Abstract
The seasonal variation and removal efficiency of antibiotic resistance genes (ARGs), including tetracycline resistance genes (tetG, tetM, and tetX) and macrolide (ermB, ermF, ereA, and mefA), were investigated in two typical swine wastewater treatment systems in both winter and summer. ARGs, class 1 integron gene, and 16S rRNA gene were quantified using real-time polymerase chain reaction assays. There was a 0.31–3.52 log variation in ARGs in raw swine wastewater, and the abundance of ARGs in winter was higher than in summer. tetM, tetX, ermB, ermF, and mefA were highly abundant. The abundance of ARGs was effectively reduced by most individual treatment process and the removal efficiencies of ARGs were higher in winter than in summer. However, when examining relative abundance, the fate of ARGs was quite variable. Anaerobic digestion reduced the relative abundance of tetX, ermB, ermF, and mefA, while lagoon treatment decreased tetM, ermB, ermF, and mefA. Sequencing batch reactor (SBR) decreased tetM, ermB, and ermF, but biofilters and wetlands did not display consistent removal efficiency on ARGs in two sampling seasons. As far as the entire treatment system is concerned, ermB and mefA were effectively reduced in both winter and summer in both total and relative abundance. The relative abundances of tetG and ereA were significantly correlated with intI1 (p < 0.01), and both tetG and ereA increased after wastewater treatment. This may pose a great threat to public health.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Antibiotic resistance is an increasing threat to global public health, not only increasing the difficulty in infection control but also reaching alarming levels worldwide (WHO 2014). Antibiotic resistance genes (ARGs) were indicated as emerging contaminants by Pruden et al. (2006) and play an important role in antibiotic resistance. According to WHO and several recent studies, many gaps exist in our knowledge about the spread of ARGs, especially regarding animal production and related environments (WHO 2014; Berendonk et al. 2015). Extensive information is thus required to reduce the risk posed by environmental ARGs (Berendonk et al. 2015).
Animal production extensively utilizes antibiotics, especially in developing countries; for example, 52 % of total antibiotic use (162,000 tons) was due to animal production in China in 2013 (Zhang et al. 2015). with growth promotion, prophylactic, and therapeutic uses. Tetracyclines and macrolides are the most commonly used antibiotics for these purposes (Barton 2000; Apley et al. 2012). Chantziaras et al. (2014) reported that the use of specific antibiotics on swine farms strongly correlated to the prevalence of antibiotic resistant bacteria. Zhu et al. (2013) found that ARGs were enriched 192 to 28,000-fold in antibiotic-treated feces compared with antibiotic-free swine manure or soil controls.
Swine wastewater is an important ARGs reservoir. Koike et al. (2007) detected tetM with a relative abundance of 1.38E−02 copies/16S rRNA in swine wastewater. Cheng et al. (2013) detected tet genes with different resistance mechanisms, showing that the abundance of ribosomal protection protein genes (tetQ, tetM, tetW, and tetO) in swine wastewater was higher than the efflux pump gene (tetG) and enzymatic modification gene (tetX). Jindal et al. (2006) found a high level of resistant rRNA (approximately 50 %) with methylation of a specific nucleotide encoded by the erm gene causing resistance to macrolides, lincosamides, and streptogramin B (MLSB) in swine wastewater. However, most studies on ARGs abundance were focused on a single season, and few studies have examined the seasonal variation of ARGs.
Animal wastewater is typically initially treated with anaerobic digestion, aerobic biological treatment, constructed wetlands, or lagoons (Nasir et al. 2012; Deng et al. 2008; Cronk 1996). Tao et al. (2014) evaluated the removal of ARGs from swine wastewater using successive anaerobic and aerobic treatments, with a mean removal efficiency of tetA, tetW, sulI and sulII, and bla TEM in the range of 33.30–97.56 %. Diehl and Lapara (2010) reported that removal of tetA, tetO, tetW, and tetX by anaerobic digestion fit a first-order kinetic model, while the abundance of tetL did not show appreciable decline. Joy et al. (2013) indicated that abundance of ermB and ermF were reduced by 50–60 % and 80–90 %, respectively, by simulated lagoon storage. However, the ARGs removal efficiency was different even with similar treatment processes. Tao et al. (2014) reported that tetW increased by 0.78 log after treatment by two-stage anaerobic digestion, while decreasing by 1.66 log on another swine farm with similar anaerobic process. Chen et al. (2010) indicated that the rate of ARGs removal was influenced by seasonal variation, as ermB was reduced by 1.9 log in a biofilter in winter while only decreasing 0.88 log in summer. Resende et al. (2014a) evaluated the dynamics of ermB and bla TEM-1 by anaerobic digestion at ambient temperature as a pilot scale in summer and winter. The removal efficiency of ermB was slightly higher in summer (93.9 %) than in winter (84.0 %), but the removal efficiency of bla TEM-1 was much higher in summer (2.2 log) than in winter (0.84 log). However, the seasonal variation of tetracycline ARGs has not, to out knowledge, been reported. Thus, the abundance and removal efficiency of ARGs may be impacted by seasonal factors; however, these variation patterns are unclear and needs further study.
Integrons are mobile genetic elements that could facilitate ARG prevalence, as some bacteria can capture ARGs housed on mobile genetic elements via horizontal transfer (Martínez 2008; Shi et al. 2013). Martínez et al. (2014) ranked the ARGs on mobile genetic elements harbored by human bacterial pathogens at the highest risk level. Amos et al. (2015) identified the class 1 integron as a molecular marker of antibiotic resistance, which has helped to better understand the transfer of ARGs in the environment.
Therefore two different swine wastewater treatment systems were chosen in this study, i.e., an “anaerobic digestion-lagoon” system and an “anaerobic digestion-aerobic biological treatment-wetland.” The objective of this study was to evaluate ARGs, class 1 integron gene, and 16S rRNA gene variations between summer and winter both before and after wastewater treatment, as well as to determine the removal efficiencies of different tetracycline and macrolide resistance genes with real-time quantitative polymerase chain reaction (qPCR) assays.
Materials and methods
Description of sampling sites
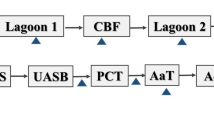
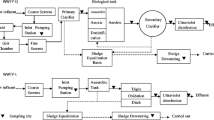
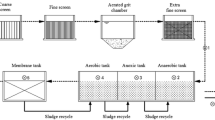
Wastewater samples were collected from two confined swine farms. Farm 1 is located in Beijing, China (116°E, 40°N) and farm 2 is located in Jiangxi Province, China (117°E, 28°N). The swine wastewater treatment flowchart of these two farms is shown in Fig. 1. More detailed information about the treatment processes of the two systems is given in Table 1.
Wastewater samples of 500 mL were collected using sterile containers at influent and effluent of each treatment unit in August 2014 and February 2015. Grab samples were collected and stored at 4 °C before transferred to the laboratory.
DNA extraction
Wastewater samples were first filtered through 0.22-μm filters. For wastewater with high suspended solid (SS higher than 5000 mg/L), 10–20 mL of raw wastewater was filtered; for relatively low SS wastewater, 40–100 mL was filtered for DNA extraction. Then total DNA was extracted from the filter using the FAST DNA extraction Kit (MP Biomedicals, USA) according to the manufacturer’s instructions. Extracted DNA was visualized and quantified using 1 % agarose gel electrophoresis and a NanoDrop 2000 (Thermo Scientific, USA), and then stored at −20 °C until use. Gene concentration was recalculated based on the volume of wastewater used for DNA extraction.
Quantification of ARGs by real-time qPCR
Tetracycline (tetG, tetM, tetX) and macrolide (ermB, ermF, ereA, and mefA) resistance genes, as well as a class 1 integron gene (intI1) and bacterial 16S rRNA genes were quantified using qPCR. The resistance mechanisms of the selected ARGs and the primers used in qPCR are listed in Table 2. The plasmids containing these specific genes were manufactured by Zhejiang Tianke Biotechnology Company (Zhejiang, China), as shown in Table S1. The standard samples were diluted to yield a series of decreasing 10-fold concentrations and subsequently used for generating standard curves. The 25 μL PCR reaction mixtures contained 12.5 μL SYBR Green qPCR Super-Mix-UDG with Rox (Invitrogen, USA), 0.5 μL each of 10 μM forward and reverse primers, 10.5 μL DNA-free water, and 1.0 μL standard plasmid or DNA extract. The cycling conditions for qPCR amplification were as follows: 50 °C for 2 min, 95 °C for 5 min; followed by 40 cycles of 95 °C for 20 s, annealing temperature for 30 s, 72 °C for 31 s, and plate reading. Melt-curve analyses were performed from 60 to 95 °C, with 0.2 °C/read. Reaction was performed using an ABI Real-time PCR system 7500 (ABI, USA). Melting curve analysis and gel electrophoresis ensured specificity. Each gene was quantified in triplicate using a standard curve and a negative control.
Statistical analysis
Independent t tests were conducted to detect significant differences between the target genes. Differences with a p value <0.05 were considered statistically significant. The Spearman rank correlation was used to assess the association between ARGs and intI1. Statistical analyses were performed using SPSS 20.0 (IBM, USA).
Results and discussion
Seasonal variation of ARGs abundance
Target genes were detected in almost all wastewater samples, as shown in Fig. 2. The abundance of each ARG and 16S rRNA gene in the influent (raw swine wastewater) of the two swine farms was similar for each sampling season. The abundance of tetM, tetX, ermB, ermF, and mefA was relatively higher than that of other detected ARGs, with mean abundances of 6.91E+10, 2.43E+11, 1.07E+11, 1.45E+11, and 7.16E+10 copies/mL in raw swine wastewater, respectively. The mean abundance of tetG and ereA was at least 2 logs lower, at 6.02E+09 and 8.27E+08 copies/mL, respectively. The reported abundance of tetracycline and macrolide resistance genes was lower than this study, including tetA of 9.1E+03, tetW of 8.0E+04 (Tao et al. 2014). tetG of 2.0E+07 copies/mL (Chen et al. 2010). ermB of 6.3E+08, ermF of 1.0E+08, and ermX of 7.9E+05 copies/mL (Chen et al. 2010). In this study, filtration of samples prior to DNA extraction may have increased the DNA extraction efficiency compared with the wet material centrifugation method adopted by other studies (Tao et al. 2014; Chen et al. 2010). In general, the abundance of ARGs in the raw swine wastewater may reflect the feeding status, antibiotic consumption, and state of antibiotic resistance contamination. The abundance of ARGs detected in these samples confirms that a more rational use of antibiotics in swine farming is essential, which is reiterated by WHO (2014).
Abundance of ARGs and 16S rRNA gene in swine wastewater treatment systems (a tetG; b tetM; c tetX; d ermB; e ermF; f ereA; g mefA; h 16S rRNA gene; *p < 0.05, significant difference between values of summer and winter; USR upflow solid reactor, PFR plug-flow reactor, SBR sequencing batch reactor, BF biofilter, WL wetland, inf influent sample, eff effluent sample)
The abundance of all the genes quantified in summer was significantly different (p < 0.05) from winter, indicating that season plays an important role in ARGs abundance. In most cases, the ARGs and 16S rRNA genes of samples were more abundant in winter than in summer. The mean difference and range of variation in target genes were 1.19 log (0.09–2.35 log) for tetG, 1.14 log (0.47–3.40 log) for tetM, 1.10 log (0.25–5.35 log) for tetX, 0.83 log (0.32–2.48 log) for ermB, 1.34 log (0.21–5.87 log) for ermF, 1.16 log (0.47–3.24 log) for ereA, 1.16 log (0.41–3.11 log) for mefA, and 1.09 log (0.29–2.34 log) for 16S rRNA gene, respectively, with the range of average variation of 0.31–3.52 log in summer and winter.
However, all detected genes in the lagoon 2 effluent from farm 1 were more abundant in summer, which was similar to the BF effluent from farm 2. Chen et al. (2010) evaluated the seasonal variation of macrolide resistance genes, including ermB and ermF, and showed that these are more abundant in winter than in summer. However, after anaerobic digestion and BF treatments, the abundance of ermB and ermF was higher in summer than in winter. These variations in ermB and ermF were inconsistent with our findings which suggested other currently unknown mechanisms may be at work in causing variation in these ARGs levels.
Veterinary antibiotics are used in many forms, including oral solutions, oral powders, premixed with feed, injections, etc. In addition, the impact on ARGs abundance in manure of antibiotics administered in feed is quite different than those administered by injection (Bibbal et al. 2007). The higher abundance of ARGs in winter than in summer may be due to increased antibiotic administration during the cold season to prevent disease (Awad et al. 2014). And the antibiotics administered between summer and winter greatly impacted the seasonal ARGs abundance. Additionally, as shown in Fig. 2, higher population of total bacteria (copies of 16S rRNA gene) was in winter. Given that animals are kept inside barns for heat preservation during the winter, decreased ventilation and less frequent cleaning may result in discharging less wastewater with a higher concentration of pollutants, such as higher concentrations of COD, ammonia, nitrogen, and bacteria.
ARGs removal efficiencies by different treatment processes
As shown in Fig. 2, in most cases, the abundance of ARGs was effectively reduced by most treatment processes, including anaerobic digestion, sequencing batch reactor (SBR), and wetland. Removal of ARGs was more efficient in winter than in summer; however, the lagoons and the biofilter increased the abundance of ARGs, especially in summer. Therefore, further research is needed, as this may be due to the proliferation of ARG carriers in summer.
The abundance of ARGs was significantly correlated with the 16S copy number (p < 0.01), and the efficiency of ARG removal was greatly influenced by the effects of treatments on total bacterial numbers. To observe effects on ARGs specifically in each treatment process, the relative abundance of ARGs in the swine wastewater treatment systems, normalized to 16S rRNA gene, is shown in Fig. 3. In raw swine wastewater, tetM and tetX were the highly abundant tetracycline resistance genes in this study. Cheng et al. (2013) reported that ribosomal protection genes (tetQ, tetM, tetW, tetO) were more abundant than other tetracycline resistance genes in swine wastewater, with 5.53 × 10−2 copies of tetM/16S rRNA. Li et al. (2015) proposed that tetM has a strong correlation with other ARGs and has been proposed as an indicator of co-occurring ARGs in environment. In this study, the relative abundance of tetM was high in raw swine wastewater and was correlated with the abundance of macrolide resistance genes, including ermB, ermF, and mefA (p < 0.05). The tetX system, encoding tetracycline inactivation enzyme, had a similar magnitude to tetM in raw swine wastewater on the two investigated farms. After anaerobic digestion, the relative abundance of tetracycline resistance genes shifted; while tetM and tetX still dominated, with tetX slightly decreased and tetM increased in most cases, the relative abundance of tetG constantly increased. Unlike the changes seen in absolute abundance of ARGs, the relative abundance of ARGs after biological treatments did not show consistent seasonal trend.
Relative abundance of ARGs (normalized to 16S rRNA gene) in swine wastewater treatment systems (a tet in summer; b tet in winter; c ARGs of macrolide in summer; d ARGs of macrolide in winter; USR upflow solid reactor, PFR plug-flow reactor, SBR sequencing batch reactor, BF biofilter, WL wetland, inf influent sample, eff effluent sample)
After anaerobic digestion, the relative abundance of tetM was increased by 0.14–0.54 log and tetX reduced by 0.11–0.96 log, while tetM in the lagoon system was reduced by 0.06–0.27 log. In most cases, the relative abundance of tetX was increased by 0.28–0.89 log after lagoon treatment. The relative abundance of tetG was constantly increased by 0.22–0.60 log and 0.03–1.31 log after anaerobic digestion and lagoon treatments, respectively. But for SBR, BF, and WL systems on farm 2, the variations of tetM, tetX, and tetG were inconsistent in summer and winter. Diehl and Lapara (2010) reported that the reductions of tetA, tetO, tetW, and tetX over a 5-day anaerobic digestion fit a first-order kinetic model, but aerobic processes had a less substantial effect on removal of tet genes compared with the anaerobic process. The increase in relative abundance of tetG after biological treatment of swine wastewater was also reported by Cheng et al. (2013) and Chen et al. (2015). Interestingly, tetG was not the dominant ARGs in raw swine wastewater, but increased in relative abundance after biological treatment, which could be due to an increase in abundance of the microbial community harboring the tetG gene. Salmonella spp. is the primary potential host of tetG, which poses a high risk to the environment in effluent discharged from swine farms (Adesiji et al. 2014). Although Salmonella is vulnerable to biological treatment, tetG still increased; therefore, Salmonella alone cannot explain the occurrence of tetG, and further study is needed into tetG carriers and antibiotic-resistant pathogens. For the total tetracycline resistance genes detected (tetG, tetM, and tetX), the effluents of the two wastewater treatment systems showed an increase in relative abundance, comparing with raw swine wastewater, especially in summer. The normalized abundance of total tet genes decreased only in the effluent sample of farm 2 during in winter.
For macrolide resistance genes as shown in Fig. 3, ermB, ermF, and mefA were the dominant ARGs in raw swine wastewater, whereas ereA showed the lowest relative abundance. ermB and mefA, encoding 23S rRNA methyltransferase and an efflux protein, respectively, showed a decline in relative abundance during the anaerobic, lagoon, and aerobic treatments on the two swine farms, both in summer and winter. ermB is one of the most common macrolide resistance genes detected in swine wastewater, with relatively high abundance of 1.2E−01 copies/16S rRNA (Joy et al. 2013; Joy et al. 2014). mefA was always detected in Streptococci and Enterococci isolated from pig carcasses (Martel et al. 2003). but its abundance has only been rarely quantified in swine wastewater. ermF is another 23S rRNA methylation gene and was always detected with high relative abundance of 2.2E−03 (Joy et al. 2014) and 6.04E−04 copies/16S rRNA (Brooks et al. 2014) in swine wastewater. ereA is not the dominant macrolide resistance genes in wastewater; Pei et al. (2007) detected ereA in dairy wastewater with a low relative abundance of 5.95E−6 copies/16S rRNA (Pei et al. 2007).
In this study, the relative abundance of ermB was effectively reduced after anaerobic digestion, lagoon, and SBR treatments, by 0.08–0.76 log, 0.24–0.83 log, and 0.65–1.71 log, respectively. In most cases, ermF was reduced after anaerobic digestion, lagoon, SBR, and constructed wetland treatments, by 0.57–0.98 log, 0.16–0.94 log, 2.16–5.56 log, and 0.32–0.55 log, respectively. Joy et al. (2014) reported that the relative abundance of ermB and ermF decreased by 0.5 log and 1.5 log, respectively, at 37 °C in a 100-mL storage reactor after 40 days. These results suggest that the variation of ARGs under real environmental conditions can be quite different from those in a controlled experiment.
The relative abundance of ereA increased after anaerobic digestion and lagoon treatment by 0.09–1.41 log and 0.06–0.40 log, respectively, but did not show a constant trend in the “SBR+BF+WL” system. While ereA may not be a prevailing ARG in animal wastewater, its relative abundance increased after biological treatment observed by Pei et al. (2007) in a study with aerobic or anaerobic incubation of dairy wastewater for 10–20 days. In most cases, mefA was effectively removed after anaerobic digestion and lagoon treatments, with reduction of 0.28–0.74 log and 0.10–0.29 log. The total relative abundance of all the macrolide resistance genes (ermB, ermF, ereA and mefA) in the effluent was lower in winter than that of the influent. However, removal of macrolide resistance genes in summer was not obvious.
Removal of ARGs by the entire treatment system
Swine wastewater is generally treated by combinations of technologies, including anaerobic and aerobic processes. The occurrence of ARGs in the two biological wastewater treatment systems tested is shown in Fig. 4a. Treatment removed more ARGs in winter than in summer in most cases; in both systems, some ARGs were only reduced in winter, including tetG and ereA. Interestingly, these two ARGs proliferated in relative abundance after biological wastewater treatment. Some ARGs may have differences in variation of abundance and relative abundance after wastewater treatment, which was also reported by Li et al. (2014). In these two wastewater treatment systems, tetM, ermB, and mefA were all effectively removed, with 0.95, 1.43, and 1.08 log removed in summer, and 1.43, 2.01, and 1.82 log removed in winter, respectively. If we compare the two wastewater treatment systems in this study, the system of farm 2 was more effective than that of farm 1, except for tetG and ereA. However, these were not the predominant ARGs for tetracycline and macrolide resistance. tetX was only removed in the wastewater treatment system of farm 2 and not affected by the lagoon system of farm 1. Barkovskii et al. (2012) indicated that tetX was a persistent ARG in swine wastewater lagoons with an abundance of 106–107 copies/mL. This study indicated that a combination of anaerobic digestion with aerobic biological and constructed wetland processes may be effective for tetX removal.
Absolute and relative logarithmic removal of target genes in swine wastewater treatment systems (a absolute abundance removal; b relative abundance removal; Farm1-S summer samples from Farm 1, Farm1-W winter samples from Farm 1, Farm2-S summer samples from Farm 2, Farm2-W winter samples from Farm 2)
The decrease in relative abundance of ARGs (normalized to 16S rRNA copy number) is shown in Fig. 4b. In most cases, the relative log removal of ARGs during the entire treatment process was higher in winter than in summer. The relative abundance of tetM was reduced only by the wastewater treatment system of farm 2. The relative abundance of tetM increased after lagoon treatment on farm 1, indicating swine wastewater stored in lagoons may increase bacterial proliferation resulting in amplification of the tetM gene, which was also reported by Barkovskii et al. (2012). The declining trend of relative abundance of tetX and ermF was greatly influenced by the season. The relative abundance of tetX and ermF was only reduced in winter, whereas their relative abundance in summer increased. The ermF variation trend between seasons was consistent with the results of Chen et al. (2010). indicating ermF could be effectively reduced in winter. Higher ambient temperatures in summer may make the removal of some kinds of antibiotic resistant bacteria (ARB) more difficult; however, this is another field that requires further research.
The abundance of ermB and mefA consistently declined in two swine wastewater treatment systems, with an average log removal of 0.97 and 0.61 in summer and 1.44 and 1.03 in winter, respectively. However, the relative abundances of tetG and ereA increased after wastewater treatment in both seasons. This could be due to proliferation of the ARB carrying tetG and ereA during the wastewater treatment process or horizontal transfer of the ARGs to other species (Li et al. 2014). The relative abundance of tetG and ereA was significantly correlated with the relative abundance of intI1 (p < 0.01), and there was also a significant correlation between the relative abundance of tetG and ereA (p < 0.01). Du et al. (2014) reported a consistent variation trend between tetG and intI1 during wastewater treatment. Integrons could facilitate the horizontal transfer of ARGs (Martínez 2008; Shi et al. 2013). Previous studies have indicated that tetG and ereA are all inducible ARGs co-occurring with the class 1 integron (Cheng et al. 2013; Shahada et al. 2007; Sung and Oh 2014). Shahada et al. (2007) indicated that tetG and class 1 integron were simultaneously harbored by Salmonella enterica serotype Typhimurium isolated from bovine. Sung and Oh (2014) reported that class 1 integron carrying cassette arrays of dfrA32-ereA-aadA2 were harbored by Proteus mirabilis and Proteus penneri belonging to Enterobacteriaceae isolated from chickens. Class 1 integron is capable of transferring resistance by conjugation between different species of Enterobacteriaceae, which have been identified in isolates from municipal wastewater treatment plant (Mokracka et al. 2012). In addition, Enterobacteriaceae are at high prevalence in animal wastewater and some species of Enterobacteriaceae are important human pathogens (Resende et al. 2014b). The antibiotic resistance of Enterobacteriaceae and the horizontal transfer of ARGs could pose public health risks and needs further study.
Conclusions
ARGs for tetracycline (tetG, tetM, and tetX) and macrolide (ermB, ermF, ereA, and mefA) were quantified in two typical swine wastewater treatment systems, i.e., an “anaerobic digestion + lagoon” system and an “anaerobic digestion + aerobic biological treatment+ wetland” system in both winter and summer. The abundance of ARGs in winter was higher than in summer, with an average variation of 0.31–3.52 log between summer and winter. tetM, tetX, ermB, ermF, and mefA were highly abundant in raw swine wastewater. ARGs abundance was effectively reduced by most of the individual treatment processes, including anaerobic digestion, SBR, and wetlands, with more efficient removal in winter than summer; however, the relative abundance of ARGs was quite variable. Anaerobic digestion decreased the relative abundance of tetX, ermB, ermF, and mefA; lagoon treatment decreased tetM, ermB, ermF, and mefA; and SBR decreased tetM, ermB, and ermF. However, biofilter and constructed wetland did not display consistent removal efficacy on ARGs over two sampling seasons. ermB and mefA were effectively reduced during both winter and summer, not only in abundance but also in relative abundance. tetG and ereA were not the dominant ARGs in raw swine wastewater but all increased after wastewater treatment. The relative abundance of tetG and ereA was significantly correlated with intI1 (p < 0.01) implying horizontal transfer of ARGs, which poses a great threat to public health. Further research is needed to determine the effective method for controlling ARGs.
References
Adesiji YO, Deekshit VK, Karunasagar I (2014) Antimicrobial-resistant genes associated with Salmonella spp. isolated from human, poultry, and seafood sources. Food Sci Nutr 2:436–442. doi:10.1002/fsn3.119
Amos GC, Gozzard E, Carter CE, et al. (2015) Validated predictive modelling of the environmental resistome. ISME J 1–10. doi:10.1038/ismej.2014.237
Apley MD, Bush EJ, Morrison RB et al (2012) Use estimates of in-feed antimicrobials in swine production in the United States. Foodborne Pathog Dis 9:272–279. doi:10.1089/fpd.2011.0983
Awad YM, Kim SC, Abd El-Azeem SAM et al (2014) Veterinary antibiotics contamination in water, sediment, and soil near a swine manure composting facility. Environ Earth Sci. doi:10.1007/s12665-013-2548-z
Barkovskii AL, Manoylov KM, Bridges C (2012) Positive and negative selection towards tetracycline resistance genes in manure treatment lagoons. J Appl Microbiol 112:907–919. doi:10.1111/j.1365-2672.2012.05252.x
Barton MD (2000) Antibiotic use in animal feed and its impact on human health. Nutr Res Rev 13:279–299. doi:10.1079/095442200108729106
Berendonk TU, Manaia CM, Merlin C, et al. (2015) Tackling antibiotic resistance: the environmental framework. Nat Rev Microbiol 1–8. doi:10.1038/nrmicro3439
Bibbal D, Dupouy V, Ferré JP et al (2007) Impact of three ampicillin dosage regimens on selection of ampicillin resistance in Enterobacteriaceae and excretion of blaTEM genes in swine feces. Appl Environ Microbiol 73:4785–4790. doi:10.1128/AEM.00252-07
Brooks JP, Adeli A, McLaughlin MR (2014) Microbial ecology, bacterial pathogens, and antibiotic resistant genes in swine manure wastewater as influenced by three swine management systems. Water Res 57:96–103. doi:10.1016/j.watres.2014.03.017
Chantziaras I, Boyen F, Callens B, Dewulf J (2014) Correlation between veterinary antimicrobial use and antimicrobial resistance in food-producing animals: a report on seven countries. J Antimicrob Chemother 69:827–834. doi:10.1093/jac/dkt443
Chen J, Yu Z, Michel FC et al (2007) Development and application of real-time PCR assays for quantification of erm genes conferring resistance to macrolides-lincosamides-streptogramin B in livestock manure and manure management systems. Appl Environ Microbiol 73:4407–4416. doi:10.1128/AEM.02799-06
Chen J, Michel FC, Sreevatsan S et al (2010) Occurrence and persistence of erythromycin resistance genes (erm) and tetracycline resistance genes (tet) in waste treatment systems on swine farms. Microb Ecol 60:479–486. doi:10.1007/s00248-010-9634-5
Chen B, Hao L, Guo X et al (2015) Prevalence of antibiotic resistance genes of wastewater and surface water in livestock farms of Jiangsu Province. China Environ Sci Pollut Res. doi:10.1007/s11356-015-4636-y
Cheng W, Chen H, Su C, Yan S (2013) Abundance and persistence of antibiotic resistance genes in livestock farms: a comprehensive investigation in eastern China. Environ Int. doi:10.1016/j.envint.2013.08.023
Cronk JK (1996) Constructed wetlands to treat wastewater from dairy and swine operations: a review. Agric Ecosyst Environ 58:97–114. doi:10.1016/0167-8809(96)01024-9
Deng L, Zheng P, Chen Z, Mahmood Q (2008) Improvement in post-treatment of digested swine wastewater. Bioresour Technol 99:3136–3145. doi:10.1016/j.biortech.2007.05.061
Diehl DL, Lapara TM (2010) Effect of temperature on the fate of genes encoding tetracycline resistance and the integrase of class 1 integrons within anaerobic and aerobic digesters treating municipal wastewater solids. Environ Sci Technol 44:9128–9133. doi:10.1021/es102765a
Du J, Ren H, Geng J et al (2014) Occurrence and abundance of tetracycline, sulfonamide resistance genes, and class 1 integron in five wastewater treatment plants. Environ Sci Pollut Res 21:7276–7284. doi:10.1007/s11356-014-2613-5
Jindal A, Kocherginskaya S, Mehboob A et al (2006) Antimicrobial use and resistance in swine waste treatment systems. Appl Environ Microbiol 72:7813–7820. doi:10.1128/AEM.01087-06
Joy SR, Bartelt-Hunt SL, Snow DD et al (2013) Fate and transport of antimicrobials and antimicrobial resistance genes in soil and runoff following land application of swine manure slurry. Environ Sci Technol. doi:10.1021/es4026358
Joy SR, Li X, Snow DD et al (2014) Fate of antimicrobials and antimicrobial resistance genes in simulated swine manure storage. Sci Total Environ 481:69–74. doi:10.1016/j.scitotenv.2014.02.027
Koike S, Krapac IG, Oliver HD et al (2007) Monitoring and source tracking of tetracycline resistance genes in lagoons and groundwater adjacent to swine production facilities over a 3-year period. Appl Environ Microbiol 73:4813–4823. doi:10.1128/AEM.00665-07
Li J, Cheng W, Xu L et al (2014) Antibiotic-resistant genes and antibiotic-resistant bacteria in the effluent of urban residential areas, hospitals, and a municipal wastewater treatment plant system. Environ Sci Pollut Res 22:4587–4596. doi:10.1007/s11356-014-3665-2
Li B, Yang Y, Ma L, et al. (2015) Metagenomic and network analysis reveal wide distribution and co-occurrence of environmental antibiotic resistance genes. ISME J 1–13. doi: 10.1038/ismej.2015.59
Martel A, Devriese LA, Decostere A, Haesebrouck F (2003) Presence of macrolide resistance genes in streptococci and enterococci isolated from pigs and pork carcasses. Int J Food Microbiol 84:27–32. doi:10.1016/S0168-1605(02)00390-2
Martínez JL (2008) Antibiotics and antibiotic resistance genes in natural environments. Science 321:365–367. doi:10.1126/science.1159483
Martínez JL, Coque TM, Baquero F (2014) What is a resistance gene? Ranking risk in resistomes. Nat Publ Gr 1–8. doi: 10.1038/nrmicro3399
Mokracka J, Koczura R, Kaznowski A (2012) Multiresistant Enterobacteriaceae with class 1 and class 2 integrons in a municipal wastewater treatment plant. Water Res 46:3353–3363. doi:10.1016/j.watres.2012.03.037
Nasir IM, Mohd Ghazi TI, Omar R (2012) Anaerobic digestion technology in livestock manure treatment for biogas production: a review. Eng Life Sci 12:258–269. doi:10.1002/elsc.201100150
Ng LK, Martin I, Alfa M, Mulvey M (2001) Multiplex PCR for the detection of tetracycline resistant genes. Mol Cell Probes 15:209–215. doi:10.1006/mcpr.2001.0363
Pei R, Cha J, Carlson KH, Pruden A (2007) Response of antibiotic resistance genes (ARG) to biological treatment in dairy lagoon water. Environ Sci Technol 41:5108–5113. doi:10.1021/es070051x
Pruden A, Pei R, Storteboom H, Carlson KH (2006) Antibiotic resistance genes as emerging contaminants: studies in northern Colorado. Environ Sci Technol 40:7445–7450. doi:10.1021/es060413l
Resende JA, Diniz CG, Silva VL et al (2014a) Dynamics of antibiotic resistance genes and presence of putative pathogens during ambient temperature anaerobic digestion. J Appl Microbiol 117:1689–1699. doi:10.1111/jam.12653
Resende JA, Silva VL, de Oliveira TLR et al (2014b) Prevalence and persistence of potentially pathogenic and antibiotic resistant bacteria during anaerobic digestion treatment of cattle manure. Bioresour Technol 153:284–291. doi:10.1016/j.biortech.2013.12.007
Shahada F, Amamoto A, Chuma T et al (2007) Antimicrobial susceptibility phenotypes, resistance determinants and DNA fingerprints of Salmonella enterica serotype Typhimurium isolated from bovine in Southern Japan. Int J Antimicrob Agents 30:150–156. doi:10.1016/j.ijantimicag.2007.03.017
Shi P, Jia S, Zhang XX et al (2013) Metagenomic insights into chlorination effects on microbial antibiotic resistance in drinking water. Water Res 47:111–120. doi:10.1016/j.watres.2012.09.046
Stokes HW, Nesbø CL, Holley M, et al (2006) Class 1 integrons potentially predating the association with tn402-like transposition genes are present in a sediment microbial community. J Bacteriol 188:5722–5730. doi:10.1128/JB.01950-05
Sung J, Oh J (2014) Distribution and characterization of integrons in Enterobacteriaceae isolates from chickens in Korea. 24:1008–1013. doi:10.4014/jmb.1404.04058
Sutcliffe J, Grebe T, Tait-Kamradt A, Wondrack L (1996) Detection of erythromycin-resistant determinants by PCR. Antimicrob Agents Chemother 40:2562–2566
Suzuki MT, Taylor LT, DeLong EF (2000) Quantitative analysis of small-subunit rRNA genes in mixed microbial populations via 5’-nuclease assays. Appl Environ Microbiol 66:4605–4614. doi:10.1128/AEM.66.11.4605-4614.2000
Tao CW, Hsu BM, Ji WT et al (2014) Evaluation of five antibiotic resistance genes in wastewater treatment systems of swine farms by real-time PCR. Sci Total Environ 496:116–121. doi:10.1016/j.scitotenv.2014.07.024
Volokhov D, Chizhikov V, Chumakov K, Rasooly A (2003) Microarray analysis of erythromycin resistance determinants. J Appl Microbiol 95:787–798. doi:10.1046/j.1365-2672.2003.02046.x
WHO (2014) Antimicrobial resistance: global report on surveillance
Zhang Q-Q, Ying G-G, Pan C-G et al (2015) A comprehensive evaluation of antibiotics emission and fate in the river basins of China: source analysis, multimedia modelling, and linkage to bacterial resistance. Environ Sci Technol 49:6772–6782. doi:10.1021/acs.est.5b00729
Zhu Y-G, Johnson TA, Su J-Q et al (2013) Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc Natl Acad Sci U S A. doi:10.1073/pnas.1222743110
Acknowledgments
This work is financially supported by Special Fund for Agro-scientific Research in the Public Interest (No. 201303091), the National Major Science & Technology Projects for Water Pollution Control and Management (No. 2015ZX07203-007), and the National Natural Science Foundation of China (No. 41501513 and No. 21577161).
Authors’ contributions
The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Responsible editor: Gerald Thouand
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 14 kb)
Rights and permissions
About this article
Cite this article
Sui, Q., Zhang, J., Tong, J. et al. Seasonal variation and removal efficiency of antibiotic resistance genes during wastewater treatment of swine farms. Environ Sci Pollut Res 24, 9048–9057 (2017). https://doi.org/10.1007/s11356-015-5891-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-015-5891-7