Abstract
Soil methane generation mainly driven by soil prokaryotic microbes can be coupled with the degradation of petroleum hydrocarbons (PHCs); however, the relationship between prokaryotic community structure and methane production activity in soil with the potential risk of PHC contamination is seldom reported. In this study, 3 soil samples (CS-1 to CS-3) in the area nearby an exploratory gas well and 5 soil samples (DC-1 to DC-5) in a drill cutting dump area were obtained from the Fuling shale gas field (Chongqing City, China). Then, the prokaryotic community structure was examined by Illumina Miseq sequencing, and the linkage between soil methane production rate (MPR) and prokaryotic community composition was analyzed. The results indicated that 2 samples (DC-4 and DC-5) collected from the drill cutting dump area had significantly higher MPR than the other samples, and a significant and positive relationship (r = 0.44, P < 0.05) was found between soil MPR and soil organic matter (OM) content. The prokaryotic community composition in the sample (DC-5) with the highest MPR was different from those in the other samples, and soil OM and MPR were the major factors significantly correlated with the prokaryotic community structure in this soil. The samples (DC-4 and DC-5) with higher MPR had a higher relative abundance of Archaea and different archaeal community structures from the other samples, and the MPR was the sole factor significantly correlated with the archaeal genus composition in this soil. Therefore, both the prokaryotic and archaeal community structures are essential in the determination of soil MPR, and the bacterial genus of Saccharibacteria and the archaeal genus of Methanolobus might be the key contributors for methane generation in this soil from the shale gas field.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Soils nearby the oil and gas fields have a high risk of petroleum hydrocarbons (PHCs) pollution, as high PHCs content can be usually detected in these soils (Liu et al. 2015; Sun et al. 2015; Wang et al. 2015). PHCs are a mixture of varied organic compounds, including alkanes, BTEX (benzene, toluene, ethylbenzene, and xylene), and polycyclic aromatic hydrocarbons (PAHs), which are toxic for ecosystem and human health due to their carcinogenicity, mutagenicity, and teratogenicity (Salanitro et al. 1997; Ritchie et al. 2001; Huang et al. 2016). Soil is considered as a major reservoir for PHCs, and PHCs in the soil can enter food chains, posing indirect threats to human health (Van-der-Oost et al. 2003; Xue and Warshawsky 2005; Gao et al. 2019). As such, soils in the region nearby the oil and gas fields, as aged soils with PHC pollution, should be very ideal objects for the study about the transformation of PHCs in soil.
At present, the investigation about the interaction between PHCs and soil microbes has been a hotspot. As soil microbes are not only the major consumers for soil PHCs but also can be used as sensitive indicators for the assessment of soil PHC contamination (Haritash and Kaushik 2009; Khan et al. 2013). In recent years, many investigators had pay attention to the responses of microbial community structure to PHC contamination in the soils within several important oilfields in China (Liao et al. 2015; Liu et al. 2015; Sun et al. 2015; Gao et al. 2016; Zhou et al. 2017; Gao et al. 2019). For example, in the soil collected from Dagang Oilfield (North China), it was reported that PHC pollution resulted in the increase of the gene copies of AlkB (alkane monooxygenase gene), while the decrease of the Nah (naphthalene dioxygenase gene) gene copies (Liu et al. 2015). Zhou et al. (2017) reported that, in the soil within the region of the Jianghan oilfield (Central China), the community structure of different microbes showed varied sensitivities to PHC contamination, with the order of bacteria > fungi > Archaea. Moreover, some bacterial genera, such as Alkanindiges, Arthrobacter, Idiomarina, Lysobacter, Massilia, Micromonospora, Mycobacterium, Oxalobacteraceae, and Pseudomonas were found to be potential PHC degraders in soil (Han et al. 2014; Li et al. 2015; Sun et al. 2015; Chen et al. 2016). Meanwhile, some fungal genera, such as Trichoderma, Fusarium, and Pestalotiopsis also, were often detected in soils contaminated by PHCs (Llado et al. 2013; Cebron et al. 2015; Kristanti and Hadibarata 2015; Andreolli et al. 2016; Zhou et al. 2017). Those studies identified some key microbes with the ability to degrade PHCs and confirmed the selection effect of PHCs on soil microbiota in the area of oilfields, which could be essential for the bioremediation and risk assessment of PHC contamination for local soils.
Methane, as the simplest alkane with a molecular structure of tetrahedron, is the main constituent of natural gas. Despite its strong greenhouse effect, methane has been considered as a clean energy source with high efficiency (Cornelissen et al. 2012; Kwietniewska and Tys 2014; Morel et al. 2019). Thus, the digestion of organic wastes or pollutants under methanogenic conditions is regarded as an ideal way, as this process can produce biogas mainly containing methane and carbon dioxide (Vergara-Fernandez et al. 2008). Furthermore, it has been confirmed that some PHCs, such as PAHs, long-chain alkane, and BTEX, can be degraded under methanogenic conditions (Ficker et al. 1999; Chang et al. 2006), and it had been reported that crude oil contamination could enhance soil methane emission (Dunfield et al. 1993; Le Mer and Roger 2001; Yang et al. 2018). The biological methane production in soil is a complex process driven by diverse prokaryotic microbes (bacteria and Archaea), whereas little information is available about the relationship between methane production activity and diversity of prokaryotic community in the soils with high risk of PHC pollution, which should be significant for the resource-based treatment of soil with PHC pollution.
In this study, soil samples were collected from the Fuling shale gas field, Chongqing City, China, which is the second-largest shale gas field in the world, and the relationship among soil properties, methane production rate (MPR), and diversity of prokaryotic community was investigated. The aim was to reveal the relationship between the MPR and the diversity of the prokaryotic community and identify the key microbes affecting MPR in this soil with a potential risk of PHC pollution, thus providing some useful microbial information for the resource-based treatment of PHC-polluted soil under methanogenic conditions.
Materials and methods
Site description and soil sampling
The Fuling shale gas field with gas storage of nearly 2.1 trillion cubic meters is the second-largest shale gas field in the world, located in Fuling District, Chongqing, China. Belonging to a subtropical monsoon moist climate, this region has a mean annual temperature of 18.1 °C and precipitation of 1072 mm, and the soils are classified as Cambisols (Chinese taxonomy) or Inceptisols (USDA taxonomy). In this study, 8 soil samples (triplicates for each sample) were obtained from this shale gas field in September 2018, and the geographic distribution for those sampling sites is presented in Fig. 1. Among those samples, 3 samples (CS-1, CS-2, and CS-3) and 5 samples (DC-1, DC-2, DC-3, DC-4, and DC-5) were obtained from the area nearby an exploratory shale gas well (CS) and a drill cutting dump (DC), respectively. At each sampling site, triplicate soil samples were collected, and each sample was obtained from a mixture of 8 soil cores (5 cm in diameter; 20 cm in depth) randomly selected within an area of 100 m2. Then, the soil samples were kept in an icebox and taken back to the lab within 24 h. After passing through a 2-mm sieve, all soil samples were stored at 4 °C and − 20 °C for the following basic properties measurement and DNA extraction, respectively.
Measurement of soil basic properties and methane production rate
Soil pH was determined with soil to water ratio of 2:5. Soil organic matter (OM) and available phosphorus (AP) were determined according to the methods of Lu (1999). After extracted from 5 g of fresh soil with 2 M L−1 KCl, ammonium and nitrate were measured by a continuous flow analyzer (SAN++, Skalar, Holland). The contents of two representative PHCs (pyrene and benzopyrene) in those soil samples were measured according to the method of Cao et al. (2009), and the detailed procedure is available in the supplementary materials (Text S1).
Soil MPR was measured under a glucose-induced and anaerobic condition mainly according to the method of Liu et al. (2008) with some modification. For each soil sample, triplicates were set up, and the treatment without the addition of soil was used as control. The detailed procedure was as follows: 20 g fresh soil, 14 mL sterilized water, and 6 mL glucose (0.1 M L−1) was added into a serum bottle of 100 mL; after sealed with rubber stopper and aluminum cover, the serum bottle was filled with pure N2 gas for 5 min to replace the air in the bottle; the serum bottle was placed in a shaker at 250 rpm for 30 min, and then was filled with pure N2 gas for 3 min to ensure an absolutely anaerobic environment; after the serum bottle was incubated at 30 °C for 72 h in dark, the concentration of methane in the serum bottle was detected by a gas chromatograph (Agilent Technologies 7890A, USA). Finally, the MPR in each soil sample was calculated according to the method of Zhang et al. (2011).
Soil DNA extraction and Illumina MiSeq sequencing
A Fast DNA® SPIN Kit for Soil (Q-BIOgene, Carlsbad, CA, USA) was used to extract the total DNA from 0.5 g of frozen soil according to the manufacturer’s protocol. The quantity and quality of the extracted DNA were checked using a 1% agarose gel electrophoresis and NanoDrop 2000 UV-vis spectrophotometer (Thermo Scientific, Wilmington, USA), respectively, and then was stored at − 20 °C before use.
In this study, the primer set of 515F (GTGCCAGCMGCCGCGGTAA) and 806R (GGACTACVSGGGTATCTAAT) was used to amplify the V4 hypervariable region of the prokaryotic 16S rRNA gene (Bates et al. 2011). After addition of the Illumina adaptors A and B to the forward and reverse primer sequences, respectively, the target gene was amplified in a 20 μL mixture with the following thermal profile: 3 min at 95 °C; 30 cycles of 30 s at 95 °C, 30 s at 62 °C, and 45 s 72 °C; 10 min at 72 °C. The PCR products were extracted and purified, and then were sequenced on an Illumina MiSeq platform (Illumina, San Diego, USA) by Majorbio Bio-Pharm Technology Co. Ltd. (Shanghai, China). The procedure of the raw data processing and operational taxonomic units (OTUs) identification was conducted according to the method described by Zhou et al. (2017) presented in the supplementary materials (Text S2). The taxonomy of each representative OTU was aligned by the RDP Classifier algorithm (http://rdp.cme.msu.edu/) based on the Silva ribosomal database (release128, http://www.arb-silva.de) using confidence threshold of 70% (Amato et al. 2013). The diversity indices of Sobs (species observed), Chao, Shannon, and Simpson were calculated by Mothur software (version v.1.41.1, http://www.mothur.org/wiki/Schloss_SOP#Alpha_diversity) under a similarity cutoff of 97% (Schloss et al. 2011). The sequencing data in this study has been submitted into the Sequence Read Archive in the NCBI (National Center for Biotechnology Information, www.ncbi.nlm.nih.gov) with the accession number PRJNA591071.
Data analysis
The quantitative difference about the properties or soil MPR between soil samples was examined by a Student-Newman-Keuls test in the one-way analysis of variance (ANOVA) using SPSS 24 (SPSS, Chicago, USA), and P < 0.05 was regarded as a significant difference. The correlation between MPR and prokaryotic diversity indices was analyzed using SPSS 24. In R software (http://www.r-project.org), the cluster analysis based on the microbial phyla was done with the method of “Jaccard distance” using the Vegan package, and the composition of the microbial genus in the samples was visualized by the heat map with the Pheatmap package. The redundancy analysis (RDA) profiles were generated based on the matrices of prokaryotic or archaeal composition and environmental variables using the CANOCO (version 5.0), and the detailed analysis steps were described in the supplementary materials (Text S3). Using OTU-based Bray-Curtis distance matrices, the correlation between prokaryotic community structures and environmental variables was evaluated by a Mantel test with 999 permutations in R software (http://www.r-project.org).
Results
Soil properties and MPR
The results of soil properties are listed in Table 1. In this region, the soil was alkaline with a narrow pH range of 8.09~8.34. Except DC-3, the samples (DC-1, DC-2, DC-4, and DC-5) from the drill cutting dump (DC) had significantly higher OM content than those samples (CS-1, CS-2, and CS-3) from the exploratory shale gas well (CS), and the OM contents in the DC-4 and DC-5 were significantly higher than all the other samples. The DC-4 and DC-5 had lower NH4+ content, which was significantly lower than that in the CS-1, while no significant difference was detected in the soil NO3− content among these soil samples. As for soil available phosphorus (AP), except DC-5, the other DC samples had significantly higher AP content than those CS samples. The result of soil MPR in these samples from the shale gas field is shown in Fig. 2. The significantly higher MPR values were detected in the DC-5 and DC-4, and the DC-1 and DC-2 showed significantly lower MPR values than the other samples.
The soil prokaryotic diversity indices
In these soil samples, a total of 344,805 available sequences were obtained from the Illumina Miseq Sequencing, and the diversity indices of prokaryotic microbes in them are presented in Table 2. The coverage of each soil sample was above 0.965, indicating that the result of Illumina Miseq Sequencing was successful and credible. Compared with the other samples, the DC-5 with the highest MPR had the lower values of Sobs and Chao, while it had a higher value of Simpson.
The prokaryotic community composition in the soil from the shale gas field
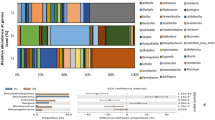
The cluster analysis and relative abundance of the prokaryotic phylum in these soil samples are shown in Fig. 3a. A total of 10 known bacterial phyla was detected in each soil sample, which occupied more than 90% of the total sequences obtained from the Illumina Miseq Sequencing. The cluster analysis indicated that those soil samples could be grouped into three clusters. In detail, four samples (DC-3, CS-2, DC-4, and CS-3) and three samples (DC-1, CS-1, and DC-2) were divided into two different clusters, whereas DC-5 formed an isolated cluster far away from the former two clusters, indicating that the DC-5 with the highest MPR had a unique phylum composition. Compared with the other samples, DC-5 had lower relative abundances of Acidobacteria, Actinobacteria, Chloroflexi, Cyanobacteria, and Planctomycetes, while had a higher relative abundance of Saccharibacteria (Fig. 3a). The heat map constructed from the dominant prokaryotic genera (top 50) in these soil samples is shown in Fig. 3b. Among these dominant microbial genera, there were 47 known bacterial genera distributed in 11 bacterial phyla, while there was only 1 archaeal genus (Crenarchaeotic). Similar to the result of cluster analysis based on the phylum level, the prokaryotic genus composition in DC-5 also was different from the other samples, and a higher relative abundance of norank Saccharibacteria was detected in the soil samples (DC-4 and DC-5) with higher MPR.
A low ratio (< 2%) of archaeal sequences to total sequences was detected in each soil sample from this shale gas field, and the soil samples (DC-4 and DC-5) with higher MPR had higher ratios of archaeal sequences to total sequences than the other samples (Fig. 4a). The relative abundance of archaeal phylum in each soil sample is shown in Fig. 4b. Four archaeal phyla of Bathyarchaeota, Euryarchaeota, Thaumarchaeota, and Woesearchaeota were detected in the soil from the shale gas region, and the soil samples (DC-4 and DC-5) with higher MPR had distinct archaeal phylum composition, which almost was solely occupied by Thaumarchaeota and Woesearchaeota, respectively (Fig. 4b).
Relationship between soil properties and prokaryotic community structure
The RDA profiles constructed from soil properties and the dominant prokaryotic OTUs (top 50) or archaeal genera in these soil samples are shown in Fig. 5. Based on the dominant OTUs composition, although a long distance was shown between the two samples (DC-4 and DC-5) with higher MPR, they were separated from the other samples, indicating their distinct prokaryotic community structure (Fig. 5a). Additionally, the result of the mental test with 999 permutations indicated that soil MPR (R2 = 0.98, P < 0.01) and OM content (R2 = 0.69, P < 0.05) were significantly correlated with the soil prokaryotic community structure (Fig. 5a and Table S3). The RDA profile based on the archaeal genus showed that, except DC-3, the soil samples from the DC field could be separated from those from the CS field (Fig. 5b). The sample with the highest MPR (DC-5) was isolated from the other samples, and the archaeal genus of Methanolobus was the most closely related to soil MPR (Fig. 5b). Furthermore, the soil MPR (R2 = 0.95, P < 0.05) was the only variable significantly correlated with soil archaeal genus composition (Fig. 5b and Table S3).
Discussion
In this study, the soil samples (DC-4 and DC-5) from the DC region had higher OM contents and higher MPR than the other samples, and a significantly positive correlation (r = 0.44, P < 0.05) was found between soil OM content and MPR (Table S1). Soil OM can supply the basic substrates for microbes to synthesize methane and has been considered as a crucial factor determining methane generation rate in both wetland and upland soils (Marschner et al. 2003; Bayer et al. 2012; Morin et al. 2014; Peng et al. 2015; Ye et al. 2016). Moreover, high OM contents were usually detected in soils with PHC pollution, and then OM content could be an important indicator for soil PHC contamination in oil and gas fields (Trofimov and Rozanova 2003; Ogboghodo et al. 2004; Zhou et al. 2017). In this study, compared with other samples, higher contents of two representative PHCs (pyrene and benzopyrene) were detected in DC-4 and DC-5 (Table S2). Therefore, the higher OM contents in DC-4 and DC-5 might be due to the PHC pollution, and the higher MPR in them confirmed that the increase in soil OM caused by PHC contamination could significantly enhance soil methane production (Yang et al. 2018).
In this study, the sample (DC-5) with the highest MPR had the lowest Sobs, Chao, and Shannon indices (Table 2). The negative correlation between soil PHCs content and bacterial diversity in the soils within several oilfields had been reported, which revealed the toxicity and selection of PHCs on soil microbes (Perez-Leblic et al. 2012; Elarbaoui et al. 2015; Abbasian et al. 2016; Zhou et al. 2017). No investigation had reported the relation between microbial diversity and MPR in those soils with PHC pollution, while Gebert and Perner (2015) found that the soil (collected from a municipal solid waste landfill region) with the highest OM and methane concentration also had the lowest bacterial diversity, which was consistent with the result in this study. Additionally, the result of this study showed that soil samples (DC-4 and DC-5) with higher MPR had higher portions of the archaeal sequence (Fig. 4a). In soil, those microbes directly participate in methane generation all belong to Archaea, which are not sensitive to both organic and inorganic pollutants due to their special structure and components in the cell wall (Valentine 2007; He et al. 2012). For example, it is ammonia-oxidizing Archaea but not ammonia-oxidizing bacteria can be responsible for the ammonia-oxidization process in some unfavorable environments, such as high temperature, low pH, low oxygen, and high salinity (Erguder et al. 2009; Hatzenpichler et al. 2008; Schauss et al. 2009). The major component is peptidoglycan in the bacterial cell wall, but it is pseudo-peptidoglycan or glycoprotein in Archaea; moreover, ester-linked lipids and ether-linked lipids are the major components in the cell membrane of bacteria and Archaea, respectively (Derosa et al. 1986; Kandler and Konig 1998). These differences in the cell structure might result in the low permeability for both nutrients and toxic pollutants in the archaeal cell, and thus ensure a decent environment for its energy metabolism and growth under the harsh environments (He et al. 2012; Valentine 2007). Consequently, in this study, the higher OM contents in these samples (DC-4 and DC-5) with higher MPR and higher contents of pyrene and benzopyrene (Fig. 2 and Table S2) might indicate their high risk of PHC contamination, which resulted in the decrease in prokaryotic diversity but the increase in the archaeal relative abundance in this soil.
In this study, those two samples (DC-4 and DC-5) with higher soil OM content and MPR all had a higher relative abundance of Saccharibacteria than the other samples (Fig. 3). This kind of bacteria has been widely detected in soil, activated sludge, sediment, and other habitats (Bond et al. 1995; Borneman and Triplett 1997; Hugenholtz et al. 2001; Pace 2009), which can utilize various monosaccharides, amino acids, and proteins, and can ferment glucose and other sugars to produce lactate, playing an important role in mineralization of organic compounds (Ariesyady et al. 2007; Nielsen et al. 2009; Nielsen et al. 2010; Albertsen et al. 2013). More importantly, although it is unsure whether this bacterial genus can utilize complex PHCs, its ability to degrade some simple PHCs in soil, such as toluene and benzene, has been confirmed (Luo et al. 2009; Xie et al. 2011). Thus, we hypothesized that Saccharibacteria might play a critical role in the degradation of OM or PHCs in this gas field soil, then supplying organic substrates for methanogens. In addition, the RDA profile constructed from the dominant OTUs indicated that samples (DC-4 and DC-5) with higher MPR had different prokaryotic community structures from the other samples, and soil OM (R2 = 0.69, P < 0.05) and MPR (R2 = 0.98, P < 0.01) were the key factors influencing the soil prokaryotic community structure (Fig. 5a and Table S3). The soil OM content is not only an indicator for assessment of soil PHC contamination but also the important carbon and energy sources for soil microbes, which has long been considered as one of the most important factors affecting soil microbial community structure (Perez-Leblic et al. 2012; Abbasian et al. 2016; Zhou et al. 2017; Gao et al. 2019). It has been previously reported that there was a significant correlation between the community structure of methanogens and soil MPR (Reim et al. 2017; Chen et al. 2018; Zhang et al. 2018), but the relationship between MPR and total prokaryotic community structure is still uncertain. For example, Wagner et al. (2017) reported that the prokaryotic community structure might control the MPR in Arctic Alaska soils, but no correlation was found between soil methane emission and prokaryotic community structure in the forest soil in Brazil (Cuer et al. 2018). Being consistent with the result of Wagner et al. (2017), our current result also found a significant correlation between the prokaryotic community structure and MPR in this soil from the shale gas field. In our opinion, besides a very small portion of Archaea in prokaryotic microbes directly participating in the methane biological synthesis, most of the prokaryotic microbes also might indirectly take part in the process of methane synthesis by supplying low molecular substrates degraded from complex organic compounds. Therefore, soil MPR ought to be tightly correlated to the prokaryotic community structure.
In this study, the RDA profile based on the archaeal genus revealed that the archaeal genus composition in DC-5 with the highest MPR was different from the other samples, and only MPR was significantly correlated with soil archaeal community structure (Fig. 5b and Table S3). Soil archaeal community should be important in determining the soil capacity of methane generation because the methanogens currently identified all belong to Archaea (Serrano-Silva et al. 2014). Ma and Lu (2010) also found archaeal community structure was an essential factor affecting methane emission in a rice paddy in Hangzhou City, China. Meanwhile, the samples (DC-4 and DC-5) with higher MPR and higher relative abundance of the archaeal phylum of Thaumarchaeota and Woesearchaeota, respectively (Fig. 4b). Presently, no methanogen has been found in these two archaeal phyla, and then, it is unsure whether these two archaeal phyla can directly participate in the process of soil methane synthesis. However, the result of RDA analysis indicated that DC-4 and DC-5 had a higher abundance of Methanolobus, which was the most closely correlated to soil MPR (Fig. 5b). Methanolobus is known as a methylotrophic methanogen that can only utilize methyl compounds to generate methane, which is the dominant methanogen in some oil or coal polluted habitats (Zhang et al. 2017; Bao et al. 2019; Okoro and Amund 2018). Therefore, our result reconfirmed that Methanolobus should be the critical methanogen directly responsible for methane production in those environments with the potential risk of PHC contamination.
Conclusions
In conclusion, in the soil from the shale gas field, OM content was significantly positively correlated with soil MPR. The sample with the highest MPR had the lowest prokaryotic diversity, and the samples with higher MPR had a higher ratio of archaeal sequences to the total prokaryotic sequences. Additionally, soil OM was a crucial parameter affecting soil prokaryotic community structure, and MPR was significantly correlated with both prokaryotic and archaeal community structure. The bacterial genus of Saccharibacteria and the archaeal genus of Methanolobus were the key microbes in determining soil MPR in the soil from the shale gas field. These results can be available for a deeper understanding of the mechanism of soil methane biological production in soil with the potential risk of PHC pollution and provide some useful microbial information for the resource-based treatment of the PHC-polluted soil.
References
Abbasian F, Lockington R, Megharaj M, Naidu R (2016) The biodiversity changes in the microbial population of soils contaminated with crude oil. Curr Microbiol 72:663–670
Albertsen M, Hugenholtz P, Skarshewski A, Nielsen KL, Tyson GW, Nielsen PH (2013) Genome sequences of rare, uncultured bacteria obtained by differential coverage binning of multiple metagenomes. Nat Biotechnol 31:533–538
Amato KR, Yeoman CJ, Kent A, Righini N, Carbonero F, Estrada A, Gaskins HR, Stumpf RM, Yildirim S, Torralba M, Gillis M, Wilson BA, Nelson KE, White BA, Leigh SR (2013) Habitat degradation impacts black howler monkey (Alouatta pigra) gastrointestinal microbiomes. ISME J 7:1344–1353
Andreolli M, Lampis S, Brignoli P, Vallini G (2016) Trichoderma longibrachiatum Evx1 is a fungal biocatalyst suitable for the remediation of soils contaminated with diesel fuel and polycyclic aromatic hydrocarbons. Environ Sci Pollut Res 23:9134–9143
Ariesyady HD, Ito T, Okabe S (2007) Functional bacterial and archaeal community structures of major trophic groups in a full-scale anaerobic sludge digester. Water Res 41:1554–1568
Bao Y, Ju YM, Huang HP, Yun JL, Guo C (2019) Potential and constraints of biogenic methane generation from coals and mudstones from Huaibei Coalfield, Eastern China. Energy Fuel 33:287–295
Bates ST, Berg-Lyons D, Caporaso JG, Walters WA, Knight R, Fierer N (2011) Examining the global distribution of dominant archaeal populations in soil. ISME J 5:908–917
Bayer C, Gomes J, Vieira FCB, Zanatta JA, Piccolo MD, Dieckow J (2012) Methane emission from soil under long-term no-till cropping systems. Soil Tillage Res 124:1–7
Bond PL, Hugenholtz P, Keller J, Blackall LL (1995) Bacterial community structures of phosphate-removing and non-phosphate-removing activated sludges from sequencing batch reactors. Appl Environ Microbiol 61:1910–1916
Borneman J, Triplett EW (1997) Molecular microbial diversity in soils from eastern Amazonia: evidence for unusual microorganisms and microbial population shifts associated with deforestation. Appl Environ Microbiol 63:2647–2653
Cao QM, Chen GZ, Liao SY (2009) Distribution correlations of polycyclic aromatic hydrocarbons with organic carbon and black carbon in surface sedimenta of three mangrove wetlands in the Shantou Wetland Demonstration Site, China. Acta Sci Circumst 29:861–868
Cebron A, Beguiristain T, Bongoua-Devisme J, Denonfoux J, Faure P, Lorgeoux C, Ouvrard S, Parisot N, Peyret P, Leyval C (2015) Impact of clay mineral, wood sawdust or root organic matter on the bacterial and fungal community structures in two aged PAH-contaminated soils. Environ Sci Pollut Res 22:13724–13738
Chang W, Um Y, Holoman TRP (2006) Polycyclic aromatic hydrocarbon (PAH) degradation coupled to methanogenesis. Biotechnol Lett 28:425–430
Chen SC, Peng JJ, Duan GL (2016) Enrichment of functional microbes and genes during pyrene degradation in two different soils. J Soils Sediments 16:417–426
Chen XP, Sun J, Wang Y, Zhang HY, He CQ, Liu XY, Bu NS, Long XE (2018) Temporal and spatial impact of Spartina alterniflora invasion on methanogens community in Chongming Island, China. J Microbiol 56:507–515
Cornelissen S, Koper M, Deng YY (2012) The role of bioenergy in a fully sustainable global energy system. Biomass Bioenergy 41:21–33
Cuer CA, Rodrigues RDR, Balieiro FC, Jesus J, Silva EP, Alves BJR, Rachid CTCC (2018) Short-term effect of Eucalyptus plantations on soil microbial communities and soil-atmosphere methane and nitrous oxide exchange. Sci Rep-UK 8:15133
Derosa M, Gambacorta A, Gliozzi A (1986) Structure, biosynthesis, and physicochemical properties of archaebacterial lipids. Microbiol Rev 50:70–80
Dunfield P, Knowles R, Dumont R, Moore TR (1993) Methane production and consumption in temperate and sub-arctic peat soils - response to temperature and pH. Soil Biol Biochem 25:321–326
Elarbaoui S, Richard M, Boufahja F, Mahmoudi E, Thomas-Guyon H (2015) Effect of crude oil exposure and dispersant application on meiofauna: an intertidal mesocosm experiment. Environ Sci Proc Imp 17:997–1004
Erguder TH, Boon N, Wittebolle L, Marzorati M, Verstraete W (2009) Environmental factors shaping the ecological niches of ammonia-oxidizing archaea. FEMS Microbiol Rev 33:855–869
Ficker M, Krastel K, Orlicky S, Edwards E (1999) Molecular characterization of a toluene-degrading methanogenic consortium. Appl Environ Microbiol 65:5576–5585
Gao PK, Tian HM, Wang YS, Li YS, Li Y, Xie JX, Zeng B, Zhou JF, Li GQ, Ma T (2016) Spatial isolation and environmental factors drive distinct bacterial and archaeal communities in different types of petroleum reservoirs in China. Sci Rep-UK 6:20174
Gao S, Liang JD, Teng TT, Zhang M (2019) Petroleum contamination evaluation and bacterial community distribution in a historic oilfield located in loess plateau in China. Appl Soil Ecol 136:30–42
Gebert J, Perner M (2015) Impact of preferential methane flow through soil on microbial community composition. Eur J Soil Biol 69:8–16
Han XM, Liu YR, Zheng YM, Zhang XX, He JZ (2014) Response of bacterial pdo1, nah, and C12O genes to aged soil PAH pollution in a coke factory area. Environ Sci Pollut Res 21:9754–9763
Haritash AK, Kaushik CP (2009) Biodegradation aspects of polycyclic aromatic hydrocarbons (PAHs): a review. J Hazard Mater 169:1–15
Hatzenpichler R, Lebedeva EV, Spieck E, Stoecker K, Richter A, Daims H, Wagner M (2008) A moderately thermophilic ammonia-oxidizing crenarchaeote from a hot spring. Proc Natl Acad Sci U S A 105:2134–2139
He JZ, Hu HW, Zhang LM (2012) Current insights into the autotrophic thanumarchaeal ammonia oxidation in acidic soils. Soil Biol Biochem 55:146–154
Huang JH, Shi YH, Zeng GM, Gu YL, Chen GQ, Shi LX, Hu Y, Tang S, Zhou JX (2016) Acyl-homoserine lactone-based quorum sensing and quorum quenching hold promise to determine the performance of biological wastewater treatments: an overview. Chemosphere 157:137–151
Hugenholtz P, Tyson GW, Webb RI, Wagner AM, Blackall LL (2001) Investigation of candidate division TM7, a recently recognized major lineage of the domain bacteria with no known pure-culture representatives. Appl Environ Microbiol 67:411–419
Kandler O, Konig H (1998) Cell wall polymers in Archaea (Archaebacteria). Cell Mol Life Sci 54:305–308
Khan MI, Cheema SA, Tang XJ, Hashmi MZ, Shen CF, Park J, Chen YX (2013) A battery of bioassays for the evaluation of phenanthrene biotoxicity in soil. Arch Environ Contam Toxicol 65:47–55
Kristanti A, Hadibarata T (2015) Biodegradation and identification of transformation products of fluorene by Ascomycete fungi. Water Air Soil Pollut 226:406
Kwietniewska E, Tys J (2014) Process characteristics, inhibition factors and methane yields of anaerobic digestion process, with particular focus on microalgal biomass fermentation. Renew Sust Energ Rev 34:491–500
Le Mer J, Roger P (2001) Production, oxidation, emission and consumption of methane by soils: a review. Eur J Soil Biol 37:25–50
Li XF, Hou LJ, Liu M, Zheng YL, Li Y, Lin XB (2015) Abundance and diversity of polycyclic aromatic hydrocarbon degradation bacteria in urban roadside soils in Shanghai. Appl Microbiol Biotechnol 99:3639–3649
Liao JQ, Wang J, Jiang DL, Wang MC, Huang Y (2015) Long-term oil contamination causes similar changes in microbial communities of two distinct soils. Appl Microbiol Biotechnol 99:10299–10310
Liu W, Lu HH, Wu W, Wei QK, Chen YX, Thies JE (2008) Transgenic Bt rice does not affect enzyme activities and microbial composition in the rhizosphere during crop development. Soil Biol Biochem 40:475–486
Liu QL, Tang JC, Bai ZH, Hecker M, Giesy JP (2015) Distribution of petroleum degrading genes and factor analysis of petroleum contaminated soil from the Dagang Oilfield, China. Sci Rep-UK 5:11068
Llado S, Gracia E, Solanas AM, Vinas M (2013) Fungal and bacterial microbial community assessment during bioremediation assays in an aged creosote-polluted soil. Soil Biol Biochem 67:114–123
Lu RK (1999) Methods of agrochemical soil analysis. China Agricultural Science Press, Beijing
Luo CL, Xie SG, Sun WM, Li XD, Cupples AM (2009) Identification of a novel toluene-degrading bacterium from the candidate phylum TM7, as determined by DNA stable isotope probing. Appl Environ Microbiol 75:4644–4647
Ma K, Lu YH (2010) Regulation of microbial methane production and oxidation by intermittent drainage in rice field soil. FEMS Microbiol Ecol 75:446–456
Marschner P, Kandeler E, Marschner B (2003) Structure and function of the soil microbial community in a long-term fertilizer experiment. Soil Biol Biochem 35:453–461
Morel X, Decharme B, Delire C, Krinner G, Lund M, Hansen BU, Mastepanov M (2019) A new process-based soil methane scheme: evaluation over Arctic field sixes with one ISBA land surface model. J Adv Model Earth Sy 11:293–326
Morin TH, Bohrer G, Frasson RPDM, Naor-Azreli L, Mesi S, Stefanik KC, Schafer KVR (2014) Environmental drivers of methane fluxes from an urban temperate wetland park. J Geophys Res-Biogeo 119:2188–2208
Nielsen PH, Kragelund C, Seviour RJ, Nielsen JL (2009) Identity and ecophysiology of filamentous bacteria in activated sludge. FEMS Microbiol Rev 33:969–998
Nielsen PH, Mielczarek AT, Kragelund C, Nielsen JL, Saunders AM, Kong YH, Hansen AA, Vollertsen J (2010) A conceptual ecosystem model of microbial communities in enhanced biological phosphorus removal plants. Water Res 44:5070–5088
Ogboghodo IA, Iruaga EK, Osemwota IO, Chokor JU (2004) An assessment of the effects of crude oil pollution on soil properties, germination and growth of maize (Zea mays) using two crude types-Forcados light and Escravos light. Environ Monit Assess 96:143–152
Okoro CC, Amund OO (2018) Microbial community structure of a low sulfate oil producing facility indicate dominance of oil degrading/nitrate reducing bacteria and methanogens. Pet Sci Technol 36:293–301
Pace NR (2009) Mapping the tree of life: progress and prospects. Microbiol Mol Biol R 73:565–576
Peng QA, Shaaban M, Hu RG, Mo YL, Wu YP, Ullah B (2015) Effects of soluble organic carbon addition on CH4 and CO2 emissions from paddy soils regulated by iron reduction processes. Soil Res 53:316–324
Perez-Leblic MI, Turmero A, Hernandez M, Hernandez AJ, Pastor J, Ball AS, Rodriguez J, Arias ME (2012) Influence of xenobiotic contaminants on landfill soil microbial activity and diversity. J Environ Manag 95:S285–S290
Reim A, Hernandez M, Klose M, Chidthaisong A, Yuttitham M, Conrad R (2017) Response of methanogenic microbial communities to desiccation stress in flooded and rain-fed paddy soil from Thailand. Front Microbiol 8:785
Ritchie GD, Still KR, Alexander WK, Nordholm AF, Wilson CL, Rossi J, Mattie DR (2001) A review of the neurotoxicity risk of selected hydrocarbon fuels. J Toxicol Environ Health B 4:223–312
Salanitro JP, Dorn PB, Huesemann MH, Moore KO, Rhodes IA, Jackson LMR, Vipond TE, Western MM, Wisniewski HL (1997) Crude oil hydrocarbon bioremediation and soil ecotoxicity assessment. Environ Sci Technol 31:1769–1776
Schauss K, Focks A, Leininger S, Kotzerke A, Heuer H, Thiele-Bruhn S (2009) Dynamics and functional relevance of ammonia-oxidizing archaea in two agricultural soils. Environ Microbiol 11:446–456
Schloss PD, Gevers D, Westcott SL (2011) Reducing the effects of PCR amplification and sequencing artifacts on 16S rRNA-based studies. PLoS One 6:e27310
Serrano-Silva N, Sarria-Guzman Y, Dendooven L, Luna-Guido M (2014) Methanogenesis and Methanotrophy in soil: a review. Pedosphere 24:291–307
Sun WM, Dong YR, Gao P, Fu MY, Ta KW, Li JW (2015) Microbial communities inhabiting oil-contaminated soils from two major oilfields in northern China: implications for active petroleum degrading capacity. J Microbiol 53:371–378
Trofimov SY, Rozanova MS (2003) Transformation of soil properties under the impact of oil pollution. Eurasian Soil Sci 36:S82–S87
Valentine DL (2007) Adaptations to energy stress dictate the ecology and evolution of the archaea. Nat Rev Microbiol 5:316–323
Van-der-Oost R, Beyer J, Vermeulen NPE (2003) Fish bioaccumulation and biomarkers in environmental risk assessment: a review. Environ Toxicol Pharmacol 13:57–149
Vergara-Fernandez A, Vargas G, Alarcon N, Velasco A (2008) Evaluation of marine algae as a source of biogas in a two-stage anaerobic reactor system. Biomass Bioenergy 32:338–344
Wagner R, Zona D, Oechel W, Lipson D (2017) Microbial community structure and soil pH correspond to methane production in Arctic Alaska soils. Environ Microbiol 19: 3398-3410
Wang J, Cao XF, Liao JQ, Huang Y, Tang XY (2015) Carcinogenic potential of PAHs in oil-contaminated soils from the main oil fields across China. Environ Sci Pollut Res 22:10902–10909
Xie SG, Sun WM, Luo CL, Cupples AM (2011) Novel aerobic benzene degrading microorganisms identified in three soils by stable isotope probing. Biodegradation 22:71–81
Xue W, Warshawsky D (2005) Metabolic activation of polycyclic and heterocyclic aromatic hydrocarbons and DNA damage: a review. Toxicol Appl Pharmacol 206:73–93
Yang JJ, Li GH, Qian Y, Zhang F (2018) Increased soil methane emissions and methanogenesis in oil contaminated areas. Land Degrad Dev 29:563–571
Ye RZ, Espe MB, Linquist B, Parikh SJ, Doane TA, Horwath WR (2016) A soil carbon proxy to predict CH4 and N2O emissions from rewetted agricultural peatlands. Agric Ecosyst Environ 220:64–75
Zhang GB, Ji Y, Ma J, Xu H, Cai ZC (2011) Case study on effects of water management and rice straw incorporation in rice fields on production, oxidation, and emission of methane during fallow and following rice seasons. Soil Res 49:238–246
Zhang YM, Yu ZS, Zhang HX, Thompson IP (2017) Microbial distribution and variation in produced water from separators to storage tanks of shale gas wells in Sichuan Basin, China. Environ Sci-Wat Res 3:340–351
Zhang WZ, Sheng R, Zhang MM, Xiong GY, Hou HJ, Li SL, Wei WX (2018) Effects of continuous manure application on methanogenic and methanotrophic communities and methane production potentials in rice paddy soil. Agric Ecosyst Environ 258:121–128
Zhou ZF, Wang MX, Zuo XH, Yao YH (2017) Comparative investigation of bacterial, fungal, and archaeal community structures in soils in a typical oilfield in Jianghan, China. Arch Environ Contam Toxicol 72:65–77
Funding
This work was supported by the Natural Science Foundation of China (41771501), the Basic and Advanced Technology Research Project of Chongqing (cstc2017jcyjAX0354), and the program of China Scholarship Council (201808505060).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Diane Purchase
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 20 kb)
Rights and permissions
About this article
Cite this article
Wang, YQ., Xiao, GQ., Cheng, YY. et al. The linkage between methane production activity and prokaryotic community structure in the soil within a shale gas field in China. Environ Sci Pollut Res 27, 7453–7462 (2020). https://doi.org/10.1007/s11356-019-07454-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-019-07454-3