Abstract
In this study, the presence, prevalence, and genotypes of Anaplasma phagocytophilum, A. ovis, and A. capra in sheep were investigated based on 16 S SSU rRNA, groEL, and gtlA gene-specific polymerase chain reaction (PCR), respectively. The sequences of the genes were used for detection of the phylogenetic position of the species. Additionally, a restriction fragment length polymorphism (RFLP) were carried out for discrimination of A. phagocytophilum and related variants (A. phagocytophilum-like 1 and 2). The prevalence of Anaplasma spp. was found as 25.8% (101/391), while it was found that A. ovis, A. phagocytophilum-like 1, and A. capra are circulating in the sheep herds in Kyrgyzstan, according to the PCRs, RFLP and the partial DNA sequencing results. The positivity rates of A. phagocytophilum-like 1, A. ovis, and A. capra genotype-1 were 6.9, 22.5, and 5.3%, respectively. A total of 32 (8.2%) sheep were found to be mix infected. Moreover, phylogenetic analyses and sequence comparison with those available in the GenBank showed that A. capra formed two distinct genetic groups (A. capra genotype-1 and A. capra genotype-2). Considering the zoonotic potential of these species, it may be necessary to make changes in the interpretation of anaplasmosis cases in animals and there is a need for further studies to determine the pathogenicity of the species/genotypes circulating in animals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Anaplasmosis is one of the emerging-tick borne diseases, and the disease affects both human and animal health. The genus Anaplasma (order Rickettsiales, family Anaplasmataceae) includes the species of A. marginale, A. centrale, A. bovis, A. platys, A. ovis, A. capra and A. phagocytophilum, the last three of which cause infection in sheep (Friedhoff 1997; Dumler et al. 2001; Liu et al. 2012).
Anaplasma capra is a tick-borne pathogen discovered for the first time in China in 2012 (Liu et al. 2012). In Northern China, Anaplasma organisms identified from asymptomatic goats considered to be pathogenic in humans and were provisionally named as Anaplasma capra in 2015 based on the molecular and phylogenetic data (Li et al. 2015; Liu et al. 2012). The clinical manifestation of the species has not been clarified, however, fever, headache, weakness, dizziness, myalgia, chills, rash, eschar, lymphadenopathy, gastrointestinal symptoms, and neck stiffness were observed in humans (Li et al. 2015). After the first detection of A. capra in goats in China, its presence has been detected in goats in seven other countries, such as France, Iran, South Korea, Kyrgyzstan, Malaysia, Spain, and Türkiye (Koh et al. 2018; Jouglin et al. 2019; Wei et al. 2020; Miranda et al. 2021; Staji et al. 2021; Altay et al. 2022a, b; Remesar et al. 2022). The novel species has been detected in humans, sheep, cattle, dog, wild animals (e.g. Korean water deer (Hydropotes inermis argyropus), forest musk deer (Moschus berezovskii), takin (Budorcas taxicolor), Persian onegar (Equus hemionus onager), Reeves’s muntjacs (Muntiacus reevesi), serows (Capricornis crispus), and ixodid tick species such as Ixodes persulcatus, Dermacentor everestianus, Haemaphysalis longicornis, H. qinghaiensis, and Rhipicephalus microplus (Li et al. 2015; Fang et al. 2015; Yang et al. 2016; Qin et al. 2018; Guo et al. 2018, 2019; Amer et al. 2019; Han et al. 2019; Lu et al. 2022). Although the existing literature may interpret A. capra as a global pathogen, researches that will contribute to the understanding of its epidemiology and genetic diversity are still required, as it is a newly defined species.
Anaplasma phagocytophilum causes human granulocytic anaplasmosis, canine granulocytic anaplasmosis, equine granulocytic anaplasmosis, and tick-borne fever, in humans, dogs, horses, and ruminants, respectively (Karshima et al. 2022). As a result of recent phylogenetic analyses based on sequences of different genes such as 16 S SSU rRNA, gltA, and groEL, two A. phagocytophilum-related variants have been identified in cattle, Cervus nippon, and ixodid ticks from Japan, and in Hyalomma asiaticum and small ruminants from China. These variants were described as A. phagocytophilum-like 1 and A. phagocytophilum-like 2, respectively (Ohashi et al. 2005; Kawahara et al. 2006; Jilintai et al. 2009; Yoshimoto et al. 2010; Kang et al. 2014; Yang et al. 2015; Ben Said et al. 2015, 2017).
Anaplasma ovis is the most prevalent Anaplasma species of sheep in the world, which also infects goats and wild ruminants (Friedhoff 1997; Dumler et al. 2001). Anaplasma ovis is transmitted by Rhipicephalus bursa and other ticks in the Old World, while Dermacentor species are vectors of A. ovis in the western United States (Friedhoff 1997). Although there is some evidence suggesting that A. ovis may cause zoonotic infections like A. phagocytophilum, these are very limited and need to be clarified. To date, A. ovis DNA has only been detected in a symptomatic human patient in Cyprus (Chochlakis et al. 2010) and an asymptomatic person in Iran (Hosseini-Vasoukolaei et al. 2014).
In this study, the presence, prevalence, and genotypes of A. phagocytophilum, A. ovis, and A. capra were investigated in sheep from Kyrgyzstan based on 16 S SSU rRNA, groEL, and gtlA gene-specific polymerase chain reaction (PCR), restriction fragment length polymorphism (RFLP) and sequencing.
Materials and methods
Collection of blood samples and DNA extraction
This study was conducted in five regions (Chuy, Talas, Jalal-Abad, Naryn, Issyk-Kul) of Kyrgyzstan (Fig. 1). Blood samples from sheep were collected between June, 2017 and September, 2018 from 34 sheep flocks. A total of 391 blood samples were taken into collection tubes with EDTA from randomly selected 22 different sheep flocks. Between 9 and 20 blood samples were collected from each flock. The animals were clinically healthy and at least 8 months age sheep and stored at -20 °C, until DNA isolation.
Total genomic DNA was extracted from EDTA-treated blood samples using commercial extraction kit (PureLink Genomic DNA kit, Cat. No.: K1820-02, Invitrogen, Carlsbad, USA), according to the manufacturer’s instructions. During the DNA extraction, positive (A. capra positive sheep blood sample, Accession number: OK267268, Altay et al. 2022b) and negative (DNase-RNase-free sterile water, Cat No.: 129,114, Qiagen®, Germany) samples were used in order to avoid false positive or negative results. Extracted total DNA samples were diluted with 200 µl DNA elution buffer and stored at -20 °C until use.
Polymerase chain reaction (PCR)
In order to investigate the presence of A. phagocytophilum and related variants (A. phagocytophilum-like 1 and 2), A. ovis, and A. capra in sheep from Kyrgyzstan, the DNA of 391 blood samples were screened for 16 S SSU rRNA, groEL, and gltA genes by PCR, respectively. The primers used in this study are listed in Table 1.
The PCR assays were performed as described before (Kawahara et al. 2006; Haigh et al. 2008; Li et al. 2015; Yang et al. 2016), and the genomic DNA of A. phagocytophilum (GenBank accession no: JF807995, Altay et al. 2014), A. ovis (HE580282, Altay et al. 2014)d capra (MW672115, Altay et al. 2022a) were used as the positive controls, and DNase-RNase-free sterile water (Cat No.: 129,114, Qiagen®, Germany) was used as the negative control in the PCRs.
PCR products were loaded on 1.6% agarose gel containing ethidium bromide and visualized under UV transilluminator. The DNA extraction, PCR, and gel electrophoresis were performed in separate compartments of the laboratory to minimize the risk of contamination.
Discrimination Anaplasma phagocytophilum and related variants (A. phagocytohilum-like 1 and 2) based restriction of 16 S SSU rRNA genes with XcmI and BsaI
A polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) was performed to discriminate between A. phagocytophilum, A. phagocytophilum-like 1 and 2. After a 641/642 bp of the 16 S SSU rRNA gene of A. phagocytophilum and/or related variants (like 1 and 2) were amplified with SSAP2f and SSAP2r primers, the PCR products were digested with XcmI and BsaI enzymes as previously described (Ben Said et al. 2017; Aktas and Colak 2021).
The expected RFLP band profiles of A. phagocytophilum digested with XcmI are 344 and 297 bp. XcmI does not cut A. phagocytophilum-like 1 and 2. On the other hand, the expected RFLP band profiles of A. phagocytophilum-like 2 digested with BsaI are 219 and 422/423 bp. BsaI does not cut A. phagocytophilum and A. phagocytophilum-like 1. In the A. phagocytophilum-like 1 and 2 mix infections, band profiles of 219, 422/423 and 641/642 bp are expected in BsaI restriction (Ben Said et al. 2017; Aktas and Colak 2021). The confirmation of RFLP results were carried out with the sequence analysis.
Sequencing and phylogenetic analysis
The 21 of A. capra, three of A. phagocytophilum-like 1, and two of A. ovis PCR positive samples were sequenced. To perform sequence analysis, the PCR products were purified from agarose gel using a commercial gel extraction kit (PCR Clean-Up & Gel Extraction Kit, GeneDireX®, Cat.No.: NA006-0300), according to the manufacturer’s recommendations. The SSAP2f/r and the inner primer pairs listed in Table 1 used for sequencing of 16 S SSU rRNA gene of A. phagocytophlum-like 1 and gltA gene of A. capra, respectively. A part of 16 S SSU rRNA gene of A. ovis were sequenced using one set of primers (16S8FE and B-GA1B) which is specific 492–498 bp fragment of the 16 S SSU rRNA gene, spanning the V1 region of Anaplasma and Ehrlichia species (Schouls et al. 1999).
Sequencing was performed using ABI 3730XL analyzer (Applied Biosystems, Foster City, CA) and BigDye Terminator v3.1 Cycle sequencing kit (Applied Biosystems, Foster City, CA).
The consensus sequences in this study were determined using the MUSCLE algorithm of MEGA-X software (Kumar et al. 2018). These consensus sequences were compared with sequences present in the GenBank to determine nucleotide similarities with the BLAST algorithm. The sequences from this study were submitted to the GenBank database and their accession numbers were obtained.
Phylogenetic analyzes of the sequences identified in this study were performed using other gltA and 16 S SSU rRNA nucleotide sequences of Anaplasma species available in the GenBank. The phylogenetic tree was carried out with maximum likelihood analysis in Mega X (Kumar et al. 2018). The best-fit model for maximum likelihood was considered as the Kimura-2 parameter model for gltA and 16 S SSU rRNA genes (Kimura 1980) using the Find Best-Fit Substitution Model in Mega X (Kumar et al. 2018). Bootstrap values were performed with 1,000 replicates (Fig. 2).
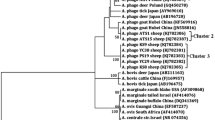
Phylogenetic tree based on the gltA gene sequences of A. capra (OM100820-OM100840) using the maximum likelihood method. Numbers at the nodes represent the bootstrap values with 1000 replicates. The evolutionary history was inferred by using the Maximum Likelihood method and Kimura 2-parameter model (Kimura 1980). Scale bar represents 0.20 substitutions per nucleotide position. Rickettsia ricketsii (Accession number: U59729) was used as an outgroup in the tree. Evolutionary analyses were conducted in MEGA X (Kumar et al. 2018)
Results
Prevalence and distribution of Anaplasma spp. in sheep
The result of PCR and RFLP of 391 samples revealed the presence of A. phagocytophilum-like 1, A. ovis and A. capra in sheep in Kyrgyzstan. The prevalence and frequency of A. phagocytophilum-like 1, A. ovis and A. capra is shown in Table 2. Overall prevalence of Anaplasma species in sheep was found to be 25.8% (101/391) by three different species-specific PCR. The most abundant species was A. ovis (88/391, 22.5%) followed by A. phagocytophilum-like 1 (27/391, 6.9%) and A. capra (21/391, 5.3%). Only one Anaplasma species was found in 69 sheep, whereas mixed infections with two or three species were detected in 32 sheep in this study. A total of 15 sheep were infected with both A. phagocytophilum-like 1 and A. ovis, eight sheep were infected with both A. ovis and A. capra whereas six sheep were infected with both A. phagocytophilum-like 1 and A. capra and three sheep were infected with the three species.
Discrimination of Anaplasma phagocytophilum and related variants A. phagocytophilum-like 1 and 2
In this study, A. phagocytophilum or related variants were detected in 27 samples by PCR (Table 2). All of the 27 PCR products were analyzed with RFLP using XcmI and BsaI restriction enzymes. A. phagocytophilum-like 1 was detected in all 27 sample by PCR-RFLP, while A. phagocytophilum and A. phagocytophilum-like 2 were not detected.
To confirm the RFLP results, randomly selected three representative samples were sequenced. These sequences were submitted to the Genbank, and deposited with accession numbers: OM540435-OM540437. The sequences were 99.83–100% similar to A. phagocytophilum-like 1 sequences available in the GenBank. The A. phagocytophilum-like 1 sequence obtained in this study were 100% identical to those of A. phagocytophilum-like 1 detected in sheep from Tunisia (KM285230), cattle from Türkiye (GU223365), goat from China (OL678408) and Sika deer (Cervus nippon) from Japan (JM055357).
Analysis of the gtlA gene sequences for determination of A. capra genotypes
All the positive samples (21 samples) were sequenced and aligned with A. capra sequences present in the GenBank and then all the sequences were deposited in the GenBank, as accession numbers: OM100820-OM100840.
The gltA gene sequences of 21 positive samples obtained in this study showed a complete consensus. However, BLAST analysis showed that the gtlA sequences of A. capra obtained in this study were found 86.01–100% similar to the 174 A. capra sequences present in the GenBank. There was a high homology (98.33–100%) between sequences obtained in this study and 27 gltA sequences of A. capra present in the GenBank. In contrast, a low homology was determined (86.01–86.24%) with 147 sequences present in GenBank. Detailed information about nucleotide similarity rates between A. capra genotypes was given in Table 3. Additionally, the sequence alignment results showed that only 0–7 nucleotides differences emerged between sequences obtained from the present study and the sequences from red deer, swamp deer (Rucervus duvaucelii), Siberian roe deer (Capreolus pygargus), takin, Reeve’s muntjac, Forest musk deer, D. everestianus, Korean water deer, cattle, and sheep, while 68–70 nucleotides differences were observed between the sequences from dog, cattle, sheep, goat, human, H. qinghaiensis H. longicornis, and R. microplus (Fig. 3).
Phylogenetic analysis
The phylogenetic analysis based on the gltA gene revealed that A. capra was separated into two clusters, and A. capra identified in this study clustered within red deer, swamp deer, Siberian roe deer, takin, Reeve’s muntjac, Forest musk deer, D. everestianus, Korean water deer, cattle, and sheep (Fig. 2).
Anaplasma phagocytophilum-like 1 variant isolated in the present study clustered a distinct group with those of previously published sequences of A. phagocytophilum-like 1 (Fig. 4).
Phylogenetic tree based on the 16 S SSU rRNA gene sequences of A. phagocytophilum- like 1 (OM540435-OM540437) using the maximum likelihood method. Numbers at the nodes represent the bootstrap values with 1000 replicates. The evolutionary history was inferred by using the Maximum Likelihood method and Kimura 2-parameter model (Kimura 1980). Scale bar represents 0.0050 substitutions per nucleotide position. Anaplasma capra (Accession number: LC432126) was used as an outgroup in the tree. Evolutionary analyses were conducted in MEGA X (Kumar et al. 2018)
In this study, we also determined a partial sequence of 16 S SSU rRNA gene of A. ovis to validate the PCR results. Two sequences of A. ovis were deposited in the GenBank under the accession numbers of OM453952 and OM453953. The BLAST and phylogenetic analysis of the sequences showed that the A. ovis sequences obtained in this study were in full compliance with the A. ovis sequences present in the Genbank (data not shown).
Discussion
Tick-borne diseases such as anaplasmosis have enormous negative effects on the livestock industry almost all over the world (Kocan et al. 2010). The prevalence of TBDs like anaplasmosis may vary according to multiple factors, including sampling seasons, differences in animal feeding and husbandry, presence and abundance of ticks and other vectors, sampling area (especially climatic and ecological factors), host resistance, and sample processing methods (Torina et al. 2008; Kocan et al. 2010; Belkahia et al. 2014). In this study, the overall prevalence of anaplasmosis in sheep was found to be 25.8% (101/391). The prevalence at the species level of A. ovis, A. phagocytophilum-like 1 and A. capra genotype-1 were determined to be 22.5, 6.9 and, 5.3%, respectively.
Anaplasma capra is a newly described species which has zoonotic character and can infect a wide range of hosts. In this study, we investigated the presence and prevalence of A. capra in sheep, and genotypes of the species were documented for the first time. The results (5.3%) in this study were compared with other countries, the prevalence of A. capra was lower than that previously found in sheep (16.3%) and goats (12.3%) from China (Yang et al. 2017), Korean water deer (13.8%) from Korea (Amer et al. 2019), dogs (12.1%) from China (Shi et al. 2019). The A. capra prevalence determined in this study was higher than in cattle (0.3%) and goats (0.3%) from Korea (Miranda et al. 2021), deer (swamp and red deer) (4.5%) from France (Jouglin et al. 2019), and cattle (0.3%) from Kyrgyzstan (Altay et al. 2022a), but was close to that found in roe deer (5.8%) from Spain (Remesar et al. 2022). This work was the first to reveal the presence of A. capra in Kyrgyzstan sheep, and it will contribute to the understanding of the epidemiology of this species in the world. However, further research is needed to determine its vectors and the pathogenicity of the novel Anaplasma species. In this study, all samples were collected from clinically healthy animals and no ticks were collected from sheep in the sampling process. The pathogenicity of A. capra is not clear among animal host, and a research conducted by Jouglin et al. (2019) showed that A. capra can persist in red deer for four months. The persistently infected animals may serve as reservoirs for vectors, and these animals are important in the epidemiology of the pathogens (Kocan et al. 2010; Brown and Barbet 2016). In this study, animals infected with A. capra were clinically healthy, and probably these animals were persistently infected with A. capra.
With the increase in the number of the hosts in the countries where A. capra is detected by molecular studies, the sequence registration rate in the GenBank of this species also increases. Thus, it is possible to compare different A. capra samples genetically. In the present study, 21 A. capra PCR positive samples were detected by the gltA gene sequences. The phylogenetic and BLAST analyses, including the A. capra sequences identified in this study and sequences present in the GenBank revealed that A. capra is divided into two different geno-groups (A. capra genotype-1 and A. capra genotype-2) (Figs. 2 and 3). A relationship between these geno-groups, the host, or the region from which they were isolated, could not be determined. While the similarity rates of 27 A. capra samples in the first group and sequences obtained in this study were 98.33–100%, the 147 A. capra sequences in the second group differ significantly from this group and the similarity rate decreases to 86.01–86.24%. Although the difference between the two groups was significant, the homology within the groups was quite high (Table 3). A. capra genotype-1 and A. capra genotype-2 are clearly distinguished from each other according to the gltA gene sequences compared to other gene sequences such as 16 S SSU rRNA and groEL (Unpublished data). We think that the naming of these two groups can be used until we reach research results that will provide a further nomenclature.
Recently based molecular studies has documented that A. phagocytophilum consists of one species and two related variants (A. phagocytophilum-like 1 and 2) (Ben Said et al. 2015, 2017). According to the results of PCR, RFLP and DNA sequence in this study, A. phagocytophilum-like 1 was found in 27 samples (6.9%). The prevalence was close to that reported in Tunisian sheep (7%) (Ben Said et al. 2017), but lower than that reported in small ruminants from Türkiye (26.5%) (Aktas et al. 2021). The phylogenetic tree based on 16 S SSU rRNA sequence revealed that samples identified in this study clustered in A. phagocytophilum-like 1 group (Fig. 4). Studies in which the presence of A. phagocytophilum related variants in farm animals were determined, stated that both variants did not cause clinical symptoms (Ben Said et al. 2015, 2017; Aktas and Colak 2021). In this study, all the animals sampled were clinically healthy, and this result was compatible with the previous studies (Aktas and Colak 2021; Noaman, 2022). When the studies are evaluated together, it can be thought that A. phagocytophilum variants do not cause clinical disease in farm animals. However, detailed studies are needed to determine its clinical effect.
Anaplasma ovis is known as the most prevalent Anaplasma species in sheep all over the world (Dumler et al. 2001). We also detected that A. ovis was the most prevalent species in sheep from Kyrgyzstan (22.5%). When the prevalence studies and the results from this study are evaluated together, it can be seen that A. ovis is an endemic species in many countries (Liu et al. 2012; Altay et al. 2014; Belkahia et al. 2014; Noaman and Sazmand 2022). Although A. ovis is generally thought to cause mild disease, it has been reported that it causes severe clinical symptoms and even death in the presence of secondary infections or predisposing factors (Friedhoff 1997; Renneker et al. 2013). Therefore, A. ovis infection should be taken into consideration more often in sheep flocks.
In conclusion, this study indicated that Anaplasma species are widespread in sheep from Kyrgyzstan with having a 25.8% prevalence. The results of this work indicate the presence of A. phagocytophilum-like 1, A. ovis, and A. capra in sheep in Kyrgyzstan for the first time. In the study, we documented that A. capra has two different genotypes. We suggest that the naming of these two groups, A. capra genotype-1 and A. capra genotype-2 can be used until we reach research results that will provide a further nomenclature. All the results show that Anaplasma species are important in sheep breeding in Kyrgyzstan, while revealing the necessity of considering genotypes in studies to be carried out on A. capra.
Data Availability
All data generated or analyzed during this study are included in this manuscript.
Code Availability
Not applicable.
References
Aktas M, Colak S (2021) Molecular detection and phylogeny of Anaplasma spp. in cattle reveals the presence of novel strains closely related to A. phagocytophilum in Turkey. Ticks Tick Borne Dis 12:101604. https://doi.org/10.1016/j.ttbdis.2020.101604
Aktas M, Ozubek S, Ulucesme MC (2021) Molecular detection and phylogeny of Anaplasma phagocytophilum and related variants in small ruminants from Turkey. Animals 11:814. https://doi.org/10.3390/ani11030814
Altay K, Dumanli N, Aktas M, Ozubek S (2014) Survey of Anaplasma Infections in Small Ruminants from East Part of Turkey. Kafkas Univ Vet Fak Derg 20:1–4. https://doi.org/10.9775/kvfd.2013.9189
Altay K, Erol U, Sahin OF, Aytmirzakizi A (2022a) () First molecular detection of Anaplasma species in cattle from Kyrgyzstan; molecular identification of human pathogenic novel genotype Anaplasma capra and Anaplasma phagocytophilum related strain. Ticks Ticks Borne Dis 13:101861. https://doi.org/10.1016/j.ttbdis.2021.101861
Altay K, Erol U, Sahin OF (2022b) The first molecular detection ofAnaplasma caprain domestic ruminants in the central part of Turkey, with genetic diversity and genotyping ofAnaplasma capra. Trop Anim Health Prod 54:1–8. https://doi.org/10.1007/s11250-022-03125-7. doi
Amer S, Kim S, Yun Y, Na KJ (2019) Novel variants of the newly emergedAnaplasma caprafrom Korean water deer (Hydropotes inermis argyropus) in South Korea. Parasit Vectors 12:1–9. https://doi.org/10.1186/s13071-019-3622-5. )
Belkahia H, Ben Said M, El Hamdi S, Yahiaoui M, Gharbi M, Daaloul-Jedidi M, Messadi L (2014) First molecular identification and genetic characterization of Anaplasma ovis in sheep from Tunisia. Small Rum Res 121:404–410. https://doi.org/10.1016/j.smallrumres.2014.07.009
Ben Said M, Belkahia H, Alberti A, Zobba R, Bousrih M, Yahiaoui M, Daaloul-Jedidi M, Mamlouk A, Gharbi M, Messadi L (2015) Molecular survey of Anaplasma species in small ruminants reveals the presence of novel strains closely related to A. phagocytophilum in Tunisia. Vector Borne Zoonotic Dis 15:580–590. https://doi.org/10.1089/vbz.2015.1796
Ben Said M, Belkahia H, El Mabrouk N, Saidani M, Ben Hassen M, Alberti A, Zobba R, Bouattour S, Bouattour A, Messadi L (2017) Molecular typing and diagnosis of Anaplasma spp. closely related to Anaplasma phagocytophilum in ruminants from Tunisia. Ticks Tick Borne Dis 8:412–422. https://doi.org/10.1016/j.ttbdis.2017.01.005
Brown WC, Barbet AF (2016) Persistent infections and immunity in ruminants to arthropod-borne bacteria in the family Anaplasmataceae. Annu Rev Anim Biosci 4:177–197. https://doi.org/10.1146/annurev-animal-022513-114206
Chochlakis D, Ioannou I, Tselentis Y, Psaroulaki A (2010) Human anaplasmosis and Anaplasma ovis variant. Emerg Infect Dis 16:1031–1032. https://doi.org/10.3201/eid1606.090175
Dumler JS, Barbet AF, Bekker CP, Dasch GA, Palmer GH, Ray SC, Rikihisa Y, Rurangirwa FR (2001) Reorganization of genera in the families Rickettsiaceae and Anaplasmataceae in the order Rickettsiales: unification of some species of Ehrlichia with Anaplasma, Cowdria with Ehrlichia and Ehrlichia with Neorickettsia, descriptions of six new species combinations and designation of Ehrlichia equi and ‘HGE agent’ as subjective synonyms of Ehrlichia phagocytophila. Int J Syst Evol Microbiol 51:2145–2165. https://doi.org/10.1099/00207713-51-6-2145
Fang LQ, Liu K, Li XL, Liang S, Yang Y, Yao HW, Sun RX, Sun Y, Chen WJ, Zuo SQ, Ma MJ, Li H, Jiang JF, Liu W, Yang XF, Gray GC, Krause PJ, Cao WC (2015) Emerging tick-borne infections in mainland China: an increasing public health threat. Lancet Infect Dis 15:1467–1479. https://doi.org/10.1016/S1473-3099(15)00177-2
Friedhoff KT (1997) Tick-borne diseases of sheep and goats caused by Babesia, Theileria or Anaplasma spp. Parassitologia 39:99–109
Guo WP, Huang B, Zhao Q, Xu G, Liu B, Wang YH, Zhou EM (2018) Human-pathogenic Anaplasma spp., and Rickettsia spp. in animals in Xi’an, China. PLoS Negl Trop Dis 12:e0006916. https://doi.org/10.1371/journal.pntd.0006916
Guo WP, Zhang B, Wang YH, Xu G, Wang X, Ni X, Zhou EM (2019) Molecular identification and characterization of Anaplasma capra and Anaplasma platys-like in Rhipicephalus microplus in Ankang, Northwest China. BMC Infect Dis 19:434. https://doi.org/10.1186/s12879-019-4075-3
Haigh JC, Gerwing V, Erdenebaatar J, Hill JE (2008) A novel clinical syndrome and detection of Anaplasma ovis in Mongolian reindeer (Rangifer tarandus). J Wildl Dis 44:569–577. https://doi.org/10.7589/0090-3558-44.3.569
Han R, Yang JF, Mukhtar MU, Chen Z, Niu QL, Lin YQ, Liu GY, Lou JX, Yin H, Liu ZJ (2019) Molecular detection of Anaplasma infections in ixodid ticks from the Qinghai-Tibet Plateau. Infect Dis Poverty 8:1–8. https://doi.org/10.1186/s40249-019-0522-z
Hosseini-Vasoukolaei N, Oshaghi MA, Shayan P, Vatandoost H, Babamahmoudi F, Yaghoobi-Ershadi MR, Telmadarraiy Z, Mohtarami F (2014) Anaplasma infection in ticks, livestock and human in Ghaemshahr, Mazandaran Province, Iran. J Arthropod Borne Dis 8:204–211
Jilintai SN, Hayakawa D, Suzuki M, Hata H, Kondo S, Matsumoto K, Yokoyama N, Inokuma H (2009) Molecular survey for Anaplasma bovis and Anaplasma phagocytophilum infection in cattle in a pastureland where sika deer appear in Hokkaido, Japan. Jpn J Infect Dis 62:73–75
Jouglin M, Blanc B, de la Cotte N, Bastian S, Ortiz K, Malandrin L (2019) First detection and molecular identification of the zoonotic Anaplasma capra in deer in France. PLoS ONE 14:0219184. https://doi.org/10.1371/journal.pone.0219184
Kang YJ, Diao XN, Zhao GY, Chen MH, Xiong Y, Shi M, Fu WM, Guo YJ, Pan B, Chen XP, Holmes EC, Gillespie JJ, Dumler SJ, Zhang YZ (2014) Extensive diversity of Rickettsiales bacteria in two species of ticks from China and the evolution of the Rickettsiales. BMC Evol Biol 14:167. https://doi.org/10.1186/s12862-014-0167-2
Karshima SN, Ahmed MI, Kogi CE, Iliya PS (2022) Anaplasma phagocytophilum infection rates in questing and host-attached ticks: a global systematic review and meta-analysis. Acta Trop 228:106299. https://doi.org/10.1016/j.actatropica.2021.106299
Kawahara M, Rikihisa Y, Lin Q, Isogai E, Tahara K, Itagaki A, Hiramitsu Y, Tajima T (2006) Novel genetic variants of Anaplasma phagocytophilum, Anaplasma bovis, Anaplasma centrale, and a novel Ehrlichia sp. in wild deer and ticks on two major islands in Japan. Appl Environ Microbiol 72:1102–1109. https://doi.org/10.1128/AEM.72.2.1102-1109.2006
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. https://doi.org/10.1007/BF01731581
Kocan KM, de la Fuente J, Blouin EF, Coetzee JF, Ewing SA (2010) The natural history of Anaplasma marginale. Vet Parasitol 167:95–107. https://doi.org/10.1016/j.vetpar.2009.09.012
Koh FX, Panchadcharam C, Sitam FT, Tay ST (2018) Molecular investigation of Anaplasma spp. in domestic and wildlife animals in Peninsular Malaysia. Vet Parasitol Reg Stud Reports 13:141–147. https://doi.org/10.1016/j.vprsr.2018.05.006
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Li H, Zheng YC, Ma L, Jia N, Jiang BG, Jiang RR, Huo QB, Wang YW, Liu HB, Chu YL, Song YD, Yao NN, Sun T, Zeng FY, Dumler JS, Jiang JF, Cao WC (2015) Human infection with a novel tick-borne Anaplasma species in China: a surveillance study. Lancet Infect Dis 15:663–670. https://doi.org/10.1016/S1473-3099(15)70051-4
Liu Z, Ma M, Wang Z, Wang J, Peng Y, Li Y, Guan G, Luo J, Yin H (2012) Molecular survey and genetic identification of Anaplasma species in goats from central and southern China. Appl Environ Microbiol 78:464–470. https://doi.org/10.1128/AEM.06848-11
Lu M, Tian J, Pan X, Qin X, Wang W, Chen J, Guo W, Li K (2022) Identification of Rickettsia spp., Anaplasma spp., and an Ehrlichia canis-like agent in Rhipicephalus microplus from Southwest and South-Central China. Ticks Tick Borne Dis 13:101884. https://doi.org/10.1016/j.ttbdis.2021.101884
Miranda EA, Han SW, Cho YK, Choi KS, Chae JS (2021) Co-Infection with Anaplasma Species and Novel Genetic Variants Detected in Cattle and Goats in the Republic of Korea. Pathogens 10:28. https://doi.org/10.3390/pathogens10010028
Noaman V (2022) Factors associated with Anaplasma phagocytophilum infection in sheep in Iran. Small Rum Res 208:106617. https://doi.org/10.1016/j.smallrumres.2022.106617
Noaman V, Sazmand A (2022) Anaplasma ovis infection in sheep from Iran: molecular prevalence, associated risk factors, and spatial clustering. Trop Anim Health Prod 54:1–11. doi: https://doi.org/10.1007/s11250-021-03007-4
Ohashi N, Inayoshi M, Kitamura K, Kawamori F, Kawaguchi D, Nishimura Y, Naitou H, Hiroi M, Masuzawa T (2005) Anaplasma phagocytophilum-infected ticks, Japan. Emerg Infect Dis 11:1780–1783. https://doi.org/10.3201/eid1111.050407
Peng Y, Wang K, Zhao S, Yan Y, Wang H, Jing J, Jian F, Wang R, Zhang L, Ning C (2018) Detection and phylogenetic characterization of Anaplasma capra: an emerging pathogen in sheep and goats in China. Front Cell Infect Microbiol 8:283. https://doi.org/10.3389/fcimb.2018.00283
Qin XR, Han FJ, Luo LM, Zhao FM, Han HJ, Zhang ZT, Liu JW, Xue ZF, Liu MM, Ma DQ, Huang YT, Sun Y, Sun XF, Li WQ, Zhao L, Yu H, Yu XJ (2018) Anaplasma species detected in Haemaphysalis longicornis tick from China. Ticks Tick Borne Dis 9:840–843. https://doi.org/10.1016/j.ttbdis.2018.03.014
Remesar S, Prieto A, Garcia-Dios D, Lopez-Lorenzo G, Martinez-Calabuig N, Diaz-Cao JM, Panadero R, Lopez CM, Fernandez G, Diez-Banos P, Morrondo P, Dia P (2022) Diversity of Anaplasma species and importance of mixed infections in roe deer from Spain. Transbound Emerg Dis 1–12. https://doi.org/10.1111/tbed.14319
Renneker S, Abdo J, Salih DE, Karagenc T, Bilgic H, Torina A, Oliva AG, Campos J, Kullmann B, Ahmed J, Seitzer U (2013) Can Anaplasma ovis in Small Ruminants be Neglected any Longer? Transbound Emerg Dis 2:105–112. https://doi.org/10.1111/tbed.12149
Schouls LM, Van De Pol I, Rijpkema SG, Schot CS (1999) Detection and identification of Ehrlichia, Borrelia burgdorferi sensu lato, and Bartonella species in Dutch Ixodes ricinus ticks. J Clin Microbiol 37:2215–2222. https://doi.org/10.1128/JCM.37.7.2215-2222.1999
Shi K, Li J, Yan Y, Chen Q, Wang K, Zhou Y, Li D, Chen Y, Yu F, Peng Y, Zhang L, Ning C (2019) Dogs as new hosts for the emerging zoonotic pathogen Anaplasma capra in China. Front Cell Infect Microbiol 9:394. https://doi.org/10.3389/fcimb.2019.00394
Staji H, Yousefi M, Hamedani MA, Tamai IA, Khaligh SG (2021) Genetic characterization and phylogenetic of Anaplasma capra in Persian onagers (Equus hemionus onager). Vet Microbiol 261:109199. https://doi.org/10.1016/j.vetmic.2021.109199
Torina A, Alongi A, Naranjo V, Estrada-Pena A, Vicente J, Scimeca S, Marino AM, Salina F, Caracappa S, de la Fuente J (2008) Prevalence and genotypes of Anaplasma species and habitat suitability for ticks in a Mediterranean ecosystem. Appl Environ Microbiol 74:7578–7584. https://doi.org/10.1128/AEM.01625-08
Wang H, Yang J, Mukhtar MZ, Liu Z, Zhang M, Wang X (2019) Molecular detection and identification of tick-borne bacteria and protozoans in goats and wild Siberian roe deer (Capreolus pygargus) from Heilongjiang Province, northeastern China. Parasit Vectors 12:296. https://doi.org/10.1186/s13071-019-3553-1
Wei W, Li J, Wang YW, Jiang BG, Liu HB, Wei R, Jiang RR, Cui XM, Li LF, Yuan TT, Wang Q, Zhao L, Xia LY, Jiang JF, Qui YF, Jia N, Cao WC, Hu YL (2020) Anaplasma platys-Like Infection in Goats, Beijing, China. Vector Borne Zoonotic Dis 20:755–762. https://doi.org/10.1089/vbz.2019.2597
Yang J, Li Y, Liu Z, Liu J, Niu Q, Ren Q, Chen Z, Guan G, Luo J, Yin H (2015) Molecular detection and characterization of Anaplasma spp. in sheep and cattle from Xinjiang, northwest China. Parasit Vectors 19:108. https://doi.org/10.1186/s13071-015-0727-3
Yang J, Liu Z, Niu Q, Liu J, Han R, Liu G, Shi Y, Luo J, Yin H (2016) Molecular survey and characterization of a novel Anaplasma species closely related to Anaplasma capra in ticks, northwestern China. Parasit Vectors 9:1–5. https://doi.org/10.1186/s13071-016-1886-6
Yang J, Liu Z, Niu Q, Liu J, Han R, Guan G, Hassan MD, Liu G, Luo J, Yin H (2017) A novel zoonotic Anaplasma species is prevalent in small ruminants: potential public health implications. Parasit Vectors 10:1–6. https://doi.org/10.1186/s13071-017-2182-9
Yang J, Liu Z, Niu Q, Mukhtar MU, Guan G, Liu G, Luo J, Yin H (2018) A novel genotype of “Anaplasma capra” in wildlife and its phylogenetic relationship with the human genotypes. Emerg Microbes Infect 7:1–4. https://doi.org/10.1038/s41426-018-0212-0
Yoshimoto K, Matsuyama Y, Matsuda H, Sakamoto L, Matsumoto K, Yokoyama N, Inokuma H (2010) Detection of Anaplasma bovis and Anaplasma phagocytophilum DNA from Haemaphysalis megaspinosain Hokkaido, Japan. Vet Parasitol 168:170–172. https://doi.org/10.1016/j.vetpar.2009.10.008
Acknowledgements
The authors would like to thank all veterinarians and technicians for their kind help during sample collection.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
Kursat Altay (DVM, PhD, Prof.) Conceptualization, Field Work, Methodology, Validation, Formal Analysis, Supervision, Writing- Original Draft Preparation, Reviewing and Editing. Ufuk EROL (DVM, PhD, Assist. Prof.) Conceptualization, Field Work, Methodology, Validation, Data Curation, Formal Analysis, Writing- Original Draft Preparation. Omer Faruk SAHIN (DVM, Res. Assist.) Field Work, Data Curation, Methodology, Formal Analysis. Ayperi AYTMIRZAKIZI (DVM, Res. Assist.) Field Work, Data Curation, Formal Analysis. Ethem Mutlu TEMIZEL (DVM, PhD, Prof.) Field Work, Data Curation, Formal Analysis. Mehmet Fatih AYDIN (DVM, PhD, Assist. Prof.) Data Curation, Methodology, Formal Analysis. Nazir DUMANLI (DVM, PhD, Prof.) Field Work, Data Curation, Methodology, Formal Analysis, Writing- Original Draft Preparation, Reviewing and Editing. Munir AKTAS (DVM, PhD, Prof.) Data Curation, Methodology, Formal Analysis, Writing- Original Draft Preparation, Reviewing and Editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
All procedures performed in studies involving animals were in accordance with the ethical standards approved by Experimentation Ethics Committee of Kyrgyz-Turkish Manas University (30.06.2017/2017–06/02) and the Sivas Cumhuriyet University Animal Experiments Local Ethics Committee (Approval number: 12.07.2021-564).
Consent to participate
The consent of all animal owners was sorted before this study was carried out.
Consent for publication
All authors have seen and approved the final version of the manuscript being submitted. They warrant that the article is the authors’ original work, has not received prior publication, and is not under consideration for publication elsewhere.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Altay, K., Erol, U., Sahin, O.F. et al. The detection and phylogenetic analysis of Anaplasma phagocytophilum-like 1, A. ovis and A. capra in sheep: A. capra divides into two genogroups. Vet Res Commun 46, 1271–1279 (2022). https://doi.org/10.1007/s11259-022-09998-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11259-022-09998-1