Abstract
Introduction
Metabolomics applications to the aquaculture research are increasing steadily. The use of standardized proton nuclear magnetic resonance (1H NMR) spectroscopy can provide the aquaculture industry with an unbiased, reproducible, and high-throughput screening tool, which can help to diagnose nutritional and disease-related metabolic disorders in farmed fish.
Objective
Standard operating procedures developed for analysing (human) plasma by 1H NMR were applied to fingerprint the metabolome in plasma samples collected from Atlantic salmon. The aim was to explore the metabolome of salmon plasma in relation to growth stage and sampling site.
Methods
A total of 72 salmon were collected from three aquaculture sites in Norway (Lat. 65, 67, and 70 °N) and over two sampling events (December 2017 and November 2018). Plasma drawn from each salmon was measured by 1H NMR and metabolites were quantified using the SigMa software. The NMR data was analysed by principal component analysis (PCA) and ANOVA-simultaneous component analysis (ASCA).
Results
Important metabolic differences were evidenced, with adult salmon having a much higher content of very low-density lipoproteins and cholesterol in their plasma, while smolts displayed significantly higher levels of propylene glycol. Overall, 24% of the metabolite variation was due to the growth stage, whereas 12% of the metabolite variation was related to the aquaculture site and practice (p < 0.001).
Conclusion
This study provides a baseline investigation of the plasma metabolome of the Atlantic salmon and demonstrates how 1H NMR metabolomics can be used in future investigations for comparing aquaculture practices and their influence on the fish metabolome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
In the era of sustainable food production, responsible aquaculture stands as a valuable solution to meet the global demand for food and animal proteins. The FAO’s state of world fisheries and aquaculture underlines the significant and rising role of fisheries and aquaculture in providing food, nutrition and employment (FAO, 2020). Aquaculture has a significant economic importance in many countries worldwide including Egypt, Chile, India, Indonesia, Vietnam, Bangladesh, and Norway (FAO, 2020). In particular, Atlantic salmon aquaculture, led by Norway and Chile, is one of the most profitable and technologically advanced fish production industries globally (FAO, 2020). Atlantic salmon, which has traditionally been harvested in the Norwegian coasts, has a significant social, cultural, and economic importance for Norwegians (Liu et al. 2011). Despite undeniable thriving of the sector, fish health management still represents a major challenge. Moreover, the aquaculture industry occasionally reports compromised gut function in farmed salmon and early research suggests that increasing levels of plant ingredients in feeds may be an important contributing factor (Penn et al. 2011).
Over the last decades, haematological tests and blood analyses have proven to be useful tools in modern intensive aquaculture systems, where several factors, including the increased use of plant-based feed in the diets of carnivorous species, may cause increased susceptibility of fish to nutritional challenges (Belghit et al. 2019; Kortner et al. 2016; Krogdahl et al. 2015; Sandnes et al. 1988). As it circulates through the entire body, blood can be considered a window to the physiological and/or pathological condition of the individual (Robinson, 2012). For this reason, blood testing has become one of the most informative screening methods in metabolomics investigations on humans and animals (livestock and aquaculture) (Alfaro & Young, 2018; Roques et al. 2020; Serkova et al. 2011; Young & Alfaro, 2016). Modern analytical techniques routinely used to analyse blood plasma and serum include chromatographic and spectroscopic methods, bioassays, and ultra-centrifugation. More recently, the use of non-destructive and untargeted analytical approaches has found extensive applications in the clinical metabolomics field. Due to the exceptional reproducibility and the ease of sample preparation and measurement, proton nuclear magnetic resonance (1H NMR) spectroscopy has become a valuable tool for blood plasma and serum-based metabolomics investigations (Dona et al. 2014). However, NMR has limited sensitivity compared to other analytical methods such as liquid chromatography- mass spectrometry (LC–MS), which can provide deeper insights into the low concentration components of the metabolome (Chaleckis et al. 2019; Xiao et al. 2012). Yet, in two aspects, 1H NMR spectroscopy has a leading edge: (1) it is unbiased and inherently quantitative and (2) its sensitivity to physical phenomena, such as diffusion, makes it a valuable analytical method for the analysis of intact lipoprotein (LP) fractions and sub-fractions (Aru et al. 2017; Cushley & Okon, 2002; Jiménez et al. 2018; Monsonis Centelles et al. 2017; Savorani et al. 2013). In order to increase reproducibility while ensuring robust quantitative information, protocols for the analysis of plasma and serum by 1H NMR have undergone extreme standardization. In particular, standard operating procedures (SOPs) have been developed for fine-tuning the analytical steps from sample collection and storage to sample measurement (Dona et al. 2014). Lately, standardization has been extended to metabolites identification and quantification and has culminated in the development of dedicated software, such as SigMa (Khakimov et al. 2020).
The analysis of blood plasma and serum by 1H NMR metabolomics has proven to be a remarkable screening method in human diagnostic medicine (Emwas et al. 2013; Erben et al. 2018; Powell et al. 2018) and for investigating important clinical conditions, such as cancer (Giskeødegård et al. 2018; Lima et al. 2018; Spratlin et al. 2009), diabetes (Lanza et al. 2010) and Parkinson’s disease (Pizarro et al. 2019). While NMR metabolomics applications in human health and medical sciences continue to feature numerous high quality research, the powerful analytical potential of 1H NMR spectroscopy has led to the expansion of blood-based metabolomics applications also to the veterinarian and animal breeding (livestock and aquaculture) fields (Başoǧlu et al. 2014; Bertram et al. 2006; Bin et al. 2018; Dove et al. 2012; Jones & Cheung, 2007; Le Moyec et al. 2014; Luck et al. 2015; Niemuth et al. 2018; Shestakova et al. 2018; Ziarrusta et al. 2018). In particular, 1H NMR metabolomics is establishing itself as an important tool for the assessment of the most relevant issues in the aquaculture industry, including fish health and alternative feed development (Alfaro & Young, 2018; Young & Alfaro, 2016). Recently, major efforts have been put into the optimization of fish feed for a more responsible aquaculture and for securing feed raw material flexibility, and specifically to introduce alternative and sustainable, mostly plant-based, feed ingredients (Aas et al. 2019; Gatlin et al. 2007; Tacon & Metian, 2008). In this context, NMR spectroscopy may serve as an efficient and high-throughput platform for investigating the fish metabolome as a function of feed composition, especially when introducing new alternative ingredients into the fish meal, such as plant proteins, insects and algae, which may impact fish metabolism and health (Krogdahl et al. 2010; Li et al. 2019; Roques et al. 2018).
In this work, the most recent SOPs developed for the analysis of human plasma and serum by 1H NMR spectroscopy were applied for the first time to analyse plasma samples from farmed Atlantic salmon (Salmo salar, L.) collected from three aquaculture sites along the Norwegian coast at different latitudes (Lat. 65°N, 67°N, and 70°N) over two sampling events (December 2017 and November 2018). The aims of the study were to (1) describe the baseline plasma metabolome of farmed Atlantic salmon as measured by standardized 1H NMR, (2) compare the spectral features of salmon plasma with a reference 1H NMR spectrum of human plasma measured using the same SOPs, and (3) describe the salmon plasma metabolome as a function of salmon growth stage and aquaculture site/practice.
2 Methods
2.1 Chemicals
Analytical grade sodium phosphate monobasic monohydrate (NaH2PO3, H2O), sodium phosphate dibasic heptahydrate (Na2HPO3, 7 H2O), deuterium oxide (D2O, 99.9%), and sodium 3-trimethylsilyl-propionate-2,2,3,3-d4 (TSP) were purchased from Sigma–Aldrich (Merk KGaA, Darmstadt, Germany).
2.2 Sampling
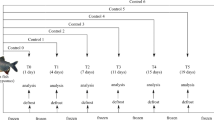
A total of 72 mixed-gender Atlantic salmon were collected along the Norwegian coast (Lat. 65°N, 67°N, and 70°N) from three commercial aquaculture farms run by three different companies, and over two sampling events (December 2017 and November 2018). Each sampling event corresponded to a specific growth stage of the salmon, namely smolt (1st sampling event, December 2017) and adult (2nd sampling event, November 2018). Smolts were supplied by different external companies. Samples collected over the two sampling events were from the same cohort, i.e., the same net pen (average net size: 140 m circumference and 40 m deep). Fish were caught from the net pen by using a sweep-net. After collection, fish were placed in a tank of about 400 l supplied with continuously running seawater. From this holding tank, fish were individually dip-netted and sacrificed by anesthetic overdose of tricaine methane sulfonate (MS222, Argent Chemical Laboratories, WA, USA). Blood was drawn from the caudal vein of each salmon and collected into sterile vacutainers containing EDTA. Plasma was obtained by centrifuging the blood samples at 4 °C and 1000 rpm for 15 min. After collection, plasma samples were immediately snap-frozen using liquid nitrogen and stored at −80 °C until NMR measurements. The experiments complied with the guidelines of the Norwegian Regulation on Animal Experimentation and European Community Directive 86/609/EEC. An overview of the experimental design is given in Fig. 1. Salmon biometries and overall diet compositions are given in Tables S1 and S2.
Experimental design. A total of 72 fish were collected along the Norwegian coast from three aquaculture sites run by three different companies. Samples from the same cohort were collected over two sampling events in December 2017 and November 2018, which corresponded to two growth stages of salmon, namely smolt and adult, respectively. Blood plasma was collected from each salmon and analysed using SOPs and 1H NMR
2.3 1H NMR spectroscopy
2.3.1 Sample preparation
Sample preparation was performed according to the SOPs described by Dona et al. (Dona et al. 2014). Briefly, for each sample, 350 μl of phosphate buffer solution (Beckonert et al. 2007) containing TSP (5 mM) and D2O (20%) were transferred to a 2 ml Eppendorf and gently mixed with equal amounts of plasma. Aliquots of 600 μl were then transferred into 5 mm O.D. SampleJet NMR tubes (Bruker BioSpin, Ettlingen, Germany).
2.3.2 1H NMR measurements
Plasma samples for metabolomics analyses were measured using SOPs largely relying on standardized and controlled data acquisition and data processing (Dona et al. 2014; Monsonis Centelles et al. 2017). Plasma samples were measured on a Bruker Avance III 600 operating at a proton’s Larmor frequency of 600.13 MHz. The spectrometer was equipped with a 5-mm broadband inverse (BBI) probe and an automated sample changer (SampleJet™, Bruker Biospin, Ettlingen, Germany). The SampleJet was equipped with a refrigerated sample storage station with cooling racks (278 K) and heating/drying station (306 K). Cooling of probe was controlled by the BCU (Bruker Cooling Unit), with temperature stability of the order of 0.01 K. Data acquisition and processing were carried out in the TopSpin software (version 3.5, Bruker, Rheinstetten, Germany). Automation of the overall measurement procedure was controlled by iconNMR™ (Bruker Biospin, Rheinstetten, Germany). Proton NMR spectra were measured using the pulse sequence for water suppression 1D NOESY (noesygppr1d, Bruker nomenclature) and 32 scans, which were collected into 131,072 data points using a spectral width of 30 ppm, a 90° pulse, and a recycle delay (d1) of 4 s. The receiver gain value was determined experimentally and kept constant for all samples (90.5). After Fourier Transformation (FT), automatic phasing and baseline correction were performed for all samples, followed by exponential filtering (line broadening LB = 0.3 Hz). The reader is referred to the manuscript by Dona et al. (2014) for further details on the instrument calibration and NMR measurements (Dona et al. 2014).
To complement our metabolomics study, we performed a side study and proof-of-concept investigation aimed at assessing the impact of increasing the experimental temperature on the metabolome of salmon plasma as measured by 1H NMR. For this purpose, three representative plasma samples were measured at seven temperatures, namely 280 K, 287 K, 294 K, 300 K, 310 K, 318 K and 324 K. Measurements were performed using the same equipment as described above. In order to quantitatively assess the metabolic variations occurring in plasma as a result of temperature increase, eight representative metabolites covering the entire spectral region from high- to low-field were quantified from the 1H NMR spectra. These metabolites include cholesterol, valine, lactate, alanine, glucose, tyrosine, phenylalanine, and formate. More details on the variable temperature experiments can be found in the Supporting Information.
2.3.3 Data pre-processing and analysis
The 1H NMR spectra were imported into the SigMa software (Khakimov et al. 2020) where the 1H NMR signals were aligned towards the alanine signal centred at 1.49 ppm and referenced to the TSP signal at 0.00 ppm. Each spectrum was scaled to its ERETIC (Electronic REference To access In vivo Concentrations) signal at 15 ppm, equivalent to 10 mM protons (Akoka et al. 1999; Dona et al. 2014). Metabolites assignment was based on the SigMa library and complemented with data from J-resolved spectroscopy (JRES) experiments and comparison with literature data (Ala-Korpela, 1995; Nicholson et al. 1995; Solanky et al. 2005). Metabolites in the 1H NMR spectra were quantified in SigMa by multivariate curve resolution (De Juan et al. 2014), while complex heterogeneous regions with more than one signal were quantified by raw sum of the spectral intensities. The metabolite table was imported into the PLS_Toolbox (version 8.8.1, Eigenvector Research, Manson, WA, USA) where principal component analysis (PCA) (Wold et al. 1987) was performed to analyse the autoscaled metabolite concentrations. A total of two outliers were identified and excluded from subsequent data analysis. The between-groups variation, corresponding to differences (1) between the sampling events and (2) sampling sites, was calculated using ANOVA simultaneous component analysis (ASCA) with permutation test (1000 permutations) (Smilde et al. 2005).
3 Results and discussion
3.1 Salmon plasma as measured by 1H NMR and SOPs
Expansions of selected regions from a representative 1H NMR spectrum of salmon plasma measured at 310 K are shown in Fig. 2a-c. More details on the impact of the experimental temperature on the salmon plasma metabolome and 1H NMR spectra can be found in the Supporting Information. Overall, the 1H NMR spectrum of salmon plasma is characterized by sharp and well-defined spectral lines, with the lactate doublet (1.32 ppm) and the methylene resonance (1.25 ppm) from lipids dominating the spectral landscape. A total of 28 metabolites including amino acids, organic acids, small sugars, and functional groups from lipids in lipoproteins (LPs) were identified in the spectra measured at 310 K by using the library for plasma metabolites implemented in the SigMa software (Khakimov et al. 2020) as well as literature data (Ala-Korpela, 1995). Metabolite assignments are given in the caption for Fig. 2. For illustrative purposes, the spectral features of salmon plasma were compared to a reference 1H NMR spectrum of human plasma collected from another study (Khakimov et al. 2021) and measured under the same SOPs and experimental conditions (310 K) as the ones described here for salmon plasma. In Fig. 3a-d, representative 1H NMR spectra of human and salmon plasma are shown, including expansions of different spectral regions and peak assignment of the major common resonances. For illustrative purposes, the 1H NMR spectra of plasma are divided into two sub-regions: the “lipid region” (0.50–5.50 ppm), where all the signals from lipids in LPs resonate (Fig. 3b and c), and the “aromatic region” (5.51–10 ppm), where protons from the aromatic residues of proteins resonate (Fig. 3d). A remarkable distortion of the baseline can be observed in the 1H NMR spectrum of human plasma (Fig. 3b and d), which arises from the broad resonances of LPs and other plasma proteins, including albumin. Albumin is one of the most important carriers of nutrients and various substances including hormones, vitamins, and enzymes through the body. Due to the very low mobility, the 1H-NMR spectrum of albumin consists of a broad envelope of overlapping resonances, even at the highest available frequency (Patel et al. 1993). In both types of plasma, the broad peaks of lipids in LPs can also be observed, with salmon plasma having the highest amount of lipids (Fig. 3b and c). In animals, including vertebrates (mammals, birds, reptiles, amphibian and fish) and invertebrates (crustaceans and insects), LPs are the main vehicle for lipid translocation (Chapman, 1980). As for mammals, lipid transport in fish is believed to consist of two loops: dietary (exogenous) lipids are transported to the liver by chylomicrons and/or other carrier proteins, while LPs, in the form of very low density lipoproteins (VLDL) as well as high density (HDL) and low density lipoproteins (LDL), transport lipids from the liver to the tissues and the storage sites (Lie et al. 1993). Compared to humans, fish have much higher contents of lipids in their blood, with average postprandial lipid values of 1940 mg/dl measured in the plasma of young rainbow trout (Babin & Vernier, 1989). Interestingly, in the 1H NMR spectra of salmon, the methyl-end groups of lipids in LPs display two distinct and well separated broad lipid signals, which resonate at 0.84 ppm (HDL) and 0.92 ppm (VLDL) (Fig. 3c). In contrast, in the 1H NMR spectrum of human plasma, the same region appears as one broad envelope of signals from lipids in all LPs, including VLDL, LDL and HDL (Fig. 3c). At lower field, the intense 1H NMR resonances stemming from the methylene groups (1.24 ppm) of lipids in LPs resonate again with much higher intensity in salmon plasma compared to human plasma (the signal is both proportional to the concentration of lipids and to the length of the carbon chain of fatty acids). In salmon plasma, the methylene resonance is asymmetric and skewed to the right, at higher field, confirming that HDL are the predominant LP particles in Atlantic salmon (Torstensen, 2001). Babin and Vernier (1989) have reviewed different aspects of plasma LPs in fish and have reported considerable values of more than 2000 mg/dl of plasma HDL in Salmo spp. (Babin & Vernier, 1989). On average, HDL values for healthy humans are around 50 mg/dl of plasma. Differently from salmon plasma, the methylene resonance in the 1H NMR spectrum of human plasma is symmetric and slightly skewed to the left, at low field, where the resonances of LDL, VLDL and chylomicrons are normally found.
Expansions of a representative 1H NMR spectrum of salmon plasma measured at 310 K. Keys: (1) -C(18)H3 of cholesterol; (2) -CH3 of fatty acids (FA) in LPs; (3) Branched chain amino acids (isoleucine, leucine, and valine); (4) Isobutyrate; (5) Propylene glycol; (6) -CH2- of FA; (7) Lactate; (8) Alanine; (9) -CH2-CH2-COOC- of FA; (10) (-CH2-)n-CH2-CH = of FA; (11) N-acetyl protons of FA; (12) -CH2-CH2-COOC- of FA; (13) -CH = CH-CH2-CH = CH- of FA; (14) Creatinine; (15) Choline; (16) trimethylamine N-oxide (TMAO); (17) Glycine; (18) -CH2N of phosphatidylcholine (PC); (19) sn1,3 CH2 of glycerol; (20) CH2OP of PC; (21) α-glucose (anomeric 1H); (22) -CH = CH- of FA; (23) Tyrosine; (24) Histidine; (25) 1-methyl histidine; (26) Phenylalanine; (27) Adenine; (28) Formate; (*) EDTA); (§) residual water
Representative 1H NMR spectra of salmon (blue line) and human (red line) blood plasma measured using the same SOPs and under the same experimental conditions. The major functional groups from lipids in LPs and resonances from metabolites are assigned. a full spectra; b lipid regions; c –CH3 and –CH2 signals from lipids in LPs; d protein regions. Keys: Cho cholesterol, Lac lactate, PC phosphatidylcholine, UFA unsaturated fatty acids (Color figure online)
When compared to human plasma, the 1H NMR spectrum of salmon plasma has a much higher content of lactate (doublet at 1.34 ppm). Lactate is an important intermediary in numerous metabolic processes as it plays a central role in the metabolism of vertebrates, including fish. When accelerated glycolysis overwhelms lactate utilization and clearance, the level of lactate increases in the blood (Gladden, 2004). Following an increased glycolytic rate the level of circulating lactate rises, for instance as a consequence of stress, acute anaerobic exercise, decreased intramuscular oxygen pressure, and decreased lactate oxidation by the liver and kidneys (Bloomer & Cole, 2009). The higher lactate content in salmon plasma may be the consequence of the handling/catching stress.
The 1H NMR resonance of olefinic protons in unsaturated fatty acids is positioned at 5.30 ppm (Fig. 3b). Again, this intensity is significantly higher in salmon plasma than in human plasma. Cold-water fish species, such as salmon, are normally rich in long chain polyunsaturated fatty acids (LC-PUFAs) with low melting point, which are indispensable to maintain cell membrane fluidity (Chapman, 1980). Even though the majority of PUFAs present in fish derive from their diet, the Atlantic salmon is known for its ability to convert alpha-linolenic acid (18:3n − 3) to eicosapentaenoic (EPA; 20:5n − 3) and docosahexaenoic (DHA; 22:6n − 3) acids, and linoleic acid (18:2n − 6) to arachidonic acid (20:4n − 6) (Zheng et al. 2009). LC-PUFAs can be found in the free form, due to a partial hydrolysis, or in conjugated form. In the latter case, esterification in phospholipids normally occurs. Together with cholesterol, phospholipids, such as phosphatidylcholine (PC), play a key role in determining the membrane fluidity. Cholesterol is one of the main biochemical constituents of the salmon plasma and the 1H NMR signal from the methyl group of the cholesterol molecule (-C (18)H3) is clearly visible at 0.66 ppm (Fig. 3c). Similarly to all vertebrates, salmon require cholesterol for growth and development as it is involved in many biological functions, including providing stability to the cell membrane. In fish, cholesterol homoeostasis is achieved by balancing dietary intake and biosynthesis with bile acid formation and excretion (Kortner et al. 2014). In most fish species, cholesterol is almost entirely present in the esterified form (Babin & Vernier, 1989). No distinct cholesterol signal is visible in the 1H NMR spectrum of human plasma.
At mid-field, in the lipid region, the 1H NMR resonance of choline (3.19 ppm) is visible in salmon plasma (Fig. 2b). Choline is an essential nutrient that is involved in several metabolic processes including lipid metabolism and acts as an indirect methyl donor in several metabolic processes. In animals, including fish, choline is necessary for the synthesis of lecithin, PC and the neurotransmitter acetylcholine. Recent studies have shown that dietary choline deficiency aggravated the intestinal apoptosis, decreased the growth performances and damaged the amino acid absorption capacity in juvenile grass carp (Ctenopharyngodon idella) (Yuan et al. 2021; Yuan et al. 2020). Krogdahl et al. (2020) have recently shown that choline and phosphatidylcholine can reduce excessive gut mucosal lipid accumulation in Atlantic salmon (Salmo salar) (Krogdahl et al. 2020). Moreover, Hansen et al. (2020) have reported an essential role of choline in preventing lipid malabsorption syndrome (excessive lipid accumulation in the proximal intestine) in Atlantic salmon (Hansen et al. 2020). It has further been demonstrated that choline supplementation helped increasing specific fish growth by 18%. At lower field, in the lipid region, the resonance of PC was detected (Fig. 2b and 3b). In Atlantic salmon, PC is one of the major phospholipids in LPs, with highest levels in HDL particles (Lie et al. 1993). In comparison no distinct 1H NMR resonances of glycerol and PC can be found in the spectrum of human plasma (Fig. 3b).
In the aromatic region of both 1H NMR spectra of human and salmon plasma (Fig. 3d), the broad signals from proteins overlap with the 1H NMR resonances stemming from several low- molecular weight metabolites including tyrosine, 1-methyl histidine, phenylalanine, and formic acid. The broad proteins signals are much more intense in the 1H NMR spectrum of human plasma, reflecting a higher protein concentration in human blood.
3.2 Metabolomics analysis
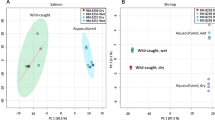
A total of 41 signals from metabolites and functional groups of lipids in LPs were identified in the spectra using the SigMa software (Khakimov et al. 2020) as well as literature data (Ala-Korpela, 1995). Of these, 25 spin systems were from known signature signals (SS), two were from unknown spin systems (SUS), and 15 were complex unresolved regions (BINS). Metabolites (SS and SUS) and functional groups from lipids (BINS) were quantified in SigMa by MCR (SS and SUS) and raw sum of spectral intensities (BINS), respectively, and subsequently analysed by PCA and ASCA. In Fig. 4a, the PCA biplot of PC1 and PC2 shows a clear separation between the two growth stages along PC1. The two sampling events correspond to different growth stages of the salmon, namely smolts and adults. As for other piscine species, the lifecycle of salmon comprises several developmental stages: ova (egg), alevins (just-hatched fish), fry (little fish ready to start eating on their own), parr (silvery small fish that develop vertical stripes and spots for camouflage; they undergo a physiological pre-adaptation to life in seawater), smolt (adapt for life in salt water), adults (salmon that has reached maturity) and kelt (salmon that has spawned). As observed in Fig. 4a, smolts had significantly higher levels of propylene glycol (PG) in their blood. PG is a synthetic diol commonly used as antifreeze, as a humectant, and in the production of polyester compounds. Although generally considered safe and biodegradable, when used in high doses or for prolonged periods, PG can be toxic and cause lactic acidosis, central nervous system depression, coma, hypoglycaemia, seizures, and haemolysis (Lim et al. 2014). In the aquaculture field, PG can be used to maintain high dietary moisture levels in fish feed, while enhancing diet palatability by improving texture, flavour, or moisture content. Previous studies on the effect of dietary PG on growth, survival, histology, and carcass composition of the Atlantic salmon have shown a higher feed conversion efficiency for fish fed the diets containing 5–15% PG (Hughes, 1988). Recently, Soaudy et al. (2021) have shown that inclusion of PG in the diet of Nile tilapia enhanced growth performance and feed utilization under cold stress (Soaudy et al. 2021). Niemuth et al. (2018) demonstrated the presence of PG in plasma of free-ranging sea turtles (Chelonia mydas) and suggested that this turtle species may be able to synthesize PG through reduction of glycerol, another important antifreeze compound (Niemuth et al. 2018). Several studies have pointed to glycerol as a cryoprotectant and important metabolite for cold adaptation in the hibernating black bear and in the arctic insect Gynaephora groenlandica (Ahlquist et al. 1984; Kukal et al. 1988). Furthermore, a wide range of microorganisms, including bacteria and yeast, are currently known to ferment sugars to PG (Saxena et al. 2010). As for the present study, high levels of PG in smolts could be ascribed to diet, but also to a physiological adaptation of fish to life in cold saltwater. Further studies are required to deeper investigate the origin and occurrence of PG in Norwegian Atlantic salmon.
Biplots of PCA performed on the relative concentrations of the metabolites identified in salmon plasma. a Biplot of PC1 vs PC2 coloured according to the sampling event. Red: 1st sampling event (smolt), green: 2nd sampling event (adult). b Biplot of PC1 vs PC3 coloured according to the sampling site; latitudes of sampling sites are given. Keys: Ace acetate, Ade adenine, Ala alanine, Cho cholesterol, For formate, GLC glucose, Gln glutamine, Glu glutamate, Gly glycine, Ile isoleucine, Lac lactate, Leu leucine, Met methionine, 1MH 1-methyl histidine, PG propylene glycol, Phe phenylalanine, Pyr pyruvate, TMAO trimethylamine N-oxide, Tyr tyrosine, Val valine (Color figure online)
As shown in Figs. 4a and 5, cholesterol, lipids, and several amino acids (AA) were more abundant in the plasma samples of adult salmon. The observed increase in the abovementioned metabolite levels most likely reflects the diet composition at the later stages (Table S2). Adult Atlantic salmon use energy reserves for gamete production and for reproductive activity. Reproduction is a metabolically expensive process and reproductive physiology is affected by a variety of endogenous signals providing information on the energy status of an organism (Bou et al. 2020). In particular, lipids play a key role in determining the timing of maturity (Bou et al. 2020). Due to their unique physiology and life cycle, lipid transport in wild salmon has been of great interest to the scientific community. Indeed, in wild salmonids and related species, an appetite reduction or fasting is associated with reproduction migration upriver to the spawning pools (Nelson & Shore, 1974). In general, during the reproduction period, lipids are mobilized from reserve tissues inside LPs and are used as energy source as well as to insure the formation of gametes (Babin & Vernier, 1989). In Fig. 6, an expansion of the spectral region containing the 1H NMR signals from methylene groups of lipids in LPs is shown. Grown up individuals collected during the second sampling event (adults, green spectra) have high VLDL suggesting that VLDL particles in plasma may be related to the physiological and metabolic changes occurring in salmon during growth and gamete formation.
Bar plots showing the relative quantification of lipids, PG and AA. For each metabolite, one-way ANOVA was performed to assess the statistical significance of the variation between the 1st and 2nd sampling event. Asterisk (*) indicates p < 0.05. Keys. Ile isoleucine, Leu leucine, Val valine, Ala alanine, Gln glutamine, Gly glycine, Tyr tyrosine, 1MH 1-methyl histidine, Phe phenylalanine
The PC1 vs. PC3 biplot (Fig. 4b) reveals a weak separation between the three different aqua-farms, with samples collected at the aqua-farm positioned at the lowest latitude clustering at low PC3 scores, while samples collected at the aqua-farm with the highest latitude are clustering at higher PC3. The scores distribution in the biplot is driven by organic acids, amino acids, nucleobase compounds, and PG, the latter being significantly higher in samples collected from site 3 (Lat. 70 °N) (Fig. 5). Total lipids were higher in samples collected from site 2 (Lat. 67 °N), which were characterized by average higher biometries at both growth stages (Table S1). The observed variability in the biometries is not surprising and can be ascribed to the fact that the present investigation is part of a broader study whose aim was to map gut disorders in salmon samples collected along the Norwegian coast, independently from fish size, sex, and diet, which therefore were not accurately standardized. ASCA conducted on the metabolite dataset shows that 24% of the metabolite variation is due to the growth stage and 12% of the metabolite variation is related to the aquaculture site/practice. Both effects were estimated as being significant (p < 0.001).
Even though the PCA models shown in Fig. 4a and b show a good separation between different sampling events and sites, a remarkable spreading of the scores is observed in the plots, which is due to interindividual metabolic differences among the fish. It should be emphasized that the samples included in the present study were collected from sea cages and were farmed under natural and non-standardized environmental conditions. The lack of standardization at the farming stage may have resulted in a more diverse cohort, which in turn led to a more inhomogeneous metabolic profile of salmon. Furthermore, it is important to stress the presence of confounding factors (biological and environmental) in the study design, which potentially may have an impact on the measured variances. Besides diet, sex is a biological factor that has an important role in defining the metabolome of salmonids (Benskin et al. 2014). In the present study, no information on the sex of the salmon included in the analysis was available. Further investigations based on controlled and standardized set up are therefore necessary to deepen the knowledge on the complex bioecology of the Atlantic salmon as well as to elucidate the impact of important biological and environmental factors, such as sex and diet, on the metabolic profile of fish.
4 Conclusions
Standardization of plasma analysis is a promising approach in the aquaculture field as it can help defining a reference metabolic baseline for healthy fish. NMR spectroscopy is an untargeted and inherently quantitative analytical method which can provide a reproducible, unbiased, and rapid overview of the bulk metabolic composition of the biological sample under investigation, including fish plasma. When combined with software for the identification and quantification of metabolites’ spectral signatures (SigMa) and multivariate data mining tools, 1H NMR becomes a powerful analytical tool that can be used for investigating species-specific nutritional requirements and disorders as well as for detecting changes in the metabolome of the species under investigation.
In this study, SOPs developed for the analysis of (human) blood plasma by 1H NMR metabolomics were applied for the first time to analyse plasma samples collected from Atlantic salmon. Multivariate analysis of the NMR data showed important metabolic differences between sampling events and growth stages as well as amongst different farming sites. Moreover, significant differences in the LP profiles of salmon at different growth stages were evidenced confirming the pivotal role covered by LPs in fish development. Our results demonstrate the great potential of standardized 1H NMR in the aquaculture field and pave the way for future research on the development of specific 1H NMR SOPs to be used as a diagnostic method for monitoring diseases and metabolic disorders in fish. Furthermore, in the era where sustainable food production has become imperative, standardized 1H NMR may serve as a screening tool for assessing the impact of alternative and more sustainable plant-based diets on the fish metabolome and their gut microbiome, a current issue in the aquaculture industry.
References
Aas, T. S., Ytrestøyl, T., & Åsgård, T. (2019). Utilization of feed resources in the production of Atlantic salmon (Salmo salar) in Norway: An update for 2016. Aquaculture Reports, 15, 100216. https://doi.org/10.1016/j.aqrep.2019.100216
Ahlquist, D. A., Nelson, R. A., Steiger, D. L., Jones, J. D., & Ellefson, R. D. (1984). Glycerol metabolism in the hibernating black bear. Journal of Comparative Physiology B, 155(1), 75–79. https://doi.org/10.1007/BF00688794
Akoka, S., Barantin, L., & Trierweiler, M. (1999). Concentration measurement by proton NMR using the ERETIC method. Analytical Chemistry, 71(13), 2554–2557. https://doi.org/10.1021/ac981422i
Ala-Korpela, M. (1995). 1H NMR spectroscopy of human blood plasma. Progress in Nuclear Magnetic Resonance Spectroscopy, 27(5–6), 475–554. https://doi.org/10.1016/0079-6565(95)01013-0
Alfaro, A. C., & Young, T. (2018). Showcasing metabolomic applications in aquaculture: A review. Reviews in Aquaculture, 10(1), 135–152. https://doi.org/10.1111/raq.12152
Aru, V., Lam, C., Khakimov, B., Hoefsloot, H. C. J., Zwanenburg, G., Lind, M. V., et al. (2017). Quantification of lipoprotein profiles by nuclear magnetic resonance spectroscopy and multivariate data analysis. TrAC—Trends in Analytical Chemistry, 94, 210–219. https://doi.org/10.1016/j.trac.2017.07.009
Babin, P. J., & Vernier, J. M. (1989). Plasma lipoproteins in fish. Journal of Lipid Research, 30(4), 467–489
Başoǧlu, A., Başpinar, N., & Coşkun, A. (2014). NMR-based metabolomic evaluation in dairy cows with displaced abomasum. Turkish Journal of Veterinary and Animal Sciences, 38(3), 325–330. https://doi.org/10.3906/vet-1310-52
Beckonert, O., Keun, H. C., Ebbels, T. M. D., Bundy, J., Holmes, E., Lindon, J. C., & Nicholson, J. K. (2007). Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nature protocols, 2(11), 2692–2703. https://doi.org/10.1038/nprot.2007.376
Belghit, I., Liland, N. S., Gjesdal, P., Biancarosa, I., Menchetti, E., Li, Y., et al. (2019). Black soldier fly larvae meal can replace fish meal in diets of sea-water phase Atlantic salmon (Salmo salar). Aquaculture, 503, 609–619. https://doi.org/10.1016/j.aquaculture.2018.12.032
Benskin, J. P., Ikonomou, M. G., Liu, J., Veldhoen, N., Dubetz, C., Helbing, C. C., & Cosgrove, J. R. (2014). Distinctive metabolite profiles in in-migrating Sockeye salmon suggest sex-linked endocrine perturbation. Environmental Science and Technology, 48(19), 11670–11678. https://doi.org/10.1021/es503266x
Bertram, H. C., Bach Knudsen, K. E., Serena, A., Malmendal, A., Nielsen, N. C., Fretté, X. C., & Andersen, H. J. (2006). NMR-based metabonomic studies reveal changes in the biochemical profile of plasma and urine from pigs fed high-fibre rye bread. British Journal of Nutrition, 95(5), 955–962. https://doi.org/10.1079/BJN20061761
Bin, P., Azad, M. A. K., Liu, G., Zhu, D., Kim, S. W., & Yin, Y. (2018). Effects of different levels of methionine on sow health and plasma metabolomics during late gestation. Food and Function, 9, 4979–4988. https://doi.org/10.1039/c8fo01477a
Bloomer, R. J., & Cole, B. J. (2009). Relationship between blood lactate and oxidative stress biomarkers following acute exercise. The Open Sports Medicine Journal, 3(1), 44–48. https://doi.org/10.2174/1874387000903010044
Bou, M., Wang, X., Todorčević, M., Østbye, T. K. K., Torgersen, J., & Ruyter, B. (2020). Lipid deposition and mobilisation in atlantic salmon adipocytes. International Journal of Molecular Sciences, 21(7). https://doi.org/10.3390/ijms21072332
Chaleckis, R., Meister, I., Zhang, P., & Wheelock, C. E. (2019). Challenges, progress and promises of metabolite annotation for LC–MS-based metabolomics. Current Opinion in Biotechnology, 55, 44–50. https://doi.org/10.1016/j.copbio.2018.07.010
Chapman, J. M. (1980). Animal lipoproteins: Chemistry, structure, and comparative aspects. Journal of Lipid Research, 21(7), 789–853
Cushley, R. J., & Okon, M. (2002). NMR studies of lipoprotein structure. Annual Review of Biophysics and Biomolecular Structure, 31(1), 177–206. https://doi.org/10.1146/annurev.biophys.31.101101.140910
De Juan, A., Jaumot, J., & Tauler, R. (2014). Multivariate curve resolution (MCR). Solving the mixture analysis problem. Analytical Methods, 6(14), 4964–4976. https://doi.org/10.1039/c4ay00571f
Dona, A. C., Jiménez, B., Schafer, H., Humpfer, E., Spraul, M., Lewis, M. R., et al. (2014). Precision high-throughput proton NMR spectroscopy of human urine, serum, and plasma for large-scale metabolic phenotyping. Analytical Chemistry, 86(19), 9887–9894. https://doi.org/10.1021/ac5025039
Dove, A. D. M., Leisen, J., Zhou, M., Byrne, J. J., Lim-Hing, K., Webb, H. D., et al. (2012). Biomarkers of whale shark health: A metabolomic approach. PLoS ONE, 7(11), e49379.https://doi.org/10.1371/journal.pone.0049379
Emwas, A. H. M., Salek, R. M., Griffin, J. L., & Merzaban, J. (2013). NMR-based metabolomics in human disease diagnosis: Applications, limitations, and recommendations. Metabolomics, 9, 1048–1072. https://doi.org/10.1007/s11306-013-0524-y
Erben, V., Bhardwaj, M., Schrotz-King, P., & Brenner, H. (2018). Metabolomics biomarkers for detection of colorectal neoplasms: A systematic review. Cancers, 10(8), 246. https://doi.org/10.3390/cancers10080246
FAO. (2020). The State of World Fisheries and Aquaculture 2020. Sustainability in action. FAO. https://doi.org/10.4060/ca9229en
Gatlin, D. M., Barrows, F. T., Brown, P., Dabrowski, K., Gaylord, T. G., Hardy, R. W., et al. (2007). Expanding the utilization of sustainable plant products in aquafeeds: A review. Aquaculture Research, 38(6), 551–579. https://doi.org/10.1111/j.1365-2109.2007.01704.x
Giskeødegård, G. F., Madssen, T. S., Euceda, L. R., Tessem, M., Moestue, S. A., & Bathen, T. F. (2018). NMR-based metabolomics of biofluids in cancer. NMR in Biomedicine, 19, e3927. https://doi.org/10.1002/nbm.3927
Gladden, L. B. (2004). Lactate metabolism: A new paradigm for the third millennium. Journal of Physiology, 558(1), 5–30. https://doi.org/10.1113/jphysiol.2003.058701
Hansen, A. K. G., Kortner, T. M., Krasnov, A., Björkhem, I., Penn, M., & Krogdahl, Å. (2020). Choline supplementation prevents diet induced gut mucosa lipid accumulation in post-smolt Atlantic salmon (Salmo salar L.). BMC Veterinary Research, 16, 32. https://doi.org/10.1186/s12917-020-2252-7
Hughes, S.G. (1988). Effect of dietary propylene glycol on growth, survival, histology, and carcass composition of Atlantic Salmon. The Progressive Fish-Culturist, 50(1), 12–15. https://doi.org/10.1577/1548-8640
Jiménez, B., Holmes, E., Heude, C., Tolson, R. F., Harvey, N., Lodge, S. L., et al. (2018). Quantitative lipoprotein subclass and low molecular weight metabolite analysis in human serum and plasma by 1H NMR spectroscopy in a multilaboratory trial. Analytical Chemistry, 90(20), 11962–11971. https://doi.org/10.1021/acs.analchem.8b02412
Jones, O. A. H., & Cheung, V. L. (2007). An introduction to metabolomics and its potential application in veterinary science. Comparative Medicine, 57(5). PMID: 17974125
Khakimov, B., Hoefsloot, H. C. J., Mobaraki, N., Aru, V., Kristensen, M., Lind, M. V, et al. (2021). Human blood lipoprotein predictions from 1H NMR spectra: protocol, model performances and cage of covariance. bioRxiv https://doi.org/10.1101/2021.02.24.432509
Khakimov, B., Mobaraki, N., Trimigno, A., Aru, V., & Engelsen, S. B. (2020). Signature Mapping (SigMa): An efficient approach for processing complex human urine 1H NMR metabolomics data. Analytica Chimica Acta, 1108, 142–151. https://doi.org/10.1016/j.aca.2020.02.025
Kortner, T. M., Björkhem, I., Krasnov, A., Timmerhaus, G., & Krogdahl, Å. (2014). Dietary cholesterol supplementation to a plant-based diet suppresses the complete pathway of cholesterol synthesis and induces bile acid production in Atlantic salmon (Salmo salar L.). British Journal of Nutrition, 111(12), 2089–2103. https://doi.org/10.1017/S0007114514000373
Kortner, T. M., Penn, M. H., Bjrkhem, I., Måsøval, K., & Krogdahl, Å. (2016). Bile components and lecithin supplemented to plant based diets do not diminish diet related intestinal inflammation in Atlantic salmon. BMC Veterinary Research, 12, 190. https://doi.org/10.1186/s12917-016-0819-0
Krogdahl, Å., Gajardo, K., Kortner, T. M., Penn, M., Gu, M., Berge, G. M., & Bakke, A. M. (2015). Soya Saponins induce enteritis in Atlantic Salmon (Salmo salar L.). Journal of Agricultural and Food Chemistry, 63(15), 3887–3902. https://doi.org/10.1021/jf506242t
Krogdahl, Å., Hansen, A. K. G., Kortner, T. M., Bjӧrkhem, I., Krasnov, A., Berge, G. M., & Denstadli, V. (2020). Choline and phosphatidylcholine, but not methionine, cysteine, taurine and taurocholate, eliminate excessive gut mucosal lipid accumulation in Atlantic salmon (Salmo salar L.). Aquaculture, 528, 735552. https://doi.org/10.1016/j.aquaculture.2020.735552
Krogdahl, Å., Penn, M., Thorsen, J., Refstie, S., & Bakke, A. M. (2010). Important antinutrients in plant feedstuffs for aquaculture: An update on recent findings regarding responses in salmonids. Aquaculture Research, 41(3), 333–344. https://doi.org/10.1111/j.1365-2109.2009.02426.x
Kukal, O., Serianni, A. S., & Duman, J. G. (1988). Glycerol metabolism in a freeze-tolerant arctic insect: An in vivo13C NMR study. Journal of Comparative Physiology B, 158(2), 175–183. https://doi.org/10.1007/BF01075831
Lanza, I. R., Zhang, S., Ward, L. E., Karakelides, H., Raftery, D., & Sreekumaran Nair, K. (2010). Quantitative metabolomics by 1H-NMR and LC-MS/MS confirms altered metabolic pathways in diabetes. PLoS ONE, 5(5), e10538. https://doi.org/10.1371/journal.pone.0010538〹
Le Moyec, L., Robert, C., Triba, M. N., Billat, V. L., Mata, X., Schibler, L., & Barrey, E. (2014). Protein catabolism and high lipid metabolism associated with long-distance exercise are revealed by plasma NMR metabolomics in endurance horses. PLoS ONE, 9(3), e90730. https://doi.org/10.1371/journal.pone.0090730〹
Li, Y., Kortner, T. M., Chikwati, E. M., Munang’andu, H. M., Lock, E. J., & Krogdahl, Å. (2019). Gut health and vaccination response in pre-smolt Atlantic salmon (Salmo salar) fed black soldier fly (Hermetia illucens) larvae meal. Fish and Shellfish Immunology, 86, 1106–1113. https://doi.org/10.1016/j.fsi.2018.12.057
Lie, Ø., Sandvin, A., & Waagbø, R. (1993). Influence of dietary fatty acids on the lipid composition of lipoproteins in farmed Atlantic salmon (Salmo salar). Fish Physiology and Biochemistry, 12(3), 249–260. https://doi.org/10.1007/BF00004372
Lim, T. Y., Poole, R. L., & Pageler, N. M. (2014). Propylene glycol toxicity in children. The Journal of Pediatric Pharmacology and Therapeutics, 19(4), 277–282. https://doi.org/10.5863/1551-6776-19.4.277
Liu, Y., Olaf Olaussen, J., & Skonhoft, A. (2011). Wild and farmed salmon in Norway—A review. Marine Policy, 35(3), 413–418. https://doi.org/10.1016/j.marpol.2010.11.007
Luck, M. M., Le Moyec, L., Barrey, E., Triba, M. N., Bouchemal, N., Savarin, P., & Robert, C. (2015). Energetics of endurance exercise in young horses determined by nuclear magnetic resonance metabolomics. Frontiers in Physiology, 6, 198. https://doi.org/10.3389/fphys.2015.00198
Monsonis Centelles, S., Hoefsloot, H. C. J., Khakimov, B., Ebrahimi, P., Lind, M. V., Kristensen, M., et al. (2017). Toward reliable lipoprotein particle predictions from NMR spectra of human blood: An interlaboratory ring test. Analytical Chemistry, 89(15), 8004–8012. https://doi.org/10.1021/acs.analchem.7b01329
Nelson, G. J., & Shore, V. G. (1974). Characterization of the serum high density lipoproteins and apolipoproteins of pink salmon. Journal of Biological Chemistry, 249(2), 536–542. https://doi.org/10.1016/S0021-9258(19)43063-9
Nicholson, J. K., Foxall, P. J. D., Spraul, M., Farrant, R. D., & Lindon, J. C. (1995). 750 MHz 1H and 1H–13C NMR spectroscopy of human blood plasma. Analytical Chemistry, 67(5), 793–811. https://doi.org/10.1021/ac00101a004
Niemuth, J. N., Harms, C. A., Macdonald, J. M., & Stoskopf, M. K. (2018). Propylene glycol in free-ranging green sea turtle plasma. Journal of Fish and Wildlife Management, 9(2), 617–623. https://doi.org/10.3996/122017-JFWM-099
Patel, S. U., Sadler, P. J., Tucker, A., & Viles, J. H. (1993). Direct detection of albumin in human blood plasma by 1H NMR spectroscopy. Complexation of Nickel2+. Journal of the American Chemical Society, 115(20), 9285–9286. https://doi.org/10.1021/ja00073a053
Penn, M. H., Bendiksen, E. A., Campbell, P., & Krogdahl, Å. (2011). High level of dietary pea protein concentrate induces enteropathy in Atlantic salmon (Salmo salar L.). Aquaculture, 310(3–4), 267–273. https://doi.org/10.1016/j.aquaculture.2010.10.040
Lima, A. R., Pinto, J., Bastos, M. de L., Carvalho, M., & Guedes de Pinho, P. (2018). NMR-based metabolomics studies of human prostate cancer tissue. Metabolomics, 14(88), 1–11. https://doi.org/10.1007/s11306-018-1384-2
Pizarro, C., Esteban-Díez, I., Espinosa, M., Rodríguez-Royo, F., & González-Sáiz, J.-M. (2019). An NMR-based lipidomic approach to identify Parkinson’s disease-stage specific lipoprotein–lipid signatures in plasma. Analyst, 144, 1334–1344. https://doi.org/10.1039/c8an01778f
Powell, K. L., Carrozzi, A., Stephens, A. S., Tasevski, V., Morris, J. M., Ashton, A. W., & Dona, A. C. (2018). Utility of metabolic profiling of serum in the diagnosis of pregnancy complications. Placenta, 66, 65–73. https://doi.org/10.1016/j.placenta.2018.04.005
Robinson, P. N. (2012). Deep phenotyping for precision medicine. Human Mutation, 33(5), 777–780. https://doi.org/10.1002/humu.22080
Roques, S., Deborde, C., Richard, N., Sergent, L., Kurz, F., Skiba-Cassy, S., et al. (2018). Characterizing alternative feeds for rainbow trout (O. mykiss) by 1H NMR metabolomics. Metabolomics, 14(55), 1–12. https://doi.org/10.1007/s11306-018-1454-5
Roques, S., Deborde, C., Richard, N., Skiba-Cassy, S., Moing, A., & Fauconneau, B. (2020). Metabolomics and fish nutrition: A review in the context of sustainable feed development. Reviews in Aquaculture, 12(1), 261–282. https://doi.org/10.1111/raq.12316
Sandnes, K., Lie, Ø., & Waagbø, R. (1988). Normal ranges of some blood chemistry parameters in adult farmed Atlantic salmon, Salmo salar. Journal of Fish Biology, 32, 129–136. https://doi.org/10.1111/j.1095-8649.1988.tb05341.x
Savorani, F., Rasmussen, M. A., Mikkelsen, M. S., & Engelsen, S. B. (2013). A primer to nutritional metabolomics by NMR spectroscopy and chemometrics. Food Research International, 54(1), 1131–1145. https://doi.org/10.1016/j.foodres.2012.12.025
Saxena, R. K., Anand, P., Saran, S., Isar, J., & Agarwal, L. (2010). Microbial production and applications of 1,2-propanediol. Indian Journal of Microbiology, 50(1), 2–11. https://doi.org/10.1007/s12088-010-0017-x
Serkova, N. J., Standiford, T. J., & Stringer, K. A. (2011). The emerging field of quantitative blood metabolomics for biomarker discovery in critical illnesses. American Journal of Respiratory and Critical Care Medicine, 184(6), 647–655. https://doi.org/10.1164/rccm.201103-0474CI
Shestakova, K., Brito, A., Mesonzhnik, N. V., Moskaleva, N. E., Kurynina, K. O., Grestskaya, N. M., et al. (2018). Rabbit plasma metabolomic analysis of Nitroproston®: A multi target natural prostaglandin based-drug. Metabolomics, 14(112), 1–12. https://doi.org/10.1007/s11306-018-1413-1
Smilde, A. K., Jansen, J. J., Hoefsloot, H. C. J., Lamers, R. J. A. N., van der Greef, J., & Timmerman, M. E. (2005). ANOVA-simultaneous component analysis (ASCA): A new tool for analyzing designed metabolomics data. Bioinformatics, 21(13), 3043–3048. https://doi.org/10.1093/bioinformatics/bti476
Soaudy, M. R., Mohammady, E. Y., Ashry, M. A., Ali, M. M., Ahmed, N. M., Hegab, M. H., et al. (2021). Possibility mitigation of cold stress in Nile tilapia under biofloc system by dietary propylene glycol: Performance feeding status, immune, physiological responses and transcriptional response of delta-9-desaturase gene. Aquaculture, 538, 736519. https://doi.org/10.1016/j.aquaculture.2021.736519
Solanky, K. S., Burton, I. W., Mackinnon, S. L., Walter, J. A., & Dacanay, A. (2005). Metabolic changes in Atlantic salmon exposed to Aeromonas salmonicida detected by 1H-nuclear magnetic resonance spectroscopy of plasma. Disease of Aquatic Organisms, 65, 107–114. https://doi.org/10.3354/dao065107
Spratlin, J. L., Serkova, N. J., & Eckhardt, S. G. (2009). Clinical applications of metabolomics in oncology: A review. Clinical Cancer Research, 15(2), 431–440. https://doi.org/10.1158/1078-0432.CCR-08-1059
Tacon, A. G. J., & Metian, M. (2008). Global overview on the use of fish meal and fish oil in industrially compounded aquafeeds: Trends and future prospects. Aquaculture, 285(1–4), 146–158. https://doi.org/10.1016/j.aquaculture.2008.08.015
Torstensen, B. E., Lie, & Hamre, K. (2001). A factorial experimental design for investigation of effects of dietary lipid content and pro- and antioxidants on lipid composition in Atlantic salmon (Salmo salar) tissues and lipoproteins. Aquaculture Nutrition, 7(4), 265–276. https://doi.org/10.1046/j.1365-2095.2001.00184.x
Wold, S., Esbensen, K., & Geladi, P. (1987). Principal component analysis. Chemometrics and Intelligent Laboratory Systems, 2(1–3), 37–52. https://doi.org/10.1016/0169-7439(87)80084-9
Xiao, J. F., Zhou, B., & Ressom, H. W. (2012). Metabolite identification and quantitation in LC-MS / MS-based metabolomics. Trends in Analytical Chemistry, 32, 1–14. https://doi.org/10.1016/j.trac.2011.08.009
Young, T., & Alfaro, A. C. (2016). Metabolomic strategies for aquaculture research: A primer. Reviews in Aquaculture, 10, 26–56. https://doi.org/10.1111/raq.12146
Yuan, Z., Feng, L., Jiang, W., Wu, P., Liu, Y., Kuang, S., et al. (2021). Dietary choline deficiency aggravated the intestinal apoptosis in association with the MAPK signalling pathways of juvenile grass carp (Ctenopharyngodon idella). Aquaculture, 532, 736046. https://doi.org/10.1016/j.aquaculture.2020.736046
Yuan, Z. H., Feng, L., Jiang, W. D., Wu, P., Liu, Y., Jiang, J., et al. (2020). Choline deficiency decreased the growth performances and damaged the amino acid absorption capacity in juvenile grass carp (Ctenopharyngodon idella). Aquaculture, 518, 734829. https://doi.org/10.1016/j.aquaculture.2019.734829
Zheng, X., Leaver, M. J., & Tocher, D. R. (2009). Long-chain polyunsaturated fatty acid synthesis in fish: Comparative analysis of Atlantic salmon (Salmo salar L.) and Atlantic cod (Gadus morhua L.) Δ6 fatty acyl desaturase gene promoters. Comparative Biochemistry and Physiology B Biochemistry and Molecular Biology, 154(3), 255–263. https://doi.org/10.1016/j.cbpb.2009.06.010
Ziarrusta, H., Mijangos, L., Picart-Armada, S., Irazola, M., Perera-Lluna, A., Usobiaga, A., et al. (2018). Non-targeted metabolomics reveals alterations in liver and plasma of gilt-head bream exposed to oxybenzone. Chemosphere, 211, 624–631. https://doi.org/10.1016/j.chemosphere.2018.08.013
Funding
Financial support provided by The Norwegian Seafood Research Foundation, FHF Project no. 901435—GutMatters). The authors wish to acknowledge the Danish Innovation Foundation for the generous support to the project entitled “COUNTERSTRIKE – Counteracting Sarcopenia with proteins and exercise – screening the CALM cohort for lipoprotein biomarkers” (IFD: 4105-00015B).
Author information
Authors and Affiliations
Contributions
VA, SBE, ÅK and PM conceived and designed research. VA, TMK and EMC conducted experiments. VA, BK and KMS analysed data. VA drafted the manuscript. All authors read, commented, and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Aru, V., Khakimov, B., Sørensen, K.M. et al. The plasma metabolome of Atlantic salmon as studied by 1H NMR spectroscopy using standard operating procedures: effect of aquaculture location and growth stage. Metabolomics 17, 50 (2021). https://doi.org/10.1007/s11306-021-01797-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-021-01797-0