Abstract
In view of the extensive potential applications of chitinase (ChiA) in various fields such as agriculture, environmental protection, medicine, and biotechnology, the development of a high-yielding strain capable of producing chitinase with enhanced activity holds significant importance. The objective of this study was to utilize the extracellular chitinase from Bacillus thuringiensis as the target, and Bacillus licheniformis as the expression host to achieve heterologous expression of ChiA with enhanced activity. Initially, through structural analysis and molecular dynamics simulation, we identified key amino acids to improve the enzymatic performance of chitinase, and the specific activity of chitinase mutant D116N/E118N was 48% higher than that of the natural enzyme, with concomitant enhancements in thermostability and pH stability. Subsequently, the expression elements of ChiA(D116N/E118N) were screened and modified in Bacillus licheniformis, resulting in extracellular ChiA activity reached 89.31 U/mL. Further efforts involved the successful knockout of extracellular protease genes aprE, bprA and epr, along with the gene clusters involved in the synthesis of by-products such as bacitracin and lichenin from Bacillus licheniformis. This led to the development of a recombinant strain, DW2△abelA, which exhibited a remarkable improvement in chitinase activity, reaching 145.56 U/mL. To further improve chitinase activity, a chitinase expression frame was integrated into the genome of DW2△abelA, resulting in a significant increas to 180.26 U/mL. Optimization of fermentation conditions and medium components further boosted shake flask enzyme activity shake flask enzyme activity, achieving 200.28 U/mL, while scale-up fermentation experiments yielded an impressive enzyme activity of 338.79 U/mL. Through host genetic modification, expression optimization and fermentation optimization, a high-yielding ChiA strain was successfully constructed, which will provide a solid foundation for the extracellular production of ChiA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chitin, also known as poly-N-acetylglucosamine (GlcNAc), is a high molecular weight polymer consisting of β-(1–4)-linked 2-acetamido-2-deoxy-D-glucose. This versatile compound forms intricate polymeric scaffolds by crosslinking with various proteins, phenols, lipids, or other carbohydrates like β-glucans, thereby composing the outer shells of shrimp, crab, insect exoskeletons, and fungal cell walls (Synowiecki and Al-Khateeb 2003; Liu et al. 2019). In nature, chitin emerges as remarkably abundant, ranking as the second most prevalent polymer after cellulose and hemicellulose (Khan et al. 2017). GlcNAc, as a complete degradation product of chitin, boasts excellent biocompatibility and safe chemical properties. It serves as a crucial component in various biological structures such as urokinase plasminogen activator, hyaluronic acid, and connective tissue substances like glycosaminoglycan and proteoglycan. These applications span across health food, medicine, and other industries (Chen et al. 2010). In agriculture, GlcNAc can enrich the beneficial microorganisms in the soil, thus fostering a conducive growth environment for plant growth and indirectly inhibiting the occurrence of plant diseases and pests (Cardozo et al. 2017). With the burgeoning market demand for GlcNAc, and the current production predominantly relies on the chitin decomposition. Presently, GlcNAc production mainly employs chemical, synthetic, and enzymatic methods. Compared with other conventional methods, enzymatic method has the advantages of mild reaction conditions, less by-products and environmental friendliness, so it is the main research direction and hot spot of green biological preparation of GlcNAc (Chen et al. 2010).
Chitinase (ChiA) is an enzyme that degrades chitin to produce GlcNAc. Chitinase can be divided into endochitinase and exochitinase according to the way and location of action of catalytic substrates (Oyeleye and Normi 2018). Endochitinase can cut the β-1, 4-glucoside bond without selectivity at the internal site, and the cleavage product is soluble low molecular weight oligomer. Exochitinase can be divided into two subclasses: β-N-acetylglucosaminidase (EC 3.2.1.30) can produce oligomers from terminal hydrolysis of chitin, and the hydrolyzed product is GlcNAc monomer; Chitinosidase (EC 3.2.1.29) can catalyze the gradual release of diacetylchitose from the non-reducing end of the polysaccharide chain (Karthik et al. 2015). According to the difference of sequence homology, chitinases can be divided into two families, namely GH18 family and GH19 family (Oyeleye and Normi 2018). The amino acid sequences of these two families have little homology (Henrissat and Bairoch 1993). They are also completely different in structure and catalytic hydrolysis mechanism. In recent years, it has also been found that some chitinases belong to the GH23 family and the GH48 family (Lipski et al. 2015; Nishitani et al. 2018). The hydrolytic activity and catalytic mechanism of chitinase of GH18 family have been extensively studied, and they are widely distributed. Most bacteria, such as Bacillus, Vibrio and Serratia, produce chitinase belonging to GH18 family (Davies and Henrissat 1995). The GH19 family is mainly found in some viruses, bacteria, nematodes, and arachnids (Rao et al. 2005; Hurtado-Guerrero and van Aalten 2007; Tsuji et al. 2010). Interest in the application value of Bacillus thuringiensis chitinase sparked in the 1970s when it was revealed that enzymes secreted by this bacterium possess the capability to hydrolyze chitin. Subsequently, it was demonstrated that B. thuringiensis produces chitinase, which contributes to its toxicity when combined with other components, including Cry protein (Guttmann and Ellar 2000). Shortly thereafter, cloning of the B. thuringiensis chitinase coding gene was reported (Thamthiankul et al. 2001; Barboza-Corona et al., 2008), which initiated the development of B. thuringiensis chitinase expression.
Chitinases have functional domains, including signal peptide sequence, catalytic domain, chitin insertion domain, fibronectin type III domain and chitin binding domain, and a short C-terminal region (Robertus and Monzingo 1999). The catalytic domain determines the catalytic hydrolysis efficiency of chitinases, and the catalytic domain of chitinases of the GH18 family consists of an eight-stranded (β/α) folded barrel structure (Davies and Henrissat 1995). The key active site of their catalytic domain structure is the “DXDXE” conserved sequence composed of 2 aspartate (D) and 1 glutamate (E). In addition, “SXGG” conserved sequences exist on the outer side of the catalytic domain to promote stable binding of the enzyme and substrate (Vaaje-Kolstad et al. 2013; Sugimoto et al. 2015). In terms of protein engineering modification of chitinase, Huang et al. (Huang et al. 2021) used Rosetta supercharge software to perform molecular docking between diacetylchitobiose deacetylase (Dac) and substrate GlcNAc, and then performed alanine scanning and site-specific saturation mutation. A mutant with good enzyme activity (DacG74A) was screened, and its specific enzyme activity reached 5162.17 U/mg, which was 70.02% higher than that of the original enzyme. Li et al. (Xu et al. 2020b) successfully expressed chitinase derived from Streptomyces diastaticus, and analyzed the amino acid sequence of the catalytic domain structure through three-dimensional modeling to find potential mutation sites, revealing the active site of chitinase, which increased the activity of the enzyme by 32% compared with the original enzyme.
In addition to short growth cycle, robust secretion ability, no obvious codon preference, clear genetic background, and rich industrial fermentation information (Schumann 2007; Harwood and Cranenburgh 2008), Bacillus boasts simple nutritional requirements, strong resistance to extreme environments, abundant enzyme systems, biosafety, and other characteristics, and can be widely used in the industrial production of various enzyme preparations, polypeptide polymers, antibiotics and other substances. It is an ideal strain for industrial production with great development potential (Veith et al. 2004; Guo et al. 2015; Azarhava et al. 2020). In recent years, Bacillus licheniformis has been widely used as an excellent expression system in the production of enzyme preparations, such as high temperature resistant α-amylase and alkaline protease (Veith et al. 2004). However, in order to express the enzyme more efficiently, the expression host and expression elements need to be optimized accordingly. In this study, chitinase derived from Bacillus thuringiensis was modified by protein engineering as the target protein, combined with the screening of promoters and signal peptides as well as the optimization of signal peptide, expression host, medium composition, and fermentation process, and finally successfully and efficiently used Bacillus licheniformis DW2 to express chitinase extracellular.
Materials and methods
Strains and culture conditions
The strains and plasmids used in this study were listed in Table S1 and Table S2. Escherichia coli DH5α was employed as the host for recombinant plasmid construction. The B. licheniformis DW2 was applied as the host for ChiA production. E. coli and B. licheniformis were cultured in Luria–Bertani (LB) medium (yeast extract 5 g/L, tryptone 10 g/L, NaCl 10 g/L) at 37 °C with 20 mg/L tetracycline. The optimal fermentation medium for ChiA production consists of soluble corn starch 30 g/L, soybean meal 70 g/L, CaCO3 10 g/L, (NH4)2SO4 2 g/L, KH2PO4 1 g/L and Na2HPO4 2 g/L. The seed culture was inoculated in a 250 mL flask with 50 mL LB medium for 8 h, then 1 mL of the culture was transferred into 30 mL of fermentation medium and cultured for 48 h at 37 °C and 230 rpm. All the fermentation experiments were repeated at least three times.
Determination of mutant amino acid sites
The amino acid sequence of ChiA was submitted to the online server SWISS-MODEL for Build Model, and the obtained PDB format file was imported into the Discovery studio 2020 software for molecular docking. Based on the optimal results of molecular docking, an alanine scanning of amino acids within a 3Å distance of the binding plane was conducted using the calculate mutant energy (binding) functional module. Amino acid sites with ΔΔG changes greater than ± 5 was identified as having high mutation potential, and subsequently, the selected amino acid sites underwent saturated mutation to determine the mutant amino acid sites.
Construction of ChiA expression strains
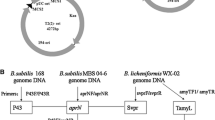
Plasmid DNA was extracted from E. coli by a standard alkaline lysis procedure using Plasmid Mini Kit I (Omega Bio-Tek, Georgia, USA). Restriction enzymes and T4 DNA ligase (TaKaRa Biotechnology Corporation, Dalian, China) were used as recommended by the manufacturer. The oligonucleotide primers (Table S3) used for PCR amplification were synthesized and purified by Sangon Biothch (Shanghai, China). The plasmid pHY300PLK was used for gene expression in B. licheniformis as previously describe (Wang et al. 2020). Chitinase gene chiA from Bacillus thuringiensis BMB171, different constitutive strong promoters (PR5, PR29, PykzA+rbs6 and P43UTR12), the signal peptide of Bacillus clausii AprE (SPAprE) and TamyL terminator were amplified using the corresponding primers in Table S3 and purified by Gel Extraction Kit (Omega Bio-Tek, Georgia, USA). Since the primers of splicing overlap extension PCR (SOE-PCR) all contained a small homologous sequence of adjacent fragments, the PCR fragments that need to be spliced were added to a reaction system, and the chimeric DNA fragments were amplified using the outermost primers. The chimeric DNA fragment was inserted into the EcoRI/XbaI–digested pHY300PLK plasmid to create a series of recombinant plasmids, so as to screen the combination with the best ChiA expression. Subsequently, the promoter P43UTR12 was combined with different signal peptides SPYwbN, SPSacC and mutated SPSacC to screen the best signal peptides for extracellular expression of ChiA.
Plasmid PykzA+rbs6-SPAprE-ChiA and P43 − UTR12-SPSacC-ChiA were used as mutagenesis template. All primers designed to introduce the site-directed mutation were synthesized and purified by Sangon Biothch (Shanghai, China) (Table S3). The PCR amplifications were carried out with 2 × KeyPo Master Mix (Vazyme, China) containing a high-fidelity enzyme, and then treated with restriction enzyme DpnI (Vazyme, China), followed by screening for positive mutants, and finally sequencing to identify mutations.
Gene knockout and integration in B. licheniformis
For gene knockout, the upstream and downstream homologous arms of the gene to be knocked out were fused into a fragment and ligated into the backbone of T2(2)-Ori to obtain the recombinant plasmids. The gene integration plasmid was added ChiA expression cassettes between the upstream and downstream homologous arms of integration sites. Gene deletion and integration were constructed though the homologous recombination system according to the previous method (Zhan et al. 2022). All the recombinant strains were verified by diagnostic PCR and DNA sequencing.
Determination of ChiA expression
At the end of each fermentation, the fermentation broth volume was assessed to ensure that the remaining fermentation broth volume was similar. The fermentation broth was then harvested by centrifugation. The supernatant was concentrated tenfold, and aliquots comprised of 10 µl of the extracellular proteins mixed with the loading dye were subjected to SDS-PAGE. SDS-PAGE was performed using a 4% acrylamide stacking gel and 12% acrylamide resolving gel. To determine the expression of ChiA more accurately, the enzyme activity of ChiA was detected. The substrate colloidal chitin and diluted crude enzyme solution of ChiA were preheated in a 60 °C constant temperature water bath for 5 min. Then, 1 mL of the crude enzyme solution was added to a reaction test tube, along with 500 µL of pH 7.0 PBS buffer and 500 µL of 2% colloidal chitin. After thorough mixing, the reaction mixture was incubated at 60 °C for 20 min. Subsequently, the mixture was immediately heated in a boiling water bath for 15 min to denature the ChiA. The mixture was then centrifuged at 12,000 rpm for 5 min, and 1 mL of the supernatant was collected. To the supernatant, 1 mL of DNS solution was added, and a 15-minute boiling water bath coloration was performed. The absorbance at 540 nm was measured to determine the ChiA activity. In the control group, the first step involved boiling the diluted crude enzyme solution of ChiA in a boiling water bath for 15 min, followed by the subsequent steps. The enzyme activity was calculated based on the difference between the experimental group and the control group. One enzymatic unit of was defined as the amount of enzyme needed to produce 1 mg of N-acetylglucose from colloidal chitin per minute at 60 °C and pH 7.0.
Effect of pH and temperature on the enzyme activity
The enzymatic hydrolysis of colloidal chitin using the crude extract of chitinase was performed at different pH conditions (ranging from 4 to 9, 50 mmol/L acetate buffer provides pH4.5-5.5; 50 mmol/L phosphate buffer provides pH6.0-7.0; 50 mmol/L Tris-HCl buffer provides pH7.5-9.0) to evaluate the optimum pH for the maximum chitinase activity. For pH stability, the crude enzyme solution was treated at the specified pH conditions for 1 h to detect residual enzyme activity. Likewise, another set of experiments was performed to evaluate the effect of temperature (ranging from 30 to 70 °C) on the chitinase activity at pH 7. For temperature stability, the crude enzyme solution was treated at a specified temperature for 1 h to detect residual enzyme activity.
Optimization of fermentation medium and fermentation process
The composition of the initial fermentation medium is soluble corn starch 45 g/L, soybean meal 100 g/L, CaCO3 1 g/L and (NH4)2SO4 1 g/L. To increase ChiA expression levels, the ratio of soybean meal to corn starch (100/45, 80/35, 70/30, 60/25, 50/22.5 g/L), the amount of CaCO3 (6, 8, 10, 12, 14 g/L), and the amount of (NH4)2SO4 (1, 2, 3, 4, 5 g/L) and potassium dihydrogen phosphate/disodium hydrogen phosphate ratio (0, 1/2, 2/4, 3/6, 6/12, 12/24 g/L) were optimized.
Recombinant B. licheniformis was cultured in the optimal ChiA production medium using 5% of a seed culture and aeration rate of 1.5 vvm in a 5 L fermenter to maximize ChiA production. The lag period (0–4 h) was stirred at 500 rpm, and then mixed with dissolved oxygen to maintain the dissolved oxygen content at about 20% (4–33 h). During the fermentation process, 600 g/L glucose was added to maintain the glucose concentration at 5 ~ 10 g/L.
Statistical analysis
All the experiments in this study were performed in triplicate, and the data are presented as the means ± standard deviations. Data were analyzed using software SPSS 18.0 (SPSS Inc., Chicago, IL, USA), by the one-way analysis of variance (ANOVA) followed by the Duncan’s multiple range test. Statistical significance was determined at the α = 0.05 level.
Results
Molecular dynamics simulation to improve specific activity of chitinase
Homology model building was performed using the SWISS-MODEL (http://swiss-model.expasy.org) online, and the chitinase ChiL from Chitiniphilus shinanonensis (PDB ID: 6kst.1) was selected as the template due to its highest amino acid sequence identity with Bacillus thuringiensis BMB171 (Accession: NZ_CM000753.1) chitinase ChiA in this study. The structure of Bacillus thuringiensis chitinase ChiA comprises 12 α helices wrapped with 13 inverted parallel β sheets and some random coils (Figure S1). The mutation sites of chitinase were predicted by Receptor-Ligand Interactions function module and Calculate Mutant Energy (binding) function module in Discovery Studio software. Amino acids within 3Å of the binding plane were scanned by alanine, and sites with ΔΔG changes exceeding ± 5 were considered as sites with high mutation potential. Then, the selected amino acid sites were saturated mutated to obtain potential mutation sites, which were N83W, D116N, E118I, E118N and A161R (Table S4).
Five recombinant strains expressing single-mutation ChiA were constructed and ChiA activity was detected. Compared with the original enzyme, D116N and E118N mutants increased the enzyme activity by 18% and 43%, respectively (Fig. 1). Subsequently, the two effective mutations were superimposed to obtain the recombinant strain DW2/PykzA+rbs6-SPAprE-ChiA(D116N/E118N). The extracellular enzyme activity of the double-mutation strain was 110% higher than that of the original strain (Fig. 1).
By performing molecular docking between the original and mutated ChiA enzymes, it is evident that the binding pocket is enlarged after mutation (Fig. 2). The amino acid side chains formed by Asp116-Asn116 and Glu118-Asn118 can form water-mediated H-bonds with the substrate chitin small molecules, which can further strengthen the existing H-bond network, resulting in tighter binding between chitinase and substrate chitin. Both the 116th and 118th mutation sites are in the sugar substrate binding crack, and Asp116-Asn116 and Glu118-Asn118 form hydrophobic planes with other sites, which greatly enhances ChiA’s ability to bind chitin. This indirectly results in higher extracellular activity of the overlapping mutant enzyme than that of the single mutant enzyme. However, when the N83 site was mutated to tryptophan and the A161 site was mutated to arginine, although they are in the sugar substrate binding crack, they are not key binding sites for the sugar moiety. Therefore, the extracellular activity of ChiA did not significantly improve, and instead decreased.
Comparison of molecular docking of the original and mutant enzymes. A: Molecular docking diagram of the original enzyme. The binding pocket of the original enzyme is small, and it is very difficult for the ligand to enter the active center. B: Molecular docking diagram of ChiA(D116N). The yellow amino acid indicates the site of the mutation, which is located near the binding pocket, which affects the size of the binding pocket after mutation, and makes it easier for the ligand to enter the active center. C: Molecular docking diagram of ChiA(E118N). The yellow amino acid indicates the site of the mutation, which is located near the binding pocket, which affects the size of the binding pocket after mutation, and makes it easier for the ligand to enter the active center. D: Molecular docking diagram of ChiA(D116N/E118N). The two yellow amino acids indicate the sites of the two mutations, the binding pocket becomes larger after the double mutation, and the ligand has easier access to the active center
Comparison of enzymological properties of original and double mutant enzyme
According to the enzyme activity and temperature curve, the enzyme activity of both the original enzyme and the double mutant enzyme increased with the gradual increase of temperature, but the enzyme activity of both decreased sharply when the temperature exceeded 60℃, indicating that the optimal reaction temperature of both the original enzyme and the double mutant enzyme was 60℃. In addition, the specific activity of the double mutant enzyme was significantly higher than that of the original enzyme at the temperature range of 30℃ to 70℃ (Fig. 3A). The thermal stability of the double mutant enzyme was consistently better than that of the original enzyme under various temperature conditions (Fig. 3B). In terms of pH, both the original enzyme and the double mutant enzyme exhibited the highest activity at pH 7 (Fig. 3C) and maintained high pH stability under both acidic and alkaline conditions (Fig. 3D). In addition, the double mutant enzyme showed better pH stability than the original enzyme at the pH range of 4 to 9 (Fig. 3D).
Comparison of enzymatic properties between wild-type enzyme and mutant. A: Effect of temperature on chitinase activity of wild type and mutant. B: Thermostability of the wild-type and mutant chitinase. C: Effect of pH on chitinase activity of wild-type and mutant. D: pH stability between wild type and mutant chitinase. Data are mean ± SD and experiments were repeated three times with similar results
Screening and optimization of chitinase expression elements in Bacillus licheniformis
Promoter is an important expression element of gene initiation transcription, which directly affects the expression level of genes to be expressed. In this study, four commonly used constitutive strong promoters Pykza+rbs6, P43 − UTR12, PR5, PR29 were selected to cooperate with signal peptide AprE to express chitinase extracellular. SDS-PAGE analysis showed that all four promoters could direct the expression of chitinase with the size of 36 kDa (Fig. 4A). The enzyme activity of chitinase mediated by promoter P43 − UTR12 was the highest, reaching 52.86 U/mL. Compared with the other three promoters (Pykza+rbs6, PR5 and PR29), the enzyme activity of chitinase was increased by 15%, 71% and 66%, respectively (Fig. 4B), which was consistent with the results of SDS-PAGE analysis.
Signal peptide is a short peptide sequence (usually 16–30 amino acids long) that exists in the N-terminal of the target protein, and its function is to guide the extracellular secretion of the target protein. Therefore, the selection of signal peptide has a great influence on the efficient secretion of foreign protein. In this study, commonly used fructanase signal peptide SacC (SPSacC) from Bacillus subtilis 168, double arginine type signal peptide YwbN (SPYwbN) from Bacillus subtilis 168 and alkaline protease signal peptide AprE (SPAprE) from Bacillus clausii were selected to construct different signal peptide-mediated chitinase expression strains, and fermentation and enzyme activity detection were carried out. SDS-PAGE analysis showed that all three signal peptides could successfully guide the extracellular expression of chitinase (Fig. 4C). When SPSacC was used, ChiA expression level was the highest, and its enzyme activity reached 76.56 U/mL, which was 44% and 23% higher than that of the other two signal peptides (SPAprE and SPYwbN), respectively (Fig. 4D).
Signal peptides are divided into N region, H region and C region. N region is rich in positively charged amino acids and interacts with the transport target protein, H region is rich in hydrophobic amino acid residues, and C region has a signal peptidase cutting site for signal peptide cutting. Significant progress has been made in secretion evolution by modifying the structure of signal peptides (Jiang et al. 2019). To enhance the secretion of ChiA by the SPSacC, the functional region of SPSacC was optimized in this study (Table S5). The amino acid composition of the N-region of SPSacC is Met-Lys-Lys-Arg, and the signal peptide SPSacC(R4K) was obtained by replacing the fourth arginine with lysine, which may affect the transport efficiency of the target protein. Signal peptide mutants SPSacC(L5K), SPSacC(L5D), SPSacC(L14D), SPSacC(L14Q) and SPSacC(L15Q) increased the N-terminal charge from + 2 to + 4 and reduced the hydrophobicity of the H-region. The expression of ChiA can be clearly detected in the crude enzyme solution of all the engineered strains modified with signal peptides by SDS-PAGE (Fig. 4E). The results of enzyme activity detection showed that the signal peptide SPSacC(R4K) had the best expression effect of chitinase among the 6 signal peptide mutants, and the activity of chitinase was up to 89.31 U/mL (Fig. 4F). Compared with the control strain, it was the only one with significantly better expression effect than the control strain.
Screening the optimal promoters and signal peptides for extracellular chitinase expression in Bacillus licheniformis. A: SDS-PAGE analysis of chitinase expression by different promoters. M means protein marker. PR5 means DW2/PR5-SPAprE-ChiA(D116N/E118N). PR29 means DW2/PR29-SPAprE-ChiA(D116N/E118N). Pykza+rbs6 means Pykza+rbs6-DW2/Pykza+rbs6-SPAprE-ChiA(D116N/E118N). P43-UTR12 means P43-UTR12-DW2/P43-UTR12-SPAprE-ChiA(D116N/E118N). B: Effects of different promoters on chitinase activity. C: SDS-PAGE analysis of different signal peptides on chitinase expression. M means protein marker. SPAprE means DW2/P43-UTR12-SPAprE-ChiA(D116N/E118N). SPYwbN means DW2/P43-UTR12-SPYwbN-ChiA(D116N/E118N). SPSacC means DW2/P43-UTR12-SPSacC-ChiA(D116N/E118N). D: Effect of different signal peptides on the synthesis of chitinase. E: SDS-PAGE analysis of chitinase expression by different SacC signal peptide mutants. M means protein marker. L5Q means DW2/P43-UTR12-SPAprE(L5Q)-ChiA(D116N/E118N). R4K means DW2/P43-UTR12-SPAprE(R4K)-ChiA(D116N/E118N). L5D means DW2/P43-UTR12-SPAprE(L5D)-ChiA(D116N/E118N). L14D means DW2/P43-UTR12-SPAprE(L14D)-ChiA(D116N/E118N). L14Q means DW2/P43-UTR12-SPAprE(L14Q)-ChiA(D116N/E118N). L15Q means DW2/P43-UTR12-SPAprE(L15Q)-ChiA(D116N/E118N). F: Effect of different SacC signal peptide mutants on the chitinase enzymatic activity
Construction of Bacillus licheniformis host bacteria with high ChiA expression
By analyzing the fermentation curve of strain DW2/P43 − UTR12-SPSacC(R4K)-ChiA(D116N/E118N) (DW2/ChiA* for short), it was found that ChiA expression decreased rather than increased in the late fermentation period, which may be due to the degradation of ChiA by extracellular protease of Bacillus licheniformis. Therefore, we analyzed the key extracellular proteases of Bacillus licheniformis, and knocked out the extracellular protease genes aprE, bprA, and epR, and successfully constructed three protease-deficient expression hosts of Bacillus licheniensis DW2△aprE△bprA△epR (DW2△abe for short) (Figure S3). Based on DW2△abe, chitinase free expression plasmid was introduced to construct recombinant strain DW2△abe/P43 − UTR12-SPSacC(R4K)-ChiA(D116N/E118N) (DW2△abe/ChiA* for short). Through analysis of fermentation curve, it was found that ChiA degradation at the late fermentation stage was significantly relieved, and the activity of ChiA at the fermentation end point reached 116.88 U/mL (Fig. 5A), which was 36% higher than that of the control strain DW2/ChiA*. This suggests that weakening extracellular enzymes is beneficial to the efficient expression of ChiA.
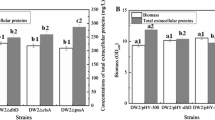
During ChiA fermentation test, we found that the fermentation solution of DW2△abe/ChiA* was accompanied by the generation of bacitracin and lichenin as by-products, and the generation of lichenin would produce a large amount of foam, which was not conducive to the production of ChiA and the later amplification culture. Therefore, on the basis of Bacillus licheniformis DW2△abe, we continued to knock out the lichenin synthase gene cluster lchAC and the bacitracin synthase gene cluster bacABC, and obtained the strain DW2△abelA (Figure S3). Similarly, chitinase free expression plasmid was introduced into the constructed host bacterium DW2△abelA and fermentation was performed. The fermentation process showed that the foam produced by strain DW2△abelA/P43 − UTR12-SPSacC(R4K)-ChiA(D116N/E118N) (DW2△abelA/ChiA* for short) was significantly reduced in the late fermentation period, and no bacitracin was produced in the supernatant of the fermentation solution, and the activity of ChiA was increased to 145.56 U/mL (Fig. 5A).
Plasmid dependent expression systems have become the first choice for foreign protein expression. However, the plasmid instability is a problem that cannot be ignored in industrial production. The plasmid-dependent expression system of ChiA may also show plasmid loss in the late fermentation period. Therefore, in order to stably express ChiA, we constructed plasmid T2::P43 − UTR12-SPSacC(R4K)-ChiA(D116N/E118N) to integrate the ChiA expression frame into the genome of Bacillus licheniformis DW2△abelA by homologous recombination. The recombinant strain was shortened to DW2△abelA::ChiA*(Figure S3). The results showed that the chitinase activity of DW2△abelA::ChiA* was 42.38 U/mL, indicating that the integrated expression of ChiA in Bacillus licheniformis was successfully realized (Fig. 5B). Then, the chitinase free expression plasmid was electrically transferred to the host bacterium DW2△abelA::ChiA* to obtain the strain DW2△abelA::ChiA*/pHY-P43 − UTR12-SPSacC(R4K)-ChiA(D116N/E118N) (DW2△abelA::ChiA*/ChiA* for short), the chitinase activity of DW2△abelA::ChiA*/ChiA* reached 180.26 U/mL (Fig. 5B).
Fermentation optimization of ChiA
To optimize the extracellular expression level of ChiA, we optimized the soybean meal media (corn starch 45 g/L, soybean meal 100 g/L, (NH4)2SO4 1 g/L, CaCO3 10 g/L) used in this study. The ratio of soybean meal to corn starch (100/45, 80/35, 70/30, 60/25, 50/22.5 g/L), the addition amount of (NH4)2SO4 (1, 2, 3, 4, 5 g/L) and the ratio of KH2PO4/Na2HPO4 (0, 1/2, 2/4, 3/6, 6/12, 12/24 g/L) were determined to improve the expression level of ChiA. The enzyme activity of strain DW2△abelA::ChiA*/ChiA* after fermentation showed that adding appropriate amount of (NH4)2SO4, optimizing the ratio of soybean meal to corn starch and adding appropriate ratio of KH2PO4/Na2HPO4 could improve the expression level of chitinase in Bacillus licheniformis (Figure S2). According to the results of orthogonal analysis, the influence priority of each factor was (NH4)2SO4 > KH2PO4/Na2HPO4 > soybean meal/corn starch (Table S6). The optimal medium was determined to be 30 g/L corn starch, 70 g/L soybean meal, CaCO3 10 g/L, (NH4)2SO4 2 g/L, KH2PO4 1 g/L, and Na2HPO4 2 g/L. Under these conditions, the enzyme activity of ChiA reached 200.28 U/mL (Table S6).
Based on optimized medium, the process optimization of 5 L fermenter was carried out. The chitinase activity of recombinant strain DW2△abelA::ChiA*/ChiA* was not significantly different from that of shaker under the condition of fermentation without feeding, which may be due to the insufficient carbon source in the medium during fermentation. In order to prolong the time of bacterial growth and chitinase synthesis in 5 L fermenter, the carbon source (glucose) was supplemented with constant flow at 12 h after fermentation. In addition, the residual sugar content was measured every 4 h, the glucose addition rate was constantly adjusted, the glucose concentration was controlled at 5–10 g/L, and the dissolved oxygen in the fermenting tank was maintained at about 20% by stirring the associated dissolved oxygen, and the pH was controlled at 6.8-7.0. Under such conditions, the activity of chitinase was the highest, reaching 338.79 U/mL (Fig. 6).
Discussion
Protein engineering is a powerful approach to enhance the properties of target enzymes, and one of the most common methods is through semi-rational design to improve substrate specificity and increase the specific activity of the target enzyme (Chica et al. 2005; Sinha and Shukla 2019). In this study, we used a semi-rational design to identify mutation sites that could enhance substrate specificity, resulting in a ChiA mutant called ChiA (D116N/E118N) with 48% higher enzyme activity than wild-type ChiA (Fig. 1). Although the ChiA (D116N/E118N) obtained in this study showed improved temperature stability and pH stability compared with wild-type ChiA (Fig. 3), the ChiA (D116N/E118N) still showed low temperature tolerance. It has been reported that the most direct way to obtain chitinase with excellent thermal stability is to clone and express from thermophilic microorganisms with high industrial applicability (Asmani et al. 2020). In addition, the chitinase mutant S244C-I319C/T259P was obtained by introducing disulfide bond and threonine substitution into chitinase using a semi-rational design based on sequence and structure. The mutant showed high specific activity at higher temperature, its half-life increased 26.3 times, and the optimal reaction temperature changed from 45℃ to 52.5℃ (Xu et al. 2020a). In future studies, it will be useful to focus on analyzing the three-dimensional structure of chitinase and the conformational changes of proteins under different temperature conditions, so that the temperature tolerance of chitinase can be improved by molecular modification.
With the rapid development of synthetic biology and metabolic engineering, more and more expression elements and gene-modified hosts have been developed to efficiently express foreign proteins and metabolites. Mao et al. screened and optimized the functional region of chitinase signal peptide, which increased the net charge of the N-terminal of the signal peptide from + 2 to + 4, and reduced the hydrophobicity of the H region, resulting in a 33% increase in the enzyme activity of chitinase Dac. Subsequently, in order to further improve the enzyme activity of chitinase, RBS Calculator was used to predict and calculate the initial translation rate of mRNA, and RBS sequence suitable for Dac expression was selected. The enzyme activity of the engineered strain constructed using this RBS sequence was 2.5 times higher than that of the initial strain (Mao et al. 2021). This study aimed to improve the expression level of ChiA by screening promoters and signal peptides. Our study found that when P43 − UTR12 promoter was used in combination with signal peptide SacC, chitinase extracellular enzyme activity was the highest (Fig. 4D). Since increasing the charge of the N-terminal of signal peptide has a positive effect on the protein secretion (Jiang et al. 2019), we also increased the positive charge of the N-terminal region of signal peptide to further modify the signal peptide SacC, and obtained an engineering strain carrying the signal peptide mutant strain SPSacC(R4K), which significantly improved the enzyme activity compared with the control strain (Fig. 4F). Our experimental results also showed that reducing the hydrophobicity of signal peptide H region was not conducive to the expression level of chitinase (Fig. 4F), which would have guided significance for the modification of signal peptide region. Bacillus has a rich extracellular protease expression system (Contesini et al. 2018), which is the main negative effect of using Bacillus as an expression system to express heterologous proteins outside the cells. Zhang et al. knocked out six protease coding genes (nprB, bpr, mpr, epr, vpr, wprA) in Bacillus subtilis WS5, and this modification significantly increased the activity of branched starch-degrading enzyme, reaching 5951.8 U/mL, the highest activity reported so far (Zhang et al. 2018). In this study, we knocked out the key extracellular protease genes aprE, bprA and epr of B. licheniformis DW2 to obtain the expression strain DW2△abe, and then proceeded to delete the by-product synthesis gene clusters lchAC and bacABC in DW2△abe to obtain the recombinant strain DW2△abelA. The fermentation results showed that ChiA activity was gradually increased (Fig. 5), foam was significantly reduced and no bacitracin was produced. Therefore, the extracellular expression level of chitinase can be improved by custom-modifying the host of B. licheniformis. In addition, other strategies can be modified for the host of B. licheniformis in the future, such as deleting other extracellular protease genes, enhancing ChiA secretion pathways, and optimizing amino acid supply based on precursor amino acids.
In our study, based on soybean meal medium, we optimized the medium components for the efficient production of chitinase (Figure S2 and Table S6). It has also been reported that the addition of 1.5% colloidal chitin and 1.25% fructose to the production medium has a significant effect on the production of chitinase. Compared with the initial fermentation conditions of the basic production medium, the total yield of chitinase was increased by 14.3 times, and the final activity was 1210.67 IU (Shivalee et al. 2018). In the follow-up study, we can also try to add an appropriate amount of colloidal chitin in the medium to achieve the effect of increasing the expression level of ChiA. In this study, we also optimized the fermentation process of a 5 L fermenter, with chitinase activity up to 338.79 U/mL under optimized conditions (Fig. 6). Although chitinase production increased after expanding fermentation to a 5 L fermenting tank, the enzyme activity did not reach the expected level, and it is necessary to further explore the optimal scaling fermentation parameters for chitinase production, such as dissolved oxygen level, liquid volume, stirring speed, inoculation volume, aeration rate, and feeding strategy. In addition, the study of the interaction between different carbon and nitrogen sources is also helpful to reveal the nutritional requirements of engineered strains and the key factors affecting the chitinase fermentation process.
Data availability
No datasets were generated or analysed during the current study.
References
Asmani KL, Bouacem K, Ouelhadj A, Yahiaoui M, Bechami S, Mechri S, Jabeur F, Taleb-Ait Menguellet K, Jaouadi B (2020) Biochemical and molecular characterization of an acido-thermostable endo-chitinase from Bacillus altitudinis KA15 for industrial degradation of chitinous waste. Carbohydr Res 495:108089. https://doi.org/10.1016/j.carres.2020.108089
Azarhava H, Bajestani MI, Jafari A, Vakilchap F, Mousavi SM (2020) Production and physicochemical characterization of bacterial poly gamma-(glutamic acid) to investigate its performance on enhanced oil recovery. Int J Biol Macromol 147:1204–1212. https://doi.org/10.1016/j.ijbiomac.2019.10.090
Barboza-Corona JE, Reyes-Rios DM, Salcedo-Hernández R, Bideshi DK (2008) Molecular and biochemical characterization of an endochitinase (ChiA-HD73) from Bacillus thuringiensis subsp. kurstaki HD-73. Mol Biotechnol 39, 29–37. https://doi.org/10.1007/s12033-007-9025-4
Cardozo FA, Gonzalez JM, Feitosa VA, Pessoa A, Rivera ING (2017) Bioconversion of α-chitin into N-acetyl-glucosamine using chitinases produced by marine-derived Aeromonas caviae isolates. World J Microbiol Biotechnol 33:201. https://doi.org/10.1007/s11274-017-2373-8
Chen JK, Shen CR, Liu CL (2010) N-acetylglucosamine: production and applications. Mar Drugs 8:2493–2516. https://doi.org/10.3390/md8092493
Chica RA, Doucet N, Pelletier JN (2005) Semi-rational approaches to engineering enzyme activity: combining the benefits of directed evolution and rational design. Curr Opin Biotechnol 16:378–384. https://doi.org/10.1016/j.copbio.2005.06.004
Contesini FJ, Melo RR, Sato HH (2018) An overview of Bacillus proteases: from production to application. Crit Rev Biotechnol 38:321–334. https://doi.org/10.1080/07388551.2017.1354354
Davies G, Henrissat B (1995) Structures and mechanisms of glycosyl hydrolases. Struct (London England: 1993) 3:853–859. https://doi.org/10.1016/s0969-2126(01)00220-9
Guo J, Cheng G, Gou XY, Xing F, Li S, Han YC, Wang L, Song JM, Shu CC, Chen SW, Chen LL (2015) Comprehensive transcriptome and improved genome annotation of Bacillus licheniformis WX-02. FEBS Lett 589:2372–2381. https://doi.org/10.1016/j.febslet.2015.07.029
Guttmann DM, Ellar DJ (2000) Phenotypic and genotypic comparisons of 23 strains from the Bacillus cereus complex for a selection of known and putative B. thuringiensis virulence factors. FEMS Microbiol Lett 188:7–13. https://doi.org/10.1016/s0378-1097(00)00200-7
Harwood CR, Cranenburgh R (2008) Bacillus protein secretion: an unfolding story. Trends Microbiol 16:73–79. https://doi.org/10.1016/j.tim.2007.12.001
Henrissat B, Bairoch A (1993) New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 293(3):781–788. https://doi.org/10.1042/bj2930781
Huang Z, Mao X, Lv X, Sun G, Zhang H, Lu W, Liu Y, Li J, Du G, Liu L (2021) Engineering diacetylchitobiose deacetylase from Pyrococcus horikoshii towards an efficient glucosamine production. Bioresour Technol 334:125241. https://doi.org/10.1016/j.biortech.2021.125241
Hurtado-Guerrero R, van Aalten DM (2007) Structure of Saccharomyces cerevisiae chitinase 1 and screening-based discovery of potent inhibitors. Chem Biol 14:589–599. https://doi.org/10.1016/j.chembiol.2007.03.015
Jiang Z, Niu T, Lv X, Liu Y, Li J, Lu W, Du G, Chen J, Liu L (2019) Secretory expression fine-tuning and directed evolution of diacetylchitobiose deacetylase by Bacillus subtilis. Appl Environ Microbiol 85. https://doi.org/10.1128/aem.01076-19
Karthik N, Binod P, Pandey A (2015) Purification and characterisation of an acidic and antifungal chitinase produced by a Streptomyces Sp. Bioresour Technol 188:195–201. https://doi.org/10.1016/j.biortech.2015.03.006
Khan FI, Rahman S, Queen A, Ahamad S, Ali S, Kim J, Hassan MI (2017) Implications of molecular diversity of chitin and its derivatives. Appl Microbiol Biotechnol 101:3513–3536. https://doi.org/10.1007/s00253-017-8229-1
Lipski A, Hervé M, Lombard V, Nurizzo D, Mengin-Lecreulx D, Bourne Y, Vincent F (2015) Structural and biochemical characterization of the β-N-acetylglucosaminidase from Thermotoga maritima: toward rationalization of mechanistic knowledge in the GH73 family. Glycobiology 25:319–330. https://doi.org/10.1093/glycob/cwu113
Liu X, Zhang J, Zhu KY (2019) Chitin in Arthropods: biosynthesis, modification, and metabolism. Adv Exp Med Biol 1142:169–207. https://doi.org/10.1007/978-981-13-7318-3_9
Mao X, Huang Z, Sun G, Zhang H, Lu W, Liu Y, Lv X, Du G, Li J, Liu L (2021) High level production of diacetylchitobiose deacetylase by refactoring genetic elements and cellular metabolism. Bioresour Technol 341:125836. https://doi.org/10.1016/j.biortech.2021.125836
Nishitani Y, Horiuchi A, Aslam M, Kanai T, Atomi H, Miki K (2018) Crystal structures of an archaeal chitinase ChiD and its ligand complexes. Glycobiology 28:418–426. https://doi.org/10.1093/glycob/cwy024
Oyeleye A, Normi YM (2018) Chitinase: diversity, limitations, and trends in engineering for suitable applications. Biosci Rep 38. https://doi.org/10.1042/bsr20180323
Rao FV, Houston DR, Boot RG, Aerts JM, Hodkinson M, Adams DJ, Shiomi K, Omura S, van Aalten DM (2005) Specificity and affinity of natural product cyclopentapeptide inhibitors against A. Fumigatus, human, and bacterial chitinases. Chem Biol 12:65–76. https://doi.org/10.1016/j.chembiol.2004.10.013
Robertus JD, Monzingo AF (1999) The structure and action of chitinases. Exs 87:125–135. https://doi.org/10.1007/978-3-0348-8757-1_9
Schumann W (2007) Production of recombinant proteins in Bacillus subtilis. Adv Appl Microbiol 62:137–189. https://doi.org/10.1016/s0065-2164(07)62006-1
Shivalee A, Lingappa K, Mahesh D (2018) Influence of bioprocess variables on the production of extracellular chitinase under submerged fermentation by Streptomyces pratensis strain KLSL55. J Genetic Eng Biotechnol 16:421–426. https://doi.org/10.1016/j.jgeb.2017.12.006
Sinha R, Shukla P (2019) Current trends in protein engineering: updates and progress. Curr Protein Pept Sci 20:398–407. https://doi.org/10.2174/1389203720666181119120120
Sugimoto H, Nakamura K, Nishino Y, Idezawa Y, Fujinuma A, Suzuki K, Watanabe T (2015) Differences in the roles of the two surface-exposed tyrosine residues, Y240 and Y481, of Serratia marcescens chitinase B during processive degradation of crystalline chitin. J Gen Appl Microbiol 61:255–261. https://doi.org/10.2323/jgam.61.255
Synowiecki J, Al-Khateeb NA (2003) Production, properties, and some new applications of chitin and its derivatives. Crit Rev Food Sci Nutr 43:145–171. https://doi.org/10.1080/10408690390826473
Thamthiankul S, Suan-Ngay S, Tantimavanich S, Panbangred W (2001) Chitinase from Bacillus thuringiensis subsp. Pakistani. Appl Microbiol Biotechnol 56:395–401. https://doi.org/10.1007/s002530100630
Tsuji H, Nishimura S, Inui T, Kado Y, Ishikawa K, Nakamura T, Uegaki K (2010) Kinetic and crystallographic analyses of the catalytic domain of chitinase from Pyrococcus furiosus- the role of conserved residues in the active site. FEBS J 277:2683–2695. https://doi.org/10.1111/j.1742-464X.2010.07685.x
Vaaje-Kolstad G, Horn SJ, Sørlie M, Eijsink VG (2013) The chitinolytic machinery of Serratia marcescens–a model system for enzymatic degradation of recalcitrant polysaccharides. FEBS J 280:3028–3049. https://doi.org/10.1111/febs.12181
Veith B, Herzberg C, Steckel S, Feesche J, Maurer KH, Ehrenreich P, Bäumer S, Henne A, Liesegang H, Merkl R, Ehrenreich A, Gottschalk G (2004) The complete genome sequence of Bacillus licheniformis DSM13, an organism with great industrial potential. J Mol Microbiol Biotechnol 7:204–211. https://doi.org/10.1159/000079829
Wang S, Wang H, Zhang D, Li X, Zhu J, Zhan Y, Cai D, Wang Q, Ma X, Wang D, Chen S (2020) Multistep metabolic engineering of Bacillus licheniformis to improve pulcherriminic acid production. Appl Environ Microbiol 86. https://doi.org/10.1128/aem.03041-19
Xu P, Ni ZF, Zong MH, Ou XY, Yang JG, Lou WY (2020a) Improving the thermostability and activity of Paenibacillus pasadenensis chitinase through semi-rational design. Int J Biol Macromol 150:9–15. https://doi.org/10.1016/j.ijbiomac.2020.02.033
Xu T, Qi M, Liu H, Cao D, Xu C, Wang L, Qi B (2020b) Chitin degradation potential and whole-genome sequence of Streptomyces diastaticus strain CS1801. AMB Express 10:29. https://doi.org/10.1186/s13568-020-0963-6
Zhan Y, Shi J, Xiao Y, Zhou F, Wang H, Xu H, Li Z, Yang S, Cai D, Chen S (2022) Multilevel metabolic engineering of Bacillus licheniformis for de novo biosynthesis of 2-phenylethanol. Metab Eng 70:43–54. https://doi.org/10.1016/j.ymben.2022.01.007
Zhang K, Su L, Wu J (2018) Enhanced extracellular pullulanase production in Bacillus subtilis using protease-deficient strains and optimal feeding. Appl Microbiol Biotechnol 102:5089–5103. https://doi.org/10.1007/s00253-018-8965-x
Funding
This work was supported by the National Key Research and Development Program of China (2021YFC2100200).
Author information
Authors and Affiliations
Contributions
ZQ, BL and WL completed all the experiments, RZ and SC designed and optimized the experiments, MX, YL, DC and XM analyzed and processed the data, ZQ, BL, RZ and SC wrote the manuscript, and all the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qi, Z., Lei, B., Xiong, M. et al. High-level production of chitinase by multi-strategy combination optimization in Bacillus licheniformis. World J Microbiol Biotechnol 40, 181 (2024). https://doi.org/10.1007/s11274-024-03995-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-024-03995-z