Abstract
Globally, coastal waters have emerged into a pool of antibiotic resistance genes and multiple antibiotic resistant microorganisms, and pathogenicity of these resistant microorganisms in terms of serotypes and virulence genes has made the environment vulnerable. The current study underscores the presence of multiple antibiotic resistant pathogenic serotypes and pathotypes of Escherichia coli, the predominant faecal indicator bacteria (FIB), in surface water and sediment samples of famous recreational beaches (Juhu, Versova, Mahim, Dadar, and Girgaon) of Mumbai. Out of 65 faecal coliforms (FC) randomly selected, 38 isolates were biochemically characterized, serotyped (for ‘O’ antigen), antibiogram-phenotyped (for 22 antimicrobial agents), and genotyped by polymerase chain reaction (for virulence factors). These isolates belonged to 16 different serotypes (UT, O141, O2, O119, O120, O9, O35, O126, O91, O128, O87, O86, R, O101, O118, and O15) out of which UT (18.4%), O141 (15.7%), and O2 (13.1%) were predominant, indicating its remarkable diversity. Furthermore, the generated antibiogram profile revealed that 95% of these isolates were multiple antibiotic resistant. More than 60% of aminoglycoside-sensitive E. coli isolates exhibited resistance to penicillin, extended penicillin, quinolone, and cephalosporin classes of antibiotic while resistance to other antibiotics was comparatively less. Antibiotic resistance (AR) indexing indicated that these isolates may have rooted from a high-risk source of contamination. Preliminary findings revealed the presence of enterotoxin-encoding genes (stx1 and stx2 specific for enterohaemorrhagic E. coli and Shiga toxin-producing E. coli, heat-stable toxin enterotoxin specific for enterotoxigenic E. coli) in pathogenic serotypes. Thus, government authorities and environmental planners should create public awareness and adopt effective measures for coastal management to prevent serious health risks associated with these contaminated coastal waters.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Swimming at recreational water sites such as the beach is one of the most popular activities during holidays. However, population explosion and inadequate infrastructure to properly treat and dispose sewage, lack of sanitary condition, poverty, and overexploitation of natural waters has resulted in the discharge of considerable quantities of untreated waste into the natural waters (Chandran et al. 2008a). Pathogenic microorganisms and toxic compounds are added to the coastal waters by sewage contamination rendering these water bodies unsafe for bathing. Microbial source tracking studies on polluted environmental waters in Southeast Queensland, Australia using nifH gene marker of Methanobrevibacter smithii detected the presence of sewage contamination (Ahmed et al. 2011). Exposure to these pathogens can occur during swimming or other recreational activities through ingestion, inhalation, or direct skin contact with polluted beach water (Praveena et al. 2015). Global estimates indicate that each year more than 120 million cases of gastrointestinal disease and 50 million cases of severe respiratory disease and 50 million cases of severe respiratory diseases are caused by swimming and bathing in waste water-polluted coastal waters (Abdelzaher et al. 2013). Antibiotics belonging to classes fluoroquinolones, quinolones, and cephalosporins are considered as first-line agents to cure these diseases (Pitout and Laupland 2008; Pitout 2012; Shepherd and Pottinger 2013), but overgrowing resistance to these agents is causing delay in appropriate therapy with subsequent increased morbidity and mortality (Schwaber and Carmeli 2007; Tumbarello et al. 2007).
Antibiotic resistance has been recognized as a global threat to humans and veterinary medicine both in developed and developing countries (Chandran et al. 2008b), but nowhere is it as stark as in India (Ganguly et al. 2011). In 2010, India was the world’s largest consumer of antibiotics for human health (Laxminarayan and Chaudhury 2016) as there is no strict monitoring programme regarding the use of antibiotics in animals and humans; more drugs are available in private clinics and medical shops than in public hospitals, even without prescription. Antibiotic pollution in the form of overuse in humans, animals, and agriculture encourages the transfer of resistance genes to human commensal and pathogenic bacteria and release of antibiotic resistant strains in the environment (Rutgersson et al. 2014). Waste generated in the form of industrial effluents and domestic sewage after treatment is discharged into the coastal waters for dilution creating a reservoir for pathogenic bacteria exhibiting resistance to multiple antimicrobials. The severity of this problem increases multifold when these bacteria account for life-threatening diseases.
For microbiological quality assessment of surface waters, traditional as culture-based methods are employed. Culture-based methods are important in investigating the microbial ecology of natural and anthropogenically impacted environment. However, with recent advancement in genomics and sequencing technologies, there is an increasing realization that comprehensive understanding of microbial ecology requires a combination of culture-based and molecular-based assays, and this is especially true for antibiotic resistance in various ecosystems. Molecular methods are extremely useful for obtaining information on diversity and presence of antibiotic-resistant genes (Luby et al. 2016). Also, advances in next generation sequencing technologies have made it possible to develop genetic-based subtyping and molecular serotyping methods for pathogen, which are more discriminatory compared to phenotypic typing methods (Fratamico et al. 2016).
E. coli, the predominant faecal indicator bacteria, have been traditionally used as indicators of faecal contamination and health status of coastal waters. Emergence of E. coli isolates with multiple antibiotic resistant phenotypes, involving co-resistance to four or more unrelated families of antibiotics, is a serious matter of concern (Maynard et al. 2003) because in spite of being commensal, some strains have acquired specific virulence traits that allow them to cause a wide spectrum of intestinal and extraintestinal infections such as diarrhoea, urinary tract infections, and both community- and hospital-acquired bacteraemia (Salvadori et al. 2004). Pathogenic strains of E. coli in clinical specimens, foods, and environmental samples can be detected by serotyping wherein according to the modified Kauffman and White scheme, E. coli is serotyped on the basis of its O (somatic), H (flagellar), and K (capsular) surface antigen profiles (Nataro and Kaper 1998). Occurrence of pathogenic serotypes like O2, O15, O25, O86, O91, O141, O157, and untypeable (UT) belonging to enterotoxigenic E. coli (ETEC), enteropathogenic E. coli (EPEC), enterohaemorrhagic E. coli (EHEC), Shiga toxin-producing E. coli (STEC), enteroaggregative E. coli (EAEC), and uropathogenic E. coli (UPEC) pathotypes have been previously reported in the waters of Cochin Estuary (Sukumaran et al. 2012; Chandran et al. 2008b) and Bhavani River (Hatha et al. 2004). Similar studies on fresh waters of famous Indian rivers like Gomti, Sarayu, and Ganga revealed that they are contaminated with multiple antimicrobial resistant diarrheagenic strains of E. coli belonging to EHEC, STEC, and ETEC pathotypes demanding regular surveillance and formulation of effective strategies to protect public health (Ram and Shanker 2005; Ram et al. 2007, 2008). However, no such study has been carried out on recreational marine waters of the metropolitan city Mumbai. Thus, the present study was designed with a view to determine the health status of famous recreational beaches of Mumbai, India affected by anthropogenic pressure. These beaches are frequently visited by the people of Mumbai as well as tourists. Hence, in this study, microbiological water quality assessment of surface water and sediment of beaches was performed and predominant faecal pollution indicator bacteria E. coli isolated were serotyped and screened for resistance to antimicrobial agents commonly used to treat infections caused by them, thereby, helping in knowing the effective treatment/medication in case of disease outbreak. Combining the data of water quality assessment, serotyping and antibiogram profile of faecal indicator bacteria (FIB) will provide a basis for robust and graded classification of beaches. The results of classification can be used to grade beaches in order to support informed personal choice, provide onsite guidance to users on relative safety, assist government authorities in the identification and promotion of effective management intervention, and provide an assessment of regulatory compliance.
Materials and methods
Description of sampling sites
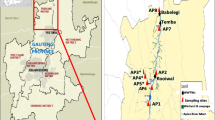
Mumbai (18°55′ N, 72°54′ E) is the most populous metropolitan city on the west coast of India and capital of the state Maharashtra. The state of Maharashtra accounts for a 653-km-long coastline with 17% sandy beaches, many of them lying within the city of Mumbai. The increase in urbanization and industrialization has led to an increase in marine discharges. The city generates 2.2 × 106 m3 day−1 of domestic sewage out of which about 2.0 × 106 m3 day−1 (partially treated or largely untreated) enters the marine waters including creeks and bays and tides, and the current brings these pollutants to the beaches, exacerbating the problem (Jayasiri et al. 2014). Also, being a metropolitan city, major industries of the country are located in and around Mumbai. Largely untreated industrial effluents and domestic sewage from the city enter the continental shelf in the region. There are a number of ports wherein the ship and cargo boarding activities contribute to marine pollution. This has resulted in the degradation of the coastal water quality and contamination of adjoining beaches and sea fronts. As a coastal city, beaches for recreation such as Girgaon, Mahim, Juhu, Aksa, Marine Drive, and Versova are few of the major attractions for tourists. Out of the nine beaches located along the city coast, five beaches namely Versova, Juhu, Mahim Bay, Dadar, and Girgaon which show wide geographic coverage and degree of beach usage for recreation and tourism activities were selected for this study. Details of sampling sites are shown in Table 1 and Fig. 1.
Collection and transportation of samples
For microbiological analyses, water and sediment samples were collected for a period of 4 months (April 2015 to July 2015) from five recreational beaches of Mumbai in presterilized polypropylene bottles (Tarsons, India) and in sterile 50-ml Falcon tubes (Tarsons, India), respectively, during high tide and low tide in triplicates. All samples were collected by taking precautions required for microbiological analysis, held in an icebox, and processed within 6–8 h of collection.
Assessment of faecal indicator bacteria
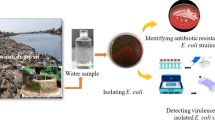
The total viable bacterial counts (TVC) from the surface water samples were estimated via the spread plating method on marine agar plates with 0.1 mL of suitable dilutions. The enumeration of total coliforms (TC) and faecal coliforms (FC) was carried out using membrane filtration according to the standard method (APHA 1998). In brief, water samples were filtered through 0.45-μm pore-sized cellulose acetate filters (Millipore) and aseptically placed on selective media plates. All of the media plates, except M-FC agar plates, were incubated at 37 °C ± 1 °C for 24–48 h and final counts of colonies were noted. M-FC agar plates were incubated at 44.5 °C ± 1 °C for 24–48 h. All trials were performed in triplicates. Typical colony morphology characteristics of different bacterial groups were noted and an initial enumeration of faecal pollution indicator bacteria was completed according to the media manufacturer’s guide. A total of 80 presumptive E. coli colonies (60 from water samples and 20 from sediment samples) showing various shades of blue on the M-FC medium were confirmed by streaking on Levine-eosin methylene blue agar (EMB Agar) and 65 isolates were identified as E. coli after streaking them on chromogenic agar (Hicrome E. coli agar) and biochemical characterization using Hi E. coli Identification Kit (Hi-Media, Mumbai, India). Purified colonies of E. coli were preserved as their 30% glycerol stock for further studies. Out of 65, 38 confirmed E. coli cultures were randomly selected and serotyped at the National Salmonella and Escherichia Centre, Kasauli, Himachal Pradesh, India. Typical colony characteristics of each bacterial group and specific media used for enumerating them are listed in Table 2.
Antibiotic resistance analysis
Purified colonies of E. coli isolated from both water and sediment samples from all the five beaches were tested for antimicrobial susceptibility. Antibiotic susceptibility testing was performed using a disk diffusion method as mentioned by Bauer et al. (1966). The isolates were challenged with 22 different antibiotics on Mueller Hinton Agar (Himedia, India), and results were interpreted based on recommendations of CLSI (Clinical and Laboratory Standards Institute, 2007). A sterile cotton swab was used to inoculate the standardized bacterial suspension matching an optical density of 0.5 McFarland standards corresponding to 108 CFU mL−1 on the surface of previously prepared sterile Mueller Hinton agar plates by rotating the plate every 60° to ensure homogenous growth, followed by addition of antibiotic-impregnated rings (Hi-Media, Mumbai, India). After 30 min, the plates were inverted and incubated at 37 °C for 18–24 h. This test was performed in duplicate for each E. coli strain and antimicrobial. In each experimental set, E. coli ATCC 25922 was used as negative control. The quality control was performed according to manufacturer’s instructions. The list of antibiotics tested and data for antimicrobial resistance of each bacterial isolate have been reported as resistant (R), intermediates (I), and sensitive (S) based on CLSI break points as shown in Table 3. The results were used to calculate the antibiotic resistance index (ARI) and multiple Antibiotic Resistance Index (MAR) for total number of isolates as:
Where, y is the number of resistant isolates, n is the number of isolates, and x is the number of antibiotics (Hinton and Linton 1983);
Where a is the number of antibiotics to which the isolates are resistant, and b is the total number of antibiotics exposed.
Statistical analysis
All statistical analyses were performed using XL-STAT (version 6.0, Addinsoft) and a p value of 0.05 was considered to be significant. Fisher’s exact test was performed to analyse associations between responses of E. coli isolates for two antimicrobials to assess the co-selection (Seigal and Catellan 1987). The significance of variation in occurrence of multiple antimicrobial resistance between the sites were also analysed using the Chi-square test (Seigal and Catellan 1987). In order to classify these occurrences in different serotypes, an agglomerative hierarchical clustering (AHC) analysis was employed, using Euclidean distance and the Ward method for the aggregation criterion.
Results
Quantitative enumeration of faecal indicator bacteria density
All the selected sampling sites exhibited bacterial contamination. Overall range (expressed in CFU 100 mL for water and CFU g−1 for sediment) of bacterial counts in dry as well as wet season is given in Table 4. In brief, the faecal coliform levels were higher in the wet season as compared to those in the dry season. In the dry season, the minimum levels of faecal coliforms were reported in water (35 × 102 CFU per 100 mL) and sediment (52 × 102 CFU g−1) of Juhu beach; whereas, maximum levels were reported in water (730 × 102 CFU per mL) and sediment (1150 × 102 CFU g−1) of Mahim beach. In wet season, the minimum levels of faecal coliforms were reported in water (20 × 102 CFU per 100 mL) and sediment (100 × 102 CFU g−1) of Girgaon beach while maximum levels were reported in the water (850 × 102 CFU mL−1) of Mahim beach and sediment (1000 × 102 CFU g−1) of Mahim and Versova beach as illustrated in Table 4.
Susceptibility to antimicrobial agents and serotyping
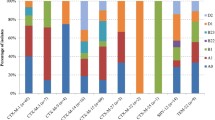
All 38 isolates confirmed as E. coli by biochemical testing were tested for susceptibility to 22 antimicrobials and serotyped. It was observed that E. coli isolated from the five sites along the Mumbai coast possess different levels of susceptibility towards antimicrobials tested. All the isolates from the study area were sensitive to amikacin, gentamicin, levofloxacin, rifampicin, and streptomycin; while, on the contrary, all of the isolates were resistant to Augmentin. Further, 97, 73.6, 68.4, 65.7, and 55.2% of the isolates were resistant to nalidixic acid, cephalothin, ampicillin/sulbactam, cefpodoxime, ampicillin, and cefepime, respectively (Table 5). Figure 2a shows the percentage (%) of E. coli isolates (%) having resistance to multiple antibiotics. One hundred per cent of the isolates were resistant to cephalothin, ampicillin/sulbactam, ampicillin, nalidixic acid, and Augmentin from site 1 (Juhu); nalidixic acid and Augmentin from site 2 (Versova) and site 4 (Dadar); Augmentin from site 3 (Mahim); and ampicillin, cefixime, nalidixic acid, and Augmentin from site 5 (Girgaon); while more than 50% of the isolates were resistant to cefazolin, cefoxitin, cefuroxime, ciprofloxacin, cefepime, and cefpodoxime from site 1 (Juhu); cephalothin, cefuroxime, cefotaxime, ampicillin, ampicillin/sulbactam, and cefpodoxime from site 2 (Versova); cefpodoxime, cefixime, nalidixic acid, ampicillin, and ampicillin/sulbactam from site 3 (Mahim); cefpodoxime, cephalothin, ciprofloxacin, ampicillin, ampicillin/sulbactam, and cefoxitin from site 4 (Dadar); and cefpodoxime, cephalothin, ampicillin/sulbactam, and ceftriazone from site 5 (Girgaon). Figure 2b shows the percentage (%) of E. coli isolates having reduced susceptibility to multiple antibiotics. Thirty-three per cent of the isolates from site 1 showed reduced susceptibility to cefazolin and tetracycline; 22 and 11% the isolates from site 2 showed reduced susceptibility to (cefazolin, cefoxitin, cefixime) and (cefoperazone, tetracycline, cefpodoxime), respectively; 30, 20 and 10% of the isolates from site 3 showed reduced susceptibility to (cephalothin, cefpodoxime), cefixime, (cefuroxime, tetracycline), respectively; 18% of the isolates from site 4 showed reduced susceptibility to cefazolin, cefotaxime, tetracycline, and cefixime while 9% showed reduced susceptibility to ampicillin/sulbactam, cefoxitin, cefoperazone, ampicillin, and chloramphenicol; and 20% of the isolates from site 5 showed reduced susceptibility to cephalothin, cefoxitin, cefuroxime, and cefoperazone. The major issue of concern here was that the isolates showing resistance to more than seven antibiotics were isolated from all the five sites with a MAR index of above 0.3. Figure 2c gives the number of isolates showing resistance to two or more antibiotics and overall MAR index and with respect to stations. AR index of all sampling sites exceeded the high risk level (0.25) and revealed that Versova has the highest AR index (0.404) followed by Juhu (0.378), Dadar (0.376), Girgaon (0.372), and Mahim (0.309); whereas, percentage of isolates resistant to eight or more antibiotics was found to be the highest in Dadar (73%) followed by Juhu (67%), Versova (56%), Mahim, and Girgaon (40%).
Susceptibility of E. coli isolates towards antimicrobial agents at different recreational beaches of Mumbai. a E. coli isolates (%) having resistance to multiple antibiotics. b E. coli isolates (%) having reduced susceptibility to multiple antibiotics. c Number of isolates showing resistance to two or more antibiotics and overall MAR index with respect to stations. (Note: Legends are common for a, b, and c)
Serotyping results indicated remarkable diversity in E. coli isolates in the way that 38 isolates exhibited 16 different serological identities predominated by UT (18.4%), O141(15.7%), O2(13.1%), O119, and O120 (7.8%) as depicted in Fig. 3. Furthermore, it was observed that dry season (April and May) encountered the highest number of E. coli (26.3%) belonging to O141 serotype followed by O35 (10.5%) while from the wet season (June and July), 31.5% of the E. coli isolates were untypeable and 26.3% of the isolates belonged to O2. MAR index and resistance pattern of 38 isolates (comprising of 16 serotypes) is represented in Table 6. Interestingly, all the pathogenic serotypes belonging to probable pathotypes (EHEC, ETEC, EPEC, STEC, EAEC, and UPEC) were multiple antibiotic resistant, and the number of antibiotics to which these isolates were resistant ranged from 2 to 14. The most common pattern of resistance observed among all serotypes was cefazolin, cephalothin, ampicillin/sulbactam, cefuroxime, ampicillin, cefixime, nalidixic acid, Augmentin, and cefpodoxime (Table 6).
Statistical analysis
The resistance to nalidixic acid was significantly associated with Augmentin (Fisher’s exact test significant at 5% level). Similarly, resistance to cephalothin, cefixime, and cefpodoxime (belonging to the cephalosporin class of antibiotic) was observed to be significantly associated with resistance to ampicillin and ampicillin/sulbactam (belonging to the penicillin and extended penicillin class of antibiotic) (Fisher’s exact test significant at 5% level). The distribution of multiantimicrobial resistance in E. coli varied significantly (X 2 p˂0.001) between the sites. However, E. coli isolated from site 1 (Juhu) were resistant to seven or more antimicrobials followed by site 2 (Versova), site 4 (Dadar), and site 5 (Girgaon) exhibiting resistance to 5–14 antimicrobials while E. coli isolates from site 3 exhibited resistance to 5–11 antimicrobials. Surprisingly, it was noted that not a single isolate was sensitive to all the antimicrobials tested.
Antibiotic clustering
Serotypes were clustered into groups according to similarity in response against different classes of antibiotics tested as shown in Fig. 4. Solid lines in the figure indicates significant dissimilarity between groups of serotypes and their response against class of antibiotics and solid lines with an asterisk at the node indicate significant similarity between two individual serotypes and their response against the class of antibiotics. Sixteen serotypes were grouped into three major clusters (cluster A, cluster B, and cluster C) which were further divided to form subclusters. The cluster A comprised of isolates belonging to serotype O2, O128, O7, O101, and UT indicating that these serotypes showed a similar resistance pattern against the class of antibiotics. Similarly, O119, O118, O120, O9, R, O86, O35, and O15 were grouped together in cluster B, and O141, O126, and O91 were grouped together in cluster C. Cluster A, cluster B and cluster C are linked together at 18 ED which is higher than the cutoff point (16 ED) thus indicating significant dissimilarity among serotypes and their responses towards a class of antibiotics. Serotypes belonging to clusters formed below the cutoff point have a significant similarity in their response against a class of antibiotics.
Discussion
Microbial pollution from sewage is becoming an increasing threat to recreational water users and coastal ecosystem health as the human population expands along global coastlines. Thus, for public health reasons, beach water assessment is of prime importance. The faecal indicator bacterial counts observed in this study are much higher than the permissible limits and support the recent findings of CSIR-NEERI, Nagpur, India (National Environmental Engineering Research Institute 2016) which states that faecal contamination at discharge points is very high. Although the sewage does get diluted when it mixes with the sea and the creek water, the FC count is still 100 to 1000 times higher than permissible limits at all the city beaches. However, this study gives the broader picture of pathogenic faecal coliforms prevailing in the coastal environment. Though a correlation of the infections due to contact polluted waters is not established, the possible associated risks have provided the basis for delineating standards for bathing and recreation waters. Hence, the Central Pollution Control Board (CPCB) specified Sea Water standards (Criteria for classification and zoning of coastal waters (sea waters)—A coastal pollution control series: COPOCS/6/ 1993) for designated best use like SWII for commercial fishing, contact recreation, and bathing activities; SWIII for industrial cooling; and SWIV for harbour water. The present study investigated the water quality at the beaches of Mumbai following the standards laid by CPCB in terms of faecal coliforms and E. coli levels indicating the poor health status of beaches. Higher and unacceptable counts of faecal coliforms in coastal waters may be attributed to the treated/untreated sewages from marine outfalls, nallahs, slum discharges, and effluents from dairy and other industries. Water quality assessment studies conducted previously in the same regions denoted the same FIB pollution levels throughout the year irrespective of tide and do not satisfy the SWII or SWIII standards of FC 100FC/100 and 500FC/100 mL, respectively. Though the Municipal Corporation of Greater Mumbai (MCGM) has diverted waste water from Love Grove Pumping Station (LGPS) through a 3.4-km-long outfall and undertaken cleanliness drive for beaches, 196 MLD of waste water is discharged in near-shore region through non-point sources; slum sanitation programmes were inadequate and therefore no improvement in microbiological water quality was witnessed (Dhage et al. 2006).
E. coli were isolated consistently from all the sites, with differences in their abundance level. Molecular-based assays and a number of other investigations have shown that pathogenic FIBs are prevalent in marine waters being impacted by sewage. These faecal-derived pathogens can cause a broad range of asymptomatic to severe gastrointestinal, respiratory, eye, nose, ear, and skin infections in people exposed through recreational use of water. Moreover, their multiple drug resistance nature is worrisome as the circulation of these microorganisms in the environment not only affects the ability to treat the infection but also the cost and duration. Present investigation revealed the presence of 95% of the isolated FIBs as MAR. Likewise, similar studies have been reported by various researchers from different types of surface waters: rivers (Watkinson et al. 2007), estuaries (Sukumaran et al. 2012), lakes (Edge and Hill 2005), and coastal waters (Maloo et al. 2014). The ability of E. coli and Enterococci to acquire antibiotic resistance by horizontal gene transfer has exacerbated the problem of antibiotic resistance in bacteria (Huddleston 2014). Active population of beach-adapted faecal bacteria may play an intermediary role in mobilizing resistance genes between environmental bacteria and pathogens (Martinez 2009). Although mutations contribute to antibiotic resistance, most pathogens acquire resistance genes through horizontal transfer of mobile genetic elements such as plasmids and transposons. Worldwide, Augmentin is commonly used to treat urinary tract infection caused by uropathogenic E. coli (Rahnama et al. 2009), but surprisingly, 100% of the isolates from this study were found to be resistant against it which can be a cause for concern. Mumbai waters also encountered the presence of pathogenic serotype O2 which is a known cause of urinary tract infections and its resistance against the widely used drug Augmentin can be fatal. According to researchers, Augmentin-resistant E. coli is usually treated with a combination of gentamicin and ceftriaxone, but 29% of the isolates from this study were resistant to ceftriaxone. In urinary tract infections, previous treatment with Augmentin is a risk factor for the development of Augmentin resistance, and its resistance can be attributed to mechanisms like β-lactamase overproduction, AmpC, cephalosporinase hyperproduction, and inhibitor-resistant penicillinases (Oteo et al. 2008). Resistance to ciprofloxacins as well as broad-spectrum first-generation quinolone antibiotics is reported to be slowly emerging in Asian, South American, and African countries (Sreela et al. 2011). Almost 97% of the isolates studied here expressed resistance to nalidixic acid, the first-generation quinolone. Antibiotics from this class inhibit the activity of bacterial DNA gyrase and DNA topoisomerase enzymes, which are essential for replication. Single nucleotide polymorphisms (SNPs) in the quinolone resistance-determining regions (QRDR) of gyrA and parC, the two genes that encode DNA gyrase and topoisomerase IV, respectively, can lead to conformational changes in these enzymes that cause them to block quinolones from binding to the DNA-substrate complex, yet still preserve their enzymatic function (Sreela et al. 2011). Resistance to penicillin and extended penicillin class of antibiotics was common among beach-isolated E. coli and can be attributed to production of β-lactamase by these organisms. Around 65–68% of the isolates from this study were found to be resistant to ampicillin and ampicillin/sulbactam which may be due to β-lactamase-producing E. coli. There are over 200 β-lactamase genes, some of which are carried on transmissible plasmids that can result in dissemination of resistance genes to pathogens (Brinas et al. 2002). High levels of ampicillin resistance in E. coli isolated from water sources can be attributed to sewage contamination (Amaya et al. 2012, Kappell et al. 2015).
Third-generation cephalosporins are broad-spectrum drugs with high intrinsic activity against Gram-negative species. Surprisingly, in the present study, it was observed that 58 and 13% of the isolates were resistant and showed reduced susceptibility to cephalosporins, respectively. The rising resistance to these drugs is worrisome because it could be a proxy for the emergence and spread of Enterobacteriaceae strains producing extended-spectrum β-lactamase (ESBL). Studies on antibiotics resistance revealed that F porin mutation in E. coli may confer resistance to the newer cephalosporins such as cefmenoxime, cefepime, ceftazidime, and cefuroxime (Moosdeen 1997).
Some of this growing resistance in E. coli and other bacteria are due to the fact that antibiotics are being overprescribed, handed out to patients who have no bacterial infections. There is also evidence that the genes that give bacteria resistance to drugs are being spread in livestock-farming operations, where antibiotics are a common ingredient in animal feed. Ciprofloxacin is one of those antibiotics and researchers have found that E. coli resistant to it are thriving in poultry farms. Overuse or misuse of antibiotics in animals for non-therapeutic use such as prophylaxis, growth promotion, or increase of feed efficiency are leading to the rise or emergence of antibiotic resistance (Paulson et al. 2015). Therefore, it is vital to limit therapeutic antibiotic use in animals and non-therapeutic use should be reduced. Spread of antibiotic resistance is amplified when this bacteria is acquired by the person recreating at beaches and when this bacteria enters his intestinal tract and passes on the resistance to other flora of the system. According to Krumperman (1983), the choice of MAR index of 0.2 to differentiate between low- and high-risk contamination is arbitrary. Indices between 0.2 and 0.25 are in a range of ambiguity, and samples in this range require careful scrutiny. Overall, indices at all stations exceeded the arbitrary level which revealed that all stations were highly polluted with faecal bacteria originating from a high-risk source. Also, MAR indexing of individual isolates ranged from 0.09 to 0.64, which is much higher than 0.25 (Table 6), indicating the high risk source of contamination.
Perelle et al. (2007) reported that food contaminated by pathogenic E. coli serotypes, including O103, represents a major public health concern. Isolation of a great number of isolates with serotypes O2, O91, O86, and O15 belonging to UPEC, EPEC, EHEC, and ETEC categories of pathotypes was a matter of concern. Literature review showed that most of the serotypes encountered in the current study are potential pathogens as they are the cause of various diseases associated with humans and animals such as birds mentioned in Table 7. It is well documented that hospital wastes are often contaminated by antimicrobial agents which even the subinhibitory concentration may promote selection and survival of resistant strains (Al-Ahmad et al. 1999; Kim et al. 2007). Occurrence of E. coli in coastal waters belonging to different serotypes can be attributed to hospital wastes released through sewage (Chandran et al. 2008b). Sukumaran et al. (2012) reported the presence of diverse serotypes in Cochin waters, and a similar observation was made in the current study wherein more diverse serotypes were isolated from site 3 (Mahim) which suggests the possible release of these organisms through waste derived from hospitals located in the vicinity. Infections caused by such serotypes may pose a significant public health risk in our country. Furthermore, serologic antigens are not directly involved in virulence but can provide crucial information about the prevailing serotypes on the environment and those involved in disease outbreak (Nataro and Kaper 1998). Pathogenecity of these serotypes also relies on the determination of virulence genes (VGs) present in their genome; however, the presence of single or multiple VGs does not necessarily indicate that the strain is pathogenic unless it expresses the appropriate combination of VGs to cause disease in the host (Sidhu et al. 2013). Thus, our future studies intend to focus on determination of virulent genes specific for ETEC, EPEC, EHEC, and STEC. Preliminary studies on this objective indicated the presence of shigatoxin genes (stx1 and stx2) specific for EHEC and STEC and heat-stable toxin gene specific for ETEC in a few of the pathogenic serotypes (data not presented here).
Emphasis should be placed on environment and coastal management in India. This will thereby provide many general opportunities to improve beach environmental management and to achieve successful management outcomes; broadening our perspective of health of beaches is required. Collaborations between researchers and environmental managers will provide correct guidance and direction for planning of management strategies.
Conclusion
Coastal waters of Mumbai have emerged into a dumping ground for the waste generated in the city. Deteriorated microbiological quality of recreational waters indicated the poor health status of the beaches monitored in this study. Elevated levels of E. coli in surface water and sediment of beaches represent a public health risk to those who frequently get exposed to contaminated waters. Moreover, the presence of E. coli with defined multiple antibiotic resistance and pathogenic serotype is a matter of concern. Further analysis of virulence factors present in marine FIBs using molecular approach will be the most promising. This highlights the need for government authorities to take effective measures (e.g. proper sanitation, disposal facility, public awareness, and waste water treatment) to reduce pollution of beaches to avoid serious consequences of a health care management within localities and visitors. Also, data presented here may be useful for decision makers and resource managers working with environmental planning and management of coastal area to implement proper treatment procedure for waste water and effluents, before discharge into the sea to avoid various human diseases outbreaks.
References
Abdelzaher AM, Solo-Gabriele HM, Phillips MC, Elmir SM, Fleming LE (2013) an alternative approach to water regulations for public health protection at bathing beaches. J Environ Pub Health Article ID 138521, 9 pages.
Ahmed W, Sidhu JP, Toze S (2011) Evaluation of the nifH gene marker of Methanobrevibacter smithii for the detection of sewage pollution in environmental waters in Southeast Queensland, Australia. Environ Sci Technol 46(1):543–550
Al-Ahmad A, Daschner F, Kümmerer K (1999) Biodegradability of cefotiam, ciprofloxacin, meropenem, penicillin G and sulfamethoxazole and inhibition of waste water bacteria. Arch Environ Con Tox 37:158–163
Amaya E, Reyes D, Paniagua M, Calderón S, Rashid MU, Colque P, Nord CE (2012) Antibiotic resistance patterns of Escherichia coli isolates from different aquatic environmental sources in León, Nicaragua. Clin Microbiol Infect 18(9):E347–E354
APHA (1998) Standard methods for the examination of water and wastewater, 20th ed. American Public Health Association, Washington, DC
Bauer AW, Kirby WMM, Sherris JC, Turck M (1966) Antibiotic susceptibility testing by a standardized single disk method. Am J Clin Pathol 45:493–496
Benenson AS (ed) (1995) Control of communicable diseases manual, sixteenth edn. United Book Press, Baltimore, pp 140–150
Boerlin P, McEwen SA, Boerlin FP, Wilson JB, Johnson RP, Gyles CL (1999) Associations between virulence factors of Shiga toxin producing Escherichia coli and disease in humans. J Clin Microbiol 37:497–503
Brinas L, Zarazaga M, Saenz Y, Ruiz-Larrea F, Torres C (2002) β-lactamases in ampicillin-resistant Escherichia coli isolates from foods, humans, and healthy animals. Antimicrob Agents Ch 46:3156–3163
Chandran A, Hatha AM, Varghese S (2008a) Increased prevalence of indicator and pathogenic bacteria in Vembanadu Lake: a function of salt water regulator, along south west coast of India. J Water Health 6(4):539–546
Chandran A, Hatha AM, Varghese S, Sheeja KM (2008b) Prevalence of multiple drug resistant Escherichia coli serotypes in a tropical estuary, India. Microbes Environ 23:153–158
Clinical and Laboratory Standards Institute (2007) Methods for antimicrobial susceptibility testing of anaerobic bacteria; approved Standard, 7th edn. CLSI document M11-A7.Wayne, PA: Clinical and Laboratory Standards Institute.
Criteria for classification and zoning of coastal waters (sea waters) (1993)—A coastal pollution control series: COPOCS/6/1993—CPCB, New Delhi.
CSIR-NEERI National Environmental Engineering Research Institute Report, 2016.
Dhage SS, Chandorkar AA, Kumar R, Srivastava A, Gupta I (2006) Marine water quality assessment at Mumbai West Coast. Environ Int 32:149–158
Dutta P, Borah MK, Sarmah R, Gangil R (2013) Isolation, histopathology and antibiogram of Escherichia coli from pigeons (Columba livia). Vet World 6:91–94
Edge TA, Hill S (2005) Occurrence of antibiotic resistance in Escherichia coli from surface waters and fecal pollution sources near Hamilton, Ontario. Can J Microbiol 51:501–505
Franke S, Harmsen D, Caprioli A, Pierard D, Wieler LH, Karch H (1995) Clonal relatedness of Shiga-like toxin-producing Escherichia coli O101 strains of human and porcine origin. J Clin Microbiol 33:3174–3178
Fratamico PM, DebRoy C, Liu Y, Needleman DS, Baranzoni GM, Feng P (2016) Advances in molecular serotyping and subtyping of Escherichia coli. Front Microbiol 7:644
Ganguly NK, Arora NK, Chandy SJ, Fairoze MN, Gill JP, Gupta U et al (2011) Rationalizing antibiotic use to limit antibiotic resistance in India. Indian J Med Res 134:281–294
Hatha AM, Chandran A, Rahiman KM (2004) Prevalence of diarrheagenic serotypes of Escherichia coli and Salmonella in the Cochin estuary. Indian J Mar Sci 33:72–77
Hinton M, Linton A (1983) Antibacterial drug resistance among Escherichia coli isolated from calves fed milk substitute. Vet Rec 112:567–568
Huddleston JR (2014) Horizontal gene transfer in the human gastrointestinal tract: potential spread of antibiotic resistance genes. Infect Drug Resist 7:167–176
Jayasiri HB, Vennila A, Purushothaman CS (2014) Spatial and temporal variability of metals in inter-tidal beach sediment of Mumbai, India. Environ Monit Assess 186:1101–1111
Kappell AD, DeNies MS, Ahuja NH, Ledeboer NA, Newton RJ, Hristova KR (2015) Detection of multi-drug resistant Escherichia coli in the urban waterways of Milwaukee, WI. Front Microbiol 6:336
Kim S, Jensen JN, Aga DS, Weber AS (2007) Tetracycline as a selector for resistant bacteria in activated sludge. Chemosphere 66:1643–1651
Krumperman PH (1983) Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of fecal contamination of foods. Appl Environ Microbiol 46:165–170
Laxminarayan R, Chaudhury RR (2016) Antibiotic resistance in India: drivers and opportunities for action. PLoS Med 13:1–7
Luby E, Ibekwe AM, Zilles J, Pruden A (2016) Molecular methods for assessment of antibiotic resistance in agricultural ecosystems: prospects and challenges. J Environ Qual 45(2):441–453
Maloo A, Borade S, Dhawde R, Gajbhiye SN, Dastager SG (2014) Occurrence and distribution of multiple antibiotic-resistant bacteria of Enterobacteriaceae family in waters of Veraval coast, India. Environ Exp Biol 12:43–50
Martinez JL (2009) The role of natural environments in the evolution of resistance traits in pathogenic bacteria. P Roy Soc B-Biol Sci 276:2521–2530
Maynard C, Fairbrother JM, Bekal S et al (2003) Antimicrobial resistance genes in enterotoxigenic Escherichia coli O149:K91 isolates obtained over a 23-year period from pigs. Antimicrob Agents Ch 47:3214–3221
Moosdeen F (1997) The evolution of resistance to cephalosporins. Clin Infect Dis 24:487–493
Nataro JP, Kaper JB (1998) Diarrheagenic Escherichia coli. Clin Microbiol Rev 11:142–201
Oteo J, Campos J, Lázaro E, Cuevas Ó, García-Cobos S, Pérez-Vázquez M (2008) Increased amoxicillin–clavulanic acid resistance in Escherichia coli blood isolates, Spain. Emerg Infect Diseases 14(8):1259–1262
Paulson JA, Zaoutis TE, Committee on Infectious Diseases (2015) Nontherapeutic use of antimicrobial agents in animal agriculture: implications for pediatrics. Pediatrics 136(6):E1670–E1677
Perelle S, Dilasser F, Grout J, Fach P (2007) Screening food raw materials for the presence of the world’s most frequent clinical cases of Shiga toxin-encoding Escherichia coli O26, O103, O111, O145 and O157. Int J Food Microbiol 113:284–288
Pitout JD (2012) Extra intestinal pathogenic Escherichia coli: an update on antimicrobial resistance, laboratory diagnosis and treatment. Expert Rev Anti Infect 10:1165–1176
Pitout JD, Laupland KB (2008) Extended-spectrum beta-lactamase-producing Enterobacteriaceae: an emerging public-health concern. Lancet Infect Dis 8:159–166
Praveena SM, Chen KS, Sharifah NSI (2015) New methods to assess fecal contamination in beach water quality. In: Finkl CW, Makowski C (eds) Environmental management and governance: advances in coastal and marine resources, coastal research library. Springer International Publishing, pp 65–81
Rahnama MS, Wagenvoort JH, van der Linden CJ (2009) Amoxicillin/clavulanate (Augmentin) resistant Escherichia coli in bacterial peritonitis after abdominal surgery—clinical outcome in ICU patients. Neth J Med 67:173–176
Ram S, Shanker R (2005) Plasmid and drug resistance of sorbitol non-fermenting cefixime-tellurite resistant Escherichia coli isolates from Gomti River. B Environ Contam Tox 75:623–628
Ram S, Vajpayee P, Shanker R (2007) Prevalence of multi antimicrobial agent resistant, shigatoxin and enterotoxin producing Escherichia coli in surface waters of river Ganga. Environ Sci Technol 41:7383–7388
Ram S, Vajpayee P, Tripathi U, Singh RL, Seth PK, Shanker R (2008) Determination of antimicrobial resistance and virulence gene signatures in surface water isolates of Escherichia coli. J Appl Microbiol 105:1899–1908
Rutgersson C, Fick J, Marathe N, Kristiansson E, Janzon A, Angelin M et al (2014) Fluoroquinolones and qnr genes in sediment, water, soil, and human fecal flora in an environment polluted by manufacturing discharges. Environ Sci Technol 48:7825–7832
Salvadori M, Coleman BL, Louie M, McEwen S, McGeer A (2004) Consumption of antimicrobial-resistant Escherichia coli contaminated well water: Human health Impact. PSI Clinical research 6–25
Schmidt H, Scheef J, Stefano Morabito S, Caprioli A et al (2000) A new Shiga toxin 2 variant (Stx2f) from Escherichia coli isolated from pigeons. Appl Environ Microbiol 66:1205–1208
Schwaber MJ, Carmeli Y (2007) Mortality and delay in effective therapy associated with extended-spectrum beta-lactamase production in Enterobacteriaceae bacteraemia: a systematic review and meta-analysis. J Antimicrob Ch 60:913–920
Seigal S, Catellan NJ Jr (1987) Nonparametric statistics for behavioral sciences. McGraw-Hill, New York
Sharma VK, Sharma M, Katoch RC, Katoch V, Dhar P (2006) Recovery of some important serotypes of E. coli from diarrhoeic and apparently healthy calves in Palam Valley of Himachal Pradesh. Int J Cow Sci 2:34–36
Shepherd AK, Pottinger PS (2013) Management of urinary tract infections in the era of increasing antimicrobial resistance. Med Clin North Am 97:737–757
Sidhu JP, Ahmed W, Hodgers L, Tozea S (2013) Occurrence of virulence genes associated with diarrheagenic pathotypes in Escherichia coli isolates from surface water. Appl Environ Microbiol 79:328–335
Sreela SN, Japheth AO, Lijek RS, Newman MJ, Okeke IN (2011) Quinolone resistance in Escherichia coli from Accra, Ghana. BMC Microbiol 11:44
Sukumaran DP, Durairaj S, Hatha AM (2012) Antibiotic resistance of Escherichia coli serotypes from Cochin Estuary. Interdisciplinary Perspectives on Infectious Diseases, Article ID 124879, 7 pages.
Tumbarello M, Sanguinetti M, Montuori E, Trecarichi EM, Posteraro B, Fiori B et al (2007) Predictors of mortality in patients with bloodstream infections caused by extended-spectrum-beta-lactamase-producing Enterobacteriaceae: importance of inadequate initial antimicrobial treatment. Antimicrob Agents Ch 51:1987–1994
Watkinson AJ, Micalizzi GR, Bates JR, Costanzo SD (2007) Novel method for rapid assessment of antibiotic resistance in Escherichia coli isolates from environmental waters by use of a modified chromogenic agar. Appl Environ Microbiol 73:2224–2229
Wieler LH, Busse B, Steinruck H, Beutin L, Weber A et al (2000) Enterohemorrhagic Escherichia coli (EHEC) strains of serogroup O118 display three distinctive clonal groups of EHEC pathogens. J Clin Microbiol 38:2162–2169
Acknowledgements
The authors are grateful to the Director, CSIR-National Institute of Oceanography (CSIR-NIO), Goa, India and CSIR-NIO, Scientist-in-Charge, Regional Centre, Mumbai for their encouragement and support. The funding for this work was provided by OLP1209. NIO contribution number: 6009.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible editor: Diane Purchase
Rights and permissions
About this article
Cite this article
Maloo, A., Fulke, A.B., Mulani, N. et al. Pathogenic multiple antimicrobial resistant Escherichia coli serotypes in recreational waters of Mumbai, India: a potential public health risk. Environ Sci Pollut Res 24, 11504–11517 (2017). https://doi.org/10.1007/s11356-017-8760-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-017-8760-8