Abstract
Transaminases, which catalyze the stereoselective transfer of an amino group between an amino donor and a prochiral ketone substrate, are interesting biocatalytic tools for the generation of optically pure chiral amines. In particular, amine transaminases (ATAs) are of industrial interest because they are capable of performing reductive amination reactions using a broad range of amine donors and acceptors. The most remarkable example of ATAs industrial application is in the production process of the anti-hyperglycaemic drug sitagliptin (Januvia®/Janumet®), which generated around 6 billion U.S. dollars of revenue to Merck in 2016. In this review, an update about the availability of microbial ATAs, discovered by both screening and database-mining approaches, or obtained by protein engineering of wild-type enzymes, will be provided. Current challenges in ATAs application and possible solutions will be also discussed. In particular, innovative biocatalytic process strategies aimed at the improvement of ATAs performances in chiral amines synthesis, e.g., using in situ product removal process strategies or flow reactors, will be presented. The progress in the industrial exploitation of these enzymes will be highlighted by selected examples of large-scale ATA-catalyzed processes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chiral amines are very important building blocks for the synthesis of several bioactive compounds and active pharmaceutical ingredients, e.g., the Alzheimer’s drug rivastigmine, the adrenergic antagonist dilevalol, and the antiretroviral drug lopinavir (Nugent and El-Shazly 2010; Paul et al. 2014). Since conventional chemical methods for the asymmetric synthesis of amines suffer from different limitations, e.g., low efficiency/selectivity and high environmental impact (Constable et al. 2007), many investigations have been carried during the last two decades to develop alternative biocatalytic routes. To this aim, different types of enzymatic activities have been explored, such as amine dehydrogenases, monoamine oxidases, imine reductases, ammonia lyases, and transaminases. Among the latter, the enzymes capable of catalyzing reductive amination reactions using, as donors, simple amines, independently from the presence of a carboxylic group in the substrate, have been denominated amine transaminases (ATAs) (Steffen-Munsberg et al. 2015; Guo and Berglund 2017). These biocatalysts are pyridoxal-5′-phosphate (PLP)-dependent enzymes, well characterized from a structural and mechanistic point of view, and their applicative potential has been already established with several examples of chiral amines synthesis starting from readily available prochiral ketones (Mathew and Yun 2012; Brundiek and Höhne 2016). However, some limitations and challenges of ATAs for large scale applications have been identified as well, such as a quite narrow substrate scope, unfavorable thermodynamic reaction equilibria, and substrate/product inhibition. These findings have largely stimulated the search of both novel enzymes and innovative process conditions, and selected examples of the results obtained so far will be given in the following.
An update on newly discovered ATAs
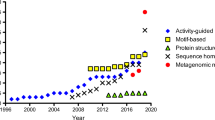
To date more than 60 microbial ATAs have been discovered (Guo and Berglund 2017), about twenty having been identified in just the last couple of years (Table 1).
New ATAs have been either isolated from wild-type microorganisms or identified in sequence databases by in silico screening and subsequently cloned and recombinantly overexpressed. Undoubtedly, sequence-based screening has proved to be a successful tool in ATAs discovery, thanks both to the huge number of new sequences and genomes available in public databases and to the more and more in-depth knowledge of the sequence/structure–function relationships of ATAs (Pavkov-Keller et al. 2016; Höhne et al. 2010; Steffen-Munsberg et al. 2015).
Among the most recent examples reported in the literature, the search of ATAs by in silico screening has given interesting results in the identification of enzymes with high process stability, and, in particular, thermostability. In fact, thermostable proteins often show better performances in industrial applications thanks to their superior stability under operational conditions and in the presence of organic solvents (Littlechild 2017). Different groups have searched for new ATAs in the genomes of thermophiles, i.e., microorganisms with an optimal growth temperature above 60 °C. In particular, three new ATAs have been recently identified from thermophilic microorganisms, specifically from Geobacillus thermodenitrificans (Chen et al. 2016), Sphaerobacter thermophilus (Mathew et al. 2016b), and Thermomicrobium roseum (Mathew et al. 2016a). All these biocatalysts showed a thermophilic behavior with optimal temperatures at 60, 65, and 80 °C, respectively, a high thermostability and activity increases after thermal treatment at 60–65 °C. As an alternative to genome mining, metagenomics, i.e., the study of the genetic material directly obtained from environmental samples, has been also exploited to find thermostable ATAs (Ferrandi et al. 2017). In this work, three new ATAs have been identified by in silico screening of a pool of metagenomic sequences from hot terrestrial environments. Among these, the B3-TA enzyme showed an outstanding thermostability, retaining 85% of activity after 5 days of incubation at 80 °C. Interestingly, despite the high sequence similarity (92%) between B3-TA and the T. roseum ATA (Mathew et al. 2016a), the latter showed a much lower thermostability by losing completely its activity after 24 h incubation at 60 °C.
As far as the substrate scope concerns, bifuctional transaminases, that accept simple and cheap mono-/diamine donors as well as a wide range of ketone acceptors as substrates, have been recently discovered by analysis of multi-sequence alignments (Slabu et al. 2016; Galman et al. 2017). In particular, the enzymes from Pseudomonas putida, P. chlororaphis, and P. aureofaciens resulted to be very efficient in the stereoselective transamination of various ketones in the presence of natural diamines as amino donors, thus providing a convenient biocatalytic access to different pharmaceutically relevant chiral amines (Galman et al. 2017).
A bioinformatic search among crystal structures with unknown function has permitted instead the discovery of an ATA from Bacillus anthracis (Ban-TA) showing active site residues different from those in related (class III) transaminases, thus making very unlikely the prediction of its function by identification of active site fingerprints (Steffen-Munsberg et al. 2016). Additionally, the use of key motifs based algorithms has recently allowed the discovery of a (R)-selective ATA from Capronia semiimersa (Iglesias et al. 2017). This enzyme has shown a good activity toward a broad range of substrates, including iso-propylamine (iPA), an amino donor suitable for industrial applications.
However, sequence-based screening suffers from some intrinsic limitations, e.g., the fact that the novel enzymes often resemble in their activity and structural features the homologues used for the search and that it doesn’t allow a quick identification of novel biocatalysts with a defined substrate scope. Thus, an alternative strategy to discover new ATAs involves the identification of microorganisms able to convert the substrate of choice, e.g., by employing a screening culture containing the target amine as sole nitrogen source. Moreover, thanks to the ever lower cost of genome sequencing, the target enzymes can be quickly identified by in silico analysis of the bacterial genomes (Wilding et al. 2015, 2016; Pavkov-Keller et al. 2016; Wu et al. 2017).
By using this approach, a (S)-ATA from Pseudomonas sp. strain AAC able to metabolize 12-aminododecanoic acid, the constituent building block of homo-nylon-12, has been identified by enriching a bacterial strain collection with 12-aminododecanoic acid (Wilding et al. 2015). Further characterization has showed that this ATA is a highly promiscuous transaminase, catalyzing amine transfers with 39 of the 50 substrates tested (Wilding et al. 2016). In a similar way, two new (R)-ATAs have been isolated from Curtobacterium pusillum and Microbacterium ginsengisoli, respectively, by using (R)-amines for enrichment culture of microorganisms from soil samples. Interestingly, structural analysis of the C. pusillum enzyme has suggested the existence a subgroup within the fold IV family of transaminases, showing significant phylogenetic distance from previously described ATAs as well as differences at the respective active sites (Pavkov-Keller et al. 2016).
Improvement of ATAs performances by protein engineering
Despite microbial world representing a extraordinary source of new enzymes, it’s unfortunately not very common to find in nature biocatalysts that show all the desirable features for industrial applications. To overcome this problem, different protein engineering approaches have been used to optimize available ATAs, a milestone having been set in 2010 with the outstanding work of Merck and Codexis in the development of a biocatalytic route for the synthesis of the drug sitagliptin (Savile et al. 2010). During the last years, thanks also to the increasing number of available crystal structures, structure-inspired design of novel transaminases has been widely applied to this aim (Table 2).
Many efforts have been spent to broad the ATA substrate scope, especially to make ATAs capable to accept bulky substrates. To this aim, specific residues in both the small (Hwang et al. 2008; Park et al. 2014; Nobili et al. 2015; Genz et al. 2016) and the large binding pocket (Cho et al. 2008; Han et al. 2015a, b, c) of ATAs active site have been changed to reduce steric constraints. For example, the small pocket of the ATA from V. fluvialis (Vf-ATA), which provides enough space mainly for a methyl group, as for example in (1) (Fig. 1) (Shin and Kim 2002), has been engineered to accept other substituents such as the side chain of (2) and (3) (Nobili et al. 2015).
In particular, Y150, V153 and F85 residues have been identified to cause a steric hindrance with these substrates. In fact, a 53-fold improvement of the specific activity toward (2) (mutant Y150M/V153A) and a 26-fold improvement of the specific activity toward (3) (mutant F85L/V153A) could be obtained by site-directed mutagenesis. These variants have been subsequently exploited in the respective transamination reactions, giving 98% conversion and > 98% enantiomeric excess (ee). In addition, the small binding pocket of Vf-ATA was further enlarged and reshaped, and the best variant (L56V/W57C/F85V/V153A) showed the highest specific activity towards (4), an amine bearing a bulky tert-butyl substituent, not accessible by any of the wild-type ATAs tested before (Genz et al. 2016). Further studies have been carried out by Pavlidis et al. (2016) to evolve the ATA from Rugeria sp. toward the stereoselective synthesis of bulky amines such as (5–7), thus allowing the identification of a specific sequence motif (Y59/Y87/Y152/T213/P423). Interestingly, one of the best variants obtained in this study was up to 8900 times more active toward (6), and 310 and 98 times more active toward (5) and (7) than the wild type, respectively. Furthermore, the variant Y59W/Y87F/Y152F/T231A has been used in the asymmetric synthesis of various amines, the best result being achieved in the obtainment of (6) with 99% conversion and > 99% ee. Further mutagenesis studies have allowed the identification of variants capable to accept also the substrates (4) and (8) (Weiß et al. 2016, 2017). Another attempt to expand ATA substrate specificity has been carried on the Halomonas elongata ATA (HEWT) by site directed mutagenesis of key residues located in the large pocket. The activity toward ortho-substituted acetophenone derivatives has been improved with the substitution W56G, while the mutation I258A has yielded an improvement in the reactivity toward para-substituted compounds (Contente et al. 2016).
The switch of ATA activity toward different functions by protein engineering has also been reported. For example, by changing the so-called “flipping” arginine, i.e., a highly conserved flexible arginine residue which enables the dual substrate recognition that permits carboxylate and hydrophobic group binding in most of known ATAs (Steffen-Munsberg et al. 2015), into an hydrophobic residue and reducing the steric hindrance of W57 and L417, Vf-ATA was converted from an amine:α-keto acid transferase into an amine:aldehyde transferase (Genz et al. 2015). Reversion or improvement of ATA enantioselectivity by structure based protein engineering have been also investigated (Svedendahl et al. 2010; Humble et al. 2012; Skalden et al. 2015). In particular, as until recently the available ATAs were predominantly (S)-selective, the need of (R)-selective ATAs has encouraged studies aimed at the switch of ATA enantioselectivity, one or two mutations in key residues being in some cases sufficient to reach this goal. For example, the substitution V328A in the Arthrobacter citreus ATA caused a shift from 98% ee (S) to 53% ee (R) in the synthesis of (9) (Svedendahl et al. 2010), while the double substitution F88A/A231F in the Chromobacterium violaceum ATA reversed the enantiomeric preference for (10) with an increase of the E value from 3.9 (S) to 63 (R) (Humble et al. 2012).
Finally, the stability of ATAs under process conditions, a challenging feature for their industrial applicability, has been recently tackled by identifying structural elements responsible for amine-donor induced inactivation (Börner et al. 2017). In this work, the combination of selected mutations in the so-called “cofactor-ring” binding motif allowed a significant improvement of the operational, solvent and thermal stability of an ATA previously discovered in a metagenomic library, and pave the way to further protein engineering studies since this structural “hotspot” is present also in other ATAs.
Development of ATA-catalyzed processes
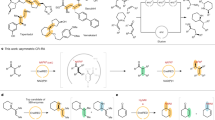
Besides enzyme inhibition and process stability, the major challenge that is often encountered in the set-up of ATA-catalyzed transaminations is the unfavorable thermodynamic equilibrium of the reaction (Brundiek and Höhne 2016). This problem could be avoided either by performing cascade reactions or by applying strategies to drive the reaction equilibrium toward amine synthesis. The use of an excess of co-substrate is the most employed method on the lab scale, but this approach may be limited by amine donor solubility and enzyme inhibition (Brundiek and Höhne 2016; Guo and Berglund 2017). Another option is to remove the product (or co-product) from the reaction by using different methods. Mild vacuum distillation with a nitrogen sweep is very efficient to remove the co-product acetone (Savile et al. 2010), but unfortunately, this strategy is applicable only if iPA is used as amino donor and this compound is not accepted by all ATAs. In situ product removal (ISPR) techniques (Lye and Woodley 1999), such as liquid/liquid or liquid/solid extraction, have proven to be effective in ketone product extraction but their applicability is generally hampered by the poor selectivity of the separation process, although some progress has been recently reported (Rehn et al. 2014; Börner et al. 2015; Satyawali et al. 2017). On the other hand, cascade systems have been largely explored (Simon et al. 2014). Coupled enzymatic reactions have been employed to convert pyruvate, the deamination product of the most commonly used amino donor alanine, into lactate by lactate dehydrogenase (LDH), together with a glucose/glucose dehydrogenase (GDH) system for cofactor regeneration (Fig. 2a) (Truppo et al. 2010).
Pyruvate could be alternatively recycled into the transamination reaction by alanine dehydrogenase (AlaDH) when employing a (S)-selective ATA (Mutti et al. 2011), or it could be converted into d-alanine by an alanine racemase to be available for (R)-selective ATAs (Richter et al. 2015). Both ATA/AlaDH and ATA/LDH/GDH systems have proven to be quite efficient and in particular the latter system has been employed also on large scale (Girardin et al. 2013). Recently, the exploitation of the innate microbial metabolism to shift transamination equilibrium toward the desired products by removing pyruvate in recombinant whole cell biocatalysts has also been proposed (Han and Shin 2014; Weber et al. 2017), while Farnberger et al. (2017) have investigated the co-expression in E. coli of ATAs together with the enzymes for co-product removal and cofactor regeneration, e.g., AlaDH and GDH. Either resting or lyophilized cells, as well as cell-free extracts, have been tested, however conversions of a model substrate didn’t exceed 77%, thus suggesting the possible need of further fine-tuning of these strategies.
Aside product/co-product removal, the use of the so-called “smart” cosubstrates, whose products spontaneously transform into end-products, could be applied to drive the asymmetric synthesis of chiral amines toward completion. In this view, diamines that spontaneously cyclize into cyclic imines (Fig. 2b) have been successfully employed to this aim (Galman et al. 2017). However, this strategy is often limited by the strict substrate specificity of ATAs (Galman et al. 2017; Payer et al. 2017).
A promising alternative to the above described solutions lies in the exploitation of immobilized ATAs in flow systems where, as the substrate is converted into the product, the latter is removed from the reaction environment, thus avoiding enzyme inhibition and shifting the balance of the reaction toward product formation. Notably, enzyme immobilization often enhances protein operational stability, allowing at the same time further implementation of the process with in-line purification or ISPR steps and devices for analysis (Bajić et al. 2017; Heintz et al. 2017). In a recent example, Planchestainer et al. (2017) have immobilized the poly-His-tagged H. elongata ATA (HEWT) on a metal derivatized epoxy-resin and synthesized a series of chiral amines in continuous flow with satisfying productivities. The packed bed reactor has been connected with an in-line purification system to achieve the easy recover of the amine product. Furthermore, it was demonstrated that the operational stability of HEWT was improved by immobilization, thus making its reuse for several cycles feasible.
Examples of large scale applications of ATAs
The wild-type or engineered ATAs and the process implementation approaches described in the previous paragraphs have been not only investigated at a laboratory scale, but, in some cases, also tested in an industrial environment in the development of synthetic routes of different bioactive molecules.
For example, ATA 117 (Codexis) has been employed to produce (12), a useful building block in the synthesis of the orexin antagonist MK-6096, a candidate for the treatment of insomnia (Fig. 3a). The bioconversion, performed in 100 L total volume starting from 4.5 kg substrate and using D-alanine as amino donor with a coupled LDH/GDH regeneration system, has successfully resulted in 74% yield and 99% ee. Interestingly, this biocatalytic reaction could be well integrated in the kg-scale nine-step chemoenzymatic synthesis of MK-6096 with 13% overall yield (Girardin et al. 2013).
Another remarkable example is the use of the ATA from V. fluvialis (Vf-ATA) for the production of (14) a key intermediate in the synthesis of the JAK2 kinase inhibitor AZD1480, developed by Astra Zeneca for the treatment of idiopathic myelofibrosis and polycythaemia rubra vera (Frodsham et al. 2013; Meadows et al. 2013) (Fig. 3b). After biocatalyst selection, process conditions were optimized on gram scale by identification of α-methyl benzyl amine as amino donor and of a biphasic system with 20% (v/v) toluene to increase substrate loading up to 0.35 M and avoid enzyme inhibition by the co-product acetophenone. Subsequent scale-up of the biotransformation has allowed the conversion of 450 g of (13) into (14) in 79% yield and 99% ee. The intermediate has been then used in the chemoenzymatic synthesis of AZD1480 on 100 L scale with a > 30% overall yield (Frodsham et al. 2013).
In a more recent example, Feng et al. (2017) have synthesized (16) (Fig. 3c), a useful building block for the production of the antibiotic besifloxacin, using a commercially available (R)-selective ATA. Under optimized reaction conditions, 0.3 kg of (15) were converted into (16) in the presence of 30% (v/v) iPA as amine donor with 80% isolated yield and > 99% ee.
While in the abovementioned examples wild-type enzymes have been employed, Merck and Codexis had to improve ATA 117 by several round of mutagenesis to catalyze the last step of the synthesis of the antidiabetic compound sitagliptin (18, Fig. 3d) (Savile et al. 2010). After widening ATA 117 active site by 12 mutations in key “hot spots” to accept the bulky substrate pro-sitagliptin ketone (17), further improvements have been necessary to meet all the process requirements such as high activity at alkaline pH and at temperatures above 45 °C and tolerance toward cosolvents and high substrate/cosubstrate concentrations. Eventually, a variant carrying 27 mutations was employed in the kg-scale synthesis of (18) using iPA as amino donor. Reaction was successfully driven toward product formation by stripping the volatile coproduct acetone, resulting in a 92% yield and > 99.95% ee.
Other interesting examples have been recently reported using commercially available ATAs, although on a smaller scale (g scale). Specifically, Pfizer successfully employed ATA 036 in the concomitant transamination and dynamic kinetic resolution of a 4-piperidone to obtain the precursor of an anti-cancer drug (Peng et al. 2014), while Merck evolved the enzyme ATA 013 to integrate a biocatalytic step in the chemoenzymatic synthesis of the antiarrhythmic drug Vernakalant (Limanto et al. 2014). In both cases good yields and excellent enantioselectivity have been achieved.
Conclusions
Recent works on ATAs further confirm the huge potential of these enzymes in the development of innovative routes to prepare chiral amines up to the industrial level. The diversity of the available biocatalysts is rapidly improving thanks to both the discovery of novel natural enzymes and the generation of engineered variants, thus facing possible issues related to both substrate specificity/selectivity and stability. Moreover, the design and development of innovative process-based strategies will help to improve the efficiency and competitiveness of ATA-catalyzed reactions and further expand their practical application and integration of current production methods.
References
Bajić M, Plazl I, Stloukal R, Žnidaršič-Plazl P (2017) Development of a miniaturized packed bed reactor with ω-transaminase immobilized in LentiKats®. Process Biochem 52:63–72. https://doi.org/10.1016/j.procbio.2016.09.021
Baud D, Jeffries JWE, Moody TS et al (2017) A metagenomics approach for new biocatalyst discovery: application to transaminases and the synthesis of allylic amines. Green Chem 19:1134–1143. https://doi.org/10.1039/C6GC02769E
Börner T, Rehn G, Grey C, Adlercreutz P (2015) A process concept for high-purity production of amines by transaminase-catalyzed asymmetric synthesis: combining enzyme cascade and membrane-assisted ISPR. Org Process Res Dev 19:793–799. https://doi.org/10.1021/acs.oprd.5b00055
Börner T, Rämisch S, Bartsch S et al (2017) Three in one: temperature, solvent and catalytic stability by engineering the cofactor-binding element of amine transaminase. ChemBioChem 18:1482–1486. https://doi.org/10.1002/cbic.201700236
Brundiek H, Höhne M (2016) Transaminases: a biosynthetic route for chiral amines. In: Applied biocatalysis: from fundamental science to industrial applications. Wiley, Weinheim, pp 199–218
Cerioli L, Planchestainer M, Cassidy J et al (2015) Characterization of a novel amine transaminase from Halomonas elongata. J Mol Catal B Enzym 120:141–150. https://doi.org/10.1016/j.molcatb.2015.07.009
Chen Y, Yi D, Jiang S, Wei D (2016) Identification of novel thermostable taurine–pyruvate transaminase from Geobacillus thermodenitrificans for chiral amine synthesis. Appl Microbiol Biotechnol 100:3101–3111. https://doi.org/10.1007/s00253-015-7129-5
Cho B-K, Park H-Y, Seo J-H et al (2008) Redesigning the substrate specificity of ω-aminotransferase for the kinetic resolution of aliphatic chiral amines. Biotechnol Bioeng 99:275–284. https://doi.org/10.1002/bit.21591
Constable DJC, Dunn PJ, Hayler JD et al (2007) Key green chemistry research areas—a perspective from pharmaceutical manufacturers. Green Chem 9:411–420. https://doi.org/10.1039/B703488C
Contente ML, Planchestainer M, Molinari F, Paradisi F (2016) Stereoelectronic effects in the reaction of aromatic substrates catalysed by Halomonas elongata transaminase and its mutants. Org Biomol Chem 14:9306–9311. https://doi.org/10.1039/C6OB01629D
Deszcz D, Affaticati P, Ladkau N et al (2015) Single active-site mutants are sufficient to enhance serine:pyruvate α-transaminase activity in an ω-transaminase. FEBS J 282:2512–2526. https://doi.org/10.1111/febs.13293
Farnberger JE, Lorenz E, Richter N et al (2017) In vivo plug-and-play: a modular multi-enzyme single-cell catalyst for the asymmetric amination of ketoacids and ketones. Microb Cell Fact 16:132. https://doi.org/10.1186/s12934-017-0750-5
Feng Y, Luo Z, Sun G et al (2017) Development of an efficient and scalable biocatalytic route to (3R)-3-aminoazepane: a pharmaceutically important intermediate. Org Process Res Dev 21:648–654. https://doi.org/10.1021/acs.oprd.7b00074
Ferrandi EE, Previdi A, Bassanini I et al (2017) Novel thermostable amine transferases from hot spring metagenomes. Appl Microbiol Biotechnol 101:4963–4979. https://doi.org/10.1007/s00253-017-8228-2
Frodsham L, Golden M, Hard S et al (2013) Use of ω-transaminase enzyme chemistry in the synthesis of a JAK2 kinase inhibitor. Org Process Res Dev 17:1123–1130. https://doi.org/10.1021/op400133d
Galman JL, Slabu I, Weise NJ et al (2017) Biocatalytic transamination with near-stoichiometric inexpensive amine donors mediated by bifunctional mono- and di-amine transaminases. Green Chem 9:285–288. https://doi.org/10.1039/C6GC02102F
Gao S, Su Y, Zhao L et al (2017) Characterization of a (R)-selective amine transaminase from Fusarium oxysporum. Process Biochem. https://doi.org/10.1016/j.procbio.2017.08.012
Genz M, Vickers C, van den Bergh T et al (2015) Alteration of the donor/acceptor spectrum of the (S)-amine transaminase from Vibrio fluvialis. Int J Mol Sci 16:26953–26963. https://doi.org/10.3390/ijms161126007
Genz M, Melse O, Schmidt S et al (2016) Engineering the amine transaminase from Vibrio fluvialis towards branched-chain substrates. ChemCatChem 8:1–5. https://doi.org/10.1002/cctc.201601007
Girardin M, Ouellet SG, Gauvreau D et al (2013) Convergent kilogram-scale synthesis of dual orexin receptor antagonist. Org Process Res Dev 17:61–68. https://doi.org/10.1021/op3002678
Guo F, Berglund P (2017) Transaminase biocatalysis: optimization and application. Green Chem 19:333–360. https://doi.org/10.1039/C6GC02328B
Han S-W, Shin J-S (2014) Metabolically driven equilibrium shift of asymmetric amination of ketones by ω-transaminase using alanine as an amino donor. Biosci Biotechnol Biochem 78:1788–1790. https://doi.org/10.1080/09168451.2014.930328
Han SW, Park ES, Dong JY, Shin JS (2015a) Mechanism-guided engineering of ω-transaminase to accelerate reductive amination of ketones. Adv Synth Catal 357:1732–1740. https://doi.org/10.1002/adsc.201500211
Han SW, Park ES, Dong JY, Shin JS (2015b) Active-site engineering of ω-transaminase for production of unnatural amino acids carrying a side chain bulkier than an ethyl substituent. Appl Environ Microbiol 81:6994–7002. https://doi.org/10.1128/AEM.01533-15
Han SW, Park ES, Dong JY, Shin JS (2015c) Expanding substrate specificity of ω-transaminase by rational remodeling of a large substrate-binding pocket. Adv Synth Catal 357:2712–2720. https://doi.org/10.1002/adsc.201500239
Heintz S, Börner T, Ringborg RH et al (2017) Development of in situ product removal strategies in biocatalysis applying scaled-down unit operations. Biotechnol Bioeng 114:600–609. https://doi.org/10.1002/bit.26191
Höhne M, Schätzle S, Jochens H et al (2010) Rational assignment of key motifs for function guides in silico enzyme identification. Nat Chem Biol 6:807–813. https://doi.org/10.1038/nchembio.447
Huang J, Xie DF, Feng Y (2017) Engineering thermostable (R)-selective amine transaminase from Aspergillus terreus through in silico design employing B-factor and folding free energy calculations. Biochem Biophys Res Commun 483:397–402. https://doi.org/10.1016/j.bbrc.2016.12.131
Humble MS, Cassimjee KE, Abedi V et al (2012) Key amino acid residues for reversed or improved enantiospecificity of an ω-transaminase. ChemCatChem 4:1167–1172. https://doi.org/10.1002/cctc.201100487
Hwang BY, Ko SH, Park HY et al (2008) Identification of ω-aminotransferase from Caulobacter crescentus and site-directed mutagenesis to broaden substrate specificity. J Microbiol Biotechnol 18:48–54
Iglesias C, Panizza P, Rodriguez Giordano S (2017) Identification, expression and characterization of an R-ω-transaminase from Capronia semiimmersa. Appl Microbiol Biotechnol 101:5677–5687. https://doi.org/10.1007/s00253-017-8309-2
Limanto J, Ashley ER, Yin J et al (2014) A highly efficient asymmetric synthesis of vernakalant. Org Lett 16:2716–2719. https://doi.org/10.1021/ol501002a
Littlechild JA (2017) Improving the “tool box” for robust industrial enzymes. J Ind Microbiol Biotechnol 44:711–720. https://doi.org/10.1007/s10295-017-1920-5
Lye GJ, Woodley JM (1999) Application of in situ product-removal techniques to biocatalytic processes. Trends Biotechnol 17:395–402. https://doi.org/10.1016/S0167-7799(99)01351-7
Mathew S, Yun H (2012) ω-Transaminases for the production of optically pure amines and unnatural amino acids. ACS Catal 2:993–1001. https://doi.org/10.1021/cs300116n
Mathew S, Deepankumar K, Shin G et al (2016a) Identification of novel thermostable ω-transaminase and its application for enzymatic synthesis of chiral amines at high temperature. RSC Adv 6:69257–69260. https://doi.org/10.1039/C6RA15110H
Mathew S, Nadarajan SP, Chung T et al (2016b) Biochemical characterization of thermostable ω-transaminase from Sphaerobacter thermophilus and its application for producing aromatic β- and γ-amino acids. Enzyme Microb Technol 87–88:52–60. https://doi.org/10.1016/j.enzmictec.2016.02.013
Meadows RE, Mulholland KR, Schürmann M et al (2013) Efficient synthesis of (S)-1-(5-fluoropyrimidin-2-yl)ethylamine using an ω-transaminase biocatalyst in a two-phase system. Org Process Res Dev 17:1117–1122. https://doi.org/10.1021/op400131h
Midelfort KS, Kumar R, Han S et al (2013) Redesigning and characterizing the substrate specificity and activity of Vibrio fluvialis aminotransferase for the synthesis of imagabalin. Protein Eng Des Sel 26:25–33. https://doi.org/10.1093/protein/gzs065
Mutti FG, Fuchs CS, Pressnitz D et al (2011) Stereoselectivity of four (R)-selective transaminases for the asymmetric amination of ketones. Adv Synth Catal 353:3227–3233. https://doi.org/10.1002/adsc.201100558
Nobili A, Steffen-Munsberg F, Kohls H et al (2015) Engineering the active site of the amine transaminase from Vibrio fluvialis for the asymmetric synthesis of aryl-alkyl amines and amino alcohols. ChemCatChem 7:757–760. https://doi.org/10.1002/cctc.201403010
Nugent TC, El-Shazly M (2010) Chiral amine synthesis: recent developments and trends for enamide reduction, reductive amination, and imine reduction. Adv Synth Catal 352:753–819. https://doi.org/10.1002/adsc.200900719
Park ES, Park SR, Han SW et al (2014) Structural determinants for the non-canonical substrate specificity of the ω-transaminase from Paracoccus denitrificans. Adv Synth Catal 356:212–220. https://doi.org/10.1002/adsc.201300786
Paul CE, Rodríguez-Mata M, Busto E et al (2014) Transaminases applied to the synthesis of high added-value enantiopure amines. Org Process Res Dev 18:788–792. https://doi.org/10.1021/op4003104
Pavkov-Keller T, Strohmeier GA, Diepold M et al (2016) Discovery and structural characterisation of new fold type IV-transaminases exemplify the diversity of this enzyme fold. Sci Rep 6:38183. https://doi.org/10.1038/srep38183
Pavlidis IV, Weiß MS, Genz M et al (2016) Identification of (S)-selective transaminases for the asymmetric synthesis of bulky chiral amines. Nat Chem 8:1076–1082. https://doi.org/10.1038/nchem.2578
Payer SE, Schrittwieser JH, Kroutil W (2017) Vicinal diamines as smart cosubstrates in the transaminase-catalyzed asymmetric amination of ketones. European J Org Chem 2017:2553–2559. https://doi.org/10.1002/ejoc.201700253
Peng Z, Wong JW, Hansen EC et al (2014) Development of a concise, asymmetric synthesis of a smoothened receptor (SMO) inhibitor: Enzymatic transamination of a 4-piperidinone with dynamic kinetic resolution. Org Lett 16:860–863. https://doi.org/10.1021/ol403630g
Planchestainer M, Contente ML, Cassidy J et al (2017) Continuous flow biocatalysis: production and in-line purification of amines by immobilised transaminase from Halomonas elongata. Green Chem 19:372–375. https://doi.org/10.1039/C6GC01780K
Rehn G, Adlercreutz P, Grey C (2014) Supported liquid membrane as a novel tool for driving the equilibrium of omega-transaminase catalyzed asymmetric synthesis. J Biotechnol 179:50–55. https://doi.org/10.1016/j.jbiotec.2014.03.022
Richter N, Farnberger JE, Pressnitz D et al (2015) A system for ω-transaminase mediated (R)-amination using l-alanine as an amine donor. Green Chem 17:2952–2958. https://doi.org/10.1039/C4GC02363C
Satyawali Y, Ehimen E, Cauwenberghs L et al (2017) Asymmetric synthesis of chiral amine in organic solvent and in-situ product recovery for process intensification: a case study. Biochem Eng J 117:97–104. https://doi.org/10.1016/j.bej.2016.11.006
Savile CK, Janey JM, Mundorff EC et al (2010) Biocatalytic asymmetric synthesis of chiral amines from ketones applied to sitagliptin manufacture. Science 329:305–309. https://doi.org/10.1126/science.1188934
Shin J, Kim B (2002) Exploring the active site of amine: pyruvate aminotransferase on the basis of the substrate structure–reactivity relationship : how the enzyme controls substrate specificity and stereoselectivity. J Org Chem 67:2848–2853. https://doi.org/10.1021/jo016115i
Simon RC, Richter N, Busto E, Kroutil W (2014) Recent developments of cascade reactions involving ω-transaminases. ACS Catal 4:129–143. https://doi.org/10.1021/cs400930v
Skalden L, Peters C, Dickerhoff J et al (2015) Two subtle amino acid changes in a transaminase substantially enhance or invert enantiopreference in cascade syntheses. ChemBioChem 16:1041–1045. https://doi.org/10.1002/cbic.201500074
Slabu I, Galman JL, Weise NJ et al (2016) Putrescine transaminases for the synthesis of saturated nitrogen heterocycles from polyamines. ChemCatChem 8:1038–1042. https://doi.org/10.1002/cctc.201600075
Steffen-Munsberg F, Vickers C, Thontowi A et al (2013) Revealing the structural basis of promiscuous amine transaminase activity. ChemCatChem 5:154–157. https://doi.org/10.1002/cctc.201200545
Steffen-Munsberg F, Vickers C, Kohls H et al (2015) Bioinformatic analysis of a PLP-dependent enzyme superfamily suitable for biocatalytic applications. Biotechnol Adv 33:566–604. https://doi.org/10.1016/j.biotechadv.2014.12.012
Steffen-Munsberg F, Matzel P, Sowa MA et al (2016) Bacillus anthracis ω-amino acid:pyruvate transaminase employs a different mechanism for dual substrate recognition than other amine transaminases. Appl Microbiol Biotechnol 100:4511–4521. https://doi.org/10.1007/s00253-015-7275-9
Svedendahl M, Branneby C, Lindberg L, Berglund P (2010) Reversed enantiopreference of an ω-transaminase by a single-point mutation. ChemCatChem 2:976–980. https://doi.org/10.1002/cctc.201000107
Truppo MD, David Rozzell J, Turner NJ (2010) Efficient production of enantiomerically pure chiral amines at concentrations of 50 g/L using transaminases. Org Process Res Dev 14:234–237. https://doi.org/10.1021/op900303q
Weber N, Gorwa-Grauslund M, Carlquist M (2017) Improvement of whole-cell transamination with Saccharomyces cerevisiae using metabolic engineering and cell pre-adaptation. Microb Cell Fact 16:3. https://doi.org/10.1186/s12934-016-0615-3
Weiß MS, Pavlidis IV, Spurr P et al (2016) Protein-engineering of an amine transaminase for the stereoselective synthesis of a pharmaceutically relevant bicyclic amine. Org Biomol Chem 14:10249–10254. https://doi.org/10.1039/C6OB02139E
Weiß MS, Pavlidis IV, Spurr P et al (2017) Amine transaminase engineering for spatially bulky substrate acceptance. ChemBioChem 18:1022–1026. https://doi.org/10.1002/cbic.201700033
Wilding M, Walsh EFA, Dorrian SJ, Scott C (2015) Identification of novel transaminases from a 12-aminododecanoic acid-metabolizing Pseudomonas strain. Microb Biotechnol 8:665–672. https://doi.org/10.1111/1751-7915.12278
Wilding M, Peat TS, Newman J, Scott C (2016) A β-alanine catabolism pathway containing a highly promiscuous ω-transaminase in the 12-aminododecanate-degrading Pseudomonas sp. strain AAC. Appl Environ Microbiol 82:3846–3856. https://doi.org/10.1128/AEM.00665-16
Wu HL, Zhang JD, Zhang CF et al (2017) Characterization of four new distinct ω-transaminases from Pseudomonas putida NBRC 14164 for kinetic resolution of racemic amines and amino alcohols. Appl Biochem Biotechnol 181:972–985. https://doi.org/10.1007/s12010-016-2263-9
Acknowledgements
This work was supported by Fondazione Cariplo, Grant Numbers 2016-0731.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ferrandi, E.E., Monti, D. Amine transaminases in chiral amines synthesis: recent advances and challenges. World J Microbiol Biotechnol 34, 13 (2018). https://doi.org/10.1007/s11274-017-2395-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-017-2395-2