Abstract
Obesity is currently an increasing public health problem. The intra-uterine environment plays a critical role in foetal development. The objective of this study is to investigate the association of obesity with modifications in the metabolic profiles of pregnant women, and their new-borns. Based on the PELAGIE cohort (Brittany, France), a sample of 321 pregnant women was divided into three groups according to their body mass index (BMI) (normal, over-weight and obese). Nuclear magnetic resonance-based metabolomics analyses were performed on maternal urine and cord-blood samples. Partial least squares regression-discriminant analysis (PLS-DA), polytomous and logistic regressions were used to differentiate the metabolic profiles of the three BMI groups after adjusting for potential confounders. Specific profiles were observed for the overweight and obese women (BMI > 25) compared to the normal-weight women: they had a decrease in urinary hippurate excretion associated with a decrease in phenylalanine and an increase in creatinine. We also showed an increase in the urinary excretion of lactate, citrate, acetate, creatine, and lysine only in obese women (BMI > 30) compared to the normal-weight women. The PLS-DA modelling did not reveal any significant difference between the cord-blood metabolic profiles of newborns according to maternal BMI—although infants born of obese women had a higher birth weight and a lower Apgar score. Our results confirmed the potential link between obesity and gut microbiota disruption (changes in urinary acids), as well as energy and amino-acid metabolism but did not reveal any disruption among newborns.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Obesity, associated with type II diabetes or cardiovascular diseases, has reached epidemic status. A third of American women of reproductive age are obese (Orsi et al. 2011). Now, epidemiological studies have shown increased birth weight in new-borns from obese mothers, associated with a higher risk of developing obesity in the future (Muhlhausler et al. 2013; Symonds et al. 2013). Epidemiological studies, experimentations on animals and placental models have shown the critical role played by the intrauterine environment in foetal development, including a direct transmission of obesity or metabolic pathologies (Dong et al. 2013; Symonds et al. 2013). For example, the male offspring of mice with gestational obesity developed excess weight, insulin resistance and hyperleptinemia at adulthood, without any post-natal obesogenic influence (Dahlhoff et al. 2014). Furthermore, several human studies based on mother–child cohorts have found an association between maternal obesity and disrupted childhood metabolism associated with cardiovascular diseases (Godfrey and Barker 2000; O’Reilly and Reynolds 2013; Wen et al. 2011).
These phenomena, known as “foetal programming” involve different mechanisms (Bouanane et al. 2010; Symonds et al. 2013; Zambrano and Nathanielsz 2013). First of all, several studies provide supporting evidence on the involvement of an epigenetic mechanism. In one human cohort, modifications in DNA methylation at the leptin and adiponectin genes have been observed in placentas exposed to maternal gestational diabetes mellitus (Bouchard et al. 2012, 2010). In another, global methylation levels in the placenta and umbilical cord-blood was higher with obesity (Norman et al. 2014).
Secondly, overexpression of certain miRNA in foetal muscle from obese mothers has been shown in a model of maternal obesity in ewes, suggesting a decrease in adipogenic markers and in inflammatory cytokines (Yan et al. 2013). In rats, the wiring of connections within the hypothalamic arcuate nuclei (HAC) - a complex energy and appetite control system—occurs in the first 3 weeks of life coincident with a peak in leptin in the new-born pups’ peripheral blood. Pups born of obese mothers have a distorted plasma leptin peak that is longer lasting and higher in amplitude (Kirk et al. 2009). This could be the sign of a perturbation in HAC development, resulting in future resistance to leptin feedback and energy balance alterations. Hormonal changes in foetuses from obese mothers—such as increased cortisol levels or decreased thyroxine (T4) levels—have also been reported as potential mechanisms involved in programmed obesity (Guzmán et al. 2006; Magyar et al. 1980; Nuermaimaiti Tuersunjiang 2013; Suter et al. 2012). Changes in placenta function have also been observed, with inflammation and changes in glucose or amino acid transport rates (Challier et al. 2008; Farley et al. 2010; Jones et al. 2009). Maternal reactive oxygen and nitrogen species, which are associated with intrauterine oxidative stress, also seem to play a role in obesity programming. Indeed, oxidative stress is suspected of being implicated in mitochondrial damage, changes in pancreas functions and dysfunctions of the electron transport chain in skeletal muscle (Shelley et al. 2009; Simmons et al. 2005). There is also supporting evidence for a link between maternal exposure to xenobiotics and chemicals, and higher Body Mass Index (BMI) in offspring (Janesick and Blumberg 2011; Karmaus et al. 2009; Smink et al. 2008).
Otherwise, some of these mechanisms (epigenetics, corticosteroids, miRNA or maternal exposure to chemicals) are investigated for the understanding of the programming of obesity for children exposed to famine in utero (Inadera 2013). Most studies have been undertaken in animal models.
Metabolomics describes the study of the metabolome, which is defined as the collective set of metabolites produced or present in a sample of interest, for example blood or other biological fluid, tissue lysate or cells. Metabolomics based on spectroscopic techniques, is able to generate “fingerprints” or metabolic profiles that can be correlated with phenotypes. The metabolic profile constitutes the ultimate step in the cellular response, and is considered the key link between genes and phenotypes (Fiehn 2002). It has come to be widely used in recent years to identify metabolic pathways modified by disease (Vinayavekhin et al. 2010). This technique has already proved its capacity to study the physiopathology of obesity in animal and human studies (Zhang et al. 2013). When it is non-targeted, variations can be shown in the levels of metabolites between different groups of population without a priori.
Metabolomics have already been used to investigate metabolic differences between pregnant and non-gravid women (Lowe and Karban 2014) as well as to analyse metabolic differences between obese and normal weight people (Xie et al. 2012a). The aim of the present study is to investigate the existence of differential urinary metabolic profiles in pregnant women according to their obesity status, and to understand whether observed changes are associated with modifications in the new-born metabolic profile. Thus, we have used nuclear magnetic resonance (NMR)-based metabolomics analysis on urinary and cord-blood samples of pregnant women and new-borns from the PELAGIE cohort (Brittany, France). To our knowledge, this is the first study to investigate the maternal programming of obesity in a human population, using metabolomics.
2 Materials and methods
2.1 Population, sample collection and obesity groups
The population was selected from the PELAGIE cohort, which includes 3421 pregnant women in Brittany (France) enrolled during early pregnancy from the general population by gynaecologists, between 2002 and 2006. Gynaecologists informed the women of the nature of the study and asked them to participate, after providing written consent. This consent was accompanied by a letter of information describing the goal of the study, the consortium, data collection procedures and follow-up after birth (via questionnaires and medical examinations). Explicit mention was made of the right to refuse to participate, and the fact that such a refusal would not have any effect on the woman’s relationship with her doctor. Both the INSERM (French National Institute of Health and Medical Research) ethics committee and the National Commission in charge of Data Protection (CNIL) approved the study procedures (Nu902076; 31 may 2002). A detailed description of this cohort is available elsewhere (Chevrier et al. 2011). At inclusion, during the first trimester of the pregnancy (4–15th week), women had to complete a self-report questionnaire including information about social and demographic characteristics, diet and lifestyle, and data on their height and weight (used to calculate BMI). They also had to return a first morning void urine sample that they collected and transferred into two vials containing nitric acid to avoid bacterial degradation. No blood collection was planned at this step of inclusion because of the absence of hospital appointment. Samples were mailed to the study laboratory in a pre-stamped package at ambient temperature, with routine delivery taking 1–3 days. Upon receipt, the 10 mL samples were frozen and stored at −20 °C. At birth, medical data on health outcomes were obtained and cord-blood samples were collected. After centrifugation, serum samples were analysed by hospital laboratories and stored at −20 °C.
Pregnant women were selected from the PELAGIE according to the following criteria: living infant at birth (n = 3322) and availability of both urinary and cord blood samples for each mother–child pair (n = 1061). In addition, 1 year of inclusion (2004) was selected, to avoid potential variability due to different storage durations of biological samples. A detailed description of this methodology is published elsewhere (Bonvallot et al. 2013). Of the 338 eligible women, those whose BMI was unavailable or below 18.5 were excluded (n = 15), because low body weight can influence metabolic profiles (Norman et al. 2014; Sarlio-Lähteenkorva et al. 2004). Finally, the population analysed included 323 women, who were classified into three groups of BMI, according to the World Health Organization (WHO)’s definition (WHO Expert Consultation 2004): the first group comprised women with a BMI of between 18.5 and <25 (normal weight), the second those with a BMI between 25 and <30 (overweight) and the third, those with a BMI of 30 or more (obese).
2.2 Metabolomics analyses
2.2.1 Urinary sample preparation
After thawing at room temperature and vortexing, 500 μL of urine were mixed with 200 μL of phosphate buffer (pH 7.39) prepared in D2O to which was added sodium 3-trimethylsilyl-1-[2,2,3,3,-2H4]-propionate (TSP, 1 mM). The phosphate buffer is used to minimize variations in chemical shift values in the acquired NMR spectra due to pH differences. TSP served as a chemical shift reference and D2O served as a field-frequency lock for the NMR spectrometer. Each sample was vortexed and centrifuged for 10 min at 8000 rpm to remove any precipitate. Then, 600 μL aliquots were transferred to standard 5 mm—NMR tubes (Norell ST 500, Landisville, NJ) for analysis.
2.2.2 Serum sample preparation
After thawing at room temperature and vortexing, 200 μL of serum were mixed with 500 μL of D2O which served as a field-frequency lock for the NMR spectrometer. Each sample was vortexed and centrifuged for 10 min at 8000 rpm to remove any precipitate. Then, 600 μL aliquots were transferred to NMR tubes for analysis.
2.2.3 Metabolomics analyses
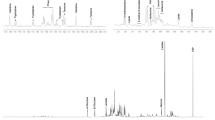
Metabolomics analyses of urine and cord-blood samples were made by NMR spectroscopy using a Bruker Avance DRX-600 operating at 600.13 MHz (Bruker Biospin, Germany) and equipped with an autosampler and an inverse 1H–13C–15N cryoprobe. All NMR spectra were phase- and baseline-corrected manually using Topspin (V2.1, Bruker Biospin, Germany). The spectral region containing residual water resonance (δ 5.515–6.600) was removed and spectra were digitized to 642 and 751 buckets corresponding to 0.01 ppm intervals using the AMIX software package (V3.9.11, Bruker Biospin, Germany). Each integrated region was divided by the total spectral intensity in order to normalise values. Spectra acquisition, pre-processing step, and metabolite identification have been described previously (Bonvallot et al. 2013).
2.2.4 Statistical analysis: metabolic profiles according to BMI groups
To investigate the association of obesity with specific metabolomic profiles, we used partial least square discriminant analyses (PLS-DA) and multivariate logistic regression models. Our objective was not to strictly build predictive models for obesity but rather to study variations in both mother and newborn metabolic profiles according to maternal obesity.
The PLS-DA approach was chosen because it handles highly collinear and noisy data, such as spectral data and it has already been used in epidemiological studies which investigated subtle health effects in relation to highly variable spectral data (Waterman et al. 2009). It allows identifying among the spectrum the most important variables which discriminate the groups of interest, using statistics such as the variable importance on projection (VIP).
Maternal urine and cord blood spectral data were analysed separately. The original digitized NMR spectral data were imported into the R software (version 3.1.1) for multivariate statistical analysis. A preliminary principal component analysis (PCA) was implemented to remove potential outliers. Data were then Pareto-scaled to reduce relative importance of large values, while partially preserving data structure (Worley and Powers 2013). PLS-DA were applied to Pareto-scaled data. In PLS-DA, linear combinations of NMR buckets are constructed to maximize covariance between the Y (BMI groups) and (NMR buckets) X matrices. Observations are then projected onto planes defined by a few of these linear combinations (also interpreted as latent variables). Various PLS-DA models were generated: a three-level model measuring the link between the metabolic profiles and the mothers’ BMI in three groups (Normal vs. Overweight vs. Obese) and four two-level models measuring the link with BMI in two groups (Normal vs. others, Obese versus others, Normal vs. Overweight, Overweight vs. Obese). The quality of the three-level models was assessed by the Q2 and the R2 parameters defined as followed:
-
\(Q2 = 1 - \frac{{PRESS_{n} }}{{RSS_{n - 1} }}\), where PRESS is the predicted residual sum of squares; RSS, the residual sum of squares and n, the number of latent variables (LV).
-
\(R2 = 1 - \frac{PRESS}{TSS}\), where TSS is the total sum of squares.
The quality of the 2-level models was assessed by the Q2, R2 and the area under the ROC curve (AUROC) criteria, following a 3–2 double-k-fold cross-validation. This consists of two nested cross-validation loops. The modelling procedure, including the cross-validation that determines the best number of Latent Variables (LV), using the lower PRESS criteria, forms the inner loop. Cross-validation for error estimation takes place in the outer loop. Double cross-validation is recommended in PLS-DA analysis (Smit et al. 2007; Szymańska et al. 2012). Here, a limited number of folds for cross validation (n = 3 in outer loop and n = 2 in inner loop) were chosen to keep an adequate number of obese mothers in each fold (n = 6 or 7). A random draw was performed to allow equal distribution of Obese and Overweight in each fold. A permutation test (1000 iterations) was conducted for each PLS-DA model to test for validity. Finally, the spectral regions (buckets) having VIP above 2 were considered as being significantly associated with the maternal BMI status and were used to identify the metabolites of interest. Kruskal–Wallis tests using crude spectral data were used to confirm the relationships between BMI groups and concentrations of metabolites in urine or in serum previously identified.
2.3 Adjustment for confounding factors
As PLS-DA did not allow for adjustment, polytomous and logistic regressions were then used to assess the association of urinary metabolic profiles with the BMI group (3 levels) after adjustment for women’s individual characteristics. For each metabolite previously identified from the PLS-DA, the corresponding buckets with VIP > 2 were simultaneously introduced in the models. Results from these analyses were reported as a global trend (direction of the association) and an adjusted p value. The literature suggested some major confounding factors. Age, educational level, alcohol and tobacco consumption, food habits (fruit and fish consumption), proportion of area covered by cereal crops in the municipality of residence during pregnancy (reflecting pesticide exposure) and Apgar score (at 5 min, only in the analysis of cord-blood) were considered as potential confounders and were retained in the model if the likelihood ratio (LR) test was statistically significant for at least one metabolite. Then, a backward selection using the LR test was performed to select confounders with an impact in term of goodness of fit. Finally, none of the confounders has been kept in the models.
3 Results
Preliminary PCA identified two outliers among the women (urinary analyses). The first one had a high concentration of urinary glucose and was identified as diabetic. The second had no specific characteristics compared to the other individuals but a high concentration of hippurate was detected in her urinary sample. The preliminary PCA among the new-borns (serum analyses) did not identify any outlier. Table 1 describes the characteristics of the 321 women finally included in the analyses. Mean maternal age was 30.3 and most of these women had a high educational level (university degree). Tobacco and alcohol consumption was limited (28.2 and 15.9 %).
Most of the women were of normal-weight (n = 256), 46 were overweight and only 19 were considered obese according to the WHO definition. Overweight and obese women had higher parity than normal weight women (p = 0.03), higher incidence of high blood pressure (p = 0.006) and higher incidence of occupational exposure to solvents (p = 0.047). Infants born of overweight or obese women had higher birth weight (p = 0.02) and those born from obese women had lower Apgar scores at 5 min (p = 0.02).
3.1 Urinary analyses of pregnant women
Obese mothers were distinguished by the first two latent variables, and overweight mothers to a slightly lesser extent. The ideal number of latent variables was one or two, depending on validation loop (mean = 1.77 in the Normal vs. Overweight vs. Obese model). Separation of the 3 levels of BMI was significant according to the Q2 (mean = 0.01, p = 0.006) and R2 (mean = 0.02, p = 0.003) criteria. The separation between normal-weight women and the rest of the population was significant according to the AUROC (mean = 0.71, p = 0.001), Q2 (mean = 0.021, p = 0.004) and R2 (mean = 0.05, p = 0.001) criteria, as well as the separation between normal-weight and overweight women (AUROC = 0.66, p = 0.003; Q2 = 0.02, p = 0.001; R2 = 0, p = 0.008). The separation between obese and the rest of the population was highly significant according to the AUROC (mean = 0.76, p = 0.001) but not to the Q2 and R2. This can be explained by the small number of obese women in the population, leading to a lower performance of the classical parameters Q2 and R2 in PLS. The separation between obese and overweight women was not significant according to any criterion. The score plot derived from the PLS-DA modelling is shown in Fig. 1. The parameters resulting from the validation procedures and permutation tests are described in Table 2.
PLS-DA score plot from the 1H NMR urinary metabolic profile from 321 pregnant women. The score plot is the projection of observations onto the first two latent variables. The PLS-DA model was run on Pareto-scaled data (N = 321, R2 = 0.02, Q2 = 0.01). Three groups of Body Mass Index (BMI). Green triangles normal weight women, blue circles overweight women, red squares obese women
Because hypertension and occupational exposure to solvent could also be associated with an increased risk in obesity, these characteristics were studied by PLS-DA modelling on urinary metabolic profile. The models did not produce any significant separation (data not shown).
Next, logistic and polytomous regressions were performed on metabolites having VIP above 2: direction of the associations and p-values using Kruskal–Wallis test were presented in Table 3. Hippurate and phenylalanine levels were lower in obese and overweight women, whereas urinary creatinine was higher. Lactate was higher in obese (though not in overweight) women, as were creatine, lysine, citrate and acetate. Two unknown signals (δ = 6.925 and δ = 2.345 ppm) appeared modified between the BMI groups.
3.2 Serum analyses of cord-blood at birth
The metabolic profiles of the cord-blood samples did not show any visual separation on the first latent variables, as shown in Fig. 2. The parameters resulting from the validation procedure and permutation test are presented in Table 4. These validation procedures did not show any significant separation according to the BMI group, irrespective of the tested model. No bucket having a VIP above 2 was found significantly associated with BMI status by the Kruskal–Wallis test.
PLS-DA score plot from the 1H NMR cord-blood metabolic profile from 321 individuals. The score plot is the projection of observations onto the first two latent variables. The PLS-DA model was run on Pareto-scaled data, (N = 324, R2 = −0.04, Q2 = −0.07). Three groups of Body Mass Index (BMI). Green triangles normal weight women, blue circles overweight women, red squares obese women
4 Discussion
Our work shows modification to seven urinary metabolites between obese, overweight, and normal-weight pregnant women, including hippurate, phenylalanine, lactate, creatine, lysine, citrate and acetate, without any measurable association with specific cord-blood metabolic profile at birth. The decrease in urinary hippurate and phenylalanine levels and the increase in creatinine levels were observed in both obese and overweight women. Increased urinary lactate, creatine, lysine, citrate and acetate levels concerned only the obese women. Hippurate, lactate and acetate are organic acids which had already been associated with obesity and type 2 diabetes, especially in studies investigating the influence of gut microbiota (Calvani et al. 2010; Dewulf et al. 2013; Phipps et al. 1998; Respondek et al. 2013; Salek et al. 2007; Waldram et al. 2009; Williams et al. 2002). In particular, the urinary excretion of hippurate has been associated with gut microbiota having a role in the metabolism of polyphenolic compounds (Lees et al. 2013). In addition, there is evidence to suggest that disruptions in gut flora and their relationship with the host are associated with the development of obesity (Gross 2013). Decreases in urinary hippurate have been observed both experimentally and in humans (Calvani et al. 2010; Friedrich et al. 2012; Salek et al. 2007).
An increase in lactate has already been shown in some studies—in urine, blood and hepatic tissue (Rull et al. 2009; Serkova et al. 2006). It has been shown that, like hippurate, lactate is linked to gut microflora, in particular to Propionibacterium (Dewulf et al. 2013; Respondek et al. 2013). Lactate is also a precursor of gluconeogenesis and an elevation could be a sign of hepatic disruptions in glucose and glycogen synthesis (Xie et al. 2012b).
Lastly, an increase in acetate levels has already been observed in the urine and blood of Zucker rat (Serkova et al. 2006; Waldram et al. 2009). Acetate is suspected to be linked with microbiome, and in particular with Bifidobacterium—which is already known to have a protective effect against obesity (Waldram et al., 2009).
It was also shown than amino-acid metabolism could be modified in the event of obesity (Lees et al. 2013). The increased urinary lysine we have observed in this study is consistent with experimental studies carried out in rats. Changes in lysine degradation were observed in obese Zucker rats compared to lean rats, and are potentially associated with impairment to energy metabolism, implying perturbations in insulin resistance (Salek et al. 2007). Nevertheless, other studies also showed a decrease in plasmatic lysine levels associated with an increase in insulin resistance in mice (Won et al. 2013) and a decrease in lysine levels in the adipose tissue of obese people (Hanzu et al. 2013). Moreover, lysine acetylation seems to play a role in some obesity-induced cancers (Lee et al. 2013). Finally, we know that protein nutrition can have a consequence on lipid metabolism (Gudbrandsen et al. 2008). Similar observations were made of phenylalanine; several authors have linked its increase with obesity (Kim et al. 2013; Wang et al. 2011; Whitehead et al. 2007). A relationship between obesity and the metabolism of branched-chain amino acids was hypothesized (Adams 2011; Kim et al. 2013). On the other hand, Won et al. showed a decrease in urinary phenylalanine concentrations in obese mice (Won et al. 2013). Despite this discrepancy regarding modifications observed in lysine and phenylalanine levels, it could be concluded that obesity has an influence on amino acid metabolism, as has already been suggested in other works (Zhou et al. 2013).
Citrate is produced from fatty acids and glucose metabolism and is an intermediate in the citrate cycle for energy production. Its regulation involves insulin and glucose. The increased levels of citrate we observed are consistent with certain studies in diabetic or obese rats (Kim et al. 2009; Li et al. 2008; Shearer et al. 2008), suggesting a link with hyperglycaemia and insulin resistance, while lowered levels have sometimes been measured (Salek et al. 2007; Schirra et al. 2008; Zhao et al. 2010). The increased citrate levels could be associated with the increase in acetate and lactate levels, suggesting an up-regulation in the citrate cycle. In addition, changes in citrate levels could be associated with renal pathologies. It is possible that an increase in the urinary secretion of citrate may be a consequence of a metabolic stress induced by hyperglycaemia and/or dyslipidaemia, or a consequence of renal dysfunctions (Salek et al. 2007).
Creatinine is a break-down product of creatine metabolism in muscles. Urinary and blood increases in creatinine and creatine have been observed in obese rodents (Calvani et al. 2010; Duggan et al. 2011; Salek et al. 2007; Schirra et al. 2008; Williams et al. 2005; Zhao et al. 2010). This phenomenon could be explained by cardiac or skeletal muscle hypertrophy as an adaptation to support the increase of body mass induced by obesity (Xie et al. 2012b). Urinary secretion of creatinine is also used to measure renal function. An increase could be the consequence of renal dysfunctions, linked with obesity (Kim et al. 1969; Proczko et al. 2013).
In recent years, studies have shown that NMR-based metabolomics on urine samples is able to identify obesity biomarkers. To the best of our knowledge, our study is the first to use metabolomics trying to investigate the mechanisms governing the transmission of obesity. Our inability to observe the effect of obese or overweight mothers on offspring metabolic profile must be put into perspective. First, the low number of obese women (n = 19) induces a low statistical power. Obesity measurement by BMI, calculated using physiological parameters (weight, height) collected by questionnaire may result in measurement errors, in particular an underestimation of obese women’s weight, leading to a weak separation of BMI subgroups. Lastly, it is possible that certain modifications to metabolic profile are not visible at birth—but appear later.
The NMR technique used does have certain advantages: the analysis is highly quantitative, measures are reproducible and required preparation is minimal (Dumas et al. 2006; Smolinska et al. 2012). The other commonly-used technique in metabolomics is mass spectrometry coupled with chromatographic techniques, the advantage of which is the possibility of detecting metabolites in lower concentrations (Werner et al. 2008; Breitling et al. 2006). It would be interesting to reproduce this kind of study using mass spectrometry, in order to confirm and improve the results of our study.
5 Concluding remarks
Metabolomics has potential for the study of obesity and the identification of biomarkers, with the possibility of a global measurement of metabolic dysfunctions in a biological matrix. This study has shown several modifications to the urinary metabolic profile of obese and overweight pregnant women when compared with normal-weight pregnant women, yet without being able to show any modification to blood metabolic profile of offspring at birth. Changes observed include modifications to amino acid metabolism, citrate cycle, and microbiome-host relationships. More studies on large human samples will be necessary to establishing strong evidence on metabolic modifications involved in the transmission of obesity.
References
Adams, S. H. (2011). Emerging perspectives on essential amino acid metabolism in obesity and the insulin-resistant state. Advances in Nutrition, 2, 445–456. doi:10.3945/an.111.000737.
Bonvallot, N., Tremblay-Franco, M., Chevrier, C., Canlet, C., Warembourg, C., Cravedi, J.-P., & Cordier, S. (2013). Metabolomics tools for describing complex pesticide exposure in pregnant women in Brittany (France). PLoS One, 8, e64433. doi:10.1371/journal.pone.0064433.
Bouanane, S., Merzouk, H., Benkalfat, N. B., Soulimane, N., Merzouk, S. A., Gresti, J., et al. (2010). Hepatic and very low-density lipoprotein fatty acids in obese offspring of overfed dams. Metabolism, 59, 1701–1709. doi:10.1016/j.metabol.2010.04.003.
Bouchard, L., Hivert, M.-F., Guay, S.-P., St-Pierre, J., Perron, P., & Brisson, D. (2012). Placental adiponectin gene DNA methylation levels are associated with mothers’ blood glucose concentration. Diabetes, 61, 1272–1280. doi:10.2337/db11-1160.
Bouchard, L., Thibault, S., Guay, S.-P., Santure, M., Monpetit, A., St-Pierre, J., et al. (2010). Leptin gene epigenetic adaptation to impaired glucose metabolism during pregnancy. Diabetes Care, 33, 2436–2441. doi:10.2337/dc10-1024.
Breitling, R., Pitt, A. R., & Barrett, M. P. (2006). Precision mapping of the metabolome. Trends in Biotechnology, 24(12), 543–548.
Calvani, R., Miccheli, A., Capuani, G., Tomassini Miccheli, A., Puccetti, C., Delfini, M., et al. (2010). Gut microbiome-derived metabolites characterize a peculiar obese urinary metabotype. International Journal of Obesity, 2005(34), 1095–1098. doi:10.1038/ijo.2010.44.
Challier, J. C., Basu, S., Bintein, T., Minium, J., Hotmire, K., Catalano, P. M., & Hauguel-de Mouzon, S. (2008). Obesity in pregnancy stimulates macrophage accumulation and inflammation in the placenta. Placenta, 29, 274–281. doi:10.1016/j.placenta.2007.12.010.
Chevrier, C., Limon, G., Monfort, C., Rouget, F., Garlantézec, R., Petit, C., et al. (2011). Urinary biomarkers of prenatal atrazine exposure and adverse birth outcomes in the PELAGIE birth cohort. Environmental Health Perspectives, 119, 1034–1041. doi:10.1289/ehp.1002775.
Dahlhoff, M., Pfister, S., Blutke, A., Rozman, J., Klingenspor, M., Deutsch, M. J., et al. (2014). Peri-conceptional obesogenic exposure induces sex-specific programming of disease susceptibilities in adult mouse offspring. Biochimica et Biophysica Acta, 1842, 304–317. doi:10.1016/j.bbadis.2013.11.021.
Dewulf, E. M., Cani, P. D., Claus, S. P., Fuentes, S., Puylaert, P. G. B., Neyrinck, A. M., et al. (2013). Insight into the prebiotic concept: lessons from an exploratory, double blind intervention study with inulin-type fructans in obese women. Gut, 62, 1112–1121. doi:10.1136/gutjnl-2012-303304.
Dong, M., Zheng, Q., Ford, S. P., Nathanielsz, P. W., & Ren, J. (2013). Maternal obesity, lipotoxicity and cardiovascular diseases in offspring. Journal of Molecular and Cellular Cardiology, 55, 111–116. doi:10.1016/j.yjmcc.2012.08.023.
Duggan, G. E., Hittel, D. S., Hughey, C. C., Weljie, A., Vogel, H. J., & Shearer, J. (2011). Differentiating short- and long-term effects of diet in the obese mouse using (1) H-nuclear magnetic resonance metabolomics. Diabetes Obesity and Metabolism, 13, 859–862. doi:10.1111/j.1463-1326.2011.01410.x.
Dumas, M.-E., Maibaum, E. C., Teague, C., Ueshima, H., Zhou, B., Lindon, J. C., et al. (2006). Assessment of analytical reproducibility of 1H NMR spectroscopy based metabonomics for large-scale epidemiological research: the INTERMAP Study. Analytical Chemistry, 78, 2199–2208. doi:10.1021/ac0517085.
Farley, D. M., Choi, J., Dudley, D. J., Li, C., Jenkins, S. L., Myatt, L., & Nathanielsz, P. W. (2010). Placental amino acid transport and placental leptin resistance in pregnancies complicated by maternal obesity. Placenta, 31, 718–724. doi:10.1016/j.placenta.2010.06.006.
Fiehn, O. (2002). Metabolomics–the link between genotypes and phenotypes. Plant Molecular Biology, 48, 155–171.
Friedrich, N., Budde, K., Wolf, T., Jungnickel, A., Grotevendt, A., Dressler, M., et al. (2012). Short-term changes of the urine metabolome after bariatric surgery. Omics A Journal of Integrative Biology, 16, 612–620. doi:10.1089/omi.2012.0066.
Godfrey, K. M., & Barker, D. J. (2000). Fetal nutrition and adult disease. American Journal of Clinical Nutrition, 71, 1344S–1352S.
Gross, M. (2013). Does the gut microbiome hold clues to obesity and diabetes? Current Biology, 23, R359–R362.
Gudbrandsen, O. A., Wergedahl, H., Liaset, B., Espe, M., Mørk, S., & Berge, R. K. (2008). Dietary single cell protein reduces fatty liver in obese Zucker rats. British Journal of Nutrition, 100, 776–785. doi:10.1017/S0007114508960906.
Guzmán, C., Cabrera, R., Cárdenas, M., Larrea, F., Nathanielsz, P. W., & Zambrano, E. (2006). Protein restriction during fetal and neonatal development in the rat alters reproductive function and accelerates reproductive ageing in female progeny. Journal of Physiology, 572, 97–108. doi:10.1113/jphysiol.2005.103903.
Hanzu, F. A., Vinaixa, M., Papageorgiou, A., Párrizas, M., Correig, X., Delgado, S., et al. (2013). Obesity rather than regional fat depots marks the metabolomic pattern of adipose tissue: An untargeted metabolomic approach. Obesity,. doi:10.1002/oby.20541.
Inadera, H. (2013). Developmental origins of obesity and type 2 diabetes: molecular aspects and role of chemicals. Environmental Health and Preventive Medicine, 18, 185–197. doi:10.1007/s12199-013-0328-8.
Janesick, A., & Blumberg, B. (2011). Endocrine disrupting chemicals and the developmental programming of adipogenesis and obesity. Birth Defects Research Part C Embryo Today: Reviews, 93, 34–50. doi:10.1002/bdrc.20197.
Jones, H. N., Woollett, L. A., Barbour, N., Prasad, P. D., Powell, T. L., & Jansson, T. (2009). High-fat diet before and during pregnancy causes marked up-regulation of placental nutrient transport and fetal overgrowth in C57/BL6 mice. The FASEB Journal, 23, 271–278. doi:10.1096/fj.08-116889.
Karmaus, W., Osuch, J. R., Eneli, I., Mudd, L. M., Zhang, J., Mikucki, D., et al. (2009). Maternal levels of dichlorodiphenyl-dichloroethylene (DDE) may increase weight and body mass index in adult female offspring. Occupational and Environmental Medicine, 66, 143–149. doi:10.1136/oem.2008.041921.
Kim, K. E., Onesti, G., Ramirez, O., Brest, A. N., & Swartz, C. (1969). Creatinine clearance in renal disease. A reappraisal. British Medical Journal, 4, 11–14.
Kim, M. J., Yang, H. J., Kim, J. H., Ahn, C.-W., Lee, J. H., Kim, K. S., & Kwon, D. Y. (2013). Obesity-related metabolomic analysis of human subjects in black soybean peptide intervention study by ultraperformance liquid chromatography and quadrupole-time-of-flight mass spectrometry. Journal of Obesity, 2013, 874981. doi:10.1155/2013/874981.
Kim, S.-H., Yang, S.-O., Kim, H.-S., Kim, Y., Park, T., & Choi, H.-K. (2009). 1H-nuclear magnetic resonance spectroscopy-based metabolic assessment in a rat model of obesity induced by a high-fat diet. Analytical and Bioanalytical Chemistry, 395, 1117–1124. doi:10.1007/s00216-009-3054-8.
Kirk, S. L., Samuelsson, A.-M., Argenton, M., Dhonye, H., Kalamatianos, T., Poston, L., et al. (2009). Maternal obesity induced by diet in rats permanently influences central processes regulating food intake in offspring. PLoS One, 4, e5870. doi:10.1371/journal.pone.0005870.
Lee, J. V., Shah, S. A., & Wellen, K. E. (2013). Obesity, cancer, and acetyl-CoA metabolism. Drug Discovery Today Disease Mechanisms, 10, e55–e61. doi:10.1016/j.ddmec.2013.03.005.
Lees, H. J., Swann, J. R., Wilson, I. D., Nicholson, J. K., & Holmes, E. (2013). Hippurate: the natural history of a mammalian-microbial cometabolite. Journal of Proteome Research, 12, 1527–1546. doi:10.1021/pr300900b.
Li, H., Xie, Z., Lin, J., Song, H., Wang, Q., Wang, K., et al. (2008). Transcriptomic and metabonomic profiling of obesity-prone and obesity-resistant rats under high fat diet. Journal of Proteome Research, 7, 4775–4783. doi:10.1021/pr800352k.
Lowe, W. L., & Karban, J. (2014). Genetics, genomics and metabolomics: new insights into maternal metabolism during pregnancy. Diabetic Medicine, 31, 254–262. doi:10.1111/dme.12352.
Magyar, D. M., Fridshal, D., Elsner, C. W., Glatz, T., Eliot, J., Klein, A. H., et al. (1980). Time-trend analysis of plasma cortisol concentrations in the fetal sheep in relation to parturition. Endocrinology, 107, 155–159. doi:10.1210/endo-107-1-155.
Muhlhausler, B. S., Gugusheff, J. R., Ong, Z. Y., & Vithayathil, M. A. (2013). Nutritional approaches to breaking the intergenerational cycle of obesity. Canadian Journal of Physiology and Pharmacology, 91, 421–428. doi:10.1139/cjpp-2012-0353.
Norman, J. D., Ferguson, M. M., & Danzmann, R. G. (2014). Transcriptomics of salinity tolerance capacity in Arctic charr (Salvelinus alpinus): A comparison of gene expression profiles between divergent QTL genotypes. Physiological Genomics, 46, 123–137. doi:10.1152/physiolgenomics.00105.2013.
O’Reilly, J. R., & Reynolds, R. M. (2013). The risk of maternal obesity to the long-term health of the offspring. Clinical Endocrinology, 78, 9–16. doi:10.1111/cen.12055.
Orsi, C. M., Hale, D. E., & Lynch, J. L. (2011). Pediatric obesity epidemiology. Current opinion in Endocrinology Diabetes and Obesity, 18, 14–22. doi:10.1097/MED.0b013e3283423de1.
Phipps, A. N., Stewart, J., Wright, B., & Wilson, I. D. (1998). Effect of diet on the urinary excretion of hippuric acid and other dietary-derived aromatics in rat. A complex interaction between diet, gut microflora and substrate specificity. Xenobiotica, 28, 527–537. doi:10.1080/004982598239443.
Proczko, M., Kaska, Ł., Kobiela, J., Stefaniak, T., Zadrożny, D., & Śledziński, Z. (2013). Bariatric surgery in morbidly obese patients with chronic renal failure, prepared for kidney transplantation–case reports. Polski Przeglad Chirurgiczny, 85, 407–411. doi:10.2478/pjs-2013-0062.
Respondek, F., Gerard, P., Bossis, M., Boschat, L., Bruneau, A., Rabot, S., et al. (2013). Short-chain fructo-oligosaccharides modulate intestinal microbiota and metabolic parameters of humanized gnotobiotic diet induced obesity mice. PLoS One, 8, e71026. doi:10.1371/journal.pone.0071026.
Rull, A., Vinaixa, M., Angel Rodríguez, M., Beltrán, R., Brezmes, J., Cañellas, N., et al. (2009). Metabolic phenotyping of genetically modified mice: An NMR metabonomic approach. Biochimie, 91, 1053–1057. doi:10.1016/j.biochi.2009.04.019.
Salek, R. M., Maguire, M. L., Bentley, E., Rubtsov, D. V., Hough, T., Cheeseman, M., et al. (2007). A metabolomic comparison of urinary changes in type 2 diabetes in mouse, rat, and human. Physiological Genomics, 29, 99–108. doi:10.1152/physiolgenomics.00194.2006.
Sarlio-Lähteenkorva, S., Silventoinen, K., Jousilahti, P., Hu, G., & Tuomilehto, J. (2004). The association between thinness and socio-economic disadvantage, health indicators, and adverse health behaviour: a study of 28 000 Finnish men and women. International Journal of Obesity, 28, 568–573. doi:10.1038/sj.ijo.0802596.
Schirra, H. J., Anderson, C. G., Wilson, W. J., Kerr, L., Craik, D. J., Waters, M. J., & Lichanska, A. M. (2008). Altered metabolism of growth hormone receptor mutant mice: a combined NMR metabonomics and microarray study. PLoS One, 3, e2764. doi:10.1371/journal.pone.0002764.
Serkova, N. J., Jackman, M., Brown, J. L., Liu, T., Hirose, R., Roberts, J. P., et al. (2006). Metabolic profiling of livers and blood from obese Zucker rats. Journal of Hepatology, 44, 956–962. doi:10.1016/j.jhep.2005.07.009.
Shearer, J., Duggan, G., Weljie, A., Hittel, D. S., Wasserman, D. H., & Vogel, H. J. (2008). Metabolomic profiling of dietary-induced insulin resistance in the high fat-fed C57BL/6 J mouse. Diabetes Obesity and Metabolism, 10, 950–958. doi:10.1111/j.1463-1326.2007.00837.x.
Shelley, P., Martin-Gronert, M. S., Rowlerson, A., Poston, L., Heales, S. J. R., Hargreaves, I. P., et al. (2009). Altered skeletal muscle insulin signaling and mitochondrial complex II-III linked activity in adult offspring of obese mice. American Journal of Physiology: Regulatory Integrative and Comparative Physiology, 297, R675–R681. doi:10.1152/ajpregu.00146.2009.
Simmons, R. A., Suponitsky-Kroyter, I., & Selak, M. A. (2005). Progressive accumulation of mitochondrial DNA mutations and decline in mitochondrial function lead to beta-cell failure. Journal of Biological Chemistry, 280, 28785–28791. doi:10.1074/jbc.M505695200.
Smink, A., Ribas-Fito, N., Garcia, R., Torrent, M., Mendez, M. A., Grimalt, J. O., & Sunyer, J. (2008). Exposure to hexachlorobenzene during pregnancy increases the risk of overweight in children aged 6 years. Acta Paediatrica, 1992(97), 1465–1469. doi:10.1111/j.1651-2227.2008.00937.x.
Smit, S., van Breemen, M. J., Hoefsloot, H. C. J., Smilde, A. K., Aerts, J. M. F. G., & de Koster, C. G. (2007). Assessing the statistical validity of proteomics based biomarkers. Analytica Chimica Acta, 592, 210–217. doi:10.1016/j.aca.2007.04.043.
Smolinska, A., Blanchet, L., Buydens, L. M. C., & Wijmenga, S. S. (2012). NMR and pattern recognition methods in metabolomics: from data acquisition to biomarker discovery: a review. Analytica Chimica Acta, 750, 82–97. doi:10.1016/j.aca.2012.05.049.
Suter, M. A., Sangi-Haghpeykar, H., Showalter, L., Shope, C., Hu, M., Brown, K., et al. (2012). Maternal high-fat diet modulates the fetal thyroid axis and thyroid gene expression in a nonhuman primate model. Molecular Endocrinology, 26, 2071–2080. doi:10.1210/me.2012-1214.
Symonds, M. E., Mendez, M. A., Meltzer, H. M., Koletzko, B., Godfrey, K., Forsyth, S., & van der Beek, E. M. (2013). Early life nutritional programming of obesity: mother-child cohort studies. Annals of Nutrition and Metabolism, 62, 137–145. doi:10.1159/000345598.
Szymańska, E., Saccenti, E., Smilde, A. K., & Westerhuis, J. A. (2012). Double-check: validation of diagnostic statistics for PLS-DA models in metabolomics studies. Metabolomics, 8, 3–16. doi:10.1007/s11306-011-0330-3.
Tuersunjiang, N., Odhiambo, J. F., Long, N. M., Shasa, D. R., Nathanielsz, P. W., & Ford, S. P. (2013). Diet reduction to requirements in obese/overfed ewes from early gestation prevents glucose/insulin dysregulation and returns fetal adiposity and organ development to control levels. American Journal of Physiology-Endocrinology and Metabolism, 305(7), E868–E878. doi:10.1152/ajpendo.00117.2013.
Vinayavekhin, N., Homan, E. A., & Saghatelian, A. (2010). Exploring disease through metabolomics. ACS Chemical Biology, 5, 91–103. doi:10.1021/cb900271r.
Waldram, A., Holmes, E., Wang, Y., Rantalainen, M., Wilson, I. D., Tuohy, K. M., et al. (2009). Top-down systems biology modeling of host metabotype-microbiome associations in obese rodents. Journal of Proteome Research, 8, 2361–2375. doi:10.1021/pr8009885.
Wang, C., Feng, R., Sun, D., Li, Y., Bi, X., & Sun, C. (2011). Metabolic profiling of urine in young obese men using ultra performance liquid chromatography and Q-TOF mass spectrometry (UPLC/Q-TOF MS). Journal of Chromatography, 879, 2871–2876. doi:10.1016/j.jchromb.2011.08.014.
Waterman, D. S., Bonner, F. W., & Lindon, J. C. (2009). Review: Spectroscopic and statistical methods in metabonomics. Bioanalysis, 1(9), 1559–1578. doi:10.4155/bio.09.143.
Wen, X., Triche, E. W., Hogan, J. W., Shenassa, E. D., & Buka, S. L. (2011). Prenatal factors for childhood blood pressure mediated by intrauterine and/or childhood growth? Pediatrics, 127, e713–e721. doi:10.1542/peds.2010-2000.
Werner, E., Heilier, J.-F., Ducruix, C., Ezan, E., Junot, C., & Tabet, J.-C. (2008). Mass Spectrometry for the Identification of the Discriminating Signals from Metabolomics: Current Status and Future Trends. Journal of Chromatography B, 871(2), 143–163.
Whitehead, T. L., Holley, A. W., Korourian, S., Shaaf, S., Kieber-Emmons, T., & Hakkak, R. (2007). (1)H nuclear magnetic resonance metabolomic analysis of mammary tumors from lean and obese Zucker rats exposed to 7,12-dimethylbenz[a]anthracene. International Journal of Molecular Medicine, 20, 573–580.
WHO Expert Consultation. (2004). Appropriate body-mass index for Asian populations and its implications for policy and intervention strategies. Lancet, 363, 157–163. doi:10.1016/S0140-6736(03)15268-3.
Williams, R. E., Eyton-Jones, H. W., Farnworth, M. J., Gallagher, R., & Provan, W. M. (2002). Effect of intestinal microflora on the urinary metabolic profile of rats: a (1)H-nuclear magnetic resonance spectroscopy study. Xenobiotica, 32, 783–794. doi:10.1080/00498250210143047.
Williams, R. E., Lenz, E. M., Evans, J. A., Wilson, I. D., Granger, J. H., Plumb, R. S., & Stumpf, C. L. (2005). A combined (1)H NMR and HPLC-MS-based metabonomic study of urine from obese (fa/fa) Zucker and normal Wistar-derived rats. Journal of Pharmaceutical and Biomedical Analysis, 38, 465–471. doi:10.1016/j.jpba.2005.01.013.
Won, E.-Y., Yoon, M.-K., Kim, S.-W., Jung, Y., Bae, H.-W., Lee, D., et al. (2013). Gender-specific metabolomic profiling of obesity in leptin-deficient ob/ob mice by 1H NMR spectroscopy. PLoS One, 8, e75998. doi:10.1371/journal.pone.0075998.
Worley, B., & Powers, R. (2013). Multivariate analysis in metabolomics. Current Metabolomics, 1, 92–107. doi:10.2174/2213235X11301010092.
Xie, B., Waters, M. J., & Schirra, H. J. (2012). Investigating potential mechanisms of obesity by metabolomics. BioMed Research International, 2012, 805683. doi:10.1155/2012/805683.
Yan, X., Huang, Y., Zhao, J.-X., Rogers, C. J., Zhu, M.-J., Ford, S. P., et al. (2013). Maternal obesity downregulates microRNA let-7 g expression, a possible mechanism for enhanced adipogenesis during ovine fetal skeletal muscle development. International Journal of Obesity, 2005(37), 568–575. doi:10.1038/ijo.2012.69.
Zambrano, E., & Nathanielsz, P. W. (2013). Mechanisms by which maternal obesity programs offspring for obesity: evidence from animal studies. Nutrition Reviews, 71(Suppl 1), S42–S54. doi:10.1111/nure.12068.
Zhang, A., Sun, H., & Wang, X. (2013). Power of metabolomics in biomarker discovery and mining mechanisms of obesity. Obesity Reviews, 14, 344–349. doi:10.1111/obr.12011.
Zhao, L.-C., Zhang, X.-D., Liao, S.-X., Gao, H.-C., Wang, H.-Y., & Lin, D.-H. (2010). A metabonomic comparison of urinary changes in Zucker and GK rats. BioMed Research International, 2010, 431894. doi:10.1155/2010/431894.
Zhou, Y., Qiu, L., Xiao, Q., Wang, Y., Meng, X., Xu, R., et al. (2013). Obesity and diabetes related plasma amino acid alterations. Clinical Biochemistry, 46, 1447–1452. doi:10.1016/j.clinbiochem.2013.05.045.
Acknowledgments
We acknowledge financial support from the EHESP School of Public Health, France. We gratefully acknowledge Marie Tremblay-Franco for helpful advice on statistics, and Sven Delaye for advice on English translation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Human and Animal Rights and Informed Consent
Gynaecologists informed the women of the nature of the study and asked them to participate, after providing written consent. This consent was accompanied by a letter of information describing the goal of the study, the consortium, data collection procedures and follow-up after birth (via questionnaires and medical examinations). Explicit mention was made of the right to refuse to participate, and the fact that such a refusal would not have any effect on the woman's relationship with her doctor. Both the INSERM (French National Institute of Health and Medical Research) ethics committee and the National Commission in charge of Data Protection (CNIL) approved the study procedures. Reference (N°902076; 31 may 2002).
Rights and permissions
About this article
Cite this article
Desert, R., Canlet, C., Costet, N. et al. Impact of maternal obesity on the metabolic profiles of pregnant women and their offspring at birth. Metabolomics 11, 1896–1907 (2015). https://doi.org/10.1007/s11306-015-0836-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11306-015-0836-1