Abstract
Microbial surfactants are natural amphiphilic compounds with high surface activities and emulsifying properties. Due to their structural diversity, low toxicity, biodegradability, and chemical stability in different conditions, these molecules are potential substitutes for chemical surfactants; their interest has grown significantly over the last decade. The current study focuses on the isolation, identification, and characterization of a lactic acid bacteria that produce two forms of biosurfactants. The OL5 strain was isolated from green olive fermentation and identified using MALDI/TOF and DNAr16S amplification. Emulsification activity and surface tension measurements were used to estimate biosurfactant production. The two biosurfactants derived from Lactiplantibacillus plantarum OL5 presented good emulsification powers in the presence of various oils. They were also shown to have the potential to reduce water surface tension from 69 mN/m to 34 mN/m and 37 mN/m within a critical micelle concentration (CMC) of 7 mg/ml and 1.8 mg/ml, respectively, for cell bound and extracellular biosurfactants. Thin layer chromatography (TLC) and FT-IR were used to analyze the composition of the two biosurfactants produced. the obtained data revealed that the two biomolecules consist of a mixture of carbohydrates, lipids and proteins. We demonstrated that they are two anionic biosurfactants with glycolipopeptide nature which are stable in extreme conditions of temperature, pH and salinity.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbial biosurfactants are a structurally diverse group of surface-active agents produced by a wide range of microorganisms from various environmental habitats, primarily bacteria, actinomycetes, yeast, and filamentous fungi, that either adhere to the cell surface or are produced extracellularly (Behzadnia et al. 2020; Wang et al. 2020). Biosurfactants bind to liquid/air or water/oil interfaces, changing surface properties and emulsifying activities (Mouafo et al. 2018; Velraeds et al. 1996). The size of biosurfactants, such as low-molecular-weight biosurfactants and high-molecular-weight biosurfactants, can be used to identify them (Morais et al. 2017). When compared to chemically synthesized counterparts, the diverse chemical structures of biosurfactants (consisting of lipids, proteins, polysaccharides and carboxyl, amino, and phosphate functional groups) provide many advantages, such as high biodegradability, low toxicity, effectiveness under extreme physical conditions(Behzadnia et al. 2020)greater ecological acceptability and the ability to be produced from renewable substrates(Makkar and Cameotra 2002)A perfect biosurfactant for use in the pharmaceutical, medical, cosmetics, and food industries must have these essential characteristics: biosurfactants should be obtained from organisms that are generally regarded as safe (GRAS) to avoid the occurrence of critical situations that may arise as a result of their pathogenic origin, as well the requirement of a minimum concentration of them to achieve specific functionality for the proposed applications(Sharma et al. 2021). Due to their remarkable potential and rules governing their safety for several uses, including the food sector, Lactobacillus (LAB) strains truly believe in the GRAS concept. Since they are not pathogenic, they have received the majority of scientific attention in order to produce biosurfactants. These surfactants generated from LAB are often multicomponent combinations of proteins, lipids, and carbohydrates (Sharma et al. 2021; Sharma and Saharan 2016). Lactobacillus biosurfactants have antimicrobial and antioxidant properties and are useful as bioemulsifiers, texture enhancers, and heavy metal quenchers. Because of their versatile functionality, biological surfactants have been proposed as an alternative to chemically synthesized surfactants (Saravanakumari and Mani 2010). Several LAB strains have been reported to produce biosurfactants as well as their functional properties. Lactobacillus fermentum, Lactobacillus paracasei,Lactiplantibacillus plantarum, Lactobacillus helveticus, Enterococcus faecium, Lactobacillus acidophilus, Lactobacillus pentosus, Lactococcus lactis, Streptococcus thermophilus, Pediococcus acidilactici, and lactobacillus sp (Anon. n.d. 2022; Ferreira et al. 2017; Morais et al. 2017; Saravanakumari and Mani 2010; Sharma et al. 2015, 2021; Sharma and Saharan 2016) Lactobacillus plantarum recently renamed as Lactiplantibacillus Plantarum (Zheng et al. 2020) is a kind of lactic acid bacteria that has ecological and metabolic adaptability, allowing it to live in a variety of ecological niches such as fermented foods, meats, plants, and the mammalian gastro-intestinal tract. It is a gram-positive heterofermentative microaerophilic bacterium that has a rod-like morphology and can be found singly or in short chains(Dadar et al. 2021). Because of their unique evolutionary history, strains of this species have a wide range of probiotic characteristics and have shown promise in the production of a wide range of primary and secondary metabolites, including lactic acid, hydrogen peroxide, bacteriocins, biosurfactants, enzymes, and other related compounds. However, efforts were made in the current study to assess the ability of a newly isolated bacterium, Lactiplantibacillus plantarum OL5, to produce two types of biosurfactants and characterize their functional properties.
Methods
Isolation of lactic acid bacteria
Lactic acid bacteria (LAB) strains were isolated from natural samples (water from the draining of fresh cheese from an industrial company, fresh camel or goat's milk, stomach and intestine of battles, vaginal swabs, and olive salting water). Stock solutions were prepared for each sample. In the case of a solid sample, 1g of each sample is suspended in 9 ml of physiological water (9‰ NaCl). If it is a liquid sample, 1mL of the sample is added to 9 ml of physiological water. From these preparations, successive dilutions (10–2–10–5) in physiological water were made. Then 100 μL of each dilution was cultured anaerobically on MRS agar plates and incubated at 37 °C for 24 h.
Identification and characterization of isolated strains
Phenotypic identification of lactic acid bacteria (LAB)
The standard identification procedures based on morphological, cultural, and biochemical features were followed in selecting all the probable isolates within the scope of the present study.
Cells were detected with Gram staining under a microscope (oil immersion, 100 ×). The Shape of the cells (cocci, bacilli, and coccobacilli) and arrangement of cells (scattered, bunches, and chain) along with the Gram reaction were observed. Further tests with catalase and oxidase have also been conducted, and the fermentation profile of the sugars of the isolated strains was carried out by the API 50 CH gallery. The selected isolates were maintained on MRS agar slants at 4 °C. Whereas stock cultures were kept in cryovials with their culture broth containing 30% sterile glycerol at − 20 °C until use.
Genotypic characterization of biosurfactant-producing LAB
Bacterial isolate which shown high biosurfactants production was selected with regard to be identified. PCR amplification of 16S rDNA gene was carried using Universal primers Forward.
Fd1 (5′AGT TTG ATC CTG GCT CAG 3′) and Reverse Rd1 (5′AAG GAG GTG ATC CAGCC3′) (Mnif et al. 2015, 2022).
The 25 μL of PCR mixture contained: 0.2 mM of deoxy nucleoside triphosphate, 1.32 μM concentration for each primer, 0.5U of DNA polymerase, 5 μL of 5 × buffer, and 1 μL (5 ng) of total DNA template. The PCR program was 94 °C for 3 min followed by 35 cycles consisting of denaturation at 94 °C for 45 s, annealing at 59 °C for 1 min and extension at 72 °C for 2 min. After amplification, the PCR product was purified (Promega Gel Extraction Kit, Biogène, Tunisia) and sequenced with ABI PRISMTM 3100 Genetic Analyzer (Applied Biosystems, USA). The obtained sequences were aligned and compared with sequences from the Gene Bank database of the National Center of Biotechnology Information (NCBI) (http:// www. ncbi. nlm. nih. gov) using the nucleotide–nucleotide blast (BLASTn) network service (Mnif et al. 2022). The phylogenetic tree was drawn by the software MEGA version 10 using the neighbor joining method.
Biosurfactant-producing LAB identification of strains by the MALDI TOF test
MALDI-TOF mass spectrometry is a technique for separating molecules transformed into ions according to their m/z ratio (where m = mass and z = charge). One of its major applications is its contribution to proteomics, aimed at identifying all the proteins contained in the cell. It allows the identification of bacteria by analyzing their total proteins (ribosomal proteins and membrane-associated proteins) based on the fact that each species has its own proteome. Once these proteomes are known and listed, they are entered and compared to a reference database. The MALDI-TOF MS analysis was performed using a Brucker apparatus (Model: Microflex LT/SH from Bruker Daltonics, Germany). The identification of the bacterial isolate was performed by the Bruker Biotyper software. This test was carried out on two colonies of OL5 sown in streaks, one on a sloping tube and the other on a Petri dish.
Screening of LAB biosurfactant producing strain
Hemolytic activity determination
The screening of biosurfactants producing strains was performed in the first place by the blood hemolysis technique. Firstly, pure colonies of the isolated strains were streaked on blood agar medium. The selection of the biosurfactant-producing strain was based on the presence of a lysis halo around the colony after 24 h incubation at 37 °C, since biosurfactants possess hemolytic activity. The strains selected as biosurfactant producers were purified by successive subcultures on solid MRS medium using the quadrant technique (Mnif et al. 2022).
Emulsifying activity determination
The emulsification index (E24) was measured using the method described for the first time by Panchal and Zajic (1978) as a qualitative test visualized by the formation of a creamy emulsion. In fact, an emulsion is defined as a heterogeneous system, composed of an immiscible liquid dispersed as microscopic droplet in another liquid continuous phase. Biosurfactants may stabilize (emulsifiers) or destabilize (demulsifies) the emulsion. To do so, equal volumes (2 mL) of aqueous sample and 2 ml of corn oil were added to the test tube and the mixture was thoroughly agitated in a vortex mixer for 2 min. Upon standing, a smooth emulsion was formed when an emulsifier was present. At that point, an emulsification index (E24%) was calculated by the succeeding equation (Varjani et al. 2007):
The residual activity was calculated using the above formula:
Surface tension (ST) measurement and critical micelle concentration (CMC) determination
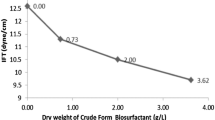
A ring-type tensiometer (Sigma 700) was used to determine the surface tension of crude glycolipopeptids biosurfactant at different concentrations, using the Du Nouy method at room temperature. Likewise, to determine the (CMC), ST was determined in function of increasing biosurfactants concentration prepared in distilled water. When an abrupt decrease of the ST was reached, the CMC was attained. It corresponds to the concentration at which biosurfactants associates into micelles. Distilled water (69 ± 0.10 mN/m) was used to verify the measurements before each reading, all measurements were taken in duplicates, and the mean value was presented in the results and discussion section.
Isolation and purification of the two biosurfactants
The chosen biosurfactant-producing LAB was inoculated in final volume of 150 ml MRS broth (Peptone (10 g), beef extract (10 g), yeast extract (5 g), dextrose (20 g), Sodium Acetate NaC2H3O2 (5 g), polysorbate 80 (1 g), KH2PO4 (2 g), ammonium citrate (2 g), magnesium sulfate heptahydrate MgSO4 (0.1 g), manganous sulfate tetrahydrate MnSO4 (0.05 g), per 1000 mL) for 48 h at 37 °C in a 250 ml Erlenmeyer flask, with an initial optical density of DO = 0.2, and fermentation was carried out in a static condition. The surface tension, the pH, and the biomass during the biosurfactant biosynthesis were determined. For the biomass determination, a volume of 1 ml of the culture was centrifuged for 10 min at 7000 rpm. The cell biomass was recovered, dried at 105 °C for 24 h, and reweighed. Dry bacterial biomass was repeatedly weighed until a constant weight was achieved. The fermentation broth was centrifuged at 10,000 rpm for 20 min at 4 °C using a centrifuge (HEHICH Zentrifugen Rotina 380 R, Germany) to remove the cell biomass. Secreted biosurfactants (EX BIOS) were precipitated from the supernatant with 6N HCl to pH 2. They were recovered after an overnight at 4 °C by centrifugation at 10,000 rpm for 20 min at 4 °C. Finally, they were suspended in 20 mM Tris–HCl buffer, pH 7. the pH of the suspension was then adjusted to 7 with 1 N sodium hydroxide before being lyophilized (Mnif et al. 2012). The removed cell biomass was washed once with deionized water and resuspended in 10 mL of PBS (0.1M of 7.0) in order to extract the cell-bound biosurfactants (CB BIOS). The cell suspensions were incubated at room temperature for 24 h with gentle agitation to release the biosurfactant. The cells were then removed by centrifugation. Subsequently, the cell-free supernatant was filtered using 0.22 μm pore size filter (Axiva, India), dialysed against demineralised water in a dialysis membrane (molecular weight cut-off 6000–8000 Dalton, Himedia, India) and freeze dried(Sharma et al. 2015).
Study of the two biosurfactants stability
The stability of the two OL5 biosurfactants (CB and EX BIOS) under different physicochemical conditions was evaluated using a solution of 10 mg m/L crude biosurfactant. The influence of pH on emulsification index (E24(%)) was estimated at different pH values ranging from 2.0 to 10.0 using glycine–HCl buffer (pH 2.0–3.0), acetate buffer (pH 4.0–5.0), phosphate buffer (pH6.0–8.0) and glycine–NaOH buffer (pH 9.0–10.0) (all at a final concentration of 20 mM (Mnif et al. 2022)). Results were expressed as Residual Activity (RA) towards the activity at neutral pH. For the determination of salt effect, different quantities of NaCl were dissolved in buffer solutions to adjust the salt concentration of test samples from 0 to 40%. Results were expressed as RA towards a negative control without salt addition. Furthermore, the heat stability of the crude biosurfactants was determined by incubating the biosurfactant solution (10 mg/ml) at different temperatures extending from 4 to 60 °C for 60 min, and then cooling at room temperature. The emulsifying activity (E24) of each sample was determined as described above. Likewise, the stability of the OL5 biosurfactants was evaluated after pre-incubation at different pH buffers, salt concentration and temperature by determining the emulsification activity (E24). Results were expressed by RA towards the activities at standard conditions without incubation. All assays were performed in triplicates.
Characterization of the two biosurfactants
Determination of ionic character T
The ionic charge of the two biosurfactants was determined using the agar diffusion method (Meylheuc et al. 2001). 20mM Cetyl Trimethyl Ammonium Bromide (CTAB) solution was used as the compound of cationic charge. The ionic charge was determined by creating two rows in 2% agar plated on Petri dishes. The wells in one row were filled with the purified biosurfactant, and the wells in the adjacent rows were filled with the cationic compounds. The biosurfactant's ionic nature was determined by monitoring the precipitation lines for 48 h at room temperature.
Compositional analysis of the two biosurfactants
The protein content was determined according to the Bradford (1976) method with Coomassie brilliant blue using bovine serum albumin as standard. The total sugar content was evaluated by the phenol–sulphuric method described by (Dubois et al. 1956) using glucose as standard and lipids were quantified from extraction with chloroform: methanol at different proportions (1:1 and 1:2, v/v)(Sharma et al. 2015). The organic extracts were then evaporated under vacuum conditions and the lipid content was estimated by gravimetric estimation.
Thin layer chromatography (TLC)
The composition of the two isolated biosurfactants (CB and EX BIOS) was determined by TLC. Briefly, after extraction of the biosurfactants 0.1 g of the freeze-dried samples was dissolved in several solvents of different polarizations, namely: methanol, chloroform and petroleum ether and analyzed by thin layer chromatography (TLC) on silica gel plates (G60, Merck, Germany). The latter were immersed in a migration solution consisting of (chloroform: methanol: acetic acid (65: 15: 2, v / v / v)). After migration, the plates were revealed by the following methods: (1) exposure to iodine vapor to detect lipid compounds, (2) spraying with Molisch's reagent to detect sugars, and (3) spraying with 1% ninhydrin solution to reveal amino acids. After spraying, the plates were heated for 20–30 min at 110 °C until the desired color appeared.
Fourier transform infrared spectroscopy (FTIR) analysis
The two biosurfactants functional groups are identified applying Fourier Transform Infrared Analysis (FTIR). A Fourier transform infrared spectrophotometer was used to record the infrared spectra (Instruments Analect fx-6 160). The purified biosurfactants (0.1 percent) were carefully mixed with KBr before being pressed into a homogenous disc (sample/KBr). The produced samples' FTIR spectra were obtained in the 400–4000 cm-1 range.
Protein molecular weight determination using sodium dodecyl sulphate polyacrylamine gel electrophoresis (SDS-PAGE)
Gel electrophoresis of both biosurfactants (CB and EX BIOS) (42µg) was performed using 17% (w/v) running gel and 4% stacking gel and run at a constant current (115 v) until the dye front attains the bottom. The gels were developed by staining with Coomassie blue R250. The molecular weight of protein of the two biosurfactants was determined by comparing it with the precision plus protein kaleidoscope standards (BIO-RAD, US).
Biosurfactants spreading capacity determination
The contact angle (CA) on various surfaces was measured by the sessile drop method using an Attention Theta 200 optical tensiometer (Biolin Scientific, Sweden) to determine the spreading ability or wettability of the two OL5 biosurfactants (CB and EX BIOS). A drop of 15 µL of sample was placed on five different surfaces: Teflon polymer (Polytetrafluoroethylene-PTFE) (highly hydrophobic), polystyrene, overhead projector transparency sheet (OHP, intermediate hydrophilic-hydrophobic), and glass slide (highly hydrophilic).Analysis of the shape of the biosurfactant droplet placed on each surface was carried out by image analysis with a high-resolution CCD camera and OneAttension software, using the Young–Laplace equation. Measurements were performed at least by triplicate and the mean value was presented in this work.
Results and discussion
Screening of biosurfactant producing LAB and biochemical characterization of the selected isolate
Many studies have been devoted to the development of biological surfactants or biosurfactants as an alternative to chemical ones, which have numerous disadvantages. biosurfactants interesting biological and surface properties have made them more advantageous than their synthetic counterparts, leading to their use in a variety of biotechnological (Banat et al. 2000; Pacwa-Płociniczak et al. 2011), pharmacological, cosmetic, agri-food, and environmental fields. To isolate lactic acid bacteria, MRS agar was used for preliminary screening of the strains. A primary screening of the selected strains (on MRS agar) was performed on blood agar based on the hemolytic properties of biosurfactants. Indeed, after 24 h at 37 °C incubation, the presence of a hemolytic zone (transparent or greenish) around the colonies is synonymous with haemolysis most likely produced by the biosurfactant molecule (Mulligan 2005). Thus, surface tension (ST), emulsifying activity (EA), and emulsification index (E24) tests were performed to confirm the production of biosurfactants by the selected strains. As a secondary screening for biosurfactant production, these tests were applied to the cell bound biosurfactant extract (CB BIOS). The obtained results are the average of two tests.
Only six strains were identified as potentially biosurfactant producing among the numerous isolates examined: LCV24 (isolated from goat milk), LCM8 (isolated from camel milk) and three strains OL3, OL5, and OL16 (isolated from olive curing water). According to the results, the analysis of the hemolytic activity, surface tension (ST), emulsifying activity, and emulsification index (E24) revealed that the OL5 strain is the most interesting to study, as a result, it was chosen for further investigations in this research work. (Table 1). The strain OL5 was identified first by morphological and microscopic analysis, and then by biochemical analysis. The colonies on MRS agar are round, bulging, and whitish. Their size ranges from 1 to 2 mm. The microscopic examination revealed that it is a gram-positive, immobile rod. The catalase test resulted in a negative result. These findings suggest that the strain is a Lactobacillus strain. The analysis of the biochemical test results of the Api galleries using the Api web software led us to the conclusion that they are most likely (99.1%) Lactiplantibacillus plantarum. These traditional microbiological methods are primarily used for bacterial identification and phenotypic typing. For some of them, however, gene amplification has completely replaced the traditional method. Among the other methods proposed, mass spectrometry enables the identification and typing of microorganisms (bacteria, viruses, fungi) from colonies or samples, either in conjunction with or without gene amplification (Courcol 2009).
Identification and characterization of isolated strains
Spectroscopic identification of the selected biosurfactant producing strain OL5
MALDI-TOF mass spectrometry has gradually replaced traditional techniques in medical microbiology laboratories due to its superior performance. According to previous research, mass spectrometry identification results (95%–97.4% correct identifications) outperform automated systems using conventional methods (75.2%–92.6%) (Courcol 2009). As a result, we used this method to identify the OL5 strain. The identification score of > 2 confirms that it is indeed Lactiplantibacillus plantarum according to the recent nomenclature. (Wardi and Prévost 2009) reported that a score greater than 2.00 indicates very good identification reliability.
Identification of the selected biosurfactant producing strain OL5 by the 16 rDNA Gene Sequencing
Using the MEGA7 program, a phylogenetic tree was constructed after sequencing the amplified DNA fragment. The 1500 bp 16S rRNA sequence amplified from isolate OL5 shared 97% identity with Lactobacillus plantarum b78, which has been deposited in GenBank under the accession number KX057646.1. According to phylogenetic analysis, strain OL5 is related to L. plantarum suebicus, L. pentosus strain 124, L. plantarum CIP, L. fabifermentans strain DSM, and Pediococcus clausseni ATCC (Fig. 1).
Growth and biosurfactant production of L. plantarum OL5
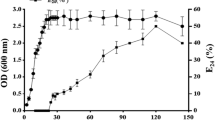
The surface tension of the medium broth was used to determine the biosurfactants produced by Lactiplantibacillus plantarum OL5. MRS medium was used in this study to produce the two biosurfactants of the OL5 strain; the pH was reduced over a 48-hour period from 6.5 to 3.94 (Fig. 2). The surface tension decreased the most during the exponential phase, from 52.36 mN/m to 38.54 mN/m (Fig. 2), and then increased marginally for the last 48 hours.
Surface tension measurement and determination of CMC
As suggested by (Satpute et al. 2010), the Du-Nouy-Ring method is based on measuring the force required to detach a ring or loop of wire from an interface or surface. The detachment force is proportional to the interfacial tension. It can be measured with an automated tensiometer, which is available from many manufacturers. The Du-Nouy-Ring assay is widely applied for screening of biosurfactant producing microbes (Anyanwu et al. 2011; Płaza et al. 2006).
Surfactants' main property is their ability to self-organize. This tendency is usually characterized by the critical micellar concentration (CMC). Below the CMC, the surfactant forms a layer on the liquid's surface, with the remainder dispersed in the solution. When the amount of surfactant is increased, the concentration increases proportionally until it reaches a limit value: the CMC. The CMC is thus the total surfactant concentration at which a constant and small number of surfactant molecules are aggregated. It is also the smallest concentration required to initiate micelle formation (Mata-Sandoval et al. 2000) the fluctuation of the surface tension of water in the presence of increasing concentrations of these surfactants (0–12 mg / ml) was determined in order to determine the CMC of the two biosurfactants of the strain OL5 (Fig. 3).
The results show that SDS reduces water surface tension from 69 mN/m to 26 ± 0.25 mN/m with a CMC of 1.86 mg/ml. The two OL5 biosurfactants reduce water surface tension from 69 mN/m to 34 ± 0.05 mN/m with a CMC of 7 mg/ml for the OL5 cell bound biosurfactant and to 37 ± 0.15 mN/m with a CMC of 1.8 mg/ml for the OL5 extracellular biosurfactant. In this regard, (Rodrigues et al. 2004) reported similar results with other lactic acid bacteria, specifically Streptococcus thermophilus A and Lactococcus lactis, whose biosurfactants reduce water surface tension from 70 mN/m to 36.0 mN/m and 37.0 mN/m, respectively. (Gudiña et al. 2015), on the other hand, demonstrated that the biosurfactant adhered to the cell membrane of Lactobacillus agilis CCUG31450 reduces water surface tension to 42.5 mN/m with a CMC in the range of 7.5 mg/mL. Similarly, (Morais et al. 2017) demonstrated that two biosurfactants produced by L. jensenii P6A and L. gasseri P65, respectively, reduce the TS of water to 43.2 mN/m and 42.5 mN/m, with CMC values of 7.1 mg/ml and 8.58 mg/ml.
Study of OL5 biosurfactants stability
The thermal stability of the two OL5 biosurfactants (CB and EX BIOS) emulsifying properties was investigated by determining their ability to emulsify corn oil after a 60-min pre-incubation at different temperatures: 4 °C, 20 °C, 40 °C and 60 °C. (Fig. 4) shows that the cell bound biosurfactant (CB BIOS) of the strain OL5 are practically stable after 1 h of incubation at the various temperatures tested. The extracellular biosurfactant (EX BIOS) appears to be less stable (87% to 100% activity retention). In this regard, (Morais et al. 2017)demonstrated that the biosurfactants produced by L. jensenii P6A and L. gasseri P65 retain their activities even after 60 min at 100 °C incubation. Likewise, the stability of two OL5 biosurfactants was investigated at pH levels ranging from 2 to 10. (Fig. 5) shows that the activity of two biosurfactants from strain OL5 remains nearly stable over a wide pH range. Indeed, varying the pH from 3 to 10 retains more than 80% of the emulsifying activity of the two biosurfactants. Moreover, Incubating the OL5 strain biosurfactants for 1 h in the presence of sodium chloride concentrations ranging from 10 to 40% results in a decrease in activity of approximately 18% to 35% in the height of the layer emulsified by OL5 cell bound biosurfactant(CB BIOS) and a decrease in activity of approximately 5% to 34% in the emulsifying activity of the OL5 extracellular biosurfactant (EX BIOS) (Fig. 6). Similarly, we noticed that the emulsifying activity of the commercial surfactant SDS decreases as NaCl concentration increases.
Determination of the CMC of the two biosurfactants of the strain OL5: extracellular (EX) and cell bound (CB) biosurfactants compared to the anionic surfactant (SDS). A Variation in surface tension and B semi-logarithmic reflection of critical micelle concentration (CMC) value from SFT (mN/m) versus biosurfactants concentration
Characterization of the two biosurfactants
Characterization of the two biosurfactants Determination of ionic character:
The ionic charge of the strain OL5's two biosurfactants (CB biosurfactant and EX biosurfactant) was determined using the double agar diffusion method, which is based on the diffusion of two soluble compounds with the same or different charge in a low concentration gel (2%).
(Fig. 7) depicts the formation of a halo around the positively charged chemical surfactant CTAB, whereas no line was formed between the strain OL5 biosurfactant and the anionic surfactant (SDS) (results not shown). This implies that both biosurfactants produced by the strain OL5 are anionic in nature (Ghazala et al. 2017; Siegmund and Wagner 1991). Similar findings have been reported in the literature, demonstrating the anionic charge of the majority of lactic acid bacteria biosurfactants, such as the xylolipid biosurfactant produced by Lactococcus lactis (Saravanakumari and Mani 2010), the glycolipid biosurfactant produced by Enterococcum faecium strain MRTL9 (Sharma et al. 2015), and the lactic acid bacteria LCM2 (Matei et al. 2017).
Compositional analysis of the two biosurfactants
Using traditional biochemical assay methods, the sugar, protein, and lipid levels of the two biosurfactants of Lactiplantibacillus plantarum OL5 strain were determined. The OL5 cell-bound biosurfactant (CB BIOS) is 78% sugars, 19.35% lipids, and 2% proteins, whereas the extracellular biosurfactant (EX BIOS) is 79% sugars, 17% lipids, and 4% proteins. These findings suggest that both biosurfactants are glycolipopeptides in nature.
Thin layer chromatography (TLC)
Thin layer chromatography (TLC) is a method of separating mixture components based on their affinity for the stationary and mobile phases. TLC was used to determine the nature of the biosurfactants produced by L. plantarum OL5 strain (Fig. 8). The presence of lipid, carbohydrate, and protein spots is indicated by the appearance of a dark yellow spot when the plate was exposed to iodine, a brown spot when exposed to sulphuric acid, and a purple spot when exposed to ninhydrin, respectively.
These findings suggest that two biosurfactants in strain OL5 (CB and EX BIOS) are heterogeneous. The two corresponding biosurfactants could be glycolipopeptides. These findings are similar to those obtained with glycolipopeptides biosurfactants produced by Lactobacillus acidophilus NCIM 2903(Satpute et al. 2019) and Lactobacillus plantarum ATCC 8014 (Behzadnia et al. 2020).
FTIR analyses
Infrared analysis of a sample is a quick qualitative method for detecting the presence of specific functional groups. Infrared spectroscopy was used to analyse the structures of two biosurfactants from strain OL5 in order to identify their functional groups. Figure 9 shows the obtained infrared spectra.
The infrared spectra of both biosurfactants of L. plantarum OL5 (CB and EX BIOS) show a strong absorption band at about 3386 cm−1 which could be assigned to the stretching vibrations of OH hydroxyl groups (of carboxyls, phenols or alcohols) (Sharma et al. 2014) indicating the presence of polysaccharides (Bakhshi et al. 2018), either carboxyls, phenols or alcohols (Kumar et al. 2012). A band at 2924 cm−1 was also detected for both biosurfactants which generally characterizes aliphatic C-H elongation vibrations (Sharma et al. 2014) indicating the presence of polysaccharides (Bakhshi et al. 2018), proteins or lipids (Kumar et al. 2012). The relatively strong absorption band detected for both types of biosurfactants between 1500 cm−1 and 1700 cm−1 was attributed to the asymmetric C = O stretching vibrations of the carboxyl groups (–COO–) that may be present in the composition of fatty acids and amino acids or to Groups N–H in proteins. The absorption bands detected at 1405 and 1041 cm−1 could be attributed to C–H bond deformation vibrations indicating the presence of an aliphatic chain (–CH3 and –CH2), indicating the presence of alkyl chains in the samples studied (Bakhshi et al. 2018; Sharma et al. 2014). Furthermore, both biosurfactants exhibit a distinct band between 1000 and 1200 cm−1. This region is dominated by ring vibrations superimposed on side group elongation vibrations (C–OH) and glucoside band vibrations (C–O–C) (Bakhshi et al. 2018).
Protein molecular weight determination using Sodium Dodecyl Sulphate Polyacrylamine Gel Electrophoresis (SDS-PAGE)
The biosurfactants SDS-PAGE revealed multiple bands with molecular weights ranging from 15 to 102 kDa, with prominent bands of 102.5 kDa for the EX BIOS and 54 kDa for the CB BIOS (Fig. 10). Similar results were reported by (Hashim et al. 2022)showing that following separation of Lactiplantibacillus plantarum biosurfactant on SDS-PAGE gel and staining with Coomassie blue, several protein bands were observed with molecular weight extending from 10 to 150 kDa. In a similar study, (Heinemann et al. 2000) used SDS-PAGE electrophoresis to determine the chemical structure of biosurfactants synthesized by Lactobacillus fermentum RC 14 after an 18-h culture on MRS medium. The electrophoresis demonstrated that the protein fraction of the analyzed surface-active compounds included fractions with molecular weights ranging from 14.0 to 94.0 kDa and higher. Likewise, (Gołek et al. 2009) showed that the protein fraction of biosurfactants synthesized by Lactobacillus casei, after 18 h of culture, on the MRS medium was constituted with molecular weights covered in 6.1–106.5 kDa.
Wetting property of the two biosurfactants
The wetting behaviour of solid surfaces can be altered by the adsorption of agents such as biosurfactants. The wetting properties of two biosurfactants (EX BIOS and CB BIOS) from L. plantarum OL5 were tested on four different surfaces: glass, polystyrene, Teflon, and transparency OHP film. In comparison to the CA values of water on the respective surfaces, the two biosurfactants showed the greatest decrease in CA on the Teflon surface from 113 to 81° ± 0.77for the cell bound biosurfactant (CB BIOS) and from 113 to 83.65° ± 4.35 for the extracellular biosurfactant (EX BIOS), followed by polystyrene from 90 to 59° ± 0.94 and 60.24° ± 2.045 respectively for the CB and EX BIOS. (Fig. 11). When the observed CA exceeds 90°, the surface is thought to have poor wetting properties and thus is hydrophobic. CA values for Lactobacillus sp. biosurfactants are rarely reported in the literature, although (Satpute et al. 2018) observed similar results. The biosurfactants produced by Lactobacillus acidophilus reduced the CA on polystyrene (from 90 to 49°) and Teflon (from 115 to 75°). Similarly, (Satpute et al. 2019) found that the highest reduction in CA was observed on polystyrene surface (from 96 to 36°) followed by Teflon (from 112 to 65°) for the biosurfactant produced by a Lactobacillus acidophilus strain. Based on these findings, it can be concluded that both biosurfactants have good wetting properties, which implies that they could be a viable industrial alternative to synthetic surfactants in a variety of sectors, including the cosmetic industry as alternatives to chemical surface-active (Ferreira et al. 2017).Another field of application for lactobacillus biosurfactants is in the medical field due to their good wetting ability as a coating substance on PDMS-based implants (Satpute et al. 2019). Future research will focus on implicating these OL5 biosurfactants in a variety of applications.
Conclusion
Due to their GRAS (Generally Recognized as Safe) status, interest in biosurfactants produced by lactic acid bacteria has increased significantly over the past decade. In the current study, we were interested in the isolation of a new biosurfactant-producing lactic acid bacterium (OL5) from olive curing water. Sequencing of 1500 bp of 16S rRNA of the OL5 strain confirmed the results of biochemical identification by API and mass spectroscopy and showed that it is Lactiplantibacillus plantarum. In addition, the isolated strain can produce two types of glycolipopeptides as biosurfactants. In conclusion, we showed that the activities of the two biosurfactants of the OL5 strain are stable under extreme temperature, pH and salinity conditions. In addition, both biosurfactants have good wetting ability, so they can be used in a wide range of industrial applications.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Velraeds MM, Mei van der HC, Reid G, Busscher HJ (1996) Inhibition of initial adhesion of uropathogenic enterococcus faecalis by biosurfactants from lactobacillus isolates. Appl Environ Microbiol https://doi.org/10.1128/aem.62.6.1958-1963.1996).
Anyanwu CU, Obi SKC, Okolo BN (2011) Lipopeptide biosurfactant production by serratia marcescens NSK-1 strain isolated from petroleum-contaminated soil. J Appl Sci Res 7:79–87
Bakhshi N, Sheikh-Zeinoddin M, Soleimanian-Zad S (2018) Production and partial characterization of a glycoprotein bioemulsifier produced by Lactobacillus Plantarum Subsp. Plantarum PTCC 1896. J Agric Sci Technol 20(1):37–49
Banat IM, Makkar RS, Cameotra SS (2000) Potential commercial applications of microbial surfactants. Appl Microbiol Biotechnol 53(5):495–508. https://doi.org/10.1007/S002530051648
Behzadnia A, Moosavi-Nasab M, Tiwari BK, Setoodeh P (2020) Lactobacillus Plantarum-Derived Biosurfactant: Ultrasound-Induced Production and Characterization. Ultrason Sonochem. https://doi.org/10.1016/j.ultsonch.2020.105037
Courcol R (2009) Quelles Utilisations de La Spectrométrie de Masse de Type MALDI-TOF En Microbiologie Médicale ? Revue Francophone Des Laboratoires 2009(416):61–64. https://doi.org/10.1016/S1773-035X(09)70251-5
Dadar M, Yi H, Fidanza M, Panigrahi P, Kollmann TR (2021) Lactiplantibacillus plantarum-nomad and ideal probiotic. Front Microbiol. https://doi.org/10.3389/fmicb.2021.712236
Dubois M, Gilles KA, Hamilton JK, Rebers PA, Smith F (1956) Colorimetric method for determination of sugars and related substances. Anal Chem 28(3):350–356. https://doi.org/10.1021/ac60111a017
Ferreira A, Vecino X, Ferreira D, Cruz JM, Moldes AB, Rodrigues LR (2017) Novel cosmetic formulations containing a biosurfactant from Lactobacillus Paracasei. Colloids Surf, B 155:522–529. https://doi.org/10.1016/j.colsurfb.2017.04.026
Ghazala I, Bouassida M, Krichen F, Benito JM, Ellouz-Chaabouni S, Haddar A (2017) Anionic lipopeptides from Bacillus Mojavensis I4 as effective antihypertensive agents: production, characterization, and identification. Eng Life Sci 17(12):1244–1253. https://doi.org/10.1002/ELSC.201700020
Gołek P, Bednarski W, Brzozowski B, Dziuba B (2009) The obtaining and properties of biosurfactants synthesized by bacteria of the genus Lactobacillus. Ann Microbiol 59(1):119–126. https://doi.org/10.1007/BF03175608/METRICS
Gudiña EJ, Fernandes EC, Teixeira JA, Rodrigues LR (2015) Antimicrobial and anti-adhesive activities of cell-bound biosurfactant from Lactobacillus Agilis CCUG31450. RSC Adv 5(110):90960–90968. https://doi.org/10.1039/C5RA11659G
Hashim ZA, Maillard JY, Wilson MJ, Waddington RJ (2022) Determining the potential use of biosurfactants in preventing endodontic infections. Eur J Oral Sci. https://doi.org/10.1111/EOS.12900
Heinemann C, van Hylckama JET, Vlieg DB, Janssen HJ, Busscher HC, van der Mei, and Gregor Reid. (2000) Purification and characterization of a surface-binding protein from Lactobacillus Fermentum RC-14 that inhibits adhesion of enterococcus Faecalis 1131. FEMS Microbiol Lett 190(1):177–180. https://doi.org/10.1111/J.1574-6968.2000.TB09282.X
Kumar CG, Mamidyala SK, Sujitha P, Muluka H, Akkenapally S (2012) Evaluation of critical nutritional parameters and their significance in the production of Rhamnolipid Biosurfactants from Pseudomonas Aeruginosa BS-161R. Biotechnol Prog 28(6):1507–1516. https://doi.org/10.1002/BTPR.1634
Makkar R, Cameotra S (2002) An update on the use of unconventional substrates for biosurfactant production and their new applications. Appl Microbiol Biotechnol 58(4):428–434. https://doi.org/10.1007/S00253-001-0924-1
Mata-Sandoval JC, Karns J, Torrents A (2000) Effect of Rhamnolipids produced by Pseudomonas Aeruginosa UG2 on the Solubilization of Pesticides. Environ Sci Technol 34(23):4923–4930
Matei, Gabi Mirela, Adrian Matei, Draghici Elena, Sorin Matei, and Elena Draghici. 2017. “Article in The EuroBiotech Journal ·.” The EuroBiotech Journal 212. doi: https://doi.org/10.24190/ISSN2564-615X/2017/03.02.
Meylheuc T, van Oss CJ, Bellon-Fontaine MN (2001) Adsorption of Biosurfactant on solid surfaces and consequences regarding the bioadhesion of Listeria Monocytogenes LO28. J Appl Microbiol 91(5):822–832. https://doi.org/10.1046/J.1365-2672.2001.01455.X
Mnif I, Chaabouni-Ellouze S, Ghribi D (2012) Optimization of the nutritional parameters for enhanced production of B. subtilis SPB1 biosurfactant in submerged culture using response surface methodology. Biotechnol Res Int. https://doi.org/10.1155/2012/795430
Mnif I, Mnif S, Sahnoun R, Maktouf S, Ayedi Y, Ellouze-Chaabouni S, Ghribi D (2015) Biodegradation of diesel oil by a novel microbial consortium: comparison between co-inoculation with biosurfactant-producing strain and exogenously added biosurfactants. Environ Sci Pollut Res 22(19):14852–14861. https://doi.org/10.1007/s11356-015-4488-5
Mnif I, Rajhi H, Bouallegue A, Trabelsi N, Ghribi D (2022) Characterization of lipopeptides biosurfactants produced by a newly isolated strain bacillus subtilis ZNI5: potential environmental application. J Polym Environ. https://doi.org/10.1007/s10924-021-02361-6
Morais IMC, Cordeiro AL, Teixeira GS, Domingues VS, Nardi RMD, Monteiro AS, Alves RJ, Siqueira EP, Santos VL (2017) Biological and physicochemical properties of biosurfactants produced by Lactobacillus Jensenii P6A and Lactobacillus Gasseri P65. Microb Cell Fact 16(1):1–15. https://doi.org/10.1186/S12934-017-0769-7/TABLES/4
Mouafo TH, Mbawala A, Ndjouenkeu R (2018) Effect of different carbon sources on biosurfactants’ production by three strains of Lactobacillus Spp. BioMed Res Int. https://doi.org/10.1155/2018/5034783
Mulligan CN (2005) Environmental applications for biosurfactants. Environ Pollut 133(2):183–198. https://doi.org/10.1016/J.ENVPOL.2004.06.009
Pacwa-Płociniczak M, Płaza GA, Piotrowska-Seget Z, Cameotra SS (2011) Environmental applications of biosurfactants: recent advances. Int J Mol Sci 12(1):633–654. https://doi.org/10.3390/IJMS12010633
Płaza GA, Zjawiony I, Banat IM (2006) Use of different methods for detection of thermophilic biosurfactant-producing bacteria from hydrocarbon-contaminated bioremediated soils. J Petrol Sci Eng 50(1):71–77. https://doi.org/10.1016/J.PETROL.2005.10.005
Rodrigues L, van der Mei HC, Teixeira J, Oliveira R (2004) Influence of biosurfactants from probiotic bacteria on formation of biofilms on voice prostheses. Appl Environ Microbiol 70(7):4408–4410
Saravanakumari P, Mani K (2010) Structural characterization of a Novel Xylolipid Biosurfactant from Lactococcus Lactis and analysis of antibacterial activity against multi-drug resistant pathogens. Biores Technol 101(22):8851–8854. https://doi.org/10.1016/J.BIORTECH.2010.06.104
Satpute SK, Banpurkar AG, Dhakephalkar PK, Banat IM, Chopade BA (2010) Methods for investigating biosurfactants and bioemulsifiers: a review. Critic Rev Biotechnol 30(2):127–144. https://doi.org/10.3109/07388550903427280
Satpute SK, Mone NS, Das P, Banpurkar AG, Banat IM (2018) Lactobacillus acidophilus derived biosurfactant as a biofilm inhibitor: a promising investigation using microfluidic approach. Appl Sci 8(9):1555. https://doi.org/10.3390/APP8091555
Satpute SK, Mone NS, Das P, Banat IM, Banpurkar AG (2019) Inhibition of pathogenic bacterial biofilms on PDMS based implants by L. Acidophilus derived biosurfactant. BMC Microbiol 19(1):1–15. https://doi.org/10.1186/S12866-019-1412-Z/FIGURES/12
Sharma D, Saharan BS (2016) Functional characterization of biomedical potential of biosurfactant produced by Lactobacillus helveticus. Biotechnol Reports 11:27–35. https://doi.org/10.1016/J.BTRE.2016.05.001
Sharma D, Saharan BS, Chauhan N, Bansal A, Procha S (2014) Production and structural characterization of lactobacillus helveticus derived biosurfactant. Sci World J. https://doi.org/10.1155/2014/493548
Sharma D, Saharan BS, Chauhan N, Procha S, Lal S (2015) Isolation and functional characterization of novel biosurfactant produced by enterococcus faecium. Springerplus 4(1):1–14. https://doi.org/10.1186/2193-1801-4-4
Sharma V, Singh D, Manzoor M, Banpurkar AG, Satpute SK, Sharma D (2021) Characterization and cytotoxicity assessment of biosurfactant derived from lactobacillus pentosus NCIM 2912. Braz J Microbiol 53:327
Siegmund I, Wagner F (1991) New method for detecting rhamnolipids excreted bypseudomonas species during growth on mineral agar. Biotechnol Tech 5(4):265–268. https://doi.org/10.1007/BF02438660
Varjani, S. J., D. P. Rana, S. Bateja, M. C. Sharma, and V. N. Upasani. 2007. “Screening and Identification of Biosurfactant (Bioemulsifier) Producing Bacteria from Crude Oil Contaminated Sites of Gujarat, India.” International Journal of Innovative Research in Science, Engineering and Technology (An ISO Certified Organization) 3297(2):9205–13.
Velraeds MMC, van der Mei HC, Reid G, Busscher HJ (1996) Inhibition of Initial adhesion of uropathogenic enterococcus faecalis by biosurfactants from Lactobacillus isolates. Appl Environ Microbiol 62(6):1958–1963. https://doi.org/10.1128/AEM.62.6.1958-1963.1996
Wang W, Ng DKS, Levin DB, Philippini RR, Silvério S, da Silva S, Martiniano E, Ingle AP, Marcelino PRF, Silva GM, Barbosa FG, César J, dos Santos. (2020) Agroindustrial byproducts for the generation of biobased products: alternatives for sustainable biorefineries. Front Ener Res. https://doi.org/10.3389/fenrg.2020.00152
Wardi, Moussaoui, and Gilles Prévost. 2009. Applications de la spectrographie de masse maldi-tof a l’identification bacterienne staphylococcus aureus leucotoxins and NETosis View Project Staphylococcus Aureus Leukotoxins: Mechanisms and Inhibition View Project.
Zheng J, Wittouck S, Salvetti E, Franz CMAP, Harris HMB, Mattarelli P, Paul W, O’toole, Bruno Pot, Peter Vandamme, Jens Walter, Koichi Watanabe, Sander Wuyts, Giovanna E. Felis, Michael G. Gänzle, and Sarah Lebeer. (2020) A taxonomic note on the genus lactobacillus: description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and Union of Lactobacillaceae and Leuconostocaceae. Int J Syst Evol Microbiol 70(4):2782–2858. https://doi.org/10.1099/IJSEM.0.004107/CITE/REFWORKS
Acknowledgements
This work was supported by the Ministry of High Education and Scientific Research- Tunisia.
Author information
Authors and Affiliations
Contributions
A.B and A.B: literature search and writing original final draft,and experimental assays M.B and N.F:strain screening experiments G.G and P.M: sds page experiments supervising and results approval J.B: contact angel experiments supervising and results approval D.G: Conceptualization, manuscript edditing and final approval All authors read and approved the final manuscript
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that no relevant conflicts of interest with respect to the content of this scientific paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ammar, A.B., Bouassida, M., Bouallegue, A. et al. Isolation and characterization of two glycolipopeptids biosurfactants produced by a Lactiplantibacillus plantarum OL5 strain isolated from green olive curing water. World J Microbiol Biotechnol 39, 308 (2023). https://doi.org/10.1007/s11274-023-03744-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03744-8