Abstract
Bacterial phenazine metabolites belong to a group of nitrogen-containing heterocyclic compounds with antimicrobial activities. In this study, a rhizosphere Pseudomonas aeruginosa strain PA1201 was isolated and identified through 16S rDNA sequence analysis and fatty acid profiling. PA1201 inhibited the growth of various pathogenic microorganisms, including Rhizotonia solani, Magnaporthe grisea, Fusarium graminearum, Xanthomonas oryzae pv. oryzae, Xanthomonas oryzae pv. oryzicola, and Staphylococcus aureus. High Performance Liquid Chromatography showed that PA1201 produced high levels of phenazine-1-carboxylic acid (PCA), a registered green fungicide ‘Shenqinmycin’ with the fermentation titers of 81.7 mg/L in pigment producing medium (PPM) and 926.9 mg/L in SCG medium containing soybean meal, corn steep liquor and glucose. In addition, PA1201 produced another antifungal metabolite, phenazine-1-carboxaminde (PCN), a derivative of PCA, with the fermentation titers of 18.1 and 489.5 mg/L in PPM and SCG medium respectively. To the best of our knowledge, PA1201 is a rhizosphere originating P. aeruginosa strain that congenitally produces the highest levels of PCA and PCN among currently reported P. aeruginosa isolates, which endows it great biotechnological potential to be transformed to a biopesticide-producing engineering strain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phyto-pathogenic fungi have long been known to constitute a widespread threat to plant species. Plant diseases caused by fungi have caused significant reduced grain production in crops, and increased risks of biodiversity loss (Fisher et al. 2012). Sheath blight (ShB) disease caused by the necrotrophic fungal pathogen Rhizoctonia solani Kühn, has been one of the most economically detrimental diseases of rice all over the world. During favorable conditions, ShB disease causes up to 50 % reduction in rice yield with an annual loss of about 6 million tons of rice grains (Zheng et al. 2013). However, highly resistant rice cultivars to ShB have not been found in breeding, mainly due to the lack of identified resistant donors in cultivated varieties (Li et al. 1995; Pinson et al. 2005; Srinivasachary and Savary 2011). Resistance to ShB is controlled by polygenes and only quantitative resistance has been identified through extensive screening of thousands of rice cultivars from various rice growing regions (Bonman et al. 1992; Ciampi et al. 2008; Kunihiro et al. 2002; Zheng et al. 2013). As a result, it is also difficult to obtain ShB-resistant rice lines through modern transgenic technologies. Until now, ShB controlling strategies have mainly relied on fungicides all over the world. Validamycin A, an anti-fungal agricultural antibiotic produced by Streptomyces hygroscopicus 5008, is an important ShB controller, which has been widely used in rice-producing areas of Asia for several decades, occupying a market of 30–50 thousand ton per year with annual revenue of billions of Chinese Yuan in China currently (Bai et al. 2006; Zhou and Zhong 2014). However, after extensive use over the years, R. solani from several areas in China has developed some resistance to Validamycin A, leading to a continuous increase in field dosage, thus affecting the integrity of the environment. Therefore, developing alternative biological fungicides may substantially address this problem.
Fungicides originating from microbes are specifically effective on the target phytopathogens with additional benefits, for example they are synthesized biologically and are therefore inherently biodegradable (Yamaguchi 1996). Isolation of antagonistic microorganisms and discovery of microbial products have been one of the most efficient ways to develop biological pesticides. For instance, Bacillus subtilis NB12 and Streptomyces triostinicus NA1 isolated from rhizosphere showed strong inhibition to mycelial growth of R. solani (Cao et al. 2013). Pseudomonas fluorescens strain Pf-5 has been demonstrated to synthesize diverse secondary metabolites, including antibiotics that are toxic to soil-borne fungi and oomycetes infecting plant roots (Hassan et al. 2010). P. chlororaphis GP72 was isolated from the rhizosphere of green peppers in China with antagonistic activities towards Coletotrichum lagenarium, Pythium ultimum, Sclerotinia sclerotiorum, Fusarium oxysporum f. sp. cucumerinum, Carposina sasakii and R. solani (Liu et al. 2007). The fungicidal activities of GP72 were attributed to the production of phenazine-1-carboxylic acid (PCA) and 2-hydroxyphenazine (2-OH-PHZ), although at relatively low yields (Huang et al. 2011). Pseudomonas aeruginosa M18 isolated from the rhizosphere produces diverse secondary antagonistic metabolites, including phenazine-1-carboxylic acid (PCA), pyoluteorin (Plt), pyocyanine (PYO), hydrogen cyanide (HCN), L-2-amino-4-methoxy-trans-3-butenoic acid (AMB), and exhibits broad-spectrum antagonistic activity against phytopathogenic fungi (Ge et al. 2004; Hu et al. 2005; Huang et al. 2004; Wei et al. 2013; Wu et al. 2011).
Phenazines refer to a large class of nitrogen-containing redox-active colored aromatic secondary metabolites produced by many bacterial species. These compounds have great potential for use in agricultural and medicinal applications due to their wide-ranging bioactivities (Chincholkar and Thomashow 2013). In 2011, PCA, the antagonistic phenazine metabolite of P. aeruginosa M18 was registered as a new biologically-synthesized fungicide, designated as ‘Shenqinmycin’, by the Ministry of Agriculture of China. The production of PCA relies on the two PCA biosynthetic gene clusters, phzA1-G1 and phzA2-G2, in M18 (Wu et al. 2011; Supplementary Fig. S1A). In addition, M18 contains two PCA decoration genes, phzS and phzM, which are responsible for the conversion of PCA to PYO. The expression of phzM is modulated by temperature in M18, which is suppressed at 28 °C, but induced at 37 °C (Huang et al. 2009). Therefore, the PCA fermentation titer of M18 was about 4 fold higher at 28 °C than that at 37 °C (Huang et al. 2009). However, the third PCA modification gene, phzH, which is involved in the conversion of PCA to phenazine-1-carboxaminde (PCN) (Chin-A-Woeng et al. 2001), was found to be inactive in M18 due to a spontaneous stop-codon generating point mutation (C–T) at the 904th nucleotide (Supplementary Fig. S1B; Wu et al. 2011). As a result, M18 does not synthesize PCN. Recent studies have revealed that PCA production in M18 is negatively regulated by several regulatory elements, including the 5′-untranslated region (5′-UTR) of the phzA1-G1 cluster, which blocks its own expression, the sRNA chaperone Hfq, the Gac/Rsm signal transduction cascade, and a quorum-sensing control regulator, QscR (Ge et al. 2004; Li et al. 2011; Ren et al. 2014; Wang et al. 2012; Zhang et al. 2005). Based on these findings, the wild type M18 strain has been engineered to maximize the PCA fermentation titer using genetic tools. As a result, several engineering strains producing high levels of PCA have been obtained, including a gacA/qscR-inactivated engineering strain, M18GQ, harboring a recombinant plasmid, pME6032Phz, with a PCA fermentation titer of 6365 mg/L in SCG medium (Zhou et al. 2010). In addition, a triple-deleted mutant M18MSU1 was constructed by deleting the 5′-UTR of phzA1-G1, phzM and phzS in strain M18, with maximum PCA yield of 4771 mg/L in SCG medium (Du et al. 2013). However, the high yield of PCA by M18GQ/pME6032Phz requires the addition of antibiotic and isopropyl-β-d-1-thiogalactopyranoside to ensure massive PCA production (Zhou et al. 2010). Furthermore, the low fermentation temperature (28 °C) is prerequisite for both engineering strains to obtain a desirable fermentation titer (Du et al. 2013; Zhou et al. 2010), which leads to some adverse effects, including unstable fermentation titer and increased production cost. Therefore, the potential of M18 to be converted to an effective PCA production factory appears to be limited, and the discovery of new bacterial strains with stable high yield of PCA may greatly relieve this problem.
Here, we report a newly identified P. aeruginosa strain PA1201 isolated from a rice rhizosphere in a Chongqing suburb of China, displaying strong antagonistic activities towards a set of pathogenic fungi and bacteria. Furthermore, P. aeruginosa strain PA1201 produced antagonistic metabolites, PCA and its derivative PCN, in large quantities, but was deficient in the production of Plt, another antagonistic compound produced by some P. aeruginosa isolates. The identification of PA1201 provides new microbial resources for agricultural antibiotic production.
Materials and methods
Microorganisms, media, and culture conditions
Bacterial strains and fungal strains used in the present study are listed in Supplementary Table S1. Unless specifically stated, P. aeruginosa strains were grown at 28 °C in PPM medium. Fungal strains were grown on potato dextrose agar (PDA) used for antifungal activity analysis. Xanthomonas strains and Staphylococcus aureus strain were grown on nutrient agar (NA) for antibacterial activity analysis. The compositions of the media used are listed in Supplementary Table S2. For medium preparation, tryptone, peptone, beef extract, and yeast extract were purchased from Sangon Biotech (China). Where required, antibiotics were used at a concentration of 25 µg/mL for rifampicin and kanamycin, 20 µg/mL for gentamycin, and 100 µg/mL for ampicillin. Bacterial growth was determined by measuring optical density at a wavelength of 600 nm (OD600).
Isolation and screening of antagonistic Pseudomonas strains
Healthy rice samples with roots were collected from a rice field in a Chongqing suburb of China in October 2011. A suspension of rice rhizosphere soil was obtained by shaking 3 g of root plus lightly adhering soil in 50 mL of Pseudomonas Agar F (PAF) broth (Supplementary Table S2) and cultured at 28 °C with shaking (200 rpm) for 24 h. 1 mL of the cultured suspension was transferred to 50 mL of DF broth (Supplementary Table S2) and cultured at 28 °C with shaking (200 rpm) for another 24 h. Again, 1 mL of the enriched culture in DF broth was transferred to 50 mL of ADF broth and cultured at 28 °C with shaking (200 rpm) for 24–30 h, followed by spraying 200 µL of the mixture onto an ADF agar plate. The plates were incubated at 28 °C for 5 days and the isolated colonies with unique colony morphology were numbered and cryopreserved in the LB broth supplemented with 20 % glycerol.
The antagonist activity of isolated bacterial strains was tested against R. solani. Briefly, the PDA plate used for antagonist screening was divided equally into 4 sections using marker pens and each freshly cultured bacterial isolate was inoculated at the center of each section, allowed to grow at 28 °C for 2 days. An agar plug (8 mm diameter) taken from an actively growing R. solani culture was then placed at the center of the PDA agar plate. Plates inoculated with R. solani agar plugs alone were used as a control. The co-culture plates were incubated at 28 °C for another 3 days. Growth inhibition of R. solani by test bacterial strains was assessed thereafter.
Phenotypic identification and characterization of strain PA1201
Colony morphology of strain PA1201 was observed on the LB agar plate cultured for 24–36 h at 28 °C. Cell morphology of strain PA1201 was observed by 120 kV biology transmission electron microscope (TEM; Tecnai G2 spirit Biotwin; FEI™, USA).
Anti-fungal activity test
Pseudomonas aeruginosa PA1201 was tested for in vitro antagonism against phytopathogic fungi listed in Supplementary Table S1. The analysis was performed on PDA medium in triplicates. PA1201 was inoculated at the center of the PDA plate and cultured at 28 °C for 2 days followed by placing two agar plugs (8 mm diameter) taken from an actively growing fungal culture at the two opposite sides of the PDA plate. Plates inoculated with fungal agar plugs alone were used as controls. The co-culture plates were incubated at 28 °C until fungal mycelia completely covered the agar surface on the control plate, followed by measuring the diameter of the inhibition zone around the PA1201 culture. Triplicate plates were tested for each pathogenic fungal strain and the inhibition rate was calculated according to the following equation: inhibition rate (%) = 100 × diameter of the inhibition zone/distance between the two fungal agar plugs.
One mL aliquots of melted PDA medium with different pH values (ranging from 4.0 to 7.0) supplemented with purified phenazine antibiotic (final concentration at 0.25 mM) were added to the well of 24 well plates. After solidification, an agar plug (4 mm diameter) taken from an actively growing R. solani culture was placed at the center of each well. The PDA agar without phenazine antibiotic supplementation was used as a control. The test plates were incubated at 28 °C until fungal mycelia completely covered the agar surface in the control wells, after which the fungal colony diameter was measured. Triplicate wells were tested for each condition. Growth inhibition of R. solani by PCA and PCN at individual pH was calculated according to the following equation: Inhibition rate (%) = 100 × [1 − (Dt − D0)/(Dc − D0)]. In the equation, Dt = the diameter of fungal colony on the agar supplemented with phenazine antibiotic (mm), D0 = the diameter of the agar plug inoculated on the agar (4 mm), and Dc = the diameter of fungal colony on the agar in control wells (15 mm).
Anti-bacterial activity analysis
Overnight cultures of the pathogenic bacterial strains listed in Supplementary Table S1 were added into pre-heated NA medium containing 0.8 % agar (kept at 40 °C) at the inoculation ratio of 1–100, followed by vigorous shaking for the preparation of test plates. One freshly prepared colony of PA1201 was inoculated at the center of the test plate and incubated at 28 °C for 2 days. Plates without PA1201 inoculation were used as a control. Triplicate plates were tested for each pathogenic bacterial strain and the diameter of the inhibition zone around PA1201 colony was recorded as the indicator of antibacterial activity.
16S rDNA sequence and phylogenetic analysis
16S rDNA of strain PA1201 was amplified from genomic DNA by a polymerase chain reaction (PCR) using Q5® High-Fidelity DNA Polymerase (New England Biolabs®) and universal primers 16S-27F and 16S-1472R (Supplementary Table S3). The amplified PCR product was purified and used for direct sequencing by Sangon Biotech (Shanghai). Closely related homologs were identified by comparing 16S rDNA gene sequence of strain PA1201 with the non-redundant database of nucleotide sequences deposited at NCBI Web server (http://www.ncbi.nlm.nih.gov), through the Basic Local Alignment Tool (BLAST) program. Phylogenetic analysis was conducted using MEGA version 6.06.
Cloning the phenazine biosynthesis genes
Gene specific primers were designed based on the genomic information of strain PAO1, which is available at the NCBI Web server (http://www.ncbi.nlm.nih.gov). The designed primers are listed in Supplementary Table S3. Oligonucleotide primers were synthesized by Sangon Biotech (Shanghai). Q5® High-Fidelity DNA Polymerase (New England Biolabs®) was used for gene amplification by PCR, using PA1201 genomic DNA as template. The amplified PCR products were purified and cloned into the T-vector of pTA2 (TOYOBO) for sequencing by Sangon Biotech (Shanghai). Homology analysis was performed using the Clustal Omega web server (http://www.ebi.ac.uk/Tools/msa/clustalo/).
Analysis of fatty acid composition
The fatty acid methyl ester (FAME) profile of the bacterial strain PA1201 cultured on TSBA plate at 28 °C for 24–36 h was analyzed by gas chromatography (Model 7890C; Agilent) using the Microbial Identification software package (Oberg et al. 2012). Four different types of reagents were used for fatty acid analysis: Reagent I (45 g sodium hydroxide, 150 mL methanol, and 150 mL distilled water), Reagent II (325 mL certified 6.0 N hydrochloric acid and 275 mL methyl alcohol), Reagent III (200 mL hexane and 200 mL methyl tert-butyl ether), and Reagent IV (10.8 g sodium hydroxide dissolved in 900 mL distilled water). The sample was prepared according to the requirements of the Sherlock microbial identification system (MIDI). Bacterial isolates were analyzed using the MIDI Microbial Identification Software (Sherlock TSBA Library version 6.10; MIDI, Inc.). Strains with a similarity index (SIM) of 0.5 or higher and a separation of 0.100 between the first and second choice are considered good library comparisons (http://www.midi-inc.com/pdf/MIS_Similarity_Index.pdf).
Production, extraction, purification, and quantitative analysis of PCA and PCN
Strain PA1201 and M18 were grown in 50 mL of PPM medium or SCG medium (Supplementary Table S2) in an Erlenmeyer flask (250 mL) at 28 and 37 °C at 200 rpm in a rotary shaker. To extract PCA and PCN, 180 µL of fermentation culture was mixed with 20 µL of 6 M HCl, followed by extraction using 540 µL chloroform as previously described (Ge et al. 2004). A three microliter aliquot of extracted PCA sample was then taken for HPLC analysis (Agilent Technologies 1260 Infinity) under the following conditions: C18 reversed-phase column (5 μm, 4.6 × 12.5 mm) eluted with acetonitrile—5 mM ammonium acetate (60:40, v/v). PCA production was quantified using peak area (A) in HPLC elute according to the following formula: PCA (mg/L) = 0.01465A − 0.34099, which was derived from a dose–peak area plot using purified PCA with a correlation co-efficiency (R2) of 0.99986. PCN production was quantified using peak area (A) in HPLC elute according to the following formula: PCN (mg/L) = 0.01848A − 0.05292, which was derived from a dose–peak area plot using purified PCN with a correlation co-efficiency (R2) of 0.99999. The results represent the means ± SD of three independent experiments.
Liquid chromatography–mass spectrometry analysis of PCA and PCN
Five microliters of the collected samples were applied to an Ultra Performance Liquid Chromatographic system (UPLC, Agilent 1290 Infinity) on a Zorbax XDB C18 reverse phase column (4.6 × 150 mm, temperature-controlled at 30 °C), and eluted with methanol—2 mM ammonium acetate (50:50, v/v) at a flow rate of 0.2 mL/min in a diode array detector (Agilent G4212A). Data was acquired in the centroid mode using Agilent MassHunter Workstation Data Acquisition Software (revision B.04).
Quantitative HCN production assay
PA1201 and M18 were grown on 8.5 cm-diameter plates of Pseudomonas isolation agar (PIA; Supplementary Table S2) for 16–24 h at 37 °C in triplicate for the quantitative analysis of the production of HCN as previously described (Carterson et al. 2004). Cyanide production is expressed as the ratio between the total amount (micromoles) of HCN and the total amount (milligrams) of bacterial protein recovered from cells grown on a petri dish.
Quantitative analysis of the production of PYO and Plt
Quantification of PYO production by P. aeruginosa strains PA1201 and M18 in the PPM medium (Supplementary Table S2) at 28 and 37 °C was carried out at pH 7.0 in chloroform as previously described (Essar et al. 1990). Briefly, 5 mL culture suspension was extracted with 3 mL chloroform followed by re-extraction with 1 mL of 0.2 mol/L HCl to obtain a pink to deep red solution. The absorbance of this solution was measured at 520 nm. The PYO concentrations, expressed as milligrams per liter, were determined by multiplying the optical density at 520 nm by 17.072.
The production of Plt by strains PA1201 and M18 in King’s Medium B (KMB; Supplementary Table S2) at 28 °C were extracted with ethyl acetate and quantified with high performance liquid chromatography (HPLC) using a C18 reverse-phase column eluted with methanol–water (70:30) at a flow rate of 2.0 mL/min as previously described (Huang et al. 2004).
Fly strain maintenance and oral infection experiment
Flies (Drosophila melanogaster strain Canton-S) were kept on standard cornmeal media at 25 °C except for infection experiments. For oral infection experiments, 1 mL of bacterial pellets of overnight cultures were collected by centrifugation, followed by washing with 1× PBS buffer and finally re-suspended in 150 µL of LBS solution (Supplementary Table S2). Bacterial suspension was then applied to Whatman filter disc (2.4 cm in diameter) covering the surface of a standard food vial containing sucrose agar. 3–5 day old adult flies were starved for 5 h and then transferred to food vials containing contaminated filter discs and maintained at 25 °C for 12 days. Dead flies were removed daily. Survival rate was calculated as percentage of living flies at each given time point. Data of 3–6 independent experiments with a minimum of 10 flies each were analyzed.
Statistical analysis
P values were determined by an unpaired Student’s t test with two groups, assuming unequal variances using the Microsoft Office excel data analysis tool. P < 0.05 was considered statistically significant.
Results
Isolation of the antagonistic bacterial strain PA1201
Eighty-nine bacterial strains with distinct colony morphologies were isolated from the rice fields in a Chongqing suburb of China in October 2011, using 1-aminocyclopropane-1-carboxylic acid as the sole carbon source. The organisms were rated for their antagonistic activity against the fungal pathogen R. solani. Strain PA1201 showed the strongest antagonistic activity against R. solani among the 10 antagonistic isolates (Fig. 1). The antagonistic analysis was further extended to a group of phytopathogens, including pathogenic fungi C. lagenarium and Fusarium oxysporum f. sp. cucumerinum, as well as pathogenic bacteria Xanthomonas oryzae pv. oryzae and Xanthomonas oryzae pv. oryzicola. Our results showed that PA1201 effectively inhibited the growth of all the tested phytopathogens (Table 1). Moreover, PA1201 also inhibited the growth of a Gram-positive bacterial pathogen S. aureus (Table 1). Collectively, these results suggested that PA1201 had a broad spectrum of antifungal and antibacterial activity.
Antagonistic bacterium PA1201 belongs to P. aeruginosa
Morphological analysis showed that PA1201 was a rod-shaped mono-flagellated Gram-negative bacterium with a cell size of approximately 1.5–2.5 µm long and 0.5–1.0 µm wide (Fig. 2A). The microbial identification (MIDI) system was used to analyze the fatty acid composition of PA1201. Our results showed that the predominant fatty acids (>10 %) of strain PA1201 were C16:1 ω7c or C16:1 ω6c (12.71 %), C16:0 (22.97 %), and C18:1 ω7c (44.13 %; Supplementary Table S4). The overall composition of fatty acids had the highest similarity to P. aeruginosa GC subgroup A (SIM index of 0.787) with a separation of 0.285 between the first and second choice (Flavimonas oryzihabitans, SIM index of 0.502). Therefore, PA1201 was tentatively identified as P. aeruginosa.
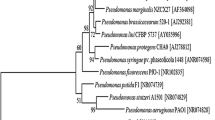
Strain PA1201 was identified as P. aeruginosa. A Transmission electron microscope of bacterial strain PA1201 grown on LB agar for 24 h. Bar 0.5 µm. B Phylogenetic tree of strain PA1201 and its related Pseudomonas species based on 16S rDNA sequences. The number at each branch point is the percentage supported by bootstrap. Bar 0.005 % sequence divergence
We also cloned and sequenced the 16S rDNA from PA1201. A blast search showed that the 16S rDNA sequence of PA1201 was 100 % identical to that of P. aeruginosa strains M18 and PAO1, and a phylogram was constructed by taking 16S rDNA sequences of related strains from the NCBI as a reference (Fig. 2B). Based on the results, the 16S rDNA sequence of this novel isolate PA1201 was submitted to the GenBank and assigned to P. aeruginosa PA1201 under accession number KM386632.
Sequence analysis of the phenazine biosynthesis genes in PA1201
Based on the genome sequence of P. aeruginosa strain PAO1, specific primers (Supplementary Table S3) were designed to clone the genes required for the biosynthesis of phenazine metabolites in strain PA1201. Subsequent sequencing revealed that PA1201 possesses two PCA biosynthesis gene clusters, phzA1-G1 and phzA2-G2, and three PCA modification genes (phzH encoding enzyme converting PCA to PCN, and phzM and phzS encoding enzymes converting PCA to PYO; Supplementary Fig. 1A). Moreover, the genomic organizations of these genes in the strain PA1201 showed significant similarity to those previously described in P. aeruginosa strains PAO1 and M18 (Wu et al. 2011). Alignment analysis showed that the products of these genes in strain PA1201 had >95 % amino acid identity with their counterparts in the type strain of P. aeruginosa PAO1 (data not shown). Importantly, the point mutation (C–T) at the 904th nucleotide in the phzH gene of M18 did not occur in PA1201 (Supplementary Fig. S1B); therefore, the PhzH protein of PA1201 consisted of 610 amino acids, 99.7 % of which were identical to the PhzH protein in PAO1 (Supplementary Fig. S2).
PCA and PCN are active antimicrobial metabolites produced by PA1201
The phenazine metabolite PCA is the main contributor to the antifungal activities of P. aerugnosa strain M18 (Huang et al. 2004; Wei et al. 2013; Wu et al. 2011), which led to the characterization of the phenazine metabolites produced by this newly isolated P. aeruginosa strain PA1201 culture grown at 28 °C in PPM medium. Two distinct peaks were detected in the crude extract of PA1201, with retention times of 1.9 min (compound a) and 2.8 min (compound b; Fig. 3A). These findings are in agreement with the retention time of the standard compounds, PCA (Fig. 3B) and PCN (Fig. 3C), respectively. The electrospray spectrum of compound a contained an ion at m/z 225.0660 and a less abundant ion at m/z 247.0478, corresponding to the protonated PCA [C13H8N2O2 + H]+ (calculated m/z 225.0569) and sodiated PCA [C13H8N2O2 + Na]+ (calculated m/z 247.0478), indicating that compound a was PCA (Fig. 3D). Additionally, the electrospray spectrum of compound b contained an ion at m/z 224.0833 and a less abundant ion at m/z 246.0652, corresponding to the protonated PCN [C13H9N3O + H]+ (calculated m/z 224.0818) and sodiated PCN [C13H9N3O +Na]+ (calculated m/z 246.0638), verifying that compound b was PCN (Fig. 3E).
PA1201 produces PCA and PCN in PPM medium. A HPLC analysis of strain PA1201 culture supernatant crude extract. B HPLC analysis and the chemical structure of PCA. C HPLC analysis and the chemical structure of PCN. D LC–MS analysis of the compound a shown in (A). E LC–MS analysis of the compound b in (A)
PA1201 produced large amounts of PCA and PCN
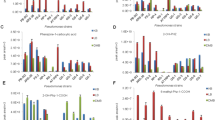
PCA production in strain M18 was tightly modulated by temperature, and fermentation at 28 °C facilitated PCA production by this strain (Huang et al. 2009). Therefore, the production profiles of phenazine metabolites by PA1201 and M18 during growth at 28 and 37 °C were analyzed and compared. In PPM medium, the cell density of PA1201 in PPM medium was slightly 28 °C (P = 0.858; Fig. 4A). However, PA1201 continuously produced higher amounts of PCA than M18 24 h post inoculation and after at 28 °C (P < 0.05), with the maximum PCA level of PA1201 reaching 81.7 mg/L at 36 h post inoculation (Fig. 4C). The PCN fermentation titer of PA1201 also increased during growth, reaching 18.1 mg/L at 48 h (Fig. 4E). As expected, reference strain M18 was unable to synthesize PCN because of the point mutation in its phzH gene (Fig. 4E; Supplementary Fig. S1). At 37 °C, PA1201 and M18 had similar growth curves in PPM medium (Fig. 4B), but the maximum fermentation titer of PCA by PA1201 was 39.6 mg/L. Even though this value was 50 % lower than that obtained at 28 °C, it was still 2.3 times greater than that produced by M18 (11.8 mg/L) (P < 0.05; Fig. 4D). Similarly, the fermentation titer of PCN by PA1201 also decreased to 1.2 mg/L in PPM medium (Fig. 4F).
Quantitative analysis of the production levels of PCA and PCN by strain PA1201 in PPM medium. Cell growth of P. aeruginosa PA1201 and M18 was monitored every 12 h at 28 (A) and 37 °C (B). PCA production curve of PA1201 and M18 at 28 (C) and 37 °C (D). PCN production curve of PA1201 and M18 at 28 (E ) and 37 °C (F). Square indicates PA1201; triangle indicates M18. The original PCA production strain M18 was used as a control. Data are presented as the means ± SD of three independent assays. The asterisks indicate P < 0.05 (T test) significant difference between data obtained for PA1201 and M18
In addition to PPM medium, SCG medium that was optimized for M18-derived engineering strains to produce PCA (Du et al. 2013) was chosen as the other fermentation medium to monitor PCA and PCN fermentation titers of PA1201. At 28 °C, the fermentation titers of PCA and PCN by PA1201 improved significantly to 926.9 mg/L and 489.5 mg/L respectively (Fig. 5A, C), while the yield of PCA by M18 was 248.9 mg/L, or 26.9 % of that by PA1201 under the same conditions (P < 0.05; Fig. 5A). However, at higher fermentation temperature (37 °C), PA1201 produced less PCA (104.8 mg/L). Nevertheless, its production was still higher than that of M18 (35.5 mg/L) in SCG medium (P < 0.05; Fig. 5B), and the maximum yield of PCN decreased to 41.9 mg/L for PA1201 in SCG medium (Fig. 5D).
Quantitative analysis of the production levels of PCA and PCN by strain PA1201 in SCG medium. PCA production curve of PA1201 and M18 at 28 (A) and 37 °C (B). PCN production curve of PA1201 and M18 at 28 (C) and 37 °C (D). Square indicates PA1201; triangle indicates M18. Data are presented as the means ± SD of three independent assays. *P < 0.05 (T test) significant difference between data obtained for PA1201 and M18
PA1201 failed to produce Plt
In addition to PCA, P. aeruginosa strain M18 also produces approximately 15 mg/L of Plt in KMB medium (Huang et al. 2004). Plt is a secondary metabolite with a resorcinol ring linked to a bichlorinated pyrrole moiety, which efficiently inhibits soilborne phytopathogenic fungi (Huang et al. 2004). Therefore, the Plt production profile of PA1201 was analyzed in this study. The growth curves of PA1201 and M18 resembled each other; however, PA1201 failed to produce any detectable level of Plt during growth at 28 °C in KMB medium. Conversely, reference strain M18 gradually increased the production of Plt, with the maximum value occurring after culture for 60 h in KMB medium (Fig. 6B). These results suggested that these two rhizosphere isolated P. aeruginosa strains, PA1201 and M18, produced specific antagonistic metabolites to combat pathogenic microorganisms.
PA1201 failed to produce Plt during growth in KMB medium at 28 °C. A Cell growth of P. aeruginosa PA1201 and M18. B Plt production curve of PA1201 and M18. Square indicates PA1201; triangle indicates M18.The original Plt production strain M18 was used as a control. Data are presented as the means ± SD of three independent assays
PA1201 was toxic to fruit flies
PA1201 was identified as an isolate of P. aeruginosa (Fig. 2; Supplementary Table S4), and most clinical isolates of P. aeruginosa are opportunistic human pathogens; therefore, we evaluated the toxicity of this bacterial isolate. The levels of two well-known virulence factors produced by many P. aeruginosa isolates, hydrogen cyanide (HCN) and pyocyanine (PYO) were measured in this study. Our results showed that PA1201 produced 30 % less HCN than M18 at 37 °C (P < 0.05) (Fig. 7A), but that it produced 50 % more PYO than M18 at 28 °C and 37 % more at 37 °C in PPM medium (P < 0.05; Fig. 7B), indicating the potential toxicity of the two rhizosphere P. aeruginosa strains. As a result, a Drosophila melanogaster oral infection model was adopted to analyze the toxicity of strain PA1201. Flies fed with PA1201 or M18 showed 100 % mortality after an 8-day feeding period or 7-day feeding period, respectively, while 90 % of the flies fed with DH5α survived after feeding for 10 days (Fig. 7C). Collectively, these results demonstrated that P. aeruginosa strains PA1201 and M18 had certain levels of toxicity towards higher organisms, even though they were originally isolated from rhizospheres.
Analysis on the production of virulence factors and toxicity of PA1201. A HCN production of strain PA1201 and M18. B PYO production of strain PA1201 and M18. C Survival rate of Drosophila melanogaster after bacterial feeding with PA1201, M18, or the control strain Escherichia coli DH5α. Data are presented as the means ± SD of three independent assays. *P < 0.05 (T test) significant difference between data obtained for PA1201 and M18
Discussion
In our study, bacterial strain PA1201 was isolated from the rice rhizosphere in central China. This organism showed antagonistic activities towards various pathogenic microorganisms, including R. solani, Magnaporthe grisea, Fusarium graminearum, Xanthomonas oryzae pv. oryzae, Xanthomonas oryzae pv. oryzicola and S. aureus. Strain PA1201 was identified as P. aeruginosa based on 16S rDNA sequence analysis and fatty acid profiling. To date, numerous florescence Pseudomonas isolates collected worldwide have been reported to have biocontrol activities, such as the PCN-producing P. chlororaphis PCL1391 (Chin-A-Woeng et al. 2000), and the 2-hydroxyphenazine- and PCA-producing P. chlororaphis GP72 (Liu et al. 2007), as well as the 1-hydroxyphenazine-producing P. aeruginosa SD12 (Dharni et al. 2012) and the PCA-producing P. aeruginosa M18 (Hu et al. 2005). Most studies of these florescence pseudomonads have focused on their biocontrol effects on crop diseases in laboratories and identification of the active antagonistic metabolites without quantification. In the present study, the fermentation titers of antimicrobial phenazines (PCA and PCN) produced by P. aeruginosa stains M18 and PA1201 were systematically compared. The following lines of evidence supported that PA1201 had better biotechnological potential than M18. First, the fermentation titer of PCA by PA1201 was significantly higher than that by M18 in both PPM and SCG medium at either 28 or 37 °C (Figs. 4C, 5A). Second, in addition to PCA, PA1201 produced large amounts of another bioactive phenazine metabolite, PCN (489.5 mg/L in SCG medium; Fig. 5C). To the best of our knowledge, PA1201 had the highest fermentation titers of PCA and PCN among the currently identified phenazine-producing wild type Pseudomonas strains.
PCA and PCN are the secondary metabolites of many plant root-colonizing bacteria, and both display antagonistic activities towards phytopathogens (Chincholkar and Thomashow 2013). Their antifungal activity towards Fusarium oxysporum f. sp. radicis-lycopersici under different pH values was compared by Chin-A-Woeng et al. (1998) in an in vitro assay. They found that the antifungal activity of PCA decreased drastically at pH 5.7 and above, while PCN remained active under the same conditions. To further verify that the pH-mediated difference in anti-fungal activity of PCA and PCN was not limited to Fusarium oxysporum f. sp. radicis-lycopersici, we extended the test to R. solani using purified PCA and PCN at pH values of 4.0–7.0. Our results confirmed that the anti-fungal activity of PCA was highly influenced by environmental pH. At pH 5.0 and below, PCA showed moderately stronger anti-Rhizotonia activity than PCN (P < 0.05), although both compounds had an inhibition rate higher than 80 % (Supplementary Fig. S3). At pH 5.5, the anti-Rhizotonia activity of PCA, but not PCN, was significantly reduced. At pH 6.0 and above, the anti-Rhizotonia activity of PCA was completed abolished, while activity of PCN persisted, with an inhibition rate of approximately 70 % (Supplementary Fig. S3). The extended potency of PCN is likely attributed to the presence of a nearly neutral carboxamide group in the place of the carboxyl group on the PCA molecule (Fig. 3B, C). Therefore, the dual-phenazine-producer PA1201 is considered a promising candidate for engineering and subsequent commercial production of the green fungicides, PCA and PCN, with the aid of chromosomal recombination techniques for in-frame marker-less gene deletion and promoter replacement.
Pseudomonads are a group of highly versatile Gram-negative bacteria, widely distributed all over the world that thrive in different ecological niches, including soil or marine habitats (Höfte and Altier 2010). The diversity of their metabolic and catabolic activities endows the genus Pseudomonas with great potential for use in plant protection and environmental control (Lo Sciuto et al. 2014). However, P. aeruginosa has long been recognized as a major nosocomial agent associated with a high level of morbidity and mortality for immune-compromised individuals and cystic fibrosis patients. Currently, clinical P. aeruginosa isolates are considered as one of the most important sources of ‘super bugs’ in clinical practice because of their persistent resistance to multiple antibiotics (Lo Sciuto et al. 2014). Although the P. aeruginosa strains PA1201 and M18 were isolated from the rhizosphere, they still produced virulence factors HCN and PYO, and exerted some toxicity toward fruit flies (Fig. 7). As a result, rhizosphere P. aeruginosa PA1201 needs to be engineered to attenuate its toxicitybefore it can be further developed as a biopesticide-producing strain. In P. aeruginosa strain M18, deletion of PCA modification genes phzS and phzM abolished PYO production, and significantly enhanced PCA yield (Du et al. 2013). In addition, the synthesis of HCN from glycine and the synthesis of phenazine metabolites were found to be metabolically linked by the PA2449 gene in P. aeruginosa strain PAO1 (Lundgren et al. 2013; Pessi and Haas 2000). Therefore, deletion of the genes required for the biosynthesis of PYO and HCN in PA1201 is likely to play a dual role by facilitating PCA and PCN production and attenuating toxicity.
References
Bai L, Li L, Xu H, Minagawa K, Yu Y, Zhang Y, Zhou X, Floss HG, Mahmud T, Deng Z (2006) Functional analysis of the validamycin biosynthetic gene cluster and engineered production of validoxylamine A. Chem Biol 13:387–397
Bonman JM, Khush GS, Nelson RJ (1992) Breeding for resistance to pests. Annu Rev Phytopathol 30:507–528
Cao QQ, Zhou DB, Zheng L, Yang M, Zhou EX (2013) Screening, identification and cultivation conditions of microbes antagonistic to rice sheath blight fungus Rhizoctonia solani. Chin J Biol Control 29(2):270–276
Carterson AJ, Morici LA, Jackson DW, Frisk A, Lizewski SE, Jupiter R, Simpson K, Kunz DA, Davis SH, Schurr JR, Hassett DJ, Schurr MJ (2004) The transcriptional regulator AlgR controls cyanide production in Pseudomonas aeruginosa. J Bacteriol 186(20):6837–6844
Chin-A-Woeng TF, Bloemberg GV, van der Bij AJ, van der Drift KMGM, Schripsema J, Kroon B, Scheffer RJ, Keel C, Bakker PAHM, Tichy HV, de Bruijn FJ, Thomas-Oates JE, Lugtenberg BJ (1998) Biocontrol by phenazine-1-carboxamide-producing Pseudomonas chlororaphis PCL1391 of tomato root rot caused by Fusarium oxysporum f. sp. radicis-lycopersici. Mol Plant Microbe Interact 11(11):1069–1077
Chin-A-Woeng TF, Bloemberg GV, Mulders IH, Dekkers LC, Lugtenberg BJ (2000) Root colonization by phenazine-1-carboxamide-producing bacterium Pseudomonas chlororaphis PCL1391 is essential for biocontrol of tomato foot and root rot. Mol Plant Microbe Interact 13(12):1340–1345
Chin-A-Woeng TF, Thomas-Oates JE, Lugtenberg BJ, Bloemberg GV (2001) Introduction of the phzH gene of Pseudomonas chlororaphis PCL1391 extends the range of biocontrol ability of phenazine-1-carboxylic acid-producing Pseudomonas spp. strains. Mol Plant Microbe Interact 14(8):1006–1015
Chincholkar SB, Thomashow L (eds) (2013) Microbial phenazines: biosynthesis, agriculture and health. Springer, Berlin
Ciampi MB, Meyer MC, Costa MJN, Zala M, McDonald BA, Ceresini PC (2008) Genetic structure of populations of Rhizoctonia solani anastomosis group-1 IA from soybean in Brazil. Phytopathology 98(8):932–941
Dharni S, Alam M, Kalani K, Abdul-Khaliq Samad A, Srivastava SK, Patra DD (2012) Production, purification, and characterization of antifungal metabolite from Pseudomonas aeruginosa SD12, a new strain obtained from tannery waste polluted soil. J Microbiol Biotechnol 22(5):674–683
Du X, Li Y, Zhou W, Zhou Q, Liu H, Xu Y (2013) Phenazine-1-carboxylic acid production in a chromosomally non-scar triple-deleted mutant Pseudomonas aeruginosa using statistical experimental designs to optimize yield. Appl Microbiol Biotechnol 97(17):7767–7778
Essar DW, Eberly L, Hadero A, Crawford IP (1990) Identification and characterization of genes for a second anthranilate synthase in Pseudomonas aeruginosa: interchangeability of the two anthranilate synthases and evolutionary implications. J Bacteriol 172:884–900
Fisher MC, Henk DA, Briggs CJ, Brownstein JS, Madoff LC, McCraw SL, Gurr SJ (2012) Emerging fungal threats to animal, plant and ecosystem health. Nature 484(7393):186–194
Ge Y, Huang X, Wang S, Zhang X, Xu Y (2004) Phenazine-1-carboxylic acid is negatively regulated and pyoluteorin positively regulated by gacA in Pseudomonas sp. M18. FEMS Microbiol Lett 237:41–47
Hassan KA, Johnson A, Shaffer BT, Ren Q, Kidarsa TA, Elbourne LD, Hartney S, Duboy R, Goebel NC, Zabriskie TM, Paulsen IT, Loper JE (2010) Inactivation of the GacA response regulator in Pseudomonas fluorescens Pf-5 has far-reaching transcriptomic consequences. Environ Microbiol 12(4):899–915
Höfte M, Altier N (2010) Fluorescent pseudomonads as biocontrol agents for sustainable agricultural systems. Res Microbiol 161(6):464–471
Hu HB, Xu YQ, Chen F, Zhang XH, Hur BK (2005) Isolation and characterization of a new fluorescent Pseudomonas strain that produces both phenazine 1-carboxylic acid and pyoluteorin. J Microbiol Biotechnol 15:86–90
Huang X, Zhu D, Ge Y, Hu H, Zhang X, Xu Y (2004) Identification and characterization of pltZ, a gene involved in the repression of pyoluteorin biosynthesis in Pseudomonas sp. M18. FEMS Microbiol Lett 232(2):197–202
Huang J, Xu Y, Zhang H, Li Y, Huang X, Ren B, Zhang X (2009) Temperature-dependent expression of phzM and its regulatory genes lasI and ptsP in rhizosphere isolate Pseudomonas sp. strain M18. Appl Environ Microbiol 75(20):6568–6580
Huang L, Chen MM, Wang W, Hu HB, Peng HS, Xu YQ, Zhang XH (2011) Enhanced production of 2-hydroxyphenazine in Pseudomonas chlororaphis GP72. Appl Microbiol Biotechnol 89(1):169–177
Kunihiro Y, Qian Q, Sato H, Teng S, Zeng DL, Fujimoto K, Zhu LH (2002) QTL analysis of sheath blight resistance in rice (Oryza Sativa L.). Yi Chuan Xue Bao 29:50–55
Li ZK, Pinson SRM, Marchetti MA, Stansel JW, Park WD (1995) Characterization of quantitative trait loci (QTLs) in cultivated rice contributing to field-resistance to sheath blight (Rhizoctonia solani). Theor Appl Genet 91:382–388
Li Y, Du X, Lu ZJ, Wu D, Zhao Y, Ren B, Huang J, Huang X, Xu Y, Xu Y (2011) Regulatory feedback loop of two phz gene clusters through 5′-untranslated regions in Pseudomonas sp. M18. PLoS ONE 6(4):e19413
Liu H, He Y, Jiang H, Peng H, Huang X, Zhang X, Thomashow LS, Xu Y (2007) Characterization of a phenazine-producing strain Pseudomonas chlororaphis GP72 with broad-spectrum antifungal activity from green pepper rhizosphere. Curr Microbiol 54(4):302–306
Lo Sciuto A, Fernández-Piñar R, Bertuccini L, Iosi F, Superti F, Imperi F (2014) The periplasmic protein TolB as a potential drug target in Pseudomonas aeruginosa. PLoS ONE 9(8):e103784
Lundgren BR, Thornton W, Dornan MH, Villegas-Peñaranda LR, Boddy CN, Nomura CT (2013) Gene PA2449 is essential for glycine metabolism and pyocyanin biosynthesis in Pseudomonas aeruginosa PAO1. J Bacteriol 195(9):2087–2100
Oberg TS, Ward RE, Steele JL, Broadbent JR (2012) Identification of plasmalogens in the cytoplasmic membrane of Bifidobacterium animalis subsp. lactis. Appl Environ Microbiol 78:880–884
Pessi G, Haas D (2000) Transcriptional control of the hydrogen cyanide biosynthetic genes hcnABC by the anaerobic regulator ANR and the quorum-sensing regulators LasR and RhlR in Pseudomonas aeruginosa. J Bacteriol 182(24):6940–6949
Pinson SRM, Capdevielle FM, Oard JH (2005) Confirming QTLs and finding additional loci conditioning sheath blight resistance in rice using recombinant inbred lines. Crop Sci 45:503–510
Ren B, Shen H, Lu ZJ, Liu H, Xu Y (2014) The phzA2-G2 transcript exhibits direct RsmA-mediated activation in Pseudomonas aeruginosa M18. PLoS ONE 9(2):e89653
Srinivasachary Willocquet L, Savary S (2011) Resistance to rice sheath blight (Rhizoctonia solani Kühn) [(teleomorph: Thanatephorus cucumeris (A.B. Frank) Donk.] disease: current status and perspectives. Euphytica 178:1–22
Wang G, Huang X, Li S, Huang J, Wei X, Li Y, Xu Y (2012) The RNA chaperone Hfq regulates antibiotic biosynthesis in the rhizobacterium Pseudomonas aeruginosa M18. J Bacteriol 194(10):2443–2457
Wei X, Huang X, Tang L, Wu D, Xu Y (2013) Global control of GacA in secondary metabolism, primary metabolism, secretion systems, and motility in the rhizobacterium Pseudomonas aeruginosa M18. J Bacteriol 195(15):3387–3400
Wu DQ, Ye J, Ou HY, Wei X, Huang X, He YW, Xu Y (2011) Genomic analysis and temperature-dependent transcriptome profiles of the rhizosphere originating strain Pseudomonas aeruginosa M18. BMC Genomics 12:438
Yamaguchi I (1996) Pesticides of microbial origin and applications of molecular biology. In: Copping LG (ed) Crop protection agents from nature: natural products and analogues. The Royal Society of Chemistry, London, pp 27–49
Zhang X, Wang S, Geng H, Ge Y, Huang X, Hu H, Xu Y (2005) Differential regulation of rsmA gene on biosynthesis of pyoluteorin and phenazine-1-carboxylic acid in Pseudomonas sp. M18. World J Microbiol Biotechnol 21:883–889
Zheng A, Lin R, Zhang D, Qin P, Xu L, Ai P, Ding L, Wang Y, Chen Y, Liu Y, Sun Z, Feng H, Liang X, Fu R, Tang C, Li Q, Zhang J, Xie Z, Deng Q, Li S, Wang S, Zhu J, Wang L, Liu H, Li P (2013) The evolution and pathogenic mechanisms of the rice sheath blight pathogen. Nat Commun 4:1424
Zhou TC, Zhong JJ (2014) Production of validamycin A from hemicellulose hydrolysate by Streptomyces hygroscopicus 5008. Bioresour Technol 175C:160–166
Zhou Q, Su J, Jiang H, Huang X, Xu Y (2010) Optimization of phenazine-1-carboxyli`c acid production by a gacA/qscR-inactivated Pseudomonas sp. M18GQ harboring pME6032Phz using response surface methodology. Appl Microbiol Biotechnol 86(6):1761–1773
Acknowledgments
This work was financially supported by National High Technology Research and Development Plan (“863” Program) of China (2012AA022107) granted to Y.-W. H.; and National “Twelfth Five-Year” Plan for Science & Technology Support (2012BAD19B0106) granted to Y.-W. H. The authors thank Prof. Mingfa Li for providing Drosophila melanogaster.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhou, L., Jiang, HX., Sun, S. et al. Biotechnological potential of a rhizosphere Pseudomonas aeruginosa strain producing phenazine-1-carboxylic acid and phenazine-1-carboxamide. World J Microbiol Biotechnol 32, 50 (2016). https://doi.org/10.1007/s11274-015-1987-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-015-1987-y