Abstract
Chiral amine diastereomers are ubiquitous in pharmaceuticals and agrochemicals1, yet their preparation often relies on low-efficiency multi-step synthesis2. These valuable compounds must be manufactured asymmetrically, as their biochemical properties can differ based on the chirality of the molecule. Herein we characterize a multifunctional biocatalyst for amine synthesis, which operates using a mechanism that is, to our knowledge, previously unreported. This enzyme (EneIRED), identified within a metagenomic imine reductase (IRED) collection3 and originating from an unclassified Pseudomonas species, possesses an unusual active site architecture that facilitates amine-activated conjugate alkene reduction followed by reductive amination. This enzyme can couple a broad selection of α,β-unsaturated carbonyls with amines for the efficient preparation of chiral amine diastereomers bearing up to three stereocentres. Mechanistic and structural studies have been carried out to delineate the order of individual steps catalysed by EneIRED, which have led to a proposal for the overall catalytic cycle. This work shows that the IRED family can serve as a platform for facilitating the discovery of further enzymatic activities for application in synthetic biology and organic synthesis.
Similar content being viewed by others
Main
Reductive amination (RA) is one of the most widely used and powerful methodologies in medicinal chemistry for the synthesis of high-value chiral amines1, enabling efficient formation of C–N bonds through the reductive coupling of carbonyls and amines2,4. The development of effective catalysts for asymmetric RA continues to be explored, including those based on metallo-5,6, organo-7 and biocatalysis3,8,9,10. Furthermore, valuable amino-containing compounds often contain multiple stereogenic centres (Fig. 1a), although total control of their chirality is more challenging, resulting in less efficient multistep syntheses11 or more-complex tandem catalysis systems12,13. Although multi-enzyme systems are highly amenable to these biomimetic tandem processes (Fig. 1b), difficulties in their assembly arise from incompatibilities in their reaction medium and reaction rates, which can lead to by-product formation and intricate reaction set-up14. To address these issues and achieve the desired reaction metrics, substantial protein engineering is often required on each enzyme component15. Discovery of a single enzyme that can control multiple stereocentres through an RA-like process would be highly desirable and enable efficient synthesis of valuable amine diastereomers using a one-pot, one-catalyst system (Fig. 1c).
a, Enantioenriched amine diastereomers are common in active pharmaceutical ingredients (APIs). b, One-pot strategies for the enzymatic synthesis of stereoenriched amine diastereomers. c, Our approach for developing a single enzyme-catalysed conjugate reduction–reductive amination (CR–RA) to prepare enantioenriched amine diastereomers. d, Identification of C=C and C=N reducing EneIRED (pIR-120) by screening 389 putative IREDs. e, Optimised reaction conditions for EneIRED-catalysed CR–RA, see Extended Data Figs. 1, 2 and Supplementary Information for further details. d.r., diastereomeric ratio; GDH, glucose dehydrogenase.
Nicotinamide-dependent enzymes are versatile biocatalysts for both asymmetric conjugate reduction (CR)16,17 and RA8,9,18,19. For RA, imine reductases (IREDs) have emerged as attractive catalysts because they possess broad substrate scope and can be engineered for industrial application20,21. IREDs are characterized as chemoselective for the reduction of C=N bonds22,23,24, although under exceptional circumstances they can reduce C=O bonds of activated carbonyl species25. Furthermore, we recently demonstrated that IREDs could be combined with ene-reductases (EREDs) in a one-pot process to reduce both the C=C and C=N bonds of cyclic α,β-unsaturated imines (ene-imines)26. We speculated that if an IRED could catalyse both of these steps, in a similar fashion to recently reported biosynthetic oxidoreductases27,28, this biocatalyst could be applied to the CR–RA of α,β-unsaturated carbonyls and allow access to enantioenriched amine diastereomers (Fig. 1c).
In pursuit of this activity, we screened both reported8,24,29,30,31 and our recently established (meta)genomic IREDs3,32 for the complete reduction of cyclic ene-imine I to amine II (Fig. 1d and Supplementary Table 1). Among the 389 IREDs screened, we observed that 262 catalysed reduction of I, with the majority behaving conventionally, that is, reducing the C=N bond only (206 enzymes, 53%). A smaller subset of IREDs were able to catalyse the reduction of both the C=C and C=N bonds of I to the diastereomerically enriched product II (44 enzymes, 11%). Furthermore, in a complementary fashion, some IREDs reduced solely the C=C bond (12 enzymes, 3%). Mapping the reaction profiles against genetic sequence indicated localized sequence–activity correlation only (Extended Data Fig. 1). One metagenomic enzyme, probably originating from an unclassified Pseudomonas species (EneIRED, Fig. 1d, see Supplementary Fig. 2 for predicted operon of EneIRED), exhibited excellent full reduction of I to II, and hence this enzyme was selected for further study.
EneIRED was examined for the ability to catalyse CR–RA of cyclohex-2-enone 1 with allylamine a, monitoring for potential reduced and coupled products 1a, 1′, 1′a including aza-conjugate addition. The reaction proceeded with high conversion, forming predominately CR–RA product 1′a and CR product 1′ without concomitant generation of direct RA product 1a. Optimization of the buffer type, pH, as well as co-solvent (Extended Data Fig. 2), increased the conversion to 1′a with 1.1 equivalents (eq.) of allylamine (61%, Supplementary Table 2) and could be scaled up using 20 eq. of a yielding the hydrochloride salt in 69% yield (Fig. 1e).
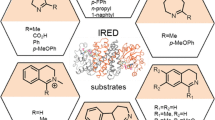
We explored the substrate scope of EneIRED using the optimized reaction conditions (Fig. 2, for a full list of substrates see Supplementary Fig. 4). EneIRED exhibited a broad substrate scope yielding CR and mono-RA products. Generally, unhindered enals and enones could be transformed with high chemoselectivity to the corresponding saturated amines. This trend is observed by comparing the reaction profiles of increasingly hindered C2-substituted but-2-enals 3–5 or decreasingly hindered amine donors a–c with cinnamaldehyde 6.
Major RA products are drawn, and their product distributions are illustrated by the donut chart, where percentage conversion = total area percentage of CR and RA products. aA non-enzymatic aza-conjugate addition product is observed. bCertain components are determined by analogy on the basis of the gas chromatography–mass spectrometry (GC–MS) spectrum; see Supplementary Table 2 for further details. conv., conversion; e.e., enantiomeric excess.
Cyclic enones with various ring sizes were all accepted by EneIRED, with 6- and 5-membered, 1 and 10, affording good conversion to the corresponding saturated amine products 1′b or 10′b without direct RA products 1b or 10b. 2-, 3- and 4,4′-methyl substituted cycloalkyl-2-enones 12–15 were also accepted by the enzyme, with 3-substituted 12 and 15 offering high conversion, chemo- and stereoselectivity to the corresponding CR–RA products 12′b and 15′b. C-3-substitution of the cyclohex-2-enone scaffold 15–22 was generally well tolerated, offering excellent conversion, chemo-, enantio- and (trans)-diastereoselectivity to the corresponding N-substituted cyclohexylamines 15′b, 16′b, 18′b and 20′b. Factors controlling stereochemistry at C-2 of the α,β-unsaturated carbonyl are discussed in Supplementary Information section 9.2.
A broad selection of amine partners was explored using 3-methyl-cyclohexenone 15 as substrate partner. Excellent conversion, chemo-, enantio- and diastereoselectivities were observed for small linear primary amines a–c, e–h. Notably, functionalized products from amines a, e, g, h could be formed efficiently, as could the secondary amine pyrrolidine i.
We were also keen to see if CR–RA products with additional stereocentres could be synthesized. (R)- or (S)-3-fluoropyrrolidine j could be coupled efficiently with 3-methylcyclohexenone 15, affording (cis)-15′j with high chemo- and diastereoselectivity (see Supplementary Information section 9 for details on the use of rac-j). Furthermore, CR–RA of cyclopropylamine b with racemic enone 23 demonstrated that the single catalyst could control three stereocentres on cyclohexylamine ring 23′b, offering excellent enantioselectivity as well as good chemo- and diastereoselectivity.
To assess the synthetic applicability of the EneIRED-catalysed CR–RA, preparative-scale syntheses were performed using 15 partnered with a, b or i, as well as 16 combined with b forming 15′a, 15’b, 15’i, and 16′b as the hydrochloride salts in 81%, 77%, 60% and 72% isolated yield respectively (Extended Data Fig. 3). The former example could be intensified to 50 mM enone 15 and 250 mM (5 eq.) amine b substrate loadings, affording 15′b in 64% isolated yield at a scale of 1.0 mmol (110 mg, total turnover number (TTN) = 640).
We next carried out mechanistic investigations to further characterize EneIRED and identify any intermediates formed during CR–RA. EneIRED effectively catalysed the reaction using reduced nicotinamide adenine dinucleotide phosphate (NADPH) (2 eq.) only. Furthermore, in the absence of EneIRED or the nicotinamide cosubstrate regeneration system, no reduction products could be detected (Extended Data Fig. 4a–c). Isotopic labelling experiments, using the in situ-generated deuterated nicotinamide cosubstrate from d-glucose-1-d1, yielded 1,3-d2-15′b from 15 and b as the hydrochloride salt confirming hydride transfer occurs at C-1 and C-3 of the unsaturated carbonyl substrate (Fig. 3a, 75% isolated yield, 91% two-dimensional incorporation; see Supplementary Figs. 4, 5 for further details).
a, Deuterium-labelling experiment. b, Amine donor-control experiments. c, Structure of EneIRED dimer in ribbon format, with monomers A and B in green and blue, respectively. d, Active site of EneIRED in complex with NADP+. Side chains of monomers A and B are shown in cylinder format, with carbon atoms in green and blue, respectively. The structure has been used to model the ene-imine ligand (the condensation product of 15 and b), shown with carbon atoms in yellow. The distance between these atoms is given in ångströms. e, Reactions of EneIRED point variants in reaction with 15 and b under standard conditions.
A time-course study is consistent with a stepwise CR–RA double-hydride transfer mechanism, in which, first, enone 15 undergoes CR to an intermediate enantioenriched ketone (R)-15′, before RA of the intermediate to the final product (1R,3R)-15′b (Extended Data Fig. 5a). Importantly, no direct RA product 15b, a potential alternative intermediate, was observed during the time course, suggesting that the reaction proceeds via ketone 15′ only. Furthermore, although 15b was inert to redox activity with EneIRED and co-substrates, ketone (R)-15′ undergoes RA with b (Extended Data Fig. 4d, e), indicating 15′ as the sole reaction intermediate.
We were also keen to probe the enzyme–substrate complex formed during the CR step. Omitting any amine donor from the reaction yielded no product of either CR or RA (Fig. 3b), suggesting that the IRED-catalysed reaction explicitly requires the presence of an amine in the catalytic cycle. Furthermore, no activity was observed when combining either enone 15 with tertiary amine donor triethylamine k or unsaturated ester methyl cyclohex-1-ene-1-carboxylate and cyclopropylamine b, suggesting that EneIRED-catalysed CR probably occurs via an ene-imine-NAD(P)H-biocatalyst complex, reminiscent of organocatalytic CR systems33 that have recently been hosted in an artificial enzyme34. To the best of our knowledge this is the first example of an enzyme that achieves CR by this type of ene-imine intermediate.
The multiple activities of EneIRED prompted us to study its structure using X-ray crystallography. Crystals of EneIRED in complex with NADP+ were obtained in the P21 space group with two molecules in the asymmetric unit, forming the now-familiar domain-swapping dimeric fold observed in the IRED family8,35 (Fig. 3c). A comparison of the monomer structure with others in the Protein DataBank using the DALI server36 revealed that the closest existing IRED structures in the database were those from Streptosporangium roseum (PDB code 5OCM, 30% seq id, r.m.s. deviation 1.6 Å over 286 Ca atoms)37, Aspergillus oryzae (5G6S, 30%, 1.6 Å)8 and Stackebrandtia nassauensis (6JIT, 30%, 2.0 Å). The most striking differences compared to other IREDs that have structures in the database were observed in the active site (Fig. 3f; see Extended Data Fig. 6 for electron density).
EneIRED possesses a tyrosine residue, Y177, at the top of the ceiling of the active site as drawn, in common with other IREDs, such as those from Streptomyces sp. GF3546 (4OQY)38, Bacillus cereus (4D3F)39, and Nocardopsis halophila (4D3S)38, which have been shown to display (S)-stereoselectivity for the reduction of model imine compound 2-methyl pyrroline. In common with those enzymes, Y177 forms a hydrogen bond with the hydroxyl group of a side chain, in this case threonine T101, which in turn H-bonds to the 2′-hydroxyl of the ribose in NADP+. However, EneIRED also possesses an additional tyrosine residue Y181 that also points into the active site towards the cosubstrate binding cleft, which is a hydrophobic leucine in both 5OCM and 6JIT. The active site also features a number of cyclic and hydrophobic amino acid side chains F185, Y269, H245 and A240 with Y129 at the rear and V244 at the front, which form a closed cavity that has been previously observed to be suitable for binding, especially of planar cyclic imines in IRED structures8,40.
We explored the role of Y177 and Y181 in CR–RA through the creation of point variants of EneIRED. Point mutation of Y181 (EneIRED-Y181A/L/F) gave mutants that exhibited lower stereoselectivity for CR–RA of 15 with b correspondent with the volume of the residue (Fig. 3e), suggesting that this position is important in providing a steric constraint for controlling the face for hydride delivery in both CR and RA steps. Both EneIRED-Y181A and the point mutation of Y177, EneIRED-Y177A, exhibited considerable reduction in the rate of CR–RA compared to the wild type, that is, a lower rate of the consumption of 15 and accumulation of 15′b (Extended Data Fig. 5b, c). Interestingly, for Y177A, only low levels of ketone (<5%) were observed during the course of the reaction, suggesting that this residue is important for CR and not RA.
A model of the enzyme active site in complex with the ene-imine formed by condensation of 15 with b was constructed using AutoDock Vina41 (Fig. 3f). In the top pose, the model suggests that the closest atom to the C-4 of the pyridinium ring of the cosubstrate, suitable for acceptance of a hydride, is the prochiral carbon atom of the C=C bond. Delivery of a hydride to this atom as shown in the model would give the experimentally observed (R)-configuration at this centre.
On the basis of our structural and mechanistic investigation, the following dual EneIRED catalytic cycle is proposed for productive CR–RA (Fig. 4a). First, the nicotinamide cosubstrate and condensation product of α,β-unsaturated carbonyl V and amine form an active-site ene-imine-NAD(P)H-EneIRED complex VI. Where substrate orientation kinetically favours the C-3 of the ene-imine orientated toward the nicotinamide hydride, CR yields the stereoenriched 1-enamine-NAD(P)+-EneIRED complex VII. Following this, the oxidized cosubstrate and prochiral 1-enamine are expelled from the enzyme, with the latter being hydrolysed in solution to form the stereoenriched carbonyl VIII. A further NAD(P)H cosubstrate binds to the enzyme together with the condensation product of the previously released carbonyl VIII and amine to form complex IX which undergoes the expected IRED-catalysed RA9, yielding the stereoenriched final product X.
a, Schematic of the proposed catalytic cycles for EneIRED-catalysed CR and RA, demonstrating the productive CR–RA reaction of α,β-unsaturated carbonyls with amines. b, Example of six-electron enzymatic CR–RA catalysed by EneIRED only, with by-product formation observed, see Supplementary Information for further details.
Finally, we sought to further extend the enzyme catalysed CR–RA by employing a conjugated dienyl-ketone. Whereas 24 was susceptible to aza-conjugate addition, EneIRED-catalysed four-electron and six-electron CR–RA of α,β,γ,δ-unsaturated enone 24 in combination with cyclopropylamine b, affording 24′b and (trans)-24″b=16′b respectively (Fig. 4b). Interestingly, six-electron CR–RA product 24″b=16′b possessed analogous diastereoselectivity and enantioselectivity to the CR–RA of the corresponding ethyl-substituted α,β-unsaturated enone 16, suggesting that reduction of 24 proceeds by a similar pathway to that of 16. This experiment suggests that EneIRED could be used to establish additional stereogenic centres during the CR–RA process.
In summary, we report the characterization of a multifunctional biocatalyst (EneIRED) that is able to catalyse CR as well as imine reduction and RA. EneIRED possesses broad substrate scope, enabling the stereoselective preparation of valuable amine diastereomers in a one-pot, one-catalyst reaction starting from simple prochiral starting materials. Mechanistic and structural studies reveal a multi-step process in which EneIRED first catalyses amine-activated CR of α,β-unsaturated carbonyl via an ene-imine-NAD(P)H-enzyme complex that is, to our knowledge, previously undescribed. We envisage that refinement of the biocatalyst though enzyme engineering will reduce the amine loading for expedient CR–RA as well as expand the substrate scope. This reaction further expands the repertoire of IREDs and emphasizes their importance in the synthesis of stereochemically defined chiral amines.
Methods
Cloning, expression and protein purification
The codon-optimized EneIRED gene sequence (TWIST Biosciences, US, GenBank accession number MW854365) was cloned into pET-28a-(+) (Supplementary Fig. 1) and used to transform chemically competent E. coli BL21 (DE3). Point variants of EneIRED were created by site-directed mutagenesis using appropriate primers, see Supplementary Information sections 1.5, 2.2. Cultivation was performed in 400 ml Terrific Broth (TB) media (Formedium, UK) supplemented with 35 μg ml−1 kanamycin in 2-l Erlenmeyer baffled flasks. Cultures were incubated at 37 °C and shaken at 200 rpm until an optical density (OD600nm) of 0.6–0.8, before gene expression was induced with isopropyl β-d-1-thiogalactopyranoside (IPTG) at a final concentration of 100 μM. Incubation was continued at 23 °C and 200 rpm for 25 h before harvesting the biomass using centrifugation. Disruption of the cells using sonication in an ice bath, then clarification by centrifugation and lyophilization afforded powdered EneIRED cell-free extract (CFE). To obtain purified enzyme, EneIRED CFE was resuspended, clarified by centrifugation and immobilized metal affinity chromatography (IMAC) was performed. Further purification of the protein for crystallography and control and mechanistic experiments was realized using gel filtration (GF) chromatography. For further details see Supplementary Information.
Biotransformations
Typical procedure for EneIRED-catalysed CR–RA: A 200 μl reaction mixture consisting of 10 mM α,β-unsaturated carbonyl, 200 mM amine, 0.2 mol% EneIRED, 0.1 mg ml−1 glucose dehydrogenase (GDH), 5.0 mol% NADP+, 30 mM glucose, 15% v/v DMSO, 100 mM glycine-NaOH buffer pH 9.0 were shaken at 900 rpm in 96-well microtiter plate at 30 °C for 18 h before being quenched with 5.0 M NaOH (aq., 20 μl) and extracted into heptane, dichloromethane or CDCl3 (2 × 200 μl). The organic fractions were combined and analysed by gas chromatography–mass spectrometry (GC–MS) before being derivatized and analysed by gas chromatography–flame-ionization detection (GC–FID) over a chiral stationary phase or 1H NMR. CR–RA reactions were undertaken in 2–5 replicates and could also be performed using EneIRED CFE (4 mg ml−1) affording comparable results.
Screening the panel of IREDs was undertaken at 100 μl total reaction volume consisting of 5 mM of ene-imine I, 12.5 mg ml−1 IRED, 0.1 mg ml−1 GDH, 1.0 mol% NADP+, 50 mM glucose in 100 mM KPi buffer pH 7.0 were shaken at 900 rpm in 384-well microtiter plate at 30 °C for 24 h before being quenched with 5.0 M NaOH (aq., 10 μl), extracted into heptane (2 × 100 μl) and analysed by GC–MS. The product distributions were mapped to the phylogenetic IRED tree (Extended Data Fig. 1) created using the Interactive Tree of Life (iTOL)42.
Time-course reactions were undertaken at 5 mM α,β-unsaturated carbonyl, 200 mM amine, 0.4 mol% EneIRED or EneIRED point variants, 0.1 mg ml−1 GDH, 10 mol% NADP+, 30 mM glucose, 15% v/v DMSO, 100 mM glycine-NaOH buffer pH 9.0 and were shaken at 250 rpm in a 2-ml Eppendorf at 30 °C. At each time point, 100 μl of the reaction was removed and basified with 5.0 M NaOH (aq., 10 μl) before extraction into heptane (2 × 200 μl) and analysis by GC–MS.
Preparative scale reactions were undertaken at 10 or 50 mM α,β-unsaturated carbonyl with 200–250 mM amine, 0.2 or 0.1 mol% EneIRED, 0.1 mg ml−1 GDH, 1.0 mol% NADP+, 30 or 150 mM glucose, 15% v/v DMSO, 100 mM glycine-NaOH buffer pH 9.0 at a volume of 30 ml or 50 ml and were shaken at 200 rpm in falcon tubes at 25 °C for 24 h before being quenched with 5.0 M NaOH (aq., 0.1 × reaction volume), clarified by centrifugation and extracted into MTBE (3 × reaction volume). The organic fractions were combined, dried with MgSO4 and concentrated under reduced pressure to remove excess amine substrate before acidification with 2.0 M HCl (dioxane, 2 eq.) and concentration in vacuo to yield the amine-HCl salt, which was analysed by NMR and GC–FID over a chiral stationary phase. For further details see Supplementary Information.
Protein crystallography data collection, structure solution and refinement
Initial screening of crystallization conditions was performed using commercially available INDEX (Hampton Research), PACT premier and CSSI/II (Molecular Dimensions) screens in 96-well sitting drop trays. Optimization was carried out in a 24-well hanging-drop format to obtain crystals for X-ray diffraction studies. For co-crystallization experiments, a 200 mM stock solution of cosubstrate NADP+ in wate was prepared.
Crystals of the wt-EneIRED-NADP+ complex were grown using wt-EneIRED concentrated to 35 mg ml−1 in 50 mM Tris buffer pH 7.5 containing 300 mM NaCl. The crystallization drop contained 0.15 μl protein: 0.15 μl mother liquor, comprising 100 mM Tris buffer pH 6.5, 25% (w/v) PEG (polyethylene glycol) 3350, and 5.0 mM NADP+. Crystals were harvested directly into liquid nitrogen with nylon CryoLoops (Hampton Research), using the mother liquor containing 10% (v/v) ethylene glycol as cryoprotectant.
The dataset described in this report was collected at the Diamond Light Source, Didcot, Oxfordshire, UK on beamline I03, at a temperature of 120 K and a wavelength of 0.97625 Å. Data were processed and integrated using XDS43 and scaled using SCALA44 included in the Xia2 processing system45. Data collection statistics are provided in (Extended Data Table 1). The crystal of EneIRED-NADP+ was obtained in space group P21, with two molecules in the asymmetric unit; the solvent content in the crystals was 42%. The structure of the EneIRED-NADP+ was solved by molecular replacement using MOLREP46 with the monomer of AtRedAm (PDB code 6EOD)47 as the model. The structure was built and refined using iterative cycles in Coot48 and REFMAC49, using local NCS restraints in the refinement cycles. Following building and refinement of the protein and water molecules in this complex, residual density was observed in the omit maps at the dimer interfaces, which could be clearly modelled as NADP+. The final structures exhibited percentage crystallographic R-factor, Rcryst, and free R-factor, Rfree, values of 19.0 and 23.0. Refinement statistics for the structures are presented in (Extended Data Table 1). The Ramachandran plot for EneIRED-NADP+ showed 96.5% of residues to be situated in the most favoured regions, 3.0% in additional allowed and 0.5% residues in outlier regions. The structure has been deposited in the Protein Databank (PDB) with accession number 7A3W
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary Information, and NMR traces are available from the Mendeley data repository (https://data.mendeley.com) at https://doi.org/10.17632/fhc429t33c.1. Sequence data have been deposited in Genbank (accession numbers MW854365, MW925135–MW925140) and the coordinate files and structure factors have been deposited in the PDB with accession number 7A3W.
References
Jarvis, L. M. The new drugs of 2019. Chem. Eng. News 98, 30–36 (2020).
Afanasyev, O. I., Kuchuk, E., Usanov, D. L. & Chusov, D. Reductive amination in the synthesis of pharmaceuticals. Chem. Rev. 119, 11857–11911 (2019).
Marshall, J. R. et al. Screening and characterization of a diverse panel of metagenomic imine reductases for biocatalytic reductive amination. Nat. Chem. 13, 140–148 (2021).
Roughley, S. D. & Jordan, A. M. The medicinal chemist’s toolbox: an analysis of reactions used in the pursuit of drug candidates. J. Med. Chem. 54, 3451–3479 (2011).
Yasukawa, T., Masuda, R. & Kobayashi, S. Development of heterogeneous catalyst systems for the continuous synthesis of chiral amines via asymmetric hydrogenation. Nat. Catal. 2, 1088–1092 (2019).
Wu, Z. et al. Secondary amines as coupling partners in direct catalytic asymmetric reductive amination. Chem. Sci. 10, 4509–4514 (2019).
Skrypai, V., Varjosaari, S. E., Azam, F., Gilbert, T. M. & Adler, M. J. Chiral Brønsted acid-catalyzed metal-free asymmetric direct reductive amination using 1-hydrosilatrane. J. Org. Chem. 84, 5021–5026 (2019).
Aleku, G. A. et al. A reductive aminase from Aspergillus oryzae. Nat. Chem. 9, 961–969 (2017).
Mayol, O. et al. A family of native amine dehydrogenases for the asymmetric reductive amination of ketones. Nat. Catal. 2, 324–333 (2019).
Yang, Y., Cho, I., Qi, X., Liu, P. & Arnold, F. H. An enzymatic platform for the asymmetric amination of primary, secondary and tertiary C(sp3)–H bonds. Nat. Chem. 11, 987–993 (2019).
Li, T. et al. Efficient, chemoenzymatic process for, anufacture of the Boceprevir bicyclic [3.1.0]proline intermediate based on amine oxidase-catalyzed desymmetrization.J. Am. Chem. Soc. 134, 6467–6472 (2012).
Zhou, J. & List, B. Organocatalytic asymmetric reaction cascade to substituted cyclohexylamines. J. Am. Chem. Soc. 129, 7498–7499 (2007).
Monti, D. et al. Cascade coupling of ene-reductases and ω-transaminases for the stereoselective synthesis of diastereomerically enriched amines. ChemCatChem 7, 3106–3109 (2015).
France, S. P., Hepworth, L. J., Turner, N. J. & Flitsch, S. L. Constructing biocatalytic cascades: in vitro and in vivo approaches to de novo multi-enzyme pathways. ACS Catal. 7, 710–724 (2017).
Huffman, M. A. et al. Design of an in vitro biocatalytic cascade for the manufacture of islatravir. Science 366, 1255–1259 (2019).
Toogood, H. S. & Scrutton, N. S. Discovery, Characterization, engineering, and applications of ene-reductases for industrial biocatalysis. ACS Catal. 8, 3532–3549 (2018).
Roth, S., Kilgore, M. B., Kutchan, T. M. & Müller, M. Exploiting the catalytic diversity of short-chain dehydrogenases/reductases: versatile enzymes from plants with extended imine substrate scope. ChemBioChem 19, 1849–1852 (2018).
Hyslop, J. F. et al. Biocatalytic synthesis of chiral N-functionalized amino acids. Angew. Chemie. Int. Ed. 57, 13821–13824 (2018).
Kato, Y., Yamada, H. & Asano, Y. Stereoselective synthesis of opine-type secondary amine carboxylic acids by a new enzyme opine dehydrogenase use of recombinant enzymes. J. Mol. Catal. B 1, 151–160 (1996).
Schober, M. et al. Chiral synthesis of LSD1 inhibitor GSK2879552 enabled by directed evolution of an imine reductase. Nat. Catal. 2, 909–915 (2019).
Bornadel, A. et al. Technical considerations for scale-up of imine-reductase-catalyzed reductive amination: a case study. Org. Process Res. Dev. 23, 1262–1268 (2019).
Hussain, S. et al. An (R)-imine reductase biocatalyst for the asymmetric reduction of cyclic imines. ChemCatChem 7, 579–583 (2015).
Yao, P., Xu, Z., Yu, S., Wu, Q. & Zhu, D. Imine reductase-catalyzed enantioselective reduction of bulky α,β-unsaturated imines en route to a pharmaceutically important morphinan skeleton. Adv. Synth. Catal. 361, 556–561 (2019).
Mitsukura, K. et al. Purification and characterization of a novel (R)-imine reductase from Streptomyces sp. GF3587. Biosci. Biotechnol. Biochem. 75, 1778–1782 (2011).
Lenz, M. et al. Asymmetric ketone reduction by imine reductases. ChemBioChem 18, 253–256 (2017).
Thorpe, T. W. et al. One-pot biocatalytic cascade reduction of cyclic enimines for the preparation of diastereomerically enriched N-heterocycles. J. Am. Chem. Soc. 141, 19208–19213 (2019).
Steiningerova, L. et al. Different reaction specificities of F420H2-dependent reductases facilitate pyrrolobenzodiazepines and lincomycin to fit their biological targets. J. Am. Chem. Soc. 142, 3440–3448 (2020).
Trenti, F. et al. Early and late steps of quinine biosynthesis. Org. Lett. 23, 1793–1797 (2021).
Mitsukura, K., Suzuki, M., Tada, K., Yoshida, T. & Nagasawa, T. Asymmetric synthesis of chiral cyclic amine from cyclic imine by bacterial whole-cell catalyst of enantioselective imine reductase. Org. Biomol. Chem. 8, 4533–4535 (2010).
France, S. P. et al. Identification of novel bacterial members of the imine reductase enzyme family that perform reductive amination. ChemCatChem 10, 510–514 (2018).
Mangas-Sanchez, J. et al. Asymmetric synthesis of primary amines catalyzed by thermotolerant fungal reductive aminases. Chem. Sci. 11, 5052–5057 (2020).
Montgomery, S. L. et al. Characterization of imine reductases in reductive amination for the exploration of structure–activity relationships. Sci. Adv. 6, 9320 (2020).
Ouellet, S. G., Walji, A. M. & Macmillan, D. W. C. Enantioselective organocatalytic transfer hydrogenation reactions using Hantzsch esters. Acc. Chem. Res. 40, 1327–1339 (2007).
Santi, N., Morrill, L. C., Świderek, K., Moliner, V. & Luk, L. Y. P. Transfer hydrogenations catalyzed by streptavidin-hosted secondary amine organocatalysts. Chem. Commun. 57, 1919–1922 (2021).
Rodríguez-Mata, M. et al. Structure and activity of NADPH-dependent reductase Q1EQE0 from Streptomyces kanamyceticus, which catalyses the R-selective reduction of an imine substrate. ChemBioChem 14, 1372–1379 (2013).
Holm, L. Benchmarking fold detection by DaliLite v.5. Bioinformatics 35, 5326–5327 (2019).
Lenz, M. et al. New imine-reducing enzymes from β-hydroxyacid dehydrogenases by single amino acid substitutions. Protein Eng. Des. Sel. 31, 109–120 (2018).
Huber, T. et al. Direct reductive amination of ketones: structure and activity of S-selective imine reductases from Streptomyces. ChemCatChem 6, 2248–2252 (2014).
Man, H. et al. Structure, activity and stereoselectivity of NADPH-dependent oxidoreductases catalysing the S-selective reduction of the imine substrate 2-methylpyrroline. ChemBioChem 16, 1052–1059 (2015).
Aleku, G. A. et al. Stereoselectivity and structural characterization of an imine reductase (IRED) from Amycolatopsis orientalis. ACS Catal. 6, 3880–3889 (2016).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2009).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Kabsch, W. XDS. Acta Crystallogr. Sect. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. Sect. D Biol. Crystallogr. 62, 72–82 (2006).
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–190 (2010).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997).
Sharma, M. et al. A mechanism for reductive amination catalyzed by fungal reductive aminases. ACS Catal. 8, 11534–11541 (2018).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
Acknowledgements
T.W.T. is grateful for a CASE award from the UK Biotechnology and Biological Sciences Research Council (BBSRC) and Pfizer (BB/M011208/1). J.R.M. acknowledges a CASE award from the Industrial Biotechnology Innovation Centre (IBioIC), BBSRC and Prozomix Ltd. A.C. was funded by grant BB/P005578/1 from the BBSRC. N.J.T. is grateful to the ERC for the award of an Advanced Grant (742987). We thank J. P. Turkenburg and S. Hart for assistance with X-ray data collection and the Diamond Light Source for access to beamline I03 under proposal number mx-9948.
Author information
Authors and Affiliations
Contributions
N.J.T., G.G., R.M.H., R.K. and D.S.B.D. devised and supervised the project. F.P. and R.E.R. managed the project. T.W.T. and A.A. performed mechanistic studies. T.W.T. and R.E.R. carried out substrate scope reactions. T.W.T. and V.H. carried out preparative scale reactions. T.W.T., R.E.R and V.H. synthesized substrates and standards. A.C., T.W.T. and G.G. performed crystallographic and docking studies. T.W.T. and R.S.H. undertook site-directed mutagenesis. J.R.M., S.J.C. and J.D.F. performed genetic identification, cloning and bioinformatics. T.W.T. and J.R.M. produced and purified the biocatalyst. N.J.T., G.G., R.M.H., R.K., D.S.B.D., S.J.C., F.P., J.D.F., A.C., R.E.R., V.H., J.R.M. and T.W.T. wrote the manuscript and generated the figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Dominic Campopiano, Sandy Schmidt and Thomas Ward for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Phylogenetic IRED tree mapped against the reaction profiles of IRED-catalysed reduction of ene-imine I.
Although the majority of the IREDs catalysed conventional imine reduction only, a small number were able to reduce both C=C and C=N bonds. Of these, EneIRED (pIR-120) possessed the highest propensity to forming the desired amine II.
Extended Data Fig. 2 Optimization of EneIRED-catalysed CR–RA reaction conditions.
Conversion to the products of CR and CR–RA were elevated in glycine-OH pH 9.0, at moderate DMSO cosolvent concentration and at higher equivalents of amine donor. Formation of the direct RA product was not observed under any conditions.
Extended Data Fig. 3 Scaled-up examples of EneIRED-catalysed CR–RA.
Several secondary and tertiary amines could be prepared including an example at elevated enone concentration and lower amine equivalents.
Extended Data Fig. 4 Control reactions and isolated reactions of potential CR–RA pathway intermediates in EneIRED-catalysed CR–RA.
a, EneIRED CR–RA of 15 and b with NADPH. b, No enzyme control reaction. c, No recycling system control reaction. d, Reactions of potential CR–RA intermediate 15b with NADP+ or NADPH. e, Reaction of potential CR–RA intermediate 15′ with b using EneIRED. f, No amine control reactions with EneIRED point variants.
Extended Data Fig. 5 Time-course studies of the CR–RA of 15 with b catalysed by wild-type EneIRED and point variants.
Both EneIRED-Y177A and EneIRED-Y181A exhibited a reduction in the rate of CR and CR–RA product formation compared to wild-type EneIRED, indicating that both residues are important for efficient catalysis. Notably, for EneIRED-Y177A the concentration of the ketone intermediate was comparatively low throughout the reaction, suggesting that Y177 is more important for CR than RA.
Extended Data Fig. 6 Active site of EneIRED highlighting electron density.
a, Side chains, with density corresponding to the refined 2Fo − Fc map (blue) at a level of 1σ. b, NADP+, with density corresponding to the Fo − Fc difference map (green) at a level of 3σ obtained from refinement in the absence of the ligand, with refined atoms included for clarity. Fo and Fc stand for the observed and calculated structure factor amplitudes, respectively.
Supplementary information
Supplementary Information
This file contains Supplementary Text, Supplementary Figures, Supplementary Discussion and Supplementary References
Rights and permissions
About this article
Cite this article
Thorpe, T.W., Marshall, J.R., Harawa, V. et al. Multifunctional biocatalyst for conjugate reduction and reductive amination. Nature 604, 86–91 (2022). https://doi.org/10.1038/s41586-022-04458-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04458-x
- Springer Nature Limited
This article is cited by
-
Chirality conferral enables the observation of hyper-Raman optical activity
Nature Photonics (2024)
-
Selective hydrogenation of nitro compounds to amines by coupled redox reactions over a heterogeneous biocatalyst
Nature Communications (2024)
-
Transaminase-catalysis to produce trans-4-substituted cyclohexane-1-amines including a key intermediate towards cariprazine
Communications Chemistry (2024)
-
Synthetic techniques for thermodynamically disfavoured substituted six-membered rings
Nature Reviews Chemistry (2024)
-
Facile access to C-N bonds via unexpected side reactions of Knoevenagel condensation and their application in high-efficiency organic solar cells
Science China Chemistry (2024)