2010) Tobacco smoke promotes lung tumorigenesis by triggering IKKbeta- and JNK1-dependent inflammation. Cancer Cell 17:89–97
Tamborero D, Rubio-Perez C, Muinos F, Sabarinathan R, Piulats JM, Muntasell A, Dienstmann R, Lopez-Bigas N, Gonzalez-Perez A (2018) A pan-cancer landscape of interactions between solid tumors and infiltrating immune cell populations. Clin Cancer Res 24:3717–3728
Tibshirani R (1997) The lasso method for variable selection in the Cox model. Stat Med 16:385–395
Tingle SJ, Severs GR, Goodfellow M, Moir JA, White SA (2018) NARCA: A novel prognostic scoring system using neutrophil-albumin ratio and Ca19-9 to predict overall survival in palliative pancreatic cancer. J Surg Oncol 118:680–686
Wang L, Tang W, Yang S, He P, Wang J, Gaedcke J, Strobel P, Azizian A, Ried T, Gaida MM et al (2020) NO(*) /RUNX3/kynurenine metabolic signaling enhances disease aggressiveness in pancreatic cancer. Int J Cancer 146:3160–3169
Wang L, Wang Y, Song Z, Chu J, Qu X (2015) Deficiency of interferon-gamma or its receptor promotes colorectal cancer development. J Interferon Cytokine Res 35:273–280
Xie F, Huang X, He C, Wang R, Li S (2022) An inflammatory response-related gene signature reveals distinct survival outcome and tumor microenvironment characterization in pancreatic cancer. Front Mol Biosci 9:876607
Yoshihara K, Shahmoradgoli M, Martinez E, Vegesna R, Kim H, Torres-Garcia W, Trevino V, Shen H, Laird PW, Levine DA et al (2013) Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun 4:2612
Zhao Z, Wang K, Tan S (2021) microRNA-211-mediated targeting of the INHBA-TGF-beta axis suppresses prostate tumor formation and growth. Cancer Gene Ther 28:514–528
Acknowledgements
We thank the Department of Hepatobiliary and Pancreatic Surgery, WeiFang People’s Hospital.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Manjiang Li wrote the article. Yuxu Wang and Yongbiao Ma processed the data analysis. Wei Ding and Futian Du designed the study and reviewed the article.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable
Consent for publication
Not applicable
Conflict of interests
The authors declare no competing interests.
Additional information
Responsible editor Lotfi Aleya
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
11356_2022_23252_MOESM1_ESM.pdf
Supplementary file1 Correlation analysis between inflammatory response‑related signature and the expression levels of immune checkpoint molecules. (A, B) PD-1. (C, D) PD-L1. (E, F) PD-L2. (G, H) CTLA-4. PD-1, programmed cell death 1; PD-L1, programmed cell death-ligand 1; PD-L2, programmed cell death-ligand 2; CTLA-4, Cytotoxic T Lymphocyte antigen 4. (PDF 995 KB)
11356_2022_23252_MOESM2_ESM.pdf
Supplementary file2 Wilcox test test and correlation analysis between inflammatory response‑related signature and the expression levels of tumor resistance genes. (A, B) MRP1. (C, D) MRP2. (E, F) MRP3. (G, H) MRP4. (I, J) MRP8. (PDF 1110 KB)
11356_2022_23252_MOESM3_ESM.pdf
Supplementary file3 Correlation analysis between prognostic model and the expression levels of tumor resistance genes. (A, B) MRP5. (C, D) MRP6. (E, F) MRP7. (G, H) MRP9. (PDF 971 KB)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, M., Ding, W., Wang, Y. et al. Development and validation of a gene signature for pancreatic cancer: based on inflammatory response–related genes. Environ Sci Pollut Res 30, 17166–17178 (2023). https://doi.org/10.1007/s11356-022-23252-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-23252-w