Abstract
As the most important member of antioxidant defense system, human Cu,Zn superoxide dismutase (hCu,Zn-SOD) protects cells against the free radicals produced by aerobic metabolism. hCu,Zn-SOD has been widely used in food, cosmetic and medicine industry due to its health benefits and therapeutic potentials. However, a more extensive application of hCu,Zn-SOD is limited by the challenge of expensive and low production of high-activity hCu,Zn-SOD in large scale. In this study, the codon-optimized hCu,Zn-SOD gene was synthesized, cloned into pET-28a( +) and transformed into Escherichia coli BL21(DE3). After induction with IPTG or lactose, hCu,Zn-SOD was highly expressed as soluble form in LB medium with 800 μM Cu2+ and 20 μM Zn2+ at 25 °C. The recombinant hCu,Zn-SOD was efficiently purified by nickel affinity chromatography. Through optimization of fed-batch fermentation conditions, 342 mg purified hCu,Zn-SOD was obtained from 1 L cultures fermented in a 3-L bioreactor. Furthermore, the recombinant hCu,Zn-SOD retained the enzymatic specific activity of 46,541 U/mg. This study has opened up an effective avenue for industrial production of hCu,Zn-SOD through microbial fermentation in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The antioxidant defense system protects living organisms against harmful effects of the oxidative by-products of aerobic metabolism. Superoxide dismutase (SOD) is the first line of defense against reactive oxygen species (ROS) such as the superoxide radical which is toxic to living cells as it oxidizes and degrades biological molecules (Johnson 2002). SOD can convert superoxide radicals to hydrogen peroxide which is further catalyzed to form H2O and O2 by catalase (CAT) and glutathione peroxidase (GSH-Px) (Kilic et al. 2014; Vats et al. 2015). When the organisms do not have sufficient SOD, ROS will accumulate massively, leading to DNA strand breakage, protein inactivation, and membrane lipid peroxidation (Kim et al. 2017). Thus, SOD has been used in clinical practice for many years. It has various medicinal properties like selective killing of cancer cells (Huang et al. 2000), anti-aging (Tolmasoff et al. 1980), anti-inflammatory (Kim et al. 2017), and protection against reperfusion damage of ischemic tissues (Fridovich 1983).
As a metalloenzyme, SOD is classified into four groups according to their metal cofactors, namely Cu,Zn-SOD, Mn-SOD, Fe-SOD and Ni-SOD (Perry et al. 2010). Cu,Zn-SOD is the leading member which plays a vital role in the antioxidant defense system. The natural content of Cu,Zn-SOD is too low to satisfy the needs of clinical applications by extracting Cu,Zn SOD from living organisms. Therefore, new strategies have been attempted to produce Cu,Zn-SOD with high yield and activity using efficient and practical production systems in recent years. So far, heterologous expression of Cu,Zn-SOD has been explored in many expression systems including Escherichia coli (Hartman et al. 1986; Hallewell et al. 1989; Marisch et al. 2013; Lin et al. 2018), Pichia pastoris (Wu et al. 2009; Yoo et al. 1999), cyanobacteria (Takeshima et al. 1994), plant cells (Park et al. 2002), insect cells (Fujii et al. 1995; Hayward et al. 2002) and mammary glands of transgenic animals (Lu et al. 2018). However, none of these systems have been able to produce recombinant Cu,Zn-SOD with satisfactory yield and activity at large industrial scale until now. Although E. coli is the most commonly used system for production of recombinant proteins, Cu,Zn-SOD expressed in this system often formed inactive inclusion bodies in previous reports, which greatly limited its yield and activity (Liu et al. 2005).
In this study, the chemically synthesized gene encoding hCu,Zn-SOD was cloned into pET-28a( +) and transformed into E. coli BL21(DE3). The recombinant hCu,Zn-SOD was expressed as soluble form with high yield and specific activity.
Materials and methods
Strains and cultivation conditions
E. coli Trans1-T1 and BL21(DE3) (TransGen Biotech, Beijing, China) were used for plasmid amplification and protein expression, respectively. E. coli strains were cultivated at 37 °C in LB medium with 100 μg/mL ampicillin or 50 μg/mL kanamycin.
Construction of the expression plasmid containing hCu,Zn-SOD gene
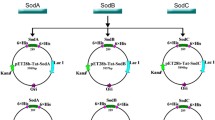
According to the reported hCu,Zn-SOD cDNA sequence (GenBank accession number: AY450286.1) and the codon usage preference of E. coli, the codon-optimized hCu,Zn-SOD gene with 6 × his-tag coding sequence at 5′ terminus (GenBank accession number: MN896904) was chemically synthesized and cloned into pEASY-Blunt Simple Cloning Vector by Shanghai Generay Biotech Co. Ltd. hCu,Zn-SOD gene was amplified from the cloning vector by PCR, and then cloned into the expression vector pET-28a( +), resulting in recombinant plasmid pET-hCu,Zn-SOD (Fig. 1). The recombinant plasmid was verified by DNA sequencing.
Expression and purification of the recombinant hCu,Zn-SOD
The recombinant expression plasmid pET-hCu,Zn-SOD was transformed into E. coli BL21(DE3) competent cells. The positive colony was cultured overnight in 10 mL LB medium supplemented with 50 μg/mL kanamycin at 37 °C and 220 rpm. 1 mL cultures were transferred to 100 mL fresh LB medium in 500-mL flasks. The cultures were incubated at 37 °C and 220 rpm until OD600 reached 0.6–0.8, and then induced with 1 mM isopropyl-β-D-1-thiogalactopyranoside (IPTG) at 15 °C, 25 °C and 37 °C, or 1 mM lactose at 25 °C. The effects of supplementation of Cu2+ /Zn2+ (200 μM/0 μM, 0 μM/5 μM, 200 μM/5 μM, 400 μM/10 μM, 800 μM/20 μM, 1200 μM/30 μM) in the induction period were also investigated. Cells were harvested by centrifugation (4 °C, 8000 rpm, 10 min). The cell pellet was resuspended in Tris–HCl (100 mM, pH 8.0) and lysed by sonication. The precipitate and supernatant were separated by centrifugation at 4 °C and 10,000 rpm for 20 min. The samples were analyzed by 15% (W/V) SDS-PAGE and Coomassie brilliant blue staining. The recombinant hCu,Zn-SOD was purified using nickel affinity chromatography column (GE Healthcare, Sweden) according to the manufacturer’s instructions. The protein concentration was detected by BCA Protein Assay Kit (Bio-Rad, USA) and the protein purity was determined by HPLC. All data are presented as means ± SD of three independent repeats.
The fed-batch fermentation process of the engineered strain
The fed-batch fermentation of the engineered strain was conducted in a 3-L bioreactor (Shanghai Baoxing Bio-Engineering Equipment Co. Ltd. China). The single colony was first grown in 100 mL LB medium supplemented with 50 μg/mL kanamycin at 37 °C and 220 rpm for 12 h as seed cultures. 100 mL seed cultures were then transferred into 1 L LB medium in a 3-L bioreactor. At the biomass accumulation stage, fermentation was carried out at 37 °C. The pH was automatically controlled at 7.0 by addition of 5 M ammonium hydroxide. When induction started, the culture temperature was kept at 37 °C or adjusted to 25 °C or 15 °C. This study focused on the effects of the following factors on the production of recombinant hCu,Zn-SOD: initial induction biomass (OD600 = 10, OD600 = 20, OD600 = 40); dissolved oxygen (DO) level (15%, 30%, 50%); constant lactose (4%,W/V) feeding rate (5 mL/h, 10 mL/h, 20 mL/h). All data are presented as means ± SD of three independent repeats.
Enzyme activity assay of the recombinant hCu,Zn-SOD
The recombinant hCu,Zn-SOD activity was detected using Total Superoxide Dismutase Assay Kit with nitroblue tetrazolium (NBT) (Beyotime Biotechnology, Shanghai, China) according to the instructions provided by the manufacturer. The detection was based on measuring the color of a formazan dye. According to the manufacturer’s instructions, the NBT/enzyme working solution and reaction starting solution were prepared beforehand. A preliminary experiment was conducted to find the optimal amount of hCu,Zn-SOD, so that the inhibition percentage of hCu,Zn-SOD lay between 30 and 70%. A certain amount of hCu,Zn-SOD (final volume was 20 μL), NBT/enzyme working solution (160 μL) and reaction starting solution (20 μL) were incubated at 37 °C for 30 min. The absorbance at 560 nm was detected. The activity of hCu,Zn-SOD was calculated according to the formula in the manufacturer’s instructions.
Results
Construction of recombinant hCu,Zn-SOD strain
The codon-optimized hCu,Zn-SOD gene with 6 × his-tag coding sequence at 5′ terminus, was chemically synthesized and cloned into E. coli expression vector pET-28a( +), resulting in recombinant plasmid pET-hCu,Zn-SOD. DNA sequencing confirmed that hCu,Zn-SOD was correctly fused with the N-terminal His-tag, and it also revealed the synthesized hCu,Zn-SOD gene was synonymous with the published one (GenBank accession number: AY450286.1). It was reported that the host strain BL21(DE3) produced the highest amount of SOD among three industrial E. coli strains (Marisch et al. 2013). Therefore, the recombinant expression plasmid pET-hCu,Zn-SOD was transformed into E. coli BL21(DE3) and the resulted engineered strain was named as BL21-hCu,Zn-SOD.
Expression and purification of the recombinant hCu,Zn-SOD at shake-flask level
Generally, the soluble expression level of the recombinant proteins is affected by the culture temperature (Francis and Page 2010). Reduced culture temperature is an increasingly popular strategy to enhance the soluble expression of the recombinant proteins. Therefore, in order to realize the soluble expression of hCu,Zn-SOD, the engineered strain BL21-hCu,Zn-SOD was induced by 1 mM IPTG at 37 °C, 25 °C and 15 °C, respectively. The recombinant hCu,Zn-SOD was expressed as soluble form at all three different temperatures, with different induction times to reach the peak level. It took 8 h to reach the peak of hCu,Zn-SOD production at 37 °C, 16 h at 25 °C and 24 h at 15 °C, respectively (Fig. 2). The highest hCu,Zn-SOD yield was achieved at 25 °C after 16 h induction (Fig. 3A and B).
Production of the recombinant hCu,Zn-SOD in E. coli at different temperatures. A SDS–PAGE analysis of hCu,Zn-SOD production at 37 °C. M: marker; 1: 0 h; 2: 4 h; 3: 8 h; 4: 16 h; 5: 24 h; 6: 32 h. B Comparison of the yields of hCu,Zn-SOD at different time points after induction at 37 °C. C SDS–PAGE analysis of hCu,Zn-SOD production at 25 °C. M: marker; 1: 0 h; 2: 4 h; 3: 8 h; 4: 16 h; 5: 24 h; 6: 32 h. D Comparison of the yields of hCu,Zn-SOD at different time points after induction at 25 °C. E SDS–PAGE analysis of hCu,Zn-SOD production at 15 °C. M: marker; 1: 0 h; 2: 8 h; 3: 16 h; 4: 24 h; 5: 32 h. F Comparison of the yields of hCu,Zn-SOD at different time points after induction at 15 °C
Results of shake-flask culture of the engineered E. coli strain producing hCu,Zn-SOD. A SDS–PAGE analysis of hCu,Zn-SOD production at different temperatures and time points after induction. M: marker; 1: 37 °C, 8 h; 2: 25 °C, 16 h; 3: 15 °C, 24 h. B Comparison of the yields of hCu,Zn-SOD at different temperatures and time points after induction. C SDS–PAGE analysis of hCu,Zn-SOD production with different inducers. M: marker; 1: no inducer; 2: lactose; 3: IPTG. D Comparison of the yields of hCu,Zn-SOD with different inducers. E Comparison of the specific activities of hCu,Zn-SOD expressed at different concentrations of Cu2+ and Zn2+. F SDS–PAGE analysis of purified hCu,Zn-SOD. M: marker; 1: purified hCu,Zn-SOD
IPTG is a highly effective inducer for lac promoter derived expression systems. IPTG is very stable, not metabolized by bacteria. Therefore, it is widely used for induced expression of recombinant proteins. However, IPTG is toxic to humans and relatively expensive, preventing its usage in large-scale production of genetically engineered drugs. Consequently, several substitutes of IPTG have been developed. To develop a safe and low-cost method for efficient production of hCu,Zn-SOD, the induction effects of lactose and IPTG on the expression of E. coli BL21- hCu,Zn-SOD were compared. The results showed that lactose achieved 87% of the induction effect of IPTG at the same concentration of 1 mM (Fig. 3C and D). Clearly, lactose can basically replace IPTG as an inducer in the induced expression of the recombinant E. coli BL21- hCu,Zn-SOD.
hCu,Zn-SOD is a metalloenzyme, requiring suitable levels of Cu2+ and Zn2+ to maintain its structural integrity and enzymatic activity. We investigated the effects of different concentrations of Cu2+ and Zn2+ in the induction medium on the specific activity of the recombinant hCu,Zn-SOD. As shown in Fig. 3E, the specific activity of hCu,Zn-SOD reached the maximum with 800 μM Cu2+ and 20 μM Zn2+, which was fivefold higher than that of control. Therefore, supplementation of Cu2+ and Zn2+ in the induction period improved hCu,Zn-SOD activity significantly.
The recombinant hCu,Zn-SOD induced under the optimized conditions was purified by nickel affinity chromatography. The purity of hCu,Zn-SOD reached about 97% by HPLC (Fig. 3F). 92 mg purified hCu,Zn-SOD was obtained from 1 L cultures at shake-flask level.
Production of the recombinant hCu,Zn-SOD through fed-batch fermentation
Compared with those of shake-flask culture mode, the dissolved oxygen (DO), pH and temperature of fed-batch fermentation mode in bioreactors are controllable, which can greatly improve both biomass production and heterologous protein expression to a great extent (Riesenberg and Guthke 1999; Lee 1996). In order to achieve higher yield of hCu,Zn-SOD, fed-batch fermentation of the engineered strain BL21-hCu,Zn-SOD was performed in a 3-L bioreactor.
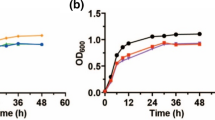
The effects of different initial induction cell densities on the expression of the recombinant hCu,Zn-SOD were investigated. As shown in Fig. 4A and B, the highest yield of hCu,Zn-SOD was achieved at the initial OD600 value of 20.
Results of fed-batch fermentation of the engineered E. coli strain producing hCu,Zn-SOD in a 3-L bioreactor. A SDS–PAGE analysis of hCu,Zn-SOD production at different initial induction biomasses. M: marker; 1: OD600 = 10; 2: OD600 = 20; 3: OD600 = 40. B Comparison of the yields of hCu,Zn-SOD at different initial induction biomasses. C SDS–PAGE analysis of hCu,Zn-SOD production at different dissolved oxygen (DO) levels. M: marker; 1: control; 2: 15%; 3: 30%; 4: 50%. D Comparison of the yields of hCu,Zn-SOD at different DO levels. E SDS–PAGE analysis of hCu,Zn-SOD production at different lactose feeding rates. M: marker; 1: 5 mL/h; 2: 10 mL/h; 3: 20 mL/h. F Comparison of the yields of hCu,Zn-SOD at different lactose feeding rates
During the fermentation process, the engineered bacteria consume a great deal of oxygen, leading to a sharp drop of the DO level. At this time, sufficient oxygen supply plays a decisive role in both biomass growth and protein expression. Raising the DO level can be achieved by increasing the airflow or enrichment of air with pure oxygen. However, in large-scale fermentation, supplying and handling pure oxygen can be highly expensive and dangerous (Liu et al. 2016). The DO level was adjusted by changing the airflow speed in this study. The effects of different DO levels on the expression of hCu,Zn-SOD were compared. The results indicated that the yield of hCu,Zn-SOD reached the maximum at DO of 30% (Fig. 4C and D).
As lactose acts as both inducer and carbon source, the best culture strategy is direct feeding of lactose after batch culture (Wurm et al. 2016; Li et al. 2006). The lactose feeding rate is crucial for protein expression. The effects of different lactose feeding rates on hCu,Zn-SOD expression were compared. The results demonstrated that the optimum lactose feeding rate was 10 mL/h (Fig. 4E and F).
342 mg hCu,Zn-SOD was purified from 1 L fed-batch cultures by nickel affinity chromatography. The yield of hCu,Zn-SOD was 3.7-fold higher than that of shake-flask level. The specific activity of the purified hCu,Zn-SOD was 46,541 U/mg.
Discussion
Recombinant protein expression in E. coli is simple, fast, inexpensive and robust, so E. coli is perhaps the most widely used bacteria for the production of high value-added proteins (Francis and Page 2010). In the past few decades, many literatures have reported heterologous expression of hCu,Zn-SOD in E. coli. As early as the mid-80s, the molecular cloning and nucleotide sequence of the cDNA for hCu,Zn-SOD were reported and the first successful expression of hCu,Zn-SOD in E. coli was realized at levels around 5% or more of total soluble cell proteins (Hallewell et al. 1985). Then, expression of hCu,Zn-SOD with activity comparable to the human erythrocyte enzyme was achieved in E. coli with the final yield of 40 mg/L (Hartman et al. 1986).
The recombinant proteins commonly form inclusion bodies when expressed in E. coli at 37 °C. Normally, low temperatures are used to increase the soluble expression level of the recombinant proteins, as it is easier for the protein chains to be folded properly with reduced protein production rate at lower temperatures (Swalley et al. 2006; Huo et al. 2010; Vasina and Baneyx 1997). The increase in protein production at temperatures lower than 37 °C was described previously either for hCu,Zn-SOD (Carrì et al. 1994) or for Cu,Zn-SOD from other organisms (Battistoni et al. 1992). In this study, the recombinant hCu,Zn-SOD was almost entirely expressed as soluble form at 37 °C, 25 °C and 15 °C. The highest hCu,Zn-SOD yield was achieved at 25 °C. However, it took twice to reach the peak of hCu,Zn-SOD yield at 25 °C as long as it did at 37 °C, perhaps due to the fact that cell processes slow down, leading to reduced rates of transcription, translation, and cell division at lower temperature. It was reported that lactose was a safer and cheaper inducer than IPTG in the induced expression of recombinant proteins in E. coli (Wurm et al. 2016). Lactose also had almost the same effect as IPTG in the induction of hCu,Zn-SOD expression in this study. It has been reported that both Cu2+ and Zn2+ are necessary to maintain the structural integrity and enzymatic activity of hCu,Zn-SOD (Nordlund et al. 2009; Li et al. 2010). Lacking Cu2+ and Zn2+ severely affects the correct folding of hCu,Zn-SOD, which can reduce activity and increase degradation (Wittung-Stafshede 2004; Rumfeldt et al. 2009). Our results indicated that the highest activity of hCu,Zn-SOD was achieved with 800 μM Cu2+ and 20 μM Zn2+, which was essentially consistent with the results of the previous reports (Lin et al. 2018).
The key to achieve high yields of recombinant proteins is fed-batch fermentation of the engineered strains. In the fed-batch fermentation process, the cell density for induction affects the cell growth and protein expression (Kamran et al. 2016). High cell density fermentation is a major bio-process engineering consideration for enhancing the yield of recombinant proteins in E. coli (Tripathi et al. 2009). The low initial induction cell density led to the reduced yield of recombinant hCu,Zn-SOD perhaps because of the less biomass produced by fermentation. However, the total yield of the protein is a function of both unit cell concentration and specific cellular protein yield. The optimal initial cell density for hCu,Zn-SOD induction was OD600 value of 20. When the initial induction cell density was raised to OD600 value of 40, the yield of hCu,Zn-SOD decreased, which probably arose from substrate inhibition, nutrient inhibition, limited oxygen transfer capacity, the formation of growth-inhibiting by-products, and limited heat dissipation.
Availability of oxygen is also a critical factor for cell growth and gene expression (Castan and Enfors 2000). Hence it is essential to keep supply of oxygen at optimal level to support cell growth and protein production. In this study, the highest yield of hCu,Zn-SOD reached at DO of 30%. When DO was increased from 30 to 50%, the yield of hCu,Zn-SOD sharply dropped. The reason for this phenomenon might be that metabolic flux was diverted more towards higher biomass formation and the high DO level led to plasmid instability. Thus a balance availability of oxygen seemed to be important for hCu,Zn-SOD production. Moreover, lactose has been reported to improve protein expression and biomass at suitable concentration, but has negative effect at excessively high concentration (Li et al. 2006). The optimal lactose feed rate for hCu,Zn-SOD production was 10 mL/h. The lactose feed rate 20 mL/h led to a decline of hCu,Zn-SOD production, probably because of the high osmotic stress or production of some other growth-inhibiting metabolites.
In this study, 342 mg purified hCu,Zn-SOD was obtained from 1 L cultures fermented under the optimized fermentation conditions. Moreover, the specific activity of the recombinant hCu,Zn-SOD reached 46,541 U/mg. This study may pave the way for the production of recombinant hCu,Zn-SOD through microbial fermentation, which has laid a foundation for industrial production of hCu,Zn-SOD in the future.
References
Battistoni A, CarrìMT MAP, Rotilio G (1992) Temperature-dependent protein folding in vivo –lower growth temperature increases yield of two genetic variants of Xenopus laevis Cu, Zn superoxide dismutase in Escherichia coli. Biochem Biophys Res Commun 186(3):1339–1344. https://doi.org/10.1016/S0006-291X(05)81553-0
Carrì MT, Battistoni A, Polizio F, Desideri A, Rotilio G (1994) Impaired copper binding by the H46R mutant of human Cu, Zn superoxide dismutase, involved in amyotrophic lateral sclerosis. FEBS Lett 356:314–316. https://doi.org/10.1016/0014-5793(94)01295-4
Castan A, Enfors SO (2000) Characteristics of a DO-controlled fed-batch culture of Escherichia coli. Bioproc Eng 22(6):509–515. https://doi.org/10.1007/s004499900094
Francis DM, Page R (2010) Strategies to optimize protein expression in E coli. Curr Protoc Protein Sci 61(1):5241–52429. https://doi.org/10.1002/0471140864.ps0524s61
Fridovich I (1983) Superoxide radical: an endogenous toxicant. Annu Rev Pharmacol Toxicol 23:239–257. https://doi.org/10.1146/annurev.pa.23.040183.001323
Fujii J, Myint T, Seo HG, Kayanoki Y, Ikeda Y, Taniguchi N (1995) Characterization of wild-type and amyotrophic lateral sclerosis-related mutant Cu, Zn-superoxide dismutases overproduced in baculovirus-infected insect cells. J Neurochem 64:1456–1461. https://doi.org/10.1046/j.1471-4159.1995.64041456.x
Hallewell RA, Laria I, Tabrizi A, Carlin G, Getzoff ED, Tainer JA, Cousens LS, Mullenbach GT (1989) Genetically engineered polymers of human CuZn superoxide dismutase. J Biol Chem 264(9):5260–5268. https://doi.org/10.1016/0014-5793(89)80289-3
Hallewell RA, Masiarz FR, Najarian RC, Puma JP, Quiroga MR, Randolph A, Sanchez-Pescador R, Scandella CJ, Smith B, Steimer KS, Mullenbach GT (1985) Human Cu/Zn superoxide dismutase cDNA: isolation of clones synthesising high levels of active or inactive enzyme from an expression library. Nucleic Acids Res 13(6):2017–2034. https://doi.org/10.1093/nar/13.6.201
Hartman JR, Geller T, Yavin Z, Bartfeld D, Kanner D, Aviv H, Gorecki M (1986) High-level expression of enzymatically active human Cu/Zn superoxide dismutase in Escherichia coli. Proc Natl Acad Sci 83(19):7142–7146. https://doi.org/10.1073/pnas.83.19.7142
Hayward LJ, Rodriguez JA, Kim JW, Tiwari A, Goto JJ, Cabelli DE, Valentine JS, Brown RHJ (2002) Decreased metallation and activity in subsets of mutant superoxide dismutases associated with familial amyotrophic lateral sclerosis. J Biol Chem 277:15923–15931. https://doi.org/10.1074/jbc.M112087200
Huang P, Feng L, Oldham EA, Keating MJ, Plunkett W (2000) Superoxide dismutase as a target for the selective killing of cancer cells. Nature 407(6802):390–395. https://doi.org/10.1038/35030140
Huo J, Shi H, Yao Q, Chen H, Wang L, Chen K (2010) Cloning and purification of recombinant silkworm dihydrolipoamide dehydrogenase expressed in Escherichia coli. Protein Expr Purif 72(1):95–100. https://doi.org/10.1016/j.pep.2010.01.014
Johnson P (2002) Antioxidant enzyme expression in health and disease: effects of exercise and hypertension. Comp Biochem Physiol C Toxicol Pharmacol 133(4):493–505. https://doi.org/10.1016/S1532-0456(02)00120-5
Kamran J, Jose MA, Zia F, Saima W, Malik ZA, Syed SY (2016) Engineered production of short chain fatty acid in Escherichia coli using fatty acid synthesis pathway. PLoS ONE 11(7):e0160035. https://doi.org/10.1371/journal.pone.0160035
Kilic N, Taslipinar YM, Guney Y, Tekin E, Onuk E (2014) An investigation into the serum thioredoxin superoxide dismutase malondialdehyde and advanced oxidation protein products in patients with breast cancer. Ann Surg Oncol 21(13):4139–4143. https://doi.org/10.1245/s10434-014-3859-3
Kim Y, Jeon YJ, Ryu K, Kim TY (2017) Zinc(II) ion promotes anti-inflammatory effects of rhSOD3 by increasing cellular association. BMB Rep 50(2):85–90. https://doi.org/10.5483/BMBRep.2017.50.2.150
Lee SY (1996) High cell-density culture of Escherichia coli. Trends Biotechnol 14(3):98–105. https://doi.org/10.1016/0167-7799(96)80930-9
Li HT, Jiao M, Chen J, Liang Y (2010) Roles of zinc and copper in modulating the oxidative refolding of bovine copper zinc superoxide dismutase. Acta Biochim Biophys Sin 42(3):183–194. https://doi.org/10.1093/abbs/gmq005
Lin F, Yan D, Chen Y, Emmanuella FE, Shi H, Han B, Zhou Y (2018) Cloning, purification and enzymatic characterization of recombinant human superoxide dismutase 1 (hSOD1) expressed in Escherichia coli. Acta Biochim Pol 65(2):235–240. https://doi.org/10.18388/abp.2017_2350
Li ZP, Zhang X, Tan TW (2006) Lactose-induced production of human soluble B lymphocyte stimulator (hsBLyS) in E coli with different culture strategies. Biotechnol Lett 28(7):477–483. https://doi.org/10.1007/s10529-006-0002-y
Liu JR, Liu JG, Zhao XY, Gu YJ (2005) Purification and renaturation of recombinant human Cu, Zn-SOD by metal-chelating affinity chromatography. Sheng Wu Gong Cheng Xue Bao 21(6):993–997. https://doi.org/10.3321/j.issn:1000-3061.2005.06.025
Liu WC, Gong T, Wang QH, Liang X, Chen JJ, Zhu P (2016) Scaling-up fermentation of pichia pastoris to demonstration-scale using new methanol-feeding strategy and increased air pressure instead of pure oxygen supplement. Sci Rep 6:18439. https://doi.org/10.1038/srep18439
Lu R, Zhang T, Wu D, He Z, Jiang L, Zhou M, Cheng Y (2018) Production of functional human Cu, Zn-SOD and EC-SOD in bitransgenic cloned goat milk. Transgenic Res 27(4):343–354. https://doi.org/10.1007/s11248-018-0080-3
Marisch K, Bayer K, Cserjan-Puschmann M, Luchner M, Striedner G (2013) Evaluation of three industrial Escherichia coli strains in fed-batch cultivations during high-level SOD protein production. Microb Cell Fact 12:1–11. https://doi.org/10.1186/1475-2859-12-58
Nordlund A, Leinartaite L, Saraboji K, Aisenbrey C, Grobner G, Zetterstrom P, Danielsson J, Logan DT, Oliveberg M (2009) Functional features cause misfolding of the ALS-provoking enzyme SOD1. Proc Natl Acad Sci 106(24):9667–9672. https://doi.org/10.1073/pnas.0812046106
Park DH, Yoon SYH, Nam HG, Park JM (2002) Expression of functional human-cytosolic Cu/Zn superoxide dismutase in transgenic tobacco. Biotechnol Lett 24:681–686. https://doi.org/10.1023/a:1015273714571
Perry JJP, Shin DS, Getzoff ED, Tainer JA (2010) The structural biochemistry of the superoxide dismutases. Biochim Biophys Acta 1804:245–262. https://doi.org/10.1016/j.bbapap.2009.11.004
Riesenberg D, Guthke R (1999) High-cell-density cultivation of microorganisms. Appl Microbiol Biotechnol 51(4):422–430. https://doi.org/10.1007/s002530051412
Rumfeldt JA, Lepock JR, Meiering EM (2009) Unfolding and folding kinetics of amyotrophic lateral sclerosis-associated mutant Cu. Zn superoxide dismutases J Mol Biol 385(1):278–298. https://doi.org/10.1016/j.jmb.2008.10.003
Swalley SE, Fulghum JR, Chambers SP (2006) Screening factors effecting a response in soluble protein expression: formalized approach using design of experiments. Anal Biochem 351(1):122–127. https://doi.org/10.1016/j.ab.2005.11.046
Takeshima Y, Takatsugu N, Sugiura M, Hagiwara H (1994) High-level expression of human superoxide dismutase in the cyanobacterium Anacystis nidulans 6301. Proc Natl Acad Sci 91(21):9685–9689. https://doi.org/10.1073/pnas.91.21.9685
Tolmasoff JM, Ono T, Cutler RG (1980) Superoxide dismutase: correlation with life-span and specific metabolic rate in primate species. Proc Natl Acad Sci 77(5):2777–2781. https://doi.org/10.1073/pnas.77.5.2777
Tripathi NK, Sathyaseelan K, Jana AM, Rao PVL (2009) High yield production of heterologous proteins with Escherichia coli. Def Sci J 59(2):137–146. https://doi.org/10.14429/dsj.59.1501
Vasina JA, Baneyx F (1997) Expression of aggregation-prone recombinant proteins at low temperatures: a comparative study of the Escherichia coli cspA and tac promoter systems. Protein Expr Purif 9(2):211–218. https://doi.org/10.1006/prep.1996.0678
Vats P, Sagar N, Singh TP, Banerjee M (2015) Association of superoxide dismutases (SOD1 and SOD2) and glutathione peroxidase 1 (GPx1) gene polymorphisms with type 2 diabetes mellitus. Free Radic Res 49(1):17–24. https://doi.org/10.3109/10715762.2014.971782
Wittung-Stafshede P (2004) Role of cofactors in folding of the blue-copper protein azurin. Inorg Chem 43(25):7926–7933. https://doi.org/10.1021/ic049398g
Wu CY, Steffen J, Eide DJ (2009) Cytosolic superoxide dismutase (SOD1) is critical for tolerating the oxidative stress of zinc deficiency in yeast. PLoS ONE 4:e7061. https://doi.org/10.1371/journal.pone.0007061
Wurm DJ, Veiter L, Ulonska S, Eggenreich B, Herwig C, Spadiut O (2016) The E coli pET expression system revisited-mechanistic correlation between glucose and lactose uptake. Appl Microbiol Biotechnol 100(20):8721–8729. https://doi.org/10.1007/s00253-016-7620-7
Yoo HY, Kim SS, Rho HM (1999) Overexpression and simple purification of human superoxide dismutase (SOD1) in yeast and its resistance to oxidative stress. J Biotechnol 68:29–35. https://doi.org/10.1016/S0168-1656(98)00188-6
Funding
This work is supported by grants from the Drug Innovation Major Project (2018ZX09711001-006) and CAMS Innovation Fund for Medical Sciences (CIFMS) (2019-I2M-005).
Author information
Authors and Affiliations
Contributions
YJL and ZP designed this project. LXL and YJL performed the experiments. All the authors analyzed the data. YJL wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yang, JL., Li, XL., Jiang, FL. et al. High-level soluble expression of human Cu,Zn superoxide dismutase with high activity in Escherichia coli. World J Microbiol Biotechnol 36, 106 (2020). https://doi.org/10.1007/s11274-020-02883-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02883-6