Abstract
A terrestrial green microalga was isolated at Ås, in Akershus County, Norway. The strain corresponded to a coccoid chlorophyte. Morphological characteristics by light and electron microscopy, in conjunction with DNA amplification and sequencing of the 18 s rDNA gene and ITS sequences, were used to identify the microalgae. The characteristics agree with those of the genus Coelastrella defined by Chodat, and formed a sister group with the recently described C. thermophila var. globulina. Coelastrella is a relatively small numbered genus that has not been observed in continental Norway before; there are no previous cultures available in collections of Norwegian strains. Gas chromatography analyses of the FAME-derivatives showed a high percentage of polyunsaturated fatty acids (44–45%) especially linolenic acid (C18:3n3; 30–34%). After the stationary phase, the cultures were able to accumulate several carotenoids as neoxanthin, pheophytin a, astaxanthin, canthaxanthin, lutein, and violaxanthin. Due to the scarcity of visual characters suitable for diagnostic purposes and the lack of DNA sequence information, there is a high possibility that species of this genus have been neglected in local environmental studies, even though it showed interesting properties for algal biotechnology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chlorophyta are an ancient and taxonomically diverse lineage of green algae with approximately 8000 described species and an estimated of 5000 still undescribed species (Hadi et al. 2016). For a long time, the classification of these organisms has been entirely based on morphological and cytological features of vegetative stages in their life cycle (Darienko et al. 2015). Unfortunately, the identification of coccoid green algae often presents a challenge for algal taxonomists due to the scarcity of visual characters suitable for diagnostic purposes and the lack of DNA sequence information (Škaloud et al. 2016).
Traditionally, the order Chlorococcales sensu lato grouped the coccoid taxa and represented one of the most diverse groups of photoautotrophic cryptogams. However, later studies have managed to transfer many taxa to other orders or classes i.e. Chlorophyceae, Trebouxiophyceae, and Prasinophyceae (Krienitz and Bock 2012). Moreover, phenotypic plasticity and the presence of cryptic species have contributed to taxonomy complication, resulting in constant reassignments in these microalgae (Eliáš et al. 2010; Malavasi et al. 2016).
Members of the subfamily Coelastroideae have been previously placed in the families Oocystaceae, Chlorellaceae, and Scotielloideae based on shape, reproduction, cell wall morphology and composition (Kalina and PunČochářová 1987), until phylogenetic molecular studies were applied. Nowadays, DNA amplification and sequencing of the 18S rDNA and internal transcribed spacer (ITS) studies placed Coelastroideae within the family Scenedesmaceae, order Sphaeropleales (Hanagata 1998, 2001; Hegewald and Hanagata 2000, 2002; Hegewald et al. 2010; Kaufnerová and Eliáš 2013; Lee et al. 2016; Ancona-Canché et al. 2017). According to AlgaeBase (Guiry and Guiry 2020), the subfamily Coelastroideae comprises seven genera: Coelastrella Chodat 1922 (16 species flagged as accepted taxonomically), Scotiellopsis Vinatzer 1975 (1 sp.), Asterarcys Comas Gonzales 1981 (1 sp.), Hariotina Dangeard 1889 (2 spp.), Dimorphococcus Braun 1855 (3 spp.), Coelastrum Nägeli 1849 (30 accepted species), and Graesiella Kalina and PunČochářová (1987) (1 sp.).
The genus Coelastrella has been known for almost 100 years and it is relatively small numbered. Coelastrella spp. are coccoid, elliptical until citriform. They occur as unicellular microalgae or in few-celled aggregations. These species are peculiar by their sculptured cell wall with 16–40 meridional ribs with or without polar thickenings (Uzunov et al. 2008; Kaufnerová and Eliáš 2013). Ultrastructurally, cells are uninucleate, present numerous conspicuous vacuoles, a single cup-shaped and parietal chloroplast, each with one pyrenoid surrounded by 2(3) starch plates. The cell wall is double layered, with an inner cellulose component and an outer trilaminar one where acetolysis-resistant material (sporopollenin) resides (Tschaikner et al. 2007a, 2008). Asexual reproduction occurs by 2–16 autospores released by rupture of parental cell wall (Guiry and Guiry 2020). Together with other morphological features like cell form, chloroplast and pyrenoid structures, the characteristic differences in the wall sculptures are useful for identification of the species (Gärtner and Ingolić 1993; Hanagata et al. 1996; Tschaikner et al. 2007a, b). The type species of the genus is Coelastrella striolata Chodat 1922.
Recently, diverse studies have reported the biotechnological interest of this genus due to pigments and fatty acid content, as well as for a potential use for bioremediation (Abe et al. 2007; Hu et al. 2013; Dimitrova et al. 2017; Kawasaki et al. 2013; Luo et al. 2016; Thao et al. 2017; Wang et al. 2019b). In the last decades, the biotechnological use of microalgae has raised the interest from the industry (Wijffels et al. 2010). Microalgae have been found to be the most promising feedstock in terms of their biomass productivity, high oil content, strong adaptive capacity to adverse environments, heavy metals, toxicants, high CO2 concentration and no competition with cultivable land (Chisti 2007). The screening of native algal species and strains is necessary for assessing the biotechnological potential of microalgae, especially in harsh environments; some of those species may be more beneficial than the commercially available strains.
In the present study, we report a terrestrial coccoid green microalga FGS-001, isolated at Ås, in Akershus County, Norway, which was identified as a Coelastrella strain, until now not described for continental Norway. Furthermore, very little is known about the distribution of this genus worldwide. The objectives of the study were (1) to increase knowledge on terrestrial Norwegian microalgae, (2) to characterize the isolated strain, (3) to determine its pigment and fatty acid composition profile, (4) to determine the phylogenetic relationships with known algal strains.
Material and methods
Microorganism isolation, medium and culture conditions
The strain FGS-001 was isolated from a foliose, land-living colony of Nostoc commune, known locally as «glye» in Norwegian language. A sample of the cyanobacterium colony was collected in summer 2016 outside the Center for Climate Regulated Plant Research (SKP), Norwegian University of Life Sciences, Campus Ås, Akershus County, South East Norway (N 59° 40′ 5.81292′′, E 10° 46′ 14.92156′′ (EU89)). The sample was washed with sterile Milli-Q water, exposed to light and enriched nutrient solution (a modified Kristalon Indigo medium) without aeration in a 50 mL conical flask.
Initially, only cyanobacteria belonging to Nostocales were visible. After 3–4 weeks of culture, green microalgae were observed under light microscope, growing inside fragments of the colony. Further isolation of the microalgae was achieved by consecutive transfers to fresh medium.
Kristalon Indigo standard nutrient solution according to producer (Yara, Norway) is composed NO3− 7.5%; NH4+ 1%; P 4.9%; K 24.7%; Mg 4.2%; S 5.7%; B 0.027%; Cu 0.004%; Fe 0.2%; Mn 0.06%; Mo 0.004%, and Zn 0.027%. A concentration was used of 0.01 g per liter of milli-Q water, enriched with 0.01 g urea and 0.002 g Opti-P 0-20-0 (Yara, Norway). The microalga was able to grow in a Tris–acetate–phosphate (TAP) (–) acetate media (see Harris 1989), and modified Provasoli nutrient media (West and McBride 1999), as well.
Finally, the isolated strain was able to grow in tubular photobioreactors (250 mL). It was cultivated under continuous illumination at a surface incident irradiance of 175 µmol m2/s at 20.0 ± 2 °C at constant aeration under filtered air flow containing 1 ± 0.2% CO2 (v/v), and at an initial pH of 7.0.
Dry matter was determined gravimetrically. Aliquots (2 mL) of samples were harvested from the photobioreactors in pre-weighed tubes by centrifugation. The supernatant was discarded, and pellets were dried at 105 °C until they reached a constant weight (Goecke et al. 2015). Growth curve was determined by linear regression of the natural log of cell biomass vs. time for the data plotted in Fig. 4.
A voucher specimen will be deposited in the Norwegian Culture Collection of Algae (NORCCA) (https://niva-cca.no/).
Optical microscopy
To obtain a detailed morphological characterization of cultured microalgae, we investigated it by different microscopical techniques.
Light microscopy observations were performed using a Leica DM5000B microscope (Leica CTR5000, Leica Microsystems Limited, Heerbrugg, Switzerland) equipped with the attached Leica camera (DC200), and microphotographs were processed with the Leica Application Suite v4.3 image program (LAS 4.3).
Additionally, to corroborate that the cells are uninucleated, samples were stained for 15 min with the nuclear marker DAPI (4′,6-diamidino-2-phenylindole; 10 µg/mL stock; Sigma-Aldrich, Saint Louis Missouri, USA), and analyzed using a UV filter under fluorescence microscopy.
Transmission electron microscopy (TEM)
Samples from two different culture ages (exponential and stationary phase) were fixed for transmission electron microscopy using chemical fixation protocols according to Olsen et al. (2015). Briefly, cells were harvested, centrifuged and prefixed in 1.25% glutaraldehyde and 2% formaldehyde in phosphate-buffered saline (PBS) solution for a minimum of 24 h. After washing several times in PBS and cacodylate buffer (0.1 M, pH 7.2), the cells were post fixed for 1 h at room temperature in 1% osmium tetroxide (OsO4), and washed again in cacodylate buffer. After dehydration through an ethanol series (15 min in 70, 90, 96%, and 4 × 15 min in 100%), cells were infiltrated and embedded in in LR-White resin (Electron Microscopy Sciences, USA). Ultrathin sections were prepared using a Leica Ultramicrotome EM UC7, counterstained with 4% aqueous uranyl acetate and 1% potassium permanganate (KMnO4) for 5 min and then washed in distilled water before examined and photographed in the transmission electron microscope (FEI Morgagni 268), using a Veleta CCD camera.
Scanning electron microscopy (SEM)
Algae were treated using the same fixing method described above for TEM. After, coverslips coated with 1 mg mL−1 poly-l-lysine were placed to allow algae to settle for 20 min. The fixed algae culture for SEM examination were washed thoroughly in 0.1 M sodium cacodylate buffer (SCB) and dehydrated with 10 min steps in ascending ethanol series (50–100%) as in Wiik-Nielsen et al. (2016). The samples were processed in a BAL-TEC Critical Point Dryer CPD 030, (BAL-TEC AG Lichtenstein), and a thin conductive coating of gold–palladium was applied to the samples using a Polar on Sputter Coater SC7640 (Quorum Technologies, UK). The coated samples were mounted on aluminum stubs, examined and photographed with a Zeiss EVO-50-EP scanning electron microscope at an accelerating voltage of 20 kV in the secondary emission mode.
DNA amplification and sequencing
For this study, the nuclear 18S rDNA gene and ITS regions of the alga were sequenced as described below.
Cells from 1 mL of algal culture were harvested by centrifugation, resuspended in 50 μL of EDTA 10 mM pH 8, and incubated at 100 °C for 10 min. Then, samples were cooled down at 4 °C for 10 min and resuspended by vortex. Finally, after 1 min of centrifugation at 10.000 rpm, 1 μL of supernatant was used to perform the PCR reactions.
The primer sets HET F (5′-ACCTGGTTGATCCTGCCAGTAGTCATAC-3′) and HET R (5′-GGTTCACCTACGGAAACCTTGTTACGACTTCA-3′) (Cavalier-Smith and Chao 2006), were used for the amplifications of the 18S rDNA regions.
The primer sets ITS1 (5′-TCCGTAGGTGAACCTGCGG-3′)/ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) (White et al. 1990), were used for the amplifications of the ITS regions.
All fragments were amplified with Q5 High-Fidelity DNA Polymerase (New England BioLabs).
Purified and concentrated PCR products were used as template for Sanger dideoxy sequencing at GATC Services (Eurofins, Germany), using the same primers sets employed in the amplification of the fragments, and 18F2 (5′-GCTCGTAGTTGGATTTCTGG-3′) (this study), in the case of 18S rDNA regions.
New sequences generated by this investigation were submitted to the nucleotide database GenBank from the National Center for Biotechnology Information (NCBI), with GenBank accession numbers MK064224 and MK040329.
Phylogenetic analysis
The identity of the isolates was assessed by a phylogenetic analysis using BEAST v1.10.4 (Suchard et al. 2018). Sequences for 18S rRNA and internal transcribed spacer (ITS) were taken from Wang et al. (2019b) and downloaded from NCBI’s GenBank (NCBI Resource Coordinators 2017) database using the respective accession numbers. Nucleotide sequences generated in this study were added to the data set, and a multiple sequence alignment was computed with MAFFT v7.310 using the L-INS-i strategy (Katoh et al. 2002; Katoh and Standley 2013). To remove uninformative sites and reduce the matrix to the loci of interest, the alignment was trimmed with trimAl (Capella-Gutiérrez et al. 2009) with the parameters -gt 0.8 -st 0.001 -cons 0.6. After manual inspection and adjustment of the alignment, the best nucleotide substitution model was determined based on AICc calculations by the modelTest function from the R package phangorn v2.5.3 (Schliep 2011; Darriba et al. 2012) using R v3.6.0 (R Core Team 2019). Thus, nucleotide substitutions in the 18S sequences were approximated with the GTR + Γ + I model (Hasegawa et al. 1985; Tavaré 1986; Yang 1994). For the ITS sequences, the GTR + Γ substitution model was used (Hasegawa et al. 1985; Tavaré 1986). Heterogeneity of the substitution rate was approximated by four discrete Γ categories in both cases. Phylogenetic trees were inferred assuming an uncorrelated relaxed molecular clock prior following a log-normal distribution (Drummond et al. 2006), assuming a Yule speciation process (Yule 1925; Gernhard 2008). Two separate Markov chain Monte Carlo (MCMC) analyses were run. Each run lasted 1.0 × 109 generations and trees were sampled every 5000 generations. The output from both MCMC chains was combined using LogCombiner v1.10.4 (Suchard et al. 2018), and 25% of the trees discarded as burn-in. The tree with maximum clade credibility was inferred with TreeAnnotator v1.10.4 (Suchard et al. 2018), rescaled to resemble mean node height, and visualized with FigTree v1.4.4 (Rambaut 2018). Chloromonas serbinowi UTEX 492 and Chlamydomonas reinhardtii UTEX 90 were used as out group with enforced monophyly for the Sphaeropleales–Chlamydomonadales split.
Analysis of pigment composition
Aliquots of 25–40 mg of freeze-dried algae material were weighed into lysis tubes (Type C, Analytik Jena, Jena, Germany) and 500 µL of ethanol (gradient grade, Merck, Darmstadt, Germany) were added. Cells were mechanically broken in a swing mill (MM 2000, Retsch, Haan, Germany) for 3 min. Afterwards the cells were centrifuged at 3000 rcf (5415R, Eppendorf, Hamburg, Germany) and the supernatant was recovered. This procedure was repeated twice until the supernatant was colorless. The combined extracts were dried under nitrogen at 40 °C in an evaporator (EVA-EC1 with metal block thermostat EC-1 V-130, both VLM, Bielefeld, Germany) and afterwards resuspended in a defined volume of ethanol and filtered through a 0.45 µm membrane filter (Chromafil Xtra PET -45/25, Macherey–Nagel, Düren, Germany). The samples were measured in an Ultra-performance liquid chromatography mass spectrometry (UPLC-MS), coupled to photodiode array detection (PDA), a UPLC-PDA-MS system (Waters, Milford, USA) with a Cortecs C18 collumn (2.7 µm, 90 Å, 3 × 100 mm, Waters, Milford, USA), with a gradient of Millipore water and acetonitrile (hypergrade for LC–MS, Merck, Darmstadt, Germany) both acidified with 0,01% formic acid (99% ULC/MS, Biosolve B.V., Valkenswaard, Netherlands). Starting conditions were 70% water, decreasing to 10% after 4 min. These conditions were kept stable further 10 min. Afterwards a washing step with 70% water was attached for 4 min. The flow velocity was 0.5 mL/min continuously. Column temperature was set at 40 °C and the spectra were measured via a PDA (2998 PDA Detector, Waters, Milford, USA) in a range of 200 to 800 nm. The mass spectrometer with electronspray ionization (ESI) (Acquity QDA, Waters, Milford, USA) was operated in positive mode with a cone voltage of 15 V and a probe temperature of 600 °C, measuring in a range of 150 to 1250 m/z.
As references standards Astaxanthin (≥ 97% from Blakeslea trispora, Sigma-Aldrich, St. Louis, USA) and Lutein (pharmaceutical secondary standard, Sigma-Aldrich, St. Louis, USA) were used. Further carotenoids were determined by comparison with literature.
For the photometric measurements (for full spectrum for chlorophyll detection), samples were prepared similarly in quantities of 5 to 10 mg, extracted in ethanol (≥ 99.5%, Ph.Eur., Carl Roth, Karlsruhe, Germany), but measured in different concentrations in a multiwellplate (96F, TPP Techno Plastic Products AG, Trasadingen, Swiss) in an microplate reader (Infinite M Plex, Tecan, Männedorf, Swiss) or alternatively in a photometer (DR 6000, Hach Lange, Düsseldorf, Germany) with a spectrum from 300 to 800 nm in steps of 2–5 nm. Due to biomass limitations, pigments were detected although not quantified.
Analysis of fatty acid composition
Conversion and extraction of algal lipids to fatty acid methyl esters (FAME) was done by the method of O’Fallon et al. (2007) with a minor modification concerning the volumes. Samples (0.3 g of freeze-dried sample) were placed into Pyrex culture tubes to which 0.3 mL of the C13:0 internal standard (0.5 mg of C13:0/mL of MeOH), 0.56 mL of 10 M KOH in water, and 4.24 mL of MeOH were added. The tubes were incubated in a 55 °C water bath for 1.5 h with vigorous hand-shaking for 5 s every 20 min to properly permeate, dissolve, and hydrolyze the sample. After cooling below room temperature in a cold tap water bath, 0.46 mL of 24 M H2SO4 in water was added. The tube was mixed by inversion and with precipitated K2SO4 present was incubated again in a 55 °C water bath for 1.5 h with hand-shaking for 5 s every 20 min. After cooling, 2.4 mL of heptane was added, and the tube was vortex-mixed for 5 min. The tube was centrifuged for 5 min in a tabletop centrifuge, and the hexane layer, containing the FAME, was placed into a gas chromatography (GC) vial. The vial was capped and placed at − 20 °C until analysis.
The fatty acid composition of the FAME was determined by capillary GC on a RT-2560, 100 m × 0.25 mm × 0.20 µm capillary column (Restek) installed on a Trace GC ULTRA gas chromatograph equipped with a Triplus autosampler with PTV inlet and a flame ionization detector, and controlled by Chromeleon 7.2 Chromatography Data System (Dionex, ThermoFisher Scientific). The initial oven temperature was 140 °C, held for 5 min, subsequently increased to 240 °C at a rate of 4 °C/min, and then held for 20 min. Helium was used as the carrier gas at a flow rate of 2 mL/min, and the column head pressure was 270 kPa. Both the injector and the detector were set at 250 °C. The injector was set in split mode and the split ratio was 20:1. Fatty acids were identified by comparing their retention times with the fatty acid methyl standards (Supelco standard FAME mixture) described previously.
Results
Light microscopy
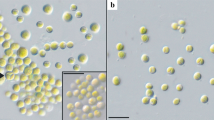
The isolated strain FGS-001 is a unicellular dark green microalga, although some cells were in aggregation. It grows in variable forms from globose or spheroidal to ellipsoidal (Fig. 1a). The cells presented variable sizes, but usually were (6–)7–10(–13) µm long (with an average of 8.42 ± 1.34 µm) and (4.5–)6–9(–11) µm wide (with an average of 7.14 ± 1.39 µm) (Fig. 1). Cell wall appears to be hyaline, and a cup-shaped chloroplast was easy to observe. A stricken and single pyrenoid was clearly visible in the vegetative cells as well as in autospores (Fig. 1a, b).
Unialgal culture of the strain FGS-001. a vegetative cells are green, unicellular and grows in variable forms from spheroidal to ellipsoidal. A single pyrenoid is visible; b cells with a smooth polar thickening were observed as well as others still surrounded by sporangium walls; c autospores in aggregation with a pyrenoid; d uninuclear cells were observed stained by DAPI under fluorescence. Scale bar: a–c = 20 µm; d = 10 µm
Meridional ribs were very difficult to observe under light microscopy. Although a smooth, apical thickening was visible in few cells (Fig. 1b).
Asexual reproduction takes place by autospores formed by successive bipartition of the protoplast to produce 4 to 16 spores within the mother cell (Fig. 1c). Autospores are elongated and polar thickenings were easier to notice, as well as their pyrenoids. They are discharged through rupture of the cell wall of the sporangium although some remain surrounded by sporangium walls (Fig. 1b, c). No sexual reproduction was observed.
Electron microscopy.
Transmission electron microphotographs of a group of vegetative and reproductive cells of Coelastrella sp. FGS-001 is shown in Fig. 2. Figure 2a is an example of the variable forms, from globose to ellipsoidal cells, which were found with variable sizes. The distribution of cell organelles is showed in detail by this technique. Vegetative cells presented a thin cell wall, where very smooth ribs were also visible at the cell wall surface (Fig. 2b). Internally, the single chloroplast proliferates throughout the cell with dense thylakoids; it presents a large cup-shaped form, which is associated with a single, prominent, spheroidal, and central pyrenoid structure with a ring of starch plates (n = 2) surrounding it, clearly visible (Fig. 2a–c). The single nucleus was located next to the pyrenoid (Fig. 2c). Occasionally, a second pyrenoid was observed by cell division (not shown).
a Transmission electron microphotographs of a group of vegetative and reproductive cells of Coelastrella sp. FGS-001 isolated from Ås, Akershus, Norway. Scale bar of 10 µm. b Young vegetative cell at exponential growth. A cup-shaped chloroplast (Ch) with dense thylakoids, the associated pyrenoid (P) structure with starch plates, and a thin cell wall, are clearly visible on the strain. Very smooth ribs are also present at the cell wall surface. c Autospore showing the single nucleus (N), vacuoles (V), thick starch plates (St), lipid droplets (L) and a thick and irregular cell wall (CW), with clear ribs of different sizes. d Autosporangia showing the formation of autospores, six cells are visible. e Cell wall detail with an outer trilaminar component. Scale bar: a = 10 µm; b–d = 2 µm; and e = 0.5 µm
The autospores presented a slightly smaller size than the vegetative cells, with a single nucleus, several vacuoles, lipid/starch droplets, thicker starch plates surrounding the pyrenoid, and a thick and irregular cell wall, with clear ribs of different sizes at the surface level (Fig. 2c). A variable number of autospores were visible at the autosporangia (Fig. 2d).
The cell wall of Coelastrella sp. FGS-001 consists of multiple layers, with an inner cellulose component and an outer apparently trilaminar one (Fig. 2e).
Scanning electron microscopy was performed at different times (4–18 days) of a living culture of the strain FGS-001. Different cell morphologies were observed ranging from globose, spheroidal until ellipsoidal. As shown in the micrographs (Fig. 3), the unicellular organism presents characteristic cell wall sculptures in form of meridional ribs, although they were more clearly visible at autospores and at younger states of the cells. The number of ribs may vary accordingly, although the difference was not statistically quantified (mainly around 10–16, with a possible maximum of 20) (Fig. 3). These ribs converged at two poles of the cells, and a smooth polar thickening was formed (Fig. 3b).
a Scanning electron microphotographs of a living culture after 4 days of growing. Different morphologies were observed ranging from spheroidal until ellipsoidal. Smooth ribs were visible at a surface level; b detail of autospores while being discharged through a rupture of the cell wall of the sporangium; c autospores or young vegetative cells of FGS-001, presenting a smooth pole thickening and several cell wall ribs at the surface. Scale bar for a, b and c = 3, 2 and 1 µm, respectively
Groups of autospores were discharged through a rupture of the cell wall of the sporangium which corresponded to the aggregations we observed under light microscopy (Fig. 3b, c).
Growth characteristics
Coelastrella sp. FGS-001 was cultured in a nutrient rich media at 175 µmol/m2/s and 20.0 ± 2 °C temperature for 10 days to observe the growth pattern in batch culture. The alga grew well in the temperature range of 15–20 °C (data not shown). Dry matter was measured (Fig. 4). The stationary phase was observed from day 8 onwards, probably due nutrient limitation in the media.
Phylogenetic associations
The nuclear 18S rDNA gene and ITS regions of the isolate FGS-001 was sequenced and compared to those of similar species in GenBank. As shown in Fig. 5, this new strain was grouped into one clade with other Coelastrella species and confirmed that the subfamily Coelastroideae are included in the monophyletic family Scenedesmaceae. We named the strain tentatively Coelastrella sp. FGS-001.
Maximum clade credibility tree of Scenedesmaceae inferred from 18S + ITS rDNA region. Numbers at nodes denote posterior probabilities (PP). Chlamydomonadales (Chla) was used as outgroup. Type species (holotype) of the genus Coelastrella is depicted with an asterisk. Original strains for species description are depicted with an empty circle. Strain FGS-001 presents a full circle. Most original strains for Coelastrella grouped together in the “core Coelastrella” as a sister group of the “Coelastrella sensu lato”, both placed into de Coelastroidea subfamily of Scenedesmaceae. Coelastroidea forms a sister group with Scenedesmoidea, except of colony-forming members of the subfamily Coelastroidea (Coe), which were included into the latter group
Coelastrella striolata (type species of the genus) and most of the available original culture strains seems to form a “core Coelastrella” group (Fig. 5). Our strain FGS-001 was placed within the “Coelastrella sensu lato” group, a sister group from the core Coelastrella group.
Pigment analysis
At the exponential phase, the main pigments of the strain FGS-001 were chlorophyll a and b, represented by a strong green color (Fig. 1b). After a few weeks at the stationary phase and without replacement of new nutrient media, the culture turned into yellowish-green until reddish-orange. This can be easily demonstrated on an agar plate at the exponential phase (Fig. 6a), and after a few weeks later of growth at the stationary phase (Fig. 6b). In the latter biomass, we were able to detect the presence of neoxanthin, pheophytin a, astaxanthin, canthaxanthin, lutein, an unknown carotenoid, and violaxanthin as the responsible pigments for this orange color (Fig. 6b, Table 2); although pigments were not quantified.
Fatty acid results
The fatty acid (FA) composition of freeze-dry biomass of the microalgal strain was analyzed using a gas chromatograph-mass spectrometer. We were able to identify 17 fatty acids after 8 and 18 days of cultivation, which were composed of saturated and unsaturated FAs with 10 to 20 carbon atoms. The FA composition is represented in Table 3.
The major FAs in Coelastrella sp. FGS-001 (see Fig. 7) were C18:3 n3 (which ranged from 30.79 to 33.45% for 8 and 18 days cultivation, respectively); C18:1 n9c (22.40% to 23.33%); C16:0 (17.84% to 18.27%), C18:2 n6c (12.31% to 7.3%), and C16:1 (10.86% to 11.54%). It was found that the total amount of FAs in the algae cells was 90.2 to 96.9 mg/g after first and second sampling, from which 27.8 ± 6.5 to 32.4 ± 4.8 mg/g corresponded to linolenic acid, the main fatty acid on the microalga (Table 3, Fig. 7).
The average percentages of total polyunsaturated fatty acids (PUFAs) were the highest (45.5% to 44.1%), in comparison with total saturated fatty acids (SFAs; 18.9% to 19.3%) and total monounsaturated fatty acids (MUFAs; 35.5% to 36.6%) (Fig. 7, after 8 days cultivation).
Slight differences in the FA percentage were observed among 8 and 18 days of cultivation (Table 3). A slight increase was observed on the FAs C16:0; C16:1; C17:0; C18:1 n9c; C18:3 n6; C20:1; C18:3 n3; C20:2, and C22:1 n9, although less than 3%. C18:2 n6c was reduced in 5% after 18 days, and it was the most notorious change in the FA profile after 10 days difference.
Discussion
In the present study, we described a terrestrial green microalga isolated at Ås, Norway. The strain corresponded to a coccoid chlorophyte. Morphological characteristics by light, fluorescence and electron microscopy (using TEM and SEM), in conjunction with sequences of 18S rDNA and ITS region, were used to identify the strain. The characteristics agree with those of the genus Coelastrella defined by Chodat (Table 1), with a sister relationship with Coelastrella thermophila var. globulina recently described by Wang et al. (2019b). A list of all the accepted species of Coelastrella and their morphological features is displayed at Supplement Table 1.
To our knowledge, Coelastrella has not been registered in continental Norway before; thus, there were no previous cultures available on collections of Norwegian strains. We agree that due to the scarcity of visual characters suitable for diagnostic purposes (see Table 1) there is a high possibility that many species of coccoid microalgae (including Coelastrella spp.) have been neglected in local environmental studies (in Norway and elsewhere). Only the last year, four new species and two new varieties have been described of Coelastrella (Kawasaki et al. 2019; Wang et al. 2019b). In Norway, there are few observations of other related green coccoid microalgae, including Dimorphococcus and Coelastrum (also members of the Coelastroideae), and Chlorococcum spp. (see Artsdatabanken 2018, https://artsdatabanken.no/; Norwegian Culture Collection of Algae, NORCCA 2018; The Culture Collection of Cryophilic Algae, CCCryo 2018; Bruteig et al. 2001). Moreover, Kol (1963) described the species Scotiella norvegica from red snow of Finse, Hordaland County (west Norway), although a phylogenetic position of this species is lacking. Many fusiform snow algae inhabiting polar areas and high alpine zones were initially believed to be a member of this genus, but further studies have pointed several of the Scotiella 'species' as zygospores of chlamydomonad-like snow algae (Procházková et al. 2018; Remias et al. 2018; Table 1).
Further north, in the Svalbard archipelago, Kim et al. (2008) investigated snow algae from northwestern Spitsbergen and reported a Scotiellopsis sp. At the same area, another group of researchers collected Scotiella sp. from supraglacial sediments (Stibal et al. 2006), and Scotiella norvegica and Scotiellopsis terrestris from wet soil and moss samples (Matuła et al. 2007). Nevertheless, over the last years, several species of Scotiella and Scotiellopsis were transferred to other genera including Coelastrella (Kaufnerová and Eliáš 2013). Recently, in Svalbard, Coelastrella aeroterrestrica, Coelastrella rubescens, Coelastrella cf. rubescens, and a Coelastrella sp., were identified by Borchhardt et al. (2017). However, apparently only strains of Chloromonas nivalis (ex Scotiella antarctica Fritsch, ex Scotiella nivalis (Chodat) Fritsch, ex Scotiella cryophila Chodat) are available from Svalbard in culture collections (see The Culture Collection of Cryophilic Algae, CCCryo 2018). This makes us conclude that there are no previous cultures available on collections of Norwegian strains.
Strain morphological identification
The strain FGS-001 grew solitary, especially under aeriation, and only small groups of cells were temporary aggregated, probably after autospores liberation. Without any aeration, the strain tended to form biofilm. Cells were also not embedded in mucilage. The family Scenedesmaceae contains numerous coenobial species of Desmodesmus, Neodesmus, and especially Scenedesmus, although some representatives of the latter genus are only known in solitary coccoid form. Other genera in this subfamily are either coenobial (e.g., Coelastrum, Hariotina) or solitary (e.g., Coelastrella, Scotiellopsis) (Fučíková et al. 2014). Thus, cell organization is a relevant taxonomic character for differentiation among Coelastroidea genera e.g., mainly solitary genera, coenobial genera, and mucilage producing genera (Table 1). Nevertheless, using light microscopy observations of living cells was almost impossible to assign any genus to the strain (Fig. 1). Furthermore, with fluorescence techniques was it possible to determine that cells are uninucleate (Fig. 1d), and that bigger structures (autosporangia) contained several cells inside (data not shown).
The apparent lack of cells with flagella, or any other type of zoospores (or potentially motile cells) in asexual reproduction, the presence of stigma, are definitive characters for differentiating the strain from Chlamydomonadales (Fig. 1, Table 1). As observed by other authors and for several Coelastrella species, the strain produced secondary carotenoids under stress or in older cultures (Fig. 6; PunČochářová and Kalina 1981; Abe et al. 2007; Hu et al. 2013; Kawasaki et al. 2019).
Scanning electron microscopy was determinant to discern the details of the cell wall surface. In this case, we could clearly see the presence of longitudinal ribs, which cover from pole to pole. They were especially defined at early stages of the cells (Fig. 3c), but they still could be observed in older and bigger cells (Fig. 3a). A similar cell wall surface has been registered by SEM for different Coelastrella species around the world, i.e. C. astaxanthina (Kawasaki et al. 2019), C. ellipsoidea and C. multistriata var. grandicosta (Gopalakrishnan et al. 2014); C. terrestris (PunČochářová and Kalina 1981; Gärtner and Ingolić 1993; Tschaikner et al. 2007a, b; Kaufnerová and Eliáš 2013; Xiao et al. 2017); C. aeroterrestrica (Uzunov et al. 2008); C. oocystiformis (PunČochářová and Kalina 1981), C. rubescens (PunČochářová and Kalina 1981; Kaufnerová and Eliáš 2013); C. saipanensis strain FACHB-2138 (Wang et al. 2019a), C. thermophila (Wang et al. 2019b), C. vacuolata (Hanagata 2001); C. multistriata var. multistriata (PunČochářová and Kalina 1981; Kalina and PunČochářová 1987; Hanagata et al. 1996); C. striolata (Kalina and PunČochářová 1987), C. yingshanensis (Wang et al. 2019b), Coelastrella sp. F50 (Hu et al. 2013); and Coelastrella sp. YC001 (Lee et al. 2016). The presence of longitudinal ribs is a clear character that helps to distinguish among genera; small thickenings at the poles, and a citriforme until globose morphology, are also considered relevant taxonomic characters (Table 1). For comparison with other close related species, i.e. other Scendesmaceae, see Kalina and PunČochářová (1987); Hanagata et al. (1996); Comas and Krienitz 1997; An et al. (1999); Hegewald et al. (2010); Eliáš et al. (2010); and Gopalakrishnan et al. (2014); or for Chlamydomonadales see Procházková et al. (2018) and Remias et al. (2018).
Using TEM, Gärtner and Ingolić (1993) and later Tschaikner et al. (2007b) showed a very similar liberation of daughter cells/autospores from an autosporangium from C. terrestris (strain N29 and strain SWK3:53 respectively, ex S. terrestris). Similar observations were made by PunČochářová and Kalina (1981) and Kaufnerová and Eliáš (2013) in C. rubescens CCALA 475, Uzunov et al. (2008) in C. aeroterrestrica, and Kawasaki et al. (2019) in C. astaxanthina and Coelastrella sp. SAG 2123. The number of daughter cells/autospores released by the sporangium is considered of taxonomic relevance as well, among these microalgae (Table 1). We observed that apparently the strain FGS-001 liberated 2 to 6 autospores (Figs. 2c and 3b).
Transmission electron microscopy was useful to observe the inner structure and distribution of organelles on the cells. The characteristic complex cell wall was also seen with this method (Fig. 2e), a taxonomic characteristic of the genus (Table 1). The different ribs types were also observed with TEM (Fig. 2d). The chloroplast form position and the clear pyrenoid complex were recorded too and appears to be as described for Coelastrella (Fig. 2b). For comparison with related or “similar” green coccoid microalgae investigated with TEM see Pickett-Heaps and Staehelin (1975), Gopalakrishnan et al. (2014) or Shebanova et al. (2017) for Desmodesmus and Scenedesmus spp., Chihara et al. (1994), Hu et al. (1998) and Feng et al. (2016) for Chlorococcum, Cardon et al. (2018) for Enallax costatus and Acutodesmus spp., Eliáš et al. (2010) for Hylodesmus, Matsuzaki et al. (2018), and Procházková et al. (2018), and Remias et al. (2018) for Chloromonas spp.
Pickett-Heaps and Staehelin (1975) showed the presence of other structures on the cell wall, like terminal spines, which is characteristic in many Scenedesmus species. Interestingly, Miller (1978) presented TEM microphotographs of zoospores of Chlorococcum oleofaciens (Utex 105), which is very useful for comparison with our strain. The authors showed the chlamydomonal-like structures, i.e. cell wall surface, stigma, flagella and a posterior nucleus, which are absent in our isolate.
Strain phylogeny
A faster and more accurate alternative for species identification and phylogenetic relationships among coccoid algal species is provided by DNA sequence comparisons (Škaloud et al. 2016). In general, our phylogenetic analysis of 18S rDNA and ITS region confirmed that the subfamily Coelastroideae are included in the monophyletic family Scenedesmaceae, although a more detailed analyses showed that the Coelastrella taxa belong to several different lineages within this family (Fig. 5). Our analyses did not clearly define a unique lineage for the Coelastrella taxa but add a new strain for such future elucidation. Similar conclusions were made by previous studies (see Hegewald et al. 2010; Kaufnerová and Eliáš 2013; Lee et al. 2016; Wang et al. 2019a).
Based on the 18S rDNA and ITS sequence phylogenetic tree phylogeny, we placed our strain into the order Sphaeropleales, as a sister order of the Chlamydomonadales, as observed previously (Hodač 2015; Watanabe and Lewis 2017; Fig. 5). Several morphological, ultrastructural and reproductive characteristics are shared (or appear to be) between these coccoid green microalgae (Table 1), therefore misidentification has been a problem for further phylogenetic studies. Several sequences available online carried misleading names and complicates the analysis of the group’s phylogeny (see Kaufnerová and Eliáš 2013; Lee et al. 2016). As mentioned, certain genera, as Scotiella, were found to be resistant stages (which presented a similar morphology to Coelastroideae: non-motile cysts with longitudinal ribs on cell surface) of other green algae as Chlamydomonadales i.e. Chloromonas spp. (Procházková et al. 2018; Remias et al. 2018). The genus are retained only for those species whose reproduction remain unknown (Hanagata 1998). Other genera have been separated from species (i.e. Scotiellopsis) by transferring them to other genera, based on new ultrastructural or molecular methods; remaining only few (old) species with unknown, too difficult to get, or none, biological material for further comparisons. Hence the importance of strain deposition on culture collections.
Similarly, as in Kaufnerová and Eliáš (2013), the type species of Coelastrella (C. striolata) and most of the available original (type culture) strains used for the description of several species of Coelastrella seems to form a “core Coelastrella” group (Fig. 5). Our strain FGS-001 grouped clearly into the subfamily Coelastroidea (family Scenedesmaceae) as well, specifically into the “Coelastrella sensu lato” group, a sister group from the core Coelastrella.
Further molecular phylogenetic investigations have raised concerns regarding the polyphyletic nature of some of those genera and further modifications are expected to happen as is the case of Chlorococcum (Kawasaki et al. 2015; Feng et al. 2016), Coelastrella (Kaufnerová and Eliáš 2013; Wang et al. 2019a), Coelastrum (Hegewald et al. 2010), and Ettlia (Pegg et al. 2015). Using 18 s rDNA sequences, Kaufnerová and Eliáš (2013), and later Wang et al. (2019a) demonstrated that the genus Coelastrella sensu lato is paraphyletic as species currently attributed to the genera Asterarcys, Scenedesmus or Ettlia are nested among taxa nominally representing Coelastrella. Clearly internal relationships among Coelastrella species are not solved yet and denser taxon sampling with more molecular markers is required to elucidate the classification of the strains (Lee et al. 2016). As mentioned by Wang et al. (2019b), it seems that 18S rDNA was too conserved to be used as a species-specific marker in this clade. We should clarify that a few original strains lacked available 18S sequences as Coelastrella compacta, C. levicostata, and C. coelastroides (see Supplement Table 1), which could also help to elucidate the phylogenetic relationships among these taxa. We contribute with a new strain isolated from northern Europe, and placed in a culture collection, which can help for further elucidation.
Recently, Wang et al. (2019a) studied the chloroplast genome sequence (cpDNA) of the Chinese strain Coelastrella saipanensis FACHB-2138. This is the first report on the cpDNA structure of the genus Coelastrella. Chloroplast genes often provide stronger phylogenetic signals, however, in the case of Scenedesmaceae, only very few chloroplast genomes are available.
Pigment analysis
Fresh cultures of the strain FGS-001 presented a strong green color due chlorophyll a and b, as main pigments, but after a few weeks under normal conditions, they turned into yellowish-green until reddish-orange after the stationary phase (Fig. 6, Table 2). HPLC–DAD for maximal extinction determination and UPLC-MS for mass detection, were used to determine those carotenoids. We detected the presence of neoxanthin, pheophytin a, astaxanthin, canthaxanthin, lutein, unknown carotenoid, and violaxanthin. A similar profile has been observed in several strains of Coelastrella as discussed below.
Abe et al. (2007) studied the pigment composition of a Japanese strain of Coelastrella striolata var. multistriata, before and after the stationary phase. The authors demonstrated the shift in pigment concentrations from chlorophyll (Chl a, b) dominance, until a carotenoid dominance (β-carotene, canthaxanthin and astaxanthin) after the stationary phase.
Hu et al. (2013) isolated a Chinese thermotolerant strain, Coelastrella sp. M-60 (Hu 2012), which under environmental stress (i.e., salt stress) produced astaxanthin (1.8% dw), β-carotene (1.4%), lutein (0.7%), canthaxanthin (0.18%), and adonirubin. Interestingly, 18S rDNA phylogeny grouped this strain with FGS-001 close to each other (Fig. 5). Recently, Karpagam et al. (2018) studied more in detail the carotenoid biosynthesis pathway in this strain.
Minyuk et al. (2017) studied the stress-induced secondary carotenogenesis in Coelastrella rubescens CCALA 475 and observed the shifts in the production of pigments (neoxanthin, violaxanthin, anteraxanthin, lutein, zeaxanthin, canthaxanthin, astaxanthin, adonixanthin, α- and β-carotene) under different culture treatments (N, P, sodium acetate, and CO2).
On one hand, the pigment profile of microalgae have been considered as a relevant taxonomical character in microalgae (Serive et al. 2017), which supported in this case the affiliation of our strain into Coelastrella spp. (Fig. 5) On the other hand, carotenoids are bioactive compounds having characteristic antioxidant, antimicrobial, antiviral, anti-tumoral, anti-inflammatory and anti-allergy effects, which give rise to health benefits (Karpagam et al. 2018).
Habitats where species of Coelastrella grow (with dehydration, temperature stress, salt stress, and high-light exposure) favors the production of pigments and fatty acids. Coelastrella species usually show strong survivability under such extreme photooxidative stresses; and therefore, they are considered good candidate species for large-scale production of natural pigments and biofuels (Wang et al. 2019b).
Fatty acid analysis
Gas chromatography analyses of the FAME-derivatives of the strain FGS-001 showed 17 fatty acids ranging from C10 until C20. The main fatty acids contained 16 and 18 carbon atoms and were identified as linolenic acid, oleic acid and palmitic acid, and palmitoleic acid (Fig. 7, Table 3).
Like other Coealastrella strains, relative fatty acid saturation the percentage of the fatty acids by saturation was high for MUFA and PUFA (Fig. 8). Fatty acid profiles have been considered as another taxonomical character, although we agree with Luo et al. (2016), that the lipid content and fatty acid composition are greatly affected by culturing conditions, growth period, and environmental situation (cf. Hu et al. 2013 and Minyuk et al. 2017).
Percentage of the fatty acids by saturation (saturated (SFA), monounsaturated (MUFA) and polyunsaturated fatty acids as PUFA) present in Coelastrella sp. FGS-001 after 8 days of cultivation with an enriched media, and comparison with other ‘Coelastrella’ isolates published worldwide. Strains F50 was isolated in China (Hu et al. 2013), strain L3 was isolated in Vietnam (Thao et al. 2017), and strain Tn1 until str. P63 were isolated in India (Minhas et al. 2016). The different strains were originally grown in different nutrient media
Thao et al. (2017) studied a strain from Vietnam identified as Coelastrella sp. L3 and registered 17 fatty acids. The main fatty acids were palmitic acid, stearic acid, and oleic acid. In another species isolated from Bulgaria (Coelastrella sp. BGV, Dimitrova et al. (2017) identified as main fatty acids palmitic acid, oleic acid, linoleic acid and α-linolenic acid. Similarly, Abe et al. (2007) analyzed C. striolata var. multistriata isolated from Japan, and found palmitic acid, oleic acid, linolenic acid, and α-linolenic acid. Palmitic acid, α-linolenic acid and linolenic acid were the main fatty acids present in C. rubescens CCALA 475, a strain from Tyrol, Austria (Minyuk et al. 2017). Coelastrella sp. F50 isolated from China presented oleic acid, palmitic acid and linoleic acid (Hu et al. 2013); and the strain Coelastrella sp. QY01 from China presented palmitic acid, linoleic acid and linolenic acid as main components (Luo et al. 2016). Although the main fatty acids are shared among the mentioned Coelastrella strains, FGS-001 is the only one with linolenic acid as the main fatty acid, instead of palmitic acid (ranked third in our isolate).
We quantified a high percentage of polyunsaturated fatty acids (44–45%) especially linolenic acid (C18:3n3; 30–34%), therefore demonstrating interesting properties for algal biotechnology. As mentioned, the major fatty acids found in the strain were C16–C18, which are commonly found in feedstock suitable for biodiesel production (i.e. oleic acid), in terms of oxidative stability and cold flow properties (Feng et al. 2016).
The genus Coelastrella demonstrates interesting properties for algal biotechnology, but clearly internal relationships among species are not solved yet. We think that a thorough investigation of all those “small and neglected” microalgae groups is necessary, not only for their evolutionary, phylogenetic and ecological implications, but for biotechnology as well. In the case of the present strain FGS-001, it would be interesting to test how it behaves on a larger scale production.
Conclusion
We investigated a coccoid green microalgae strain isolated from a terrestrial environment at Ås, Norway, and used several microscopical and molecular techniques for its identification. Coelastroideae is a subfamily in Scenedesmaceae still on the way to be clarified its phylogenetic relationships. The characteristics agree with those of the genus Coelastrella defined by Chodat in 1922, and the strain formed a sister group with the recently described C. thermophila var. globulina. The fatty acid analyses of the algal biomass showed a high percentage of polyunsaturated fatty acids especially from linolenic acid (30–34%). While the pigment analysis showed the presence of carotenoids like neoxanthin, pheophytin a, astaxanthin, canthaxanthin, lutein, and violaxanthin; the major fatty acids found in the strain were C16–C18, which are commonly found in feedstock suitable for biodiesel production. Therefore, the strain demonstrates interesting properties for algal biotechnology. Coelastrella spp. grow in habitats of high dehydration, temperature stress, salt stress, and high-light exposure, which apparently favors the production of such pigments and fatty acids.
References
Abe K, Hattori H, Hirano M (2007) Accumulation and antioxidant activity of secondary carotenoids in the aerial microalga Coelastrella striolata var. multistriata. Food Chem 100:656–661. https://doi.org/10.1016/j.foodchem.2005.10.026
An SS, Friedl T, Hegewald E (1999) Phylogenetic relationships of Scenedesmus and Scenedesmus-like coccoid green algae as inferred from ITS-2 rDNA sequence comparisons. Plant Biol 1:418–428. https://doi.org/10.1111/j.1438-8677.1999.tb00724.x
Ancona-Canché K, López-Adrián S, Espinosa-Aguilar M, Garduño-Solórzano G, Toledano-Thompson T, Narváez-Zapata J, Valdez-Ojeda R (2017) Molecular phylogeny and morphologic data of strains of the genus Coelastrella (Chlorophyta, Scenedesmaceae) from a tropical region in North America (Yucatan Peninsula). Bot Sci 95(3):527–537. https://doi.org/10.17129/botsci.1201
Artsdatabanken (2018) Trondheim, Norway. https://artskart.artsdatabanken.no/. Accessed 27 Nov 2018
Borchhardt N, Baum C, Mikhailyuk T, Karsten U (2017) Biological soil crusts of Arctic Svalbard—water availability as potential controlling factor for microalgal biodiversity. Front Microbiol 8:1485. https://doi.org/10.3389/fmicb.2017.01485
Bruteig IE, Thomsen MG, Altin D (2001) Vekstrespons hos tre aerofyttiske algar på tilførsel av nitrogen. NINA Oppdragsmeld 680:1–19
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25(15):1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Cardon ZG, Peredo EL, Dohnalkova AC, Gershone HL, Bezanilla M (2018) A model suite of green algae within the Scenedesmaceae for investigating contrasting desiccation tolerance and morphology. J Cell Sci 131:jcs212233. https://doi.org/10.1242/jcs.212233
Cavalier-Smith T, Chao E-Y (2006) Phylogeny and megasystematics of phagotrophic heterokonts (Kingdom Chromista). J Mol Evol 62(4):388–420. https://doi.org/10.1007/s00239-004-0353-8
Chihara M, Nakayama T, Inouye I, Kodama M (1994) Chlorococcum littorale, a new marine green coccoid alga (Chlorococcales, Chlorophyceae). Arch Protistenkd 144:227–235. https://doi.org/10.1016/S0003-9365(11)80133-8
Chisti Y (2007) Biodiesel from microalgae. Biotechnol Adv 25:294–306. https://doi.org/10.1016/j.biotechadv.2007.02.001
Chodat R (1922) Matériaux pour l'histoire des algues de la Suisse. Bull Soc Bot Geneve Sér 2(13):66–114 (in French)
Comas AA, Krienitz L (1997) Comparative LM-and SEM-studies on Coelastrum (Chlorophyta, Chlorococcales) under culture conditions. Algol Stud 87:87–98. https://doi.org/10.1127/algol_stud/87/1997/87
Darienko T, Gustavs L, Eggert A, Wolf W, Pröschold T (2015) Evaluating the species boundaries of green microalgae (Coccomyxa, Trebouxiophyceae, Chlorophyta) using integrative taxonomy and DNA barcoding with further implications for the species identification in environmental samples. PLoS ONE 10:e0127838. https://doi.org/10.1371/journal.pone.0127838
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: More models, new heuristics and parallel computing. Nat Methods 9(8):772–772. https://doi.org/10.1038/nmeth.2109
Dimitrova P, Marinova G, Alexandrov S, Iliev I, Pilarski P (2017) Biochemical characteristics of a newly isolated strain Coelastrella sp. BGV cultivated at different temperatures and light intensities. Annuaire de l’Université de Sofia “St. Kliment Ohridski” Faculte de Biologie 102:139–114
Drummond AJ, Ho SYW, Phillips MJ, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLoS Biol 4(5):e88. https://doi.org/10.1371/journal.pbio.0040088
Eliáš M, Němcová Y, Škaloud P, Neustupa J, Kaufnerová V, Šejnohová L (2010) Hylodesmus singaporensis gen. et sp. nov., a new autosporic subaerial green alga (Scenedesmaceae, Chlorophyta) from Singapore. Int J Syst Evol Microbiol 60:1224–1235. https://doi.org/10.1099/ijs.0.012963-0
Feng J, Guo Y, Zhang X, Wang G, Lv J, Liu Q, Xie S (2016) Identification and characterization of a symbiotic alga from soil bryophyte for lipid profiles. Biol Open 5:1317–1323. https://doi.org/10.1242/bio.019992
Fučíková K, Lewis PO, Lewis LA (2014) Putting incertae sedis taxa in their place: a proposal for ten new families and three new genera in Sphaeropleales (Chlorophyceae, Chlorophyta). J Phycol 50:14–25. https://doi.org/10.1111/jpy.12118
Gärtner G, Ingolić E (1993) Zur Morphologie und taxonomie einiger bodenalgen (Unterfamilie Scotiellocystoideae, Chlorellaceae) aus der Algensammlung in Innsbruck (ASIB, Austria). Arch Protistenkd 143:101–112. https://doi.org/10.1016/S0003-9365(11)80279-4(in German)
Gernhard T (2008) The conditioned reconstructed process. J Theor Biol 253(4):769–778. https://doi.org/10.1016/j.jtbi.2008.04.005
Goecke F, Jerez CG, Zachleder V, Figueroa FL, Řezanka T, Bišová K, Vitová M (2015) Use of lanthanides to alleviate the effects of metal ion-deficiency in Desmodesmus quadricauda (Sphaeropleales, Chlorophyta). Front Microbiol 6:2. https://doi.org/10.3389/fmicb.2015.00002
Gopalakrishnan KK, Novis PM, Visnovsky G (2014) Alpine scenedesmaceae from New Zealand: new taxonomy. N Z J Bot 52(1):84–99. https://doi.org/10.1080/0028825X.2013.859628
Guiry MD, Guiry GM (2020) AlgaeBase. World-wide electronic publication, National University of Ireland, Galway. https://www.algaebase.org. Accessed 29 May 2020
Hadi SIIA, Santana H, Brunale PPM, Gomes TG, Oliveira MD, Matthiensen A, Oliveira MEC, Silva FCP, Brasil BSAF (2016) DNA Barcoding green microalgae isolated from Neotropical inland waters. PLoS ONE 11:e0149284. https://doi.org/10.1371/journal.pone.0149284
Hanagata N (1998) Phylogeny of the subfamily Scotiellocystoideae and related taxa inferred from 18S rRNA gene sequence data. J Phycol 34:1049–1054. https://doi.org/10.1046/j.1529-8817.1998.341049.x
Hanagata N (2001) New species of Coelastrella and Scenedesmus (Chlorophyceae, Chlorophyta). J Jpn Bot 76:129–136
Hanagata N, Karube I, Chihara M (1996) Bark-inhabiting green algae in Japan (1) Scenedesmus komarekii and Coelastrella multistriata var. multistriata (Scotiellocystoideae, Chlorellaceae, Chlorophyceae). J Jpn Bot 71:87–97
Harris EH (1989) The Chlamydomonas sourcebook. Academic Press Inc, San Diego
Hasegawa M, Kishino H, Yano T-A (1985) Dating of the human–ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22(2):160–174. https://doi.org/10.1007/BF02101694
Hegewald E, Hanagata N (2000) Phylogenetic studies on Scenedesmaceae (Chlorophyta). Algol Stud 100:29–49. https://doi.org/10.1127/algol_stud/100/2000/29
Hegewald E, Hanagata N (2002) Validation of the new combinations of Coelastrella and Neodesmus and the description of the new subfamily Desmodesmoideae of the Scenedesmaceae (Chlorophyta). Algol Stud 105:7–9. https://doi.org/10.1127/algol_stud/105/2002/7
Hegewald E, Wolf M, Keller A, Friedl T, Krienitz L (2010) ITS2 sequence-structure phylogeny in the Scenedesmaceae with special reference to Coelastrum (Chlorophyta, Chlorophyceae), including the new genera Comasiella and Pectinodesmus. Phycologia 49(4):325–335. https://doi.org/10.2216/09-61.1
Hodač L (2015) Green algae in soil: assessing their biodiversity and biogeography with molecular-phylogenetic methods based on cultures. PhD Thesis, Georg-August-Universität Göttingen, Germany, p 185
Hu C-W (2012) Isolation and characterization of a new thermotolerant pigment-producing microalga: salt stress enhances pigment and oil biosynthesis in Coelastrella sp. F50. Master Thesis, National Sun Yat-sen University, China, p 32
Hu C-W, Chuang L-T, Yu P-C, Chen C-NN (2013) Pigment production by a new thermotolerant microalga Coelastrella sp. F50. Food Chem 138:2071–2078. https://doi.org/10.1016/j.foodchem.2012.11.133
Hu Q, Kurano N, Kawachi M, Iwasaki I, Miyachi S (1998) Ultrahigh-cell-density culture of a marine green alga Chlorococcum littorale in a flat-plate photobioreactor. Appl Microbiol Biotechnol 49:655–662. https://doi.org/10.1007/s002530051228
Kalina T, PunČochářová M (1987) Taxonomy of the subfamily Scotiellocystoideae Fott 1976 (Chlorellaceae, Chlorophyceae). Algol Stud 45:473–521
Karpagam R, Jawaharraj K, Ashokkumar B, Sridhar J, Varalakshmi P (2018) Unraveling the lipid and pigment biosynthesis in Coelastrella sp. M-60: genomics-enabled transcript profiling. Algal Res 29:277–289. https://doi.org/10.1016/j.algal.2017.11.031
Katoh K, Misawa K, Kuma K-i, Miyata T (2002) MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30(14):3059–3066. https://doi.org/10.1093/nar/gkf436
Katoh K, Standley DM (2013) MAFFT Multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(4):772–780. https://doi.org/10.1093/molbev/mst010
Kaufnerová V, Eliáš M (2013) The demise of the genus Scotiellopsis Vinatzer (Chlorophyta). Nova Hedwig 97(3–4):415–428. https://doi.org/10.1127/0029-5035/2013/0116
Kawasaki S, Mizuguchi K, Sato M, Kono T, Shimizu H (2013) A novel astaxanthin-binding photooxidative stress-inducible aqueous carotenoprotein from a eukaryotic microalga isolated from asphalt in midsummer. Plant Cell Physiol 54(7):1027–1040. https://doi.org/10.1093/pcp/pct080
Kawasaki Y, Nakada T, Tomita M (2015) Taxonomic revision of oil-producing green algae, Chlorococcum oleofaciens (Volvocales, Chlorophyceae), and its relatives. J Phycol 51:1000–1016. https://doi.org/10.1111/jpy.12343
Kawasaki S, Yoshida R, Ohkoshi K, Toyoshima H (2019) Coelastrella astaxanthina sp. nov. (Sphaeropleales, Chlorophyceae), a novel microalga isolated from an asphalt surface in midsummer in Japan. Phycol Res 67:1–8. https://doi.org/10.1111/pre.12412
Kim GH, Klochkova TA, Kang SH (2008) Notes on freshwater and terrestrial algae from Ny-Ålesund, Svalbard (high Arctic Sea area). J Environ Biol 29(4):485–491
Kol E (1963) On the red snow of Finse (Norway). Ann Hist Nat Mus Nat Hung 55:155–160
Krienitz L, Bock C (2012) Present state of the systematics of planktonic coccoid green algae of inland waters. Hydrobiologia 698:295–326. https://doi.org/10.1007/s10750-012-1079-z
Lee H-G, Song HJ, Kim D-S, Cho CH, La H-J, Oh H-M, Yoon HS (2016) Unique mitochondrial genome structure of the green algal strain YC001 (Sphaeropleales, Chlorophyta), with morphological observations. Phycologia 55(1):72–78. https://doi.org/10.2216/15-71.1
Luo L, He H, Yang C, Wen S, Zeng G, Wu M, Zhou Z, Lou W (2016) Nutrient removal and lipid production by Coelastrella sp. in anaerobically and aerobically treated swine wastewater. Bioresour Technol 216:135–141. https://doi.org/10.1016/j.biortech.2016.05.059
Malavasi V, Škaloud P, Rindi F, Tempesta S, Paoletti M, Pasqualetti M (2016) DNA-based taxonomy in ecologically versatile microalgae: A re-evaluation of the species concept within the coccoid green algal genus Coccomyxa (Trebouxiophyceae, Chlorophyta). PLoS ONE 11(3):e0151137. https://doi.org/10.1371/journal.pone.0151137
Matsuzaki R, Nozaki H, Kawachi M (2018) Taxonomic revision of Chloromonas nivalis (Volvocales, Chlorophyta) strains, with the new description of two snow-inhabiting Chloromonas species. PLoS ONE 13(3):e0193603. https://doi.org/10.1371/journal.pone.0193603
Matuła J, Pietryka M, Richter D, Wojtuń B (2007) Cyanoprokaryota and algae of Arctic terrestrial ecosystems in the Hornsund area, Spitsbergen. Pol Polar Res 28(4):283–315
Miller DH (1978) Cell wall chemistry and ultrastructure of Chlorococcum oleofaciens (Chlorophyta). J Phycol 14(2):189–194. https://doi.org/10.1111/j.1529-8817.1978.tb02447.x
Minhas AK, Hodgson P, Barrow CJ, Sashidhar B, Adholeya A (2016) The isolation and identification of new microalgal strains producing oil and carotenoid simultaneously with biofuel potential. Bioresour Technol 211:556–565. https://doi.org/10.1016/j.biortech.2016.03.121
Minyuk GS, Chelebieva ES, Chubchikova IN, Dantsyuk NV, Drobetskaya IV, Sakhon EG, Chekanov KA, Solovchenko AE (2017) Stress-induced secondary carotenogenesis in Coelastrella rubescens (Scenedesmaceae, Chlorophyta), a producer of value-added keto-carotenoids. Algae 32(3):245–259. https://doi.org/10.4490/algae.2017.32.8.6
NCBI Resource Coordinators (2017) Database resources of the national center for biotechnology information. Nucleic Acids Res 45(D1):D12–D17. https://doi.org/10.1093/nar/gkv1290
Norwegian Culture Collection of Algae, NORCCA. https://niva-cca.no/. Accessed 27 Nov 2018
O’Fallon JV, Busboom JR, Nelson ML, Gaskins CT (2007) A direct method for fatty acid methyl ester synthesis: application to wet meat tissues, oils, and feedstuffs. J Anim Sci 85:1511–1521. https://doi.org/10.2527/jas.2006-491
Olsen AB, Hjortaas M, Tengs T, Hellberg H, Johansen R (2015) First description of a new disease in rainbow trout (Oncorhynchus mykiss (Walbaum)) similar to heart and skeletal muscle inflammation (HSMI) and detection of a gene sequence related to Piscine Orthoreovirus (PRV). PLoS ONE 10(7):e0131638. https://doi.org/10.1371/journal.pone.0131638
Pegg C, Wolf M, Alanagreh L, Portman R, Buchheim MA (2015) Morphological diversity masks phylogenetic similarity of Ettlia and Haematococcus (Chlorophyceae). Phycologia 54(4):385–397. https://doi.org/10.2216/15-015.1
Pickett-Heaps JD, Staehelin LA (1975) The ultrastructure of Scenedesmus (Chlorophyceae). II. Cell division and colony formation. J Phycol 11:186–202. https://doi.org/10.1111/j.1529-8817.1975.tb02766.x
Procházková L, Remias D, Řezanka T, Nedbalová L (2018) Chloromonas nivalis subsp. tatrae, subsp. nov. (Chlamydomonadales, Chlorophyta): re-examination of a snow alga from the High Tatra Mountains (Slovakia). Fottea 18(1):1–18. https://doi.org/10.5507/fot.2017.010
PunČochářová M, Kalina T (1981) Taxonomy of the genus Scotiellopsis Vinatzer (Chlorococcales, Chlorophyta). Algol Stud 27:119–147. https://doi.org/10.1127/algol_stud/27/1981/119
R Core Team (2019) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/. Accessed 25 Oct 2019
Rambaut A (2018) Figtree v1.4.4. https://github.com/rambaut/figtree
Remias D, Procházková L, Holzinger A, Nedbalová L (2018) Ecology, cytology and phylogeny of the snow alga Scotiella cryophila K-1 (Chlamydomonas, Chlorophyta) from the Austrian Alps. Phycologia 57(5):581–592. https://doi.org/10.2216/18-45.1
Schliep KP (2011) Phangorn: phylogenetic analysis in R. Bioinformatics 27(4):592–593. https://doi.org/10.1093/bioinformatics/btq706
Serive B, Nicolau E, Bérard J-B, Kaas R, Pasquet V, Picot L, Cadoret J-P (2017) Community analysis of pigment patterns from 37 microalgae strains reveals new carotenoids and porphyrins characteristic of distinct strains and taxonomic groups. PLoS ONE 12(2):e0171872. https://doi.org/10.1371/journal.pone.0171872
Škaloud P, Friedl T, Hallmann C, Beck A, Dal Grande F (2016) Taxonomic revision and species delimitation of coccoid green algae currently assigned to the genus Dictyochloropsis (Trebouxiophyceae, Chlorophyta). J Phycol 52:599–617. https://doi.org/10.1111/jpy.12422
Shebanova A, Ismagulova T, Solovchenko A, Baulina O, Lobakova E, Ivanova A, Moiseenko A, Shaitan K, Polshakov V, Nedbal L, Gorelova O (2017) Versatility of the green microalga cell vacuole function as revealed by analytical transmission electron microscopy. Protoplasma 254(3):1323–1340. https://doi.org/10.1007/s00709-016-1024-5
Stibal M, Šabacká M, Kaštovská K (2006) Microbial communities on glacier surfaces in Svalbard: impact of physical and chemical properties on abundance and structure of cyanobacteria and algae. Microb Ecol 52:644–654. https://doi.org/10.1007/s00248-006-9083-3
Suchard MA, Lemey P, Baele G, Ayres DL, Drummond AJ, Rambaut A (2018) Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol 4(1):vey016. https://doi.org/10.1093/ve/vey016
Tavaré S (1986) Some probabilistic and statistical problems in the analysis of DNA sequences. Lect Math Life Sci 17:57–86
Thao TY, Linh DTN, Si VC, Carter TW, Hill RT (2017) Isolation and selection of microalgal strains from natural water sources in Viet Nam with potential for edible oil production. Mar Drugs 15:194. https://doi.org/10.3390/md15070194
The Culture Collection of Cryophilic Algae. https://cccryo.fraunhofer.de/web/infos/welcome/. Accessed 27 Nov 2018
Tschaikner A, Ingolić E, Gärtner G (2007a) Observations in a new isolate of Coelastrella terrestris (Reisigl) Hegewald & Hanagata (Chlorophyta, Scenedesmaceae) from Alpine soil (Tyrol, Austria). Phyton (Horn, Austria) 46(2):237–245
Tschaikner A, Ingolić E, Stoyneva MP, Gärtner G (2007b) Autosporulation in the soil alga Coelastrella terrestris (Chlorophyta, Scenedesmaceae, Scenedesmoideae). Phytol Balc 13(1):29–34
Tschaikner A, Gärtner G, Kofler W (2008) Coelastrella aeroterrestrica sp. Nov. (Chlorophyta, Scenedesmoideae)—a new, obviously often overlooked aeroterrestrial species. Algol Stud 128(1):11–20. https://doi.org/10.1127/1864-1318/2008/0128-0011
Uzunov BA, Stoyneva MP, Gärtner G, Kofler W (2008) First record of Coelastrella species (Chlorophyta: Scenedesmaceae) in Bulgaria. Ber Nat-Med Verein, Innsbruck 95:27–34
Vinatzer G (1975) Neue bodenalgen aus den dolomiten. Plant Syst Evol 123:213–235. https://doi.org/10.1007/BF00989405(in German)
Wang Q, Song H, Liu X, Zhu H, Hu Z, Liu G (2019a) Deep genomic analysis of Coelastrella saipanensis (Scenedesmaceae, Chlorophyta): Comparative chloroplast genomics of Scenedesmaceae. Eur J Phycol 54(1):52–65. https://doi.org/10.1080/09670262.2018.1503334
Wang Q, Song H, Liu X, Liu B, Hu Z, Liu G (2019b) Morphology and molecular phylogeny of coccoid green algae Coelastrella sensu lato (Scenedesmaceae, Sphaeropeales), including the description of three new species and two new varieties. J Phycol. https://doi.org/10.1111/jpy.12915
Watanabe S, Lewis LA (2017) Phylogenetic interpretation of light and electron microscopic features of selected members of the phylogroup Moewusinia (Chlorophyceae), with new generic taxonomy. Phycologia 56(3):329–353. https://doi.org/10.2216/16-64.1
West JA, McBride DL (1999) Long term and diurnal carpospore discharge patterns in the Ceramiaceae, Rhodomelaceae and Delesseriaceae (Rhodophyta). Hydrobiologia 398–399:101–114. https://doi.org/10.1023/A:1017025815001
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR protocols: a guide to methods and applications. Academic press, Cambridge, pp 315–322
Wiik-Nielsen J, Mo TA, Kolstad H, Mohammad SN, Hytterød S, Powell MD (2016) Morphological diversity of Paramoeba perurans trophozoites and their interaction with Atlantic salmon, Salmo salar L., gills. J Fish Dis 39(9):1113–1123. https://doi.org/10.1111/jfd.12444
Wijffels RH, Barbosa MJ, Eppink MHM (2010) Microalgae for the production of bulk chemicals and biofuels. Biofuel Bioprod Biorefin 4(3):287–295. https://doi.org/10.1002/bbb.215
Wright ES (2016) Using DECIPHER v2.0 to analyze big biological sequence data in R. R J 8(1):352–359
Xiao Y, Xing Z-f, Zhou F, Wang M (2017) Comparison of the imaging effects of Cryo-scanning electron microscopy and conventional scanning electron microscopy on aquatic plants. J Chin Electron Microsc Soc 36(2):173–176 (in Chinese)
Yang Z (1994) Maximum likelihood phylogenetic estimation from DNA sequences with variable rates over sites: approximate methods. J Mol Evol 39(3):306–314. https://doi.org/10.1007/BF00160154
Yule GU (1925) A mathematical theory of evolution, based on the conclusions of Dr. J. C. Willis, F.R.S. Philos Trans R Soc B 213(402–410):21–87
Acknowledgements
The SEM and TEM imaging were performed at the Imaging Centre Campus Ås, Department of Plant Sciences, Norwegian University of Life Sciences. Special thanks to Hilde Kolstad and Lene Cecilie Hermansen. The fatty acid profile analysis was performed at LabTek, Department of Animal and Aquacultural Sciences (IHA), NMBU. We thank (in loving memory of) Vilém Zachleder and Milada Vítová for comments on the manuscript. Professor Antje Labes and Inga Koopmann for help with the pigment analysis (Flensburg University of Applied Sciences). Mallikarjuna Rao Kovi for helping with the accession numbers. J. Noda was supported by the Czech National Program of Sustainability I (LO1416). J. Noda thanks Roman Sobotka and the Laboratory of Photosynthesis of the Centre Algatech in Třeboň. H.R. Gislerød thanks for support from Department of Plant Sciences, Faculty of Biosciences, Norwegian University of Life Sciences for conducting this work.
Funding
Open Access funding provided by Norwegian University of Life Sciences. Department of Plant Sciences, Faculty of Biosciences, Norwegian University of Life Sciences, Czech National Program of Sustainability I (LO1416).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Informed consent
All authors agreed on the publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11274_2020_2897_MOESM1_ESM.pdf
Supplementary file1 (PDF 116 kb)—Supplement Table 1. Survey of some morphological features of Coelastrella species according to literature. (Inattachment)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Goecke, F., Noda, J., Paliocha, M. et al. Revision of Coelastrella (Scenedesmaceae, Chlorophyta) and first register of this green coccoid microalga for continental Norway. World J Microbiol Biotechnol 36, 149 (2020). https://doi.org/10.1007/s11274-020-02897-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02897-0