Abstract
This study evaluated decabromodiphenyl ether (BDE-209) anaerobic debromination and bacterial community changes in mangrove sediment. BDE-209 debromination rates were enhanced with zerovalent iron compared to without zerovalent iron in the sediment. BDE-209 debromination rates in microcosms constructed with sediments collected in autumn were higher than in microcosms constructed with sediments collected in spring and were higher at the Bali sampling site than the Guandu sampling site. The intermediate products resulting from the reductive debromination of BDE-209 in sediment were nona-BDE (BDE-206, BDE-207), octa-BDEs (BDE-196, BDE-197), hepta-BDEs (BDE-183, BDE-184, BDE-191), hexa-BDEs (BDE-137, BDE-138, BDE-154, BDE-157), penta-BDEs (BDE-85, BDE-99, BDE-100, BDE-126), tetra-BDEs (BDE-47, BDE-49, BDE-66, BDE-77), tri-BDEs (BDE-17, BDE-28), and di-BDEs (BDE-15). Fifty bacterial genera associated with BDE-209 debromination were identified. Overall, 12 of the 50 bacterial genera were reported to be involved in dehalogenation of aromatic compounds. These bacteria have high potential to be BDE-209 debromination bacteria. Different combinations of bacterial community composition exhibit different abilities for BDE-209 anaerobic debromination.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Polybrominated diphenyl ethers (PBDEs) are the most widely used brominated flame retardants (de Wit 2002). PBDEs have been dispersed into air, water, sewage sludge, sediment, and human and animal tissues (de Wit 2002; Schreiber et al. 2010; Chen et al. 2013). PBDEs disrupt thyroid hormone production, lead to hepatic toxicity, and lead to developmental neurotoxicity in humans and mammals (Tseng et al. 2008). PBDEs constitute a major concern for human health and the main targets in environmental remediation.
Microbial degradation is believed to be a major process that cleans up PBDE-contaminated sediment. The fate of decabromodiphenyl ether (BDE-209) in anoxic sediment, where it may be subject to anaerobic reductive debromination (Gerecke et al. 2005), is thus of great interest. To enhance the efficiency of biodegradation, three remedial strategies, namely, natural attenuation, bioaugmentation, and biostimulation, have been proposed (Yu et al. 2005). Higher brominated PBDEs are difficult to be debrominated by the anaerobic strains and a long reaction time is needed (Smidt and deVos 2004). The addition of brij 30, brij 35, rhamnolipid, surfactin, vitamin B12, zerovalent iron, acetate, lactate, and pyruvate were shown to influence BDE-209 anaerobic debromination; zerovalent iron yielded the highest BDE-209 anaerobic debromination rate in sediment (Huang et al. 2014).
Mangrove ecosystems, the dominant intertidal estuarine wetlands along the coastlines of tropical and subtropical regions, are closely tied to human activities and are subject to pollutants contamination. The anaerobic degradation of various pollutants in mangrove sediments has been observed (Chang et al. 2008, 2009; Li et al. 2010; Andrade et al. 2012). The Guandu and Bali mangroves are the largest mangroves in subtropical Taiwan. Both mangroves are on the bank of the Tanshui River, which is one of the most heavily contaminated rivers in northern Taiwan (Liu et al. 2000). The concentration and anaerobic degradation of nonylphenol and polycyclic aromatic hydrocarbons in mangrove sediment have been demonstrated previously (Chang et al. 2008, 2009), but little is known about anaerobic debromination of BDE-209 in mangrove sediment.
The climatic characteristics of subtropical regions foster diverse microbial communities. Metagenomic studies with next-generation sequencing (NGS) allow microbial diversity to be studied in environmental samples (Sun et al. 2015; Yang et al. 2015a, b). However, little is known about the microbial communities involved in BDE-209 anaerobic debromination in mangrove sediment. The aims of this study were to compare the debromination of BDE-209 in mangrove sediment from two sampling sites, to evaluate the effects of various factors on BDE-209 debromination, and to explore changes in the bacterial community in the mangrove sediment.
Material and methods
Chemicals
An analytical of standard solutions containing PBDE congeners (BDE-15, BDE-17, BDE-28, BDE-47, BDE-49, BDE-66, BDE-71, BDE-77, BDE-85, BDE-99, BDE-100, BDE-119, BDE-126, BDE-137, BDE-138, BDE-153, BDE-154, BDE-157, BDE-183, BDE-184, BDE-191, BDE-196, BDE-197, BDE-206, BDE-207, BDE-209) were purchased from Wellington Laboratories (Guelph, Canada). BDE-209 (99%) and all other chemicals were purchased from Sigma-Aldrich (St. Louis, MO). Solvents were purchased from Mallinckrodt, Inc.
Sample collection
Sediment samples were collected from sampling sites in our previous study (Chang et al. 2009). Samples were taken from the Guandu sampling site (25.11° 6843′ N, 121.46° 4153′ E) and Bali sampling site (25.15° 8613′ N, 121.43° 5575′ E) in northern Taiwan. They were collected from deep sediment (>15 cm) during two different seasons: autumn (October 2014) and spring (March 2015) during low tide with a soil core. All sediment samples were taken randomly, in triplicate, from an area of approximately 1 m2 at the center of each mangrove sediment site. The sediment samples were stored in glass bottles on ice in a cooler and transported back to the laboratory within 4 h of sampling. For the sediment samples taken from Guandu and Bali mangroves in the spring, temperatures are 18.1 and 20.5 °C, bacterial counts are 2.2 × 105 and 3.6 × 106 CFU/g, and salinities are 1.1 and 16.5%, respectively. For those from Guandu and Bali mangroves in the autumn, temperatures are 25.1 and 27.6 °C, bacterial counts are 9.2 × 105 and 8.6 × 106 CFU/g, and salinities are 3.1 and 28.7%, respectively. For the sediment samples taken from Guandu in the spring, the concentrations of the PBDE congeners (BDE-17, BDE-28, BDE-47, BDE-49, BDE-66, BDE-71, BDE-77, BDE-85, BDE-99, BDE-100, BDE-119, BDE-126, BDE-138, BDE-153, BDE-154, BDE-183, BDE-191, BDE-196, BDE-197, BDE-206, BDE-207, BDE-209) were 132, 43.9, 848, 92.6, 57.7, 242, 1.47, 138, 934, 560, 14.0, 142, 54.0, 322, 205, 195, 30.6, 387, 269, 9309, 7599, and 250,000 ng/kg, respectively. For those from Bali in the spring, the concentrations of the PBDE congeners (BDE-17, BDE-28, BDE-47, BDE-49, BDE-66, BDE-71, BDE-77, BDE-85, BDE-99, BDE-100, BDE-119, BDE-126, BDE-138, BDE-153, BDE-154, BDE-183, BDE-191, BDE-196, BDE-197, BDE-206, BDE-207, BDE-209) were 15.4, 26.2, 225, 143, 26.9, 3.27, 2.27, 7.28, 205, 44.3, 2.29, 0.323, 6.64, 79.6, 49.9, 211, 33.2, 542, 336, 23,563, 30,777, and 640,000 ng/kg, respectively.
Experimental design
The experiments were performed using 125-mL serum bottles containing 45 mL river water, 5 g sediment, and 10 mg/kg of BDE-209. We measured BDE-209 anaerobic debromination with or without zerovalent iron (1 g/L) from the Guandu and Bali sampling sites in the spring and autumn. Inoculated controls were incubated without zerovalent iron with shaking at 30 °C in the dark. Sterile controls were autoclaved at 121 °C for 30 min on three consecutive days.
All experiments were conducted in an anaerobic glove box (Forma Scientific, Model 1025S/N, USA) filled with N2 (85%), H2 (10%), and CO2 (5%) gas. Bottles were capped with butyl rubber stoppers and crimp seals, wrapped in aluminum foil to prevent photolysis, and then incubated without shaking at 30 °C in the dark. Each treatment was performed in triplicate. Samples were periodically collected to measure residual BDE-209 and the bacterial community.
BDE-209 analysis
BDE-209 extraction and analysis were performed as described in our previous study (Huang et al. 2014). BDE-209 was extracted twice from whole bottles by use of hexane and acetone (9:1) and then extracted again for 20 min with use of a Branson 5200 ultrasonic cleaner. Extracts were analyzed using a gas chromatograph (Hewlett Packard 6890) equipped with an electron capture detector (GC-ECD) and Stx-500 capillary column (0.25 mm i.d. × 0.1 μm film thickness × 30 m, Restek). The initial column temperature was set at 170 °C, increased by 10 °C/min to 300 °C, and then increased by 2.5 °C/min to 340 °C. Injector and detector temperatures were set at 350 and 370 °C, respectively. The recovery percentage for BDE-209 was 94.9% and the detection limit was 0.05 mg/L.
Intermediate products of BDE-209 in the sediment samples were measured, as previously described (Chen et al. 2013). All collected sediment samples were mixed, evaporated to near dryness under a gentle stream of nitrogen, and measured by high-resolution gas chromatography and high-resolution mass spectrometry (HRGC/HRMS) (Agilent GC 6890/VG) with a 15-m DB-5HT column (0.25 mm i.d. × 0.1 μm film thickness; J&W Scientific, Folsom, CA). The PBDE congeners (BDE-15, BDE-17, BDE-28, BDE-47, BDE-49, BDE-66, BDE-71, BDE-77, BDE-85, BDE-99, BDE-100, BDE-119, BDE-126, BDE-137, BDE-138, BDE-153, BDE-154, BDE-157, BDE-183, BDE-184, BDE-191, BDE-196, BDE-206, BDE-207, BDE-209) were selected for calculating total PBDEs in sediment samples (Chen et al. 2013). The limits of detection for all the congeners ranged from 0.01 to 0.2 mg/mL. The recovery percentages for the PBDE congeners ranged between 91.7 and 99.5%.
DNA extraction, PCR, and pyrosequencing
Total DNA was extracted by using the PowerSoil DNA Isolation Kit (MO BIO Laboratories) for each experimental sample. Partial sequences containing the V5-V8 variable regions of the 16S ribosomal RNA (rRNA) gene were amplified from the extracted DNA. The sequence of the 5′ primer included a 454 pyrosequencing adaptor, a unique 4-mer tag for each sample and 787F (5′-ATTAGATACCCNGGTAG-3′) for 16S rRNA genes. The sequence for the 3′ primer included a 454 pyrosequencing adaptor and 1391R (5′-ACGGGCGGTGWGTRC-3′) for 16S rRNA genes. PCR was performed as previously described (Yang et al. 2015a). Pyrosequencing was performed at the Genome Center, National Yang-Ming University, Taiwan, with the GS Junior System (Roche Diagnostics Corp., CT, USA).
Data analysis
The BDE-209 remaining percentage [%] = (residue substrate concentration / initial substrate concentration) × 100 was calculated, as was the BDE-209 debromination rates [%] = [1 − (residue substrate concentration/initial substrate concentration)] × 100.
The 16S rRNA gene sequences from pyrosequencing were analyzed using the RDPipeline in the Ribosomal Database Project web site (Cole et al. 2009). Firstly, the Ribosomal Database Project (RDP) Pipeline Initial Processing steps were used to match the raw reads to experimental samples, trimming off the tag and primer portions and removing sequences of low quality. Second, the trimmed sequences over 400 bp were applied to the UCHIME (Edgar et al. 2011) to remove chimeric sequences. Third, sequences pass the preprocessing steps were used to perform Aligner (sequence alignment), complete linkage clustering (sequence clustering), Shannon index and Chao1 estimator (α-diversity evaluation), and RDP classifier (assignment of taxonomic groups). Ninety-five percent similarities (the most stringent condition provided by RDPipeline) were used for all of these analyses. Spearman correlation coefficients of sequence frequencies and BDE-209 remaining percentage data were computed by R. For testing H0, ρ = 0, bacterial genera with p < 0.05 were considered BDE-209 degradation-associated bacteria. Cluster analysis was performed by the heatmap3 package of R. Differences of bacterial diversity between experimental settings were compared by the Wilcoxon rank sum test in R.
Results and discussion
Comparison of BDE-209 concentrations in mangrove sediments
The BDE-209 concentrations in the spring and autumn were 0.25 and 0.22 mg/kg at the Guandu sampling site; they were 0.64 and 0.32 mg/kg at the Bali sampling site. The BDE-209 concentrations in the sediments of the two sampling sites were higher in the spring than in the autumn. The temperature of the sediments was higher in autumn than spring. Temperature is the key factor affecting the seasonal variation of pollutant concentrations in water environments (Xu et al. 2006). The Bali sampling site had higher BDE-209 concentrations than the Guandu sampling site. The Bali sampling site was in the Bali mangrove, which was likely affected by vehicle exhaust deposition and the discharge of industrial, livestock, and household wastewater (Chang et al. 2009).
Comparison of BDE-209 anaerobic debromination in mangrove sediments
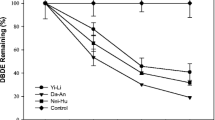
The BDE-209 concentrations in sterile controls were first examined at the end of a 75-day incubation period. The remaining percentage of BDE-209 in sediment ranged from 89.1 to 97.6%. It revealed that the BDE-209 anaerobic debromination occurring in all of the following experiments was due to microbial action. The effects of the addition of zerovalent iron on anaerobic debromination of BDE-209 in the sediment are shown in Fig. 1. The debromination rates of BDE-209 without zerovalent iron in spring and autumn were 40.1 and 51.3% in the Guandu sediments and 71.2 and 80.9% in the Bali sediments, respectively. The debromination rates of BDE-209 with zerovalent iron in spring and autumn were 58.1 and 74.3% in the Guandu sediments and 85.1 and 95.5% in the Bali sediments, respectively. BDE-209 debromination rates were higher with zerovalent iron than without it in the sediments. The addition of zerovalent iron could reduce PBDEs to less brominated compounds by anaerobic microbes (Kim et al. 2011). As well, sorption could play a role in the BDE-209 debromination process with the addition of zerovalent iron (Shih and Tai 2010).
Comparison of BDE-209 anaerobic debromination in the sediments from the Guandu and Bali sampling sites after incubation for 75 days in the sediments. a Guandu spring. b Bali spring. c Guandu autumn, d Bali autumn. Symbols: filled triangles, sterile sediment with zerovalent iron; inverted filled triangles, sterile sediment without zerovalent iron; filled circles, non-sterile sediment with zerovalent iron; empty circles, non-sterile sediment without zerovalent iron
The results also revealed that BDE-209 debromination rates in microcosms constructed with sediments collected in autumn were higher than in microcosms constructed with sediments collected in spring, and higher in Bali sediments than in Guandu sediments. The salinity at the Guandu sampling site in spring and autumn were 1.1 and 3.1%, respectively. The salinity at the Bali sampling site in spring and autumn were 16.5 and 28.7%, respectively. Salinity may affect organic pollutant degradation in environments (Castillo-Carvajal et al. 2014). Also, the concentration of BDE-209 was detected in the sediments of the two sampling sites. The adaptation process enhanced BDE-209 anaerobic debromination (Huang et al. 2014). The BDE-209 could be debrominated without zerovalent iron. Microorganisms adapt to site-specific conditions, resulting in varied but optimal biodegradation capacity (Chang et al. 2009).
Identification of intermediate products from BDE-209 debromination in the sediment
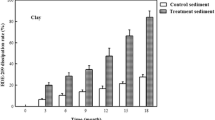
Firstly, bromide ion release was assessed in the BDE-209 debromination process after incubation for 75 days in the sediments. As shown in Fig. 2, the bromide ion concentrations increased to 1.5 and 2.4 mg/L without zerovalent iron and to 4.0 and 5.2 mg/L with zerovalent ion in the Guandu and Bali sediments collected in spring, respectively. The bromide ion concentrations increased to 3.3 and 4.8 mg/L without zerovalent iron and to 5.6 and 7.7 mg/L with zerovalent ion in the Guandu and Bali sediments collected in autumn, respectively. The results indicated that bromide ions were produced from BDE-209 debromination. The bromide ion concentrations were higher with zerovalent iron than without zerovalent iron and higher from the Bali sediments than the Guandu sediments.
The BDE-209 biotransformation in the sediment was also monitored by HRGC/HRMS. The concentrations of the PBDE congeners (BDE-15, BDE-17, BDE-28, BDE-47, BDE-49, BDE-66, BDE-77, BDE-85, BDE-99, BDE-100, BDE-126, BDE-137, BDE-138, BDE-154, BDE-157, BDE-183, BDE-184, BDE-191, BDE-196, BDE-197, BDE-206, BDE-207, BDE-209) were 4.7, 3.0, 28.9, 17.1, 3.5, 2.2, 1.3, 1.74, 14.7, 4.7, 1.5, 2.2, 2.4, 7.2, 0.99, 33.5, 12.7, 8.4, 263.6, 223.9, 5574, 7663, and 175,882 μg/kg, respectively. Compared to the concentrations of PBDE congeners in original sediments, the PBDE congeners (BDE-15, BDE-17, BDE-28, BDE-47, BDE-49, BDE-66, BDE-77, BDE-85, BDE-99, BDE-100, BDE-126, BDE-137, BDE-138, BDE-154, BDE-157, BDE-183, BDE-184, BDE-191, BDE-196, BDE-197, BDE-206, BDE-207) may be the intermediate from the reductive debromination of BDE-209 in the sediment. BDE-209 parent starting material was converted into nona-BDE (BDE-206 and BDE-207). BDE-207 was converted by meta-substitution into octa-BDE (BDE-197), by meta-substitution into BDE-184, by ortho-substitution into BDE-137, by ortho-substitution into BDE-85, by ortho-substitution into BDE-66, by meta-substitution into BDE-28, and by ortho-substitution into BDE-15. BDE-206 was converted by meta-substitution into BDE-196; BDE-196 was converted by meta-substitution into BDE-183 and by ortho-substitution into BDE-191. BDE-191 was converted by ortho-substitution into BDE-157, by ortho substitution into BDE-126, and by meta-substitution into BDE-77. BDE-183 was converted into BDE-138 and BDE-154. BDE-138 was converted by meta-substitution into BDE-85, BDE-66, BDE-28, and BDE-15. BDE-154 was converted into BDE-99 and BDE-100. BDE-99 was converted by para-substitution into BDE-49 and by meta-substitution into BDE-17. BDE-100 was converted by ortho-substitution into BDE-47. BDE-47 was converted by ortho-substitution into BDE-28 and by para-substitution into BDE-17.
From the experiments, BDE-209 with ortho, meta, and para specificity dominated the debromination in the sediment. BDE-209 could be biotransformed via a debromination process to nona-BDE (BDE-206, BDE-207), octa-BDEs (BDE-196, BDE-197), hepta-BDEs (BDE-183, BDE-184, BDE-191), hexa-BDEs (BDE-137, BDE-138, BDE-154, BDE-157), penta-BDEs (BDE-85, BDE-99, BDE-100, BDE-126), tetra-BDEs (BDE-47, BDE-49, BDE-66, BDE-77), tri-BDEs (BDE-17, BDE-28), and di-BDE (BDE-15) in the sediment. A proposed reductive debromination pathway for the biotransformation of BDE-209 in the sediment is simulated in Fig. 3.
Keum and Li (2005) revealed that BDE-209 was debrominated to di-BDE (BDE-7, BDE-8, BDE-15) by zerovalent iron treatment. Kim et al. (2011) showed that reductive debromination of BDE-209 with zerovalent iron produced various intermediates ranging from nona-BDEs to tri-BDEs (BDE-28, BDE-30) which were then treated with Sphingomonas sp. PH-07 strain to mono-BDEs. Huang et al. (2014) showed successive biotransformation of BDE-209 to BDE-3 via a reductive debromination process by BDE-adapted sediment. Our results revealed that different ecosystems present different microbial communities, resulting in various debromination rates and intermediate products from BDE-209 in the sediment. As degradation rates are expected to vary greatly between individual anaerobic systems and prevailing redox conditions (sludge, sediment, soil) (Gerecke et al. 2005). The capability of degradation appears to be dependent on several factors such as microbial community structure, salinity, redox potential, total organic carbon, and presence of other pollutants (Stiborova et al. 2015).
Diversity of microbial community in BDE-209 anaerobic debromination experiments
In total, 126,933 16S rRNA gene sequences were produced from 28 experimental samples. Analysis with an RDP classifier revealed 31 phyla, 50 classes, 74 orders, 166 families, and 379 genera of known bacteria in the samples. The number of sequences, OTUs, and biological classification groups of the bacteria as well as the diversity indexes are in Table 1. Comparison of diversity indexes by the Wilcoxon rank sum test indicates that the microbial diversity of the sediment sample at day 0 was higher than that of the experimental samples at days 15, 45, and 75 (p = 0.0064). The microbial diversity of the experiment samples from the Guandu sampling site in autumn was higher than that in spring (p = 0.0081). However, the microbial diversity of experimental samples from the Bali sampling site in autumn and the Bali sampling site in spring exhibited no difference. The microbial diversity of the experimental samples with and without zerovalent iron from both the Guandu and Bali sampling sites exhibited no difference. These results suggest that the addition of zerovalent iron did not change microbial diversity in the experiments. Table 1 shows that there is a huge decline in overall diversity after addition of BDE-209. The effects of organic pollutants on microbial community structure in natural communities are complex. This complexity is reflected by the diverse physical, chemical, and biological factors (Battin et al. 2008). BDE-209 entering the sediment may affect microbial communities not only by reducing the diversity, but they may also change the composition of the microbial community (Zhang et al. 2012).
The major bacterial classes identified in BDE-209 anaerobic debromination were Gammaproteobacteria (62.8% of 126,933 sequences), Clostridia (5.2%), Flavobacteriia (4.3%), Alphaproteobacteria (3.5%), Betaproteobacteria (3.2%), Deltaproteobacteria (2.4%), Actinobacteria (1.3%), and Bacilli (1.2%) (Fig. 4a). The five archaeal classes (1.1% of 126,933 sequences) found in the sediment samples at day 0 were largely decreased in the experimental samples at days 15, 45, and 75 (Fig. 4b). Cluster analysis revealed that the microbial community compositions are highly diverse between sampling sites, sampling seasons, and experimental settings (Fig. 5).
Major bacterial (a) and archaea (b) classes in each experiment. G and B represent the Guandu and Bali sampling sites, respectively. S and A represent spring and autumn, respectively. F and N represent with zerovalent iron and without zerovalent iron, respectively. The numbers 0, 15, 45, and 75 represent days 0, 15, 45, and 75, respectively
Cluster analysis of bacterial community compositions between experiments. G and B represent the Guandu and Bali sampling sites, respectively. S and A represent spring and autumn, respectively. F and N represent with zerovalent iron and without zerovalent iron, respectively. The numbers 0, 15, 45, and 75 represent days 0, 15, 45 and 75, respectively
Bacteria associated with BDE-209 anaerobic debromination
To identify bacteria associated with anaerobic debromination of PBDEs, the proportion of the 16S rRNA gene sequences from each bacterial genus in each sample and the BDE-209 remaining percentages in the debromination experiments were used to compute Spearman correlation coefficients. Bacteria with sequence proportions exhibiting a negative correlation (p < 0.05) with the BDE-209 remaining percentages (increasing with BDE-209 debromination) were selected. Twelve, 12, 9, and 12 bacterial genera in samples with zerovalent iron from the Guandu sediment in spring, Bali sediment in spring, Guandu sediment in autumn, and Bali sediment in autumn exhibiting negative correlations (p < 0.05) with BDE-209 debromination were identified (Fig. 6a, c, e, g). Seven, 10, 12, and 11 bacterial genera in samples without zerovalent iron from the Guandu sediment in spring, Bali sediment in spring, Guandu sediment in autumn, and Bali sediment in autumn exhibiting negative correlations (p < 0.05) with BDE-209 debromination were identified (Fig. 6b, d, f, h). Overall, 50 bacterial genera associated with BDE-209 debromination were identified in the eight experimental settings. Twelve of the 50 bacterial genera were previously found to be involved in dehalogenation of aromatic compounds, and six of the 50 bacterial genera including Bacillus, Brevibacillus, Clostridium XI, Clostridium XlVa, Clostridium_ss, and Pseudomonas were reported to be involved in debromination of aromatic compounds (Shih et al. 2012; Lu et al. 2013; Shi et al. 2013; Tang et al. 2014). Another six of the 50 bacterial genera, Anaeromyxobacter, Burkholderia, Mycobacterium, Rhodobacter, Sedimentibacter, and Shewanella, were reported to be involved in dechlorination or deiodination of aromatic compounds (Jesenska et al. 2000; He and Sanford 2002; Redwood et al. 2008; Pandey et al. 2011; Lohner and Spormann 2013; Oba et al. 2014). These results suggest that these bacteria have high potential to be BDE-209 debromination bacteria. Different combinations of bacterial community compositions exhibit different abilities for BDE-209 anaerobic debromination.
Bacterial genera exhibiting a negative correlation with BDE-209 remaining percentage in the experiments. G and B represent the Guandu and Bali sampling sites, respectively. S and A represent spring and autumn, respectively. F and N represent with zerovalent iron and without zerovalent iron, respectively. The numbers 0, 15, 45, and 75 represent day 0, 15, 45, and 75, respectively. The red asterisks and at symbols indicate bacterial genera that have been reported to be involved in reductive debromination and dehalogenation of aromatic hydrocarbons, respectively
Commonly and differentially distributed bacteria associated with BDE-209 anaerobic debromination in different settings
The results in Fig. 1 indicate that BDE-209 debromination rates were higher in autumn than in spring, and higher with zerovalent iron addition than without zerovalent iron. To identify commonly and differentially distributed bacteria associated with BDE-209 anaerobic debromination in different settings, Venn diagram analysis was conducted. For the sediments from the Guandu sampling site (Fig. 7a), 32 bacterial genera were divided into 9 groups. Eight, four, two, and eight bacterial genera were differentially distributed in the spring sample with zerovalent iron, autumn sample with zerovalent iron, spring sample without zerovalent iron, and autumn sample without zerovalent iron, respectively. For the sediments from the Bali sampling site (Fig. 7b), 31 bacterial genera were divided into 9 groups. Six, five, five, and three bacterial genera were differentially distributed in the spring sample with zerovalent iron, autumn sample with zerovalent iron, spring sample without zerovalent iron, and autumn sample without zerovalent iron, respectively. These differentially distributed bacteria groups provide insights into the different BDE-209 debromination rates in different experimental settings. Overall, 30 of the 50 bacterial genera in sediments from Guandu and Bali sampling sites, such as Ensifer, Gracilimonas, Ideonella, Marinobacter, Prolixibacter, and Sulfurimonas (Li et al. 2012; Hausler et al. 2014; Yousuf et al. 2014; Al-Mailem et al. 2015; Li et al. 2016; Vavourakis et al. 2016) were reported to be salinity tolerant extremophiles (Fig. 7). Eleven of the 30 bacterial genera including Bacillus, Brevibacillus, Clostridium XI, Clostridium XlVa, Clostridium_ss, Pseudomonas, Burkholderia, Mycobacterium, Rhodobacter, Sedimentibacter, and Shewanella were reported to be involved in dehalogenation of aromatic compounds. Therefore, study of the BDE-209 anaerobic debromination ability of bacteria from mangrove sediments may be useful for applications in bioremediation of saline environments.
Comparison of BDE-209 debromination-associated bacteria between experiments with and without zerovalent iron from the Guandu (a) and Bali (b) sampling sites. G and B represent the Guandu and Bali sampling sites, respectively. S and A represent spring and autumn, respectively. F and N represent with zerovalent iron and without zerovalent iron, respectively. The numbers 0, 15, 45, and 75 represent day 0, 15, 45, and 75, respectively. The red asterisks indicate bacterial genera that have been reported to be salinity tolerant extremophiles. The red at symbols indicate bacterial genera that have been reported to be involved in reductive dehalogenation of aromatic hydrocarbons
Conclusions
Microbial degradation of BDE-209 is a major process that results in the decontamination of mangrove sediments. The addition of zerovalent iron enhanced BDE-209 anaerobic debromination in the sediment. Anaerobic debromination of BDE-209 could be biotransformed to di-BDEs (BDE-15) in the sediment. A reductive debromination pathway for biotransformation of BDE-209 in the sediment is proposed in Fig. 3. Fifty bacterial genera associated with BDE-209 debromination were identified, and 12 of the 50 bacterial genera were previously found to be involved in dehalogenation of aromatic compounds. These bacteria have high potential to be BDE-209 debromination bacteria. Moreover, 30 of the 50 bacterial genera were reported to be salinity tolerant extremophiles. Eleven of the 30 bacterial genera were reported to be involved in dehalogenation of aromatic compounds. Bacteria from mangrove sediments may be applicable in the bioremediation of saline environments.
References
Al-Mailem DM, Eliyas M, Khanafer M, Radwan SS (2015) Biofilms constructed for the removal of hydrocarbon pollutants from hypersaline liquids. Extremophiles 19:189–196. doi:10.1007/s00792-014-0698-x
Andrade LL, Leite DCA, Ferreir EM, Ferreir LQ, Paula GR, Maguire MJ, Hubert CRJ, Peixoto RS, Domingues RMCP, Rosado AS (2012) Microbial diversity and anaerobic hydrocarbon degradation potential in an oil-contaminated mangrove sediment. BMC Microbiol 12:186–195. doi:10.1186/1471-2180-12-186
Battin TJ, Kaplan LA, Findlay S, Hopkinson CS, Marti E, Packman AI, Newbold JD, Sabater F (2008) Biophysical controls on organic carbon fluxes in fluvial networks. Nat Geosci 1:95–100. doi:10.1038/ngeo101
Castillo-Carvajal LC, Sanz-Martín JL, Barragán-Huerta BE (2014) Biodegradation of organic pollutants in saline wastewater by halophilic microorganisms: a review. Environ Sci Pollut Res Int 21:9578–9588. doi:10.1007/s11356-014-3036-z
Chang BV, Chang IT, Yuan SY (2008) Anaerobic degradation of phenanthrene and pyrene in mangrove sediment. Bull Environ Contam Toxicol 80:145–149. doi:10.1007/s00128-007-9333-1
Chang BV, Lu ZJ, Yuan SY (2009) Anaerobic degradation of nonylphenol in subtropical mangrove sediments. J Hazard Mater 165:162–167. doi:10.1016/j.jhazmat.2008.09.085
Chen CY, Tien CJ, Sun YM, Hsieh CY, Lee CC (2013) Influence of water quality parameters on occurrence of polybrominated diphenyl ether in sediment and sediment to biota accumulation. Chemosphere 90:2420–2427. doi:10.1016/j.chemosphere.2012.10.073
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Mohideen AS, McGarrell DM, Marsh T, Garrity GM, Tiedje JM (2009) The ribosomal database project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37:D141–D145. doi:10.1093/nar/gkn879
De Wit CA (2002) An overview of brominated flame retardants in the environment. Chemosphere 46:583–624. doi:10.1016/S0045-6535(01)00225-9
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. doi:10.1093/bioinformatics/btr381
Gerecke AC, Hartmann PC, Heeb NV, Kohler HPE, Giger W, Schmid P, Zennegg M, Kohler M (2005) Anaerobic degradation of decabromodiphenyl ether. Environ Sci Technol 39:1078–1083. doi:10.1021/es048634j
Hausler S, Weber M, Siebert C, Holtappels M, Noriega-Ortega BE, De Beer D, Ionescu D (2014) Sulfate reduction and sulfide oxidation in extremely steep salinity gradients formed by freshwater springs emerging into the Dead Sea. FEMS Microbiol Ecol 90:956–969. doi:10.1111/1574-6941.12449
He Q, Sanford RA (2002) Induction characteristics of reductive dehalogenation in the ortho-halophenol-respiring bacterium, Anaeromyxobacter dehalogenans. Biodegradation 13:307–316. doi:10.1023/A:1022342421909
Huang HW, Chang BV, Lee CC (2014) Reductive debromination of decabromodiphenyl ether by anaerobic microbes from river sediment. Int Biodeterior Biodegrad 87:60–65. doi:10.1016/j.ibiod.2013.10.011
Jesenska A, Sedlacek I, Damborsky J (2000) Dehalogenation of haloalkanes by Mycobacterium tuberculosis H37Rv and other mycobacteria. Appl Environ Microbiol 66:219–222. doi:10.1128/AEM.66.1.219-222.2000
Keum YS, Li QX (2005) Reductive debromination of polybrominated diphenyl ethers by zerovalent iron. Environ Sci Technol 39:2280–2286. doi:10.1021/es048846g
Kim YM, Murugesan K, Chang YY, Kim EJ, Chang YS (2011) Degradation of polybrominated diphenyl ethers by a sequential treatment with nanoscale zero valent iron and aerobic biodegradation. J Chem Technol Biotechnol 87:216–224. doi:10.1002/jctb.2699
Li CH, Wong YS, Tam NFY (2010) Anaerobic biodegradation of polycyclic aromatic hydrocarbons with amendment of iron(III) in mangrove sediment slurry. Biores Technol 101:8083–8092. doi:10.1016/j.biortech.2010.06.005
Li H, Zhang Q, Wang XL, Ma XY, Lin KF, Liu YD, JD G, SG L, Shi L, Lu Q, Shen TT (2012) Biodegradation of benzene homologues in contaminated sediment of the East China Sea. Bioresour Technol 124:129–136. doi:10.1016/j.biortech.2012.08.033
Li Y, Li X, Liu Y, Wang ET, Ren C, Liu W, Xu H, Wu H, Jiang N, Li Y, Zhang X, Xie Z (2016) Genetic diversity and community structure of rhizobia nodulating Sesbania cannabina in saline-alkaline soils. Syst Appl Microbiol 39:195–202. doi:10.1016/j.syapm.2016.02.004
Liu C, Wang SK, Lu YB (2000) Chemical characterization of river sediment in Taiwan. Toxicol Environ Chem 76:205–218. doi:10.1080/02772240009358929
Lohner ST, Spormann AM (2013) Identification of a reductive tetrachloroethene dehalogenase in Shewanella sediminis. Philos Trans R Soc Lond Ser B Biol Sci 368:20120326–20120335. doi:10.1098/rstb.2012.0326
Lu M, Zhang ZZ, Wu XJ, Xu YX, Su XL, Zhang M, Wang JX (2013) Biodegradation of decabromodiphenyl ether (BDE-209) by a metal resistant strain, Bacillus cereus JP12. Bioresour Technol 149:8–15. doi:10.1016/j.biortech.2013.09.040
Oba Y, Futagami T, Amachi S (2014) Enrichment of a microbial consortium capable of reductive deiodination of 2,4,6-triiodophenol. J Biosci Bioeng 117:310–317. doi:10.1016/j.jbiosc.2013.08.011
Pandey J, Heipieper HJ, Chauhan A, Arora PK, Prakash D, Takeo M, Jain RK (2011) Reductive dehalogenation mediated initiation of aerobic degradation of 2-chloro-4-nitrophenol (2C4NP) by Burkholderia sp. strain SJ98. Appl Microbiol Biotechnol 92:597–607. doi:10.1007/s00253-011-3254-y
Redwood MD, Deplanche K, Baxter-Plant VS, Macaskie LE (2008) Biomass-supported palladium catalysts on Desulfovibrio desulfuricans and Rhodobacter sphaeroides. Biotechnol Bioeng 99:1045–1054. doi:10.1002/bit.21689
Schreiber T, Gassmann K, Gotz C, Hubenthal U, Moors M, Krause G, Merk HF, Nguyen NH, Scanlan TS, Abel J, Rose CR, Fritsche E (2010) Polybrominated diphenyl ethers induce developmental neurotoxicity in a human in vitro model: evidence for endocrine disruption. Environ Health Perspect 118:572–578. doi:10.1289/ehp.0901435
Shi G, Yin H, Ye J, Peng H, Li J, Luo C (2013) Aerobic biotransformation of decabromodiphenyl ether (BDE-209) by Pseudomonas aeruginosa. Chemosphere 93:1487–1493. doi:10.1016/j.chemosphere.2013.07.044
Shih YH, Tai YT (2010) Reaction of decabrominated diphenyl ether by zerovalent iron nanoparticles. Chemosphere 78:1200–1206. doi:10.1016/j.chemosphere.2009.12.061
Shih YH, Chou HL, Peng YH (2012) Microbial degradation of 4-monobrominated diphenyl ether with anaerobic sludge. J Hazard Mater 213–214:341–346. doi:10.1016/j.jhazmat.2012.02.009
Smidt H, deVos WM (2004) Anaerobic microbial dehalogenation. Annu Rev Microbiol 58:43–73. doi:10.1146/annurev.micro.58.030603.123600
Stiborova H, Vrkoslavova J, Pulkrabova J, Poustka J, Hajslova J, Demnerova K (2015) Dynamics of brominated flame retardants removal in contaminated wastewater sewage sludge under anaerobic conditions. Sci Total Environ 533:439–445. doi:10.1016/j.scitotenv.2015.06.131
Sun WM, Li JW, Jiang L, Sun ZL, MY F, Peng XT (2015) Profiling microbial community structures across six large oilfields in China and the potential role of dominant microorganisms in bioremediation. Appl Microbiol Biotechnol 99:8751–8764. doi:10.1007/s00253-015-6748-1
Tang S, Bai J, Yin H, Ye J, Peng H, Liu Z, Dang Z (2014) Tea saponin enhanced biodegradation of decabromodiphenyl ether by Brevibacillus brevis. Chemosphere 114:255–261. doi:10.1016/j.chemosphere.2014.05.009
Tseng LH, Li MH, Tsai SS, Lee CW, Pan MH, Yao WJ, Hsu PC (2008) Developmental exposure to decabromodiphenyl ether (PBDE-209): effects on thyroid hormone and hepatic enzyme activity in male mouse offspring. Chemosphere 70:640–647. doi:10.1016/j.chemosphere.2007.06.078
Vavourakis CD, Ghai R, Rodriguez-Valera F, Sorokin DY, Tringe SG, Hugenholtz P, Muyzer G (2016) Metagenomic insights into the uncultured diversity and physiology of microbes in four hypersaline soda lake brines. Front Microbiol 7:211. doi:10.3389/fmicb.2016.00211
Xu J, Wang P, Guo W, Dong J, Wang L, Dai S (2006) Seasonal and spatial distribution of nonylphenol in Lanzhou Reach of Yellow River in China. Chemosphere 65:1445–1451. doi:10.1016/j.chemosphere.2006.04.042
Yang CW, Huang HW, Chao WL, Chang BV (2015a) Bacterial communities associated with aerobic degradation of polybrominated diphenyl ethers from river sediments. Environ Sci Pollut Res 22:3810–3819. doi:10.1007/s11356-014-3626-9
Yang Y, Wang Z, He T, Dai Y, Xie S (2015b) Sediment bacterial communities associated with anaerobic biodegradation of bisphenol-A. Microb Ecol 70:97–104. doi:10.1007/s00248-014-0551-x
Yousuf B, Kumar R, Mishra A, Jha B (2014) Differential distribution and abundance of diazotrophic bacterial communities across different soil niches using a gene-targeted clone library approach. FEMS Microbiol Lett 360:117–125. doi:10.1111/1574-6968.12593
Yu KSH, Wong AHY, Yau KWY, Wong YS, Tam NFY (2005) Natural attenuation, biostimulation and bioaugmentation on biodegradation of polycyclic aromatic hydrocarbons (PAHs) in mangrove sediments. Mar Pollut Bull 51:1071–1077. doi:10.1016/j.marpolbul.2005.06.006
Zhang W, Zhang M, An S, Lin K, Li H, Cui C, Fu R, Zhu J (2012) The combined effect of decabromodiphenyl ether (BDE-209) and copper (Cu) on soil enzyme activities and microbial community structure. Environ Toxicol Pharmacol 34:358–369. doi:10.1016/j.etap.2012.05.009
Acknowledgements
This research was supported by the Ministry of Science and Technology, Taiwan, Republic of China (grant no. MOST 104-2313-B-031-001-MY3).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible editor: Diane Purchase
Rights and permissions
About this article
Cite this article
Yang, CW., Lee, CC., Ku, H. et al. Bacterial communities associated with anaerobic debromination of decabromodiphenyl ether from mangrove sediment. Environ Sci Pollut Res 24, 5391–5403 (2017). https://doi.org/10.1007/s11356-016-8259-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-016-8259-8