Abstract
Purpose
To identify a novel biomarker that can predict biochemical recurrence (BCR) after radical prostatectomy.
Methods
The gene expression profile of SAMD5 in prostate cancer was explored based on the oncomine database and The Cancer Genomic Atlas (TCGA). The follow-up information and clinical pathologic variables were extracted from the following cohort study: TCGA_prostate carcinoma. And then, survival analysis was conducted using the Kaplan–Meier plot and Cox’s proportional hazard regression model. Furthermore, another independent cohort study: Taylor prostate, was also acquired to validate the predictive effect of SAMD5 on BCR. In addition, the expression profile of SAMD5 in other cancer types was investigated using TCGA dataset.
Results
SAMD5 mRNA was shown to be up-regulated in multiple microarray datasets of prostate cancer with the strict statistic criteria: p < 0.01 and fold change ≥ 2. In TCGA_PCa cohort study, high expression of SAMD5 was a risk factor for patients on post-operative BCR (HR 2.181, 95%CI 1.199–3.966, p = 0.011) and this predictive ability was independent of Gleason score and pathologic T stage (HR 2.018, 95%CI 1.102–3.698, p = 0.023). In another validating cohort study, the statistic trend was similar, and the pooled analysis by combining the two cohort study further confirmed its prognostic effect.
Conclusion
SAMD5 mRNA was overexpressed in prostate cancer and had powerful prognostic ability on predicting post-operative BCR, independent of Gleason score and pathologic T stage. Its high expression was associated with poor prognosis after RP.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Prostate cancer is the second most frequently diagnosed cancer in males worldwide and its mortality is ranked fifth among males with tumor [1]. Radical prostatectomy (RP) is the principal therapy for localized prostate cancer. And approximately 30% of patients will undergo post-operative disease relapse, initially in the form of increased serum prostate-specific antigen (PSA) value. Some patients with this biochemical recurrence (BCR) would progress to metastatic and resistant to androgen-deprivation therapy, known as lethal castration-resistant prostate cancer. Clinical variables such as Gleason score, serum PSA, TNM stage, and margin status have been applied in the prediction of BCR after RP [2]. Nevertheless, patients with the same clinical indicators evolved into diverse clinical consequences. Many groups devoted to developing molecular markers such as serum alkaline phosphatase [3] to enhance the predictive ability of clinical indicators. Although these researches were impressive, their clinical application still needs further validation.
In the Human Protein Atlas database [4], sterile alpha motif domain containing 5/SAMD5 was shown to be expressed in prostate cancer via Immunohistochemistry technology (supplementary materials: Fig. S1) and was recently reported to be engaged with EphA5 via SAM–SAM domain interaction and involved in Receptor Tyrosine kinase signaling [5]. Down-regulation of SAMD5 in small cell lung carcinoma cell lines was associated with inhibition of cell proliferation [6]. And in rectal cancer, the expression level of SAMD5 was shown to be associated with the response to chemo-radiotherapy [7]. These studies suggested that SAMD5 was relevant to tumorigenesis and development, and would probably be applied in clinical practice. However, the differential expression between prostate cancer and normal prostatic tissue and the clinical predictive effect of SAMD5 in prostate cancer have not been investigated.
In this study, we demonstrated that SAMD5 was overexpressed in prostate cancer and aberrantly expressed in other tumor types. Based on two cohort studies with follow-up data: The Cancer Genomic Atlas/TCGA [8] and Taylor prostate dataset [9], SAMD5 was shown to be a useful indicator to predict biochemical recurrence after radical prostatectomy, and its high expression was associated with poor post-operative prognosis.
Materials and methods
Data acquisition and treatment
The differential expression analysis of SAMD5 between prostate cancer and normal prostate tissue was conducted using microarray data from the oncomine database [10] (https://www.oncomine.org/resource/login.html). The RNA-sequencing data of The Cancer Genomic Atlas/TCGA dataset were available on the website of Gene Expression Profiling Interactive Analysis/GEPIA [11] (http://gepia.cancer-pku.cn/index.html). And the GEPIA online tool for differential expression analysis was easy to access and based on limma R page. The statistic significant criteria were set as p < 0.01 or adjusted p < 0.01 in the above database, respectively.
The clinical data and gene expression data of TCGA prostate cancer dataset (up to Oct 19, 2018) were obtained from TCGA official website (https://portal.gdc.cancer.gov/). The downloaded data type of gene expression was fragments per kilobase of exon per million fragments mapped (FPKM), then this data type was converted to transcripts per million (TPM) by a bioinformatics engineer [12]. The exclusion criteria of PCa patients were used as follows: (1) pathologic result is not prostate adenocarcinoma, (2) patients with clinical data but not biochemical recurrence data, (3) patients whose vital clinical information involving American Joint Committee on Cancer (AJCC) TNM stage [13] is missed. At last, 345 patients, both having clinical data and gene expression data, were obtained in our study for survival analysis. Then Taylor dataset with information of gene expression profile, the clinical pathologic data, and follow-up data was downloaded from the cbioportal website [14, 15] (http://www.cbioportal.org/datasets). And 139 patients were included in our study as shown in supplementary material.
Statistic analysis
Chi-square test was conducted in SPSS 23.0 and p < 0.05 was considered of statistical significance. In survival analysis, Kaplan–Meier Curve method and univariable Cox’s proportional hazard regression model was applied for univariable analysis, while the multivariable Cox regression model was used for multiple factor survival analysis. The test criterion was set as p < 0.05. The graphs were obtained in Graphpad prism 7.0.
The meta-analysis of hazard ratios (HR) was conducted in STATA/SE 15.0 for windows. χ2 and I2 were used to evaluate the statistical heterogeneity among the two cohort studies. Heterogeneity was considered low when I2 was less than 50% and Pheterogeneity > 0.1 was deemed as no statistical significance. Furthermore, Fixed effect model was applied to archive pooled HRs with 95%CI, when heterogeneity was lower than 25%. Otherwise, the random effect model was used if I2 was between 25 and 50%.
Gene ontology and functional enrichment analyses were performed in Metascape [16] (http://metascape.org/gp/index.html#/main/step1).
Results
Aberrant expression of SAMD5 mRNA in prostate cancer
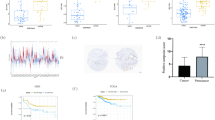
The differential expression of SAMD5 between prostate cancer and normal prostate tissue was investigated in the oncomine database. Under the threshold of p < 0.01 and Fold change ≥ 2, SAMD5 mRNA was found to be significantly up-regulated in 6 microarray datasets: Tomlins prostate [17], Varambally prostate [18], Lapointe prostate [19], Arredouani prostate [20], Grasso prostate [21], and Luo prostate [22]. Two datasets with maximal sample size were chosen to exhibit the overexpression of SAMD5 in prostate cancer, as shown in Fig. 1. Besides, in Taylor prostate dataset which was used for survival analysis, SAMD5 was overexpressed with p = 8.31e–9 and fold change = 1.163, and the result in TCGA_PCa dataset explored with an online tool: GEPIA showed the similar aberrant overexpression of SAMD5 with adjusted p value = 5.66e–27 and log2 (fold change) = 1.005.
Overexpression of SAMD5 in prostate cancer. a In Lapointe prostate dataset, differential expression of SAMD5 between normal prostatic tissues (n = 38) and prostate cancer (n = 62); b In Grasso prostate dataset, differential expression of SAMD5 between normal prostatic tissues (n = 28) and prostate cancer (n = 59); **represents p < 0.01
Patients characteristics
Two prospective cohort studies: TCGA_PCa and Taylor prostate were applied for survival analysis in this study. Patients characteristics are shown in Table 1. Forty-nine patients underwent post-operative biochemical recurrence in TCGA_PCa dataset, while in Taylor prostate dataset, the number was 35.
The relationship between the expression levels of SAMD5 and clinical pathologic parameters
Patients included in this study were divided into two groups: high expression group and low expression group according to their expression value of SAMD5 using the median as the cut-off point. As shown in Table 2, the χ2 test showed that there was no significant correlation between expression levels of SAMD5 and clinical pathologic parameters in TCGA_PCa dataset. The relationship between Gleason score, pathologic T stage, and expression levels of SAMD5 in Taylor prostate dataset still did not get statistical significance.
Predictive effect of SAMD5 expression on post-operative BCR
In TCGA_PCa dataset, overexpression of SAMD5 mRNA in prostate cancer was associated with poor prognosis of BCR (Fig. 2a, HR 2.174, 95%CI 1.242–3.807, Log-rank test p = 0.0088). Besides expression level of SAMD5, Gleason score, pathologic T stage, and AJCC TNM Stage revealed their predictive ability on BCR in univariable survival analysis (all p < 0.05). All these predictive factors with statistical significance in univariable survival analysis were entered into multivariable Cox regression model by Forward stepwise strategy, and as shown in Table 3 the predictive ability of SAMD5 was independent of Gleason score and pathologic T stage.
In order to validate this predictive effect, the other prospective cohort study with 138 patients was taken into the univariable and multivariable survival analyses. The result revealed that high expression of SAMD5 in prostate cancer was a risk factor of BCR for patients after RP (HR1.712); however, this did not get statistical significance (Fig. 2b; Table 3).
Meta-analysis of predictive effect in two datasets
In light of the inconsistent results of predictive effect, we conducted a meta-analysis for the two cohort studies to increase sample size and statistic power. The pooled analysis of two studies with 483 patients revealed that high expression of SAMD5 in prostate cancer led to the argumentation of the risk of post-operative BCR. The pooled Hazard ratio (Fig. 3a) of univariable Cox regression model was 1.96 (95%CI 1.25–3.09, p = 0.003) with extremely low heterogeneity (χ2 = 0.27, p = 0.602, I2 = 0.0%); Moreover, the pooled HR (Fig. 3b) of multivariable Cox regression model was also statistically significant (HR 1.74, 95%CI 1.09–2.77, p = 0.019) with extremely low heterogeneity (χ2 = 0.56, p = 0.456, I2 = 0.0%).
The meta-analysis of predictive effect in two cohort studies. a The HRs calculated from univariable Cox regression model in two datasets were pooled together. b The HRs calculated from multivariable Cox regression model in two datasets were pooled together. The variables entering into the model were expression level of SAMD5 (high or low expression), Gleason score (< 7, 7, or > 7), and pathologic T stage (T2, T3 or T4). I–V method represents the fixed effect model, while the D + L method represents the random effect model
In the section of sensitivity analysis, the pooled HRs kept constantly using the fixed effect model or random effect model (Fig. 3). Then we conducted another type of sensitivity analysis, removing a specific study and calculating the pooled HRs of the remaining study. The result (Fig. S2) revealed that the pooled HRs were stable too.
Gene ontology (GO) and functional enrichment analyses of SAMD5
The GO and functional enrichment analyses using Metascape showed that SAMD5 was involved in mitogen-activated protein kinase (MAPK) signaling pathway and can positively activate JNK kinase (Table 4).
Aberrant expression of SAMD5 in multiple cancer types
The pan-cancers analysis using GEPIA online tool with test criterion of adj. p < 0.01 revealed that compared with adjacent normal tissues, SAMD5 was overexpressed in prostate cancer and Stomach adenocarcinoma, while it was down-regulated in bladder urothelial carcinoma, breast invasive cancer, cervical squamous cell carcinoma and endocervical adenocarcinoma, cholangiocarcinoma, head and neck squamous cell carcinoma, kidney renal clear cell carcinoma, kidney renal papillary cell carcinoma, liver hepatocellular carcinoma, lung adenocarcinoma, lung squamous cell carcinoma, thyroid carcinoma, and uterine corpus endometrial carcinoma (Fig. 4).
Aberrant expression of SAMD5 in multiple cancer types. Left box in a specific tumor represents tumor group (T), while the right box represents normal group (N). BLCA bladder urothelial carcinoma, BRCA breast invasive carcinoma, CESC cervical squamous cell carcinoma and endocervical adenocarcinoma, CHOL cholangio carcinoma, HNSC head and neck squamous cell carcinoma, KIRC kidney renal clear cell carcinoma, KIRP kidney renal papillary cell carcinoma, LIHC liver hepatocellular carcinoma, LUAD lung adenocarcinoma, LUSC lung squamous cell carcinoma, PRAD prostate adenocarcinoma, STAD stomach adenocarcinoma, THCA Thyroid carcinoma, and UCEC Uterine Corpus Endometrial Carcinoma. *Represents p < 0.01
Discussion
In present clinical practice, the indication for post-operative adjuvant therapy was confined to pathological TNM staging as pT3 and pN+, positive surgical margins, and Gleason score ≥ 7 [2]. However, some patients without these signs still developed to BCR. The risk stratification needs to be updated.
In this study, we demonstrated the striking overexpression of SAMD5 in prostate cancer using microarray data and RNA-seq data (Fig. 1). Secondly, the association of expression level of SAMD5 and clinical pathologic parameters was explored and the result showed that they had no apparent correlation. Table 2 also shows the baseline characteristics of the two groups: Low expression group and high expression group. And Chi-square test evaluates covariates balance between these two groups. There are no obvious covariates imbalance between two groups. To investigate the predictive effect of SAMD5 expression level on post-operative BCR, these variables including Gleason score, pT, pN, and TNM stage were taken into Cox regression model using TCGA follow-up data, and the result revealed that the expression level of SAMD5 can robustly predict BCR after RP, and its predictive effect was independent of Gleason score and pathologic T stage. Then, to validate its predictive effect, we found another post-operative cohort study: Taylor prostate dataset and Cox regression analysis showed that high expression of SAMD5 was a risk factor on BCR. Unfortunately, it did not get statistical significance (Table 3).
In view of the similar statistical trend, we further conducted a meta-analysis to increase sample size and test power. We performed literature search in PubMed, Web of Science, and Cochrane Library with the following strategy: (SAMD5 OR Sterile Alpha Motif Domain Containing 5) AND (prostatic neoplasms OR prostate cancer) by the time up to November 1, 2018. No previous study about this issue was found showing the innovation of our research. The pooled HRs originated from univariable and multivariable Cox regression models exhibited that high expression of SAMD5 in prostate cancer was a powerful predictive indicator on prognosis after RP, and this effect was independent of Gleason score and pathologic T stage.
The molecular function of SAMD5 remains largely unknown. Previous studies mostly based on cell lines and their results were inconsistent. Different from small cell lung carcinoma cell lines [6], Tomoki Yagai et al. [23] reported that knockdown of SAMD5 in cholangiocarcinoma cell lines led to enhancement of cell proliferation. Our result in the tissue level suggested that the expression profile in tumor tissues was heterogeneous (Fig. 4) and indicated that SAMD5 may play diverse function depending on the cell type. A recent research reported that SAMD5 was engaged by EphA5 via SAM–SAM domain interaction and involved in Receptor Tyrosine kinase signaling [5]. Our findings (Table 4) did not contradict the previous result. MAPK signaling is part of the downstream of RTK signaling [24], the functional enrichment findings of SAMD5 binding to MAPK signaling proteins and positively activating JUN kinases/JNK may offer a new clue for further research on SAMD5. However, the precise mechanism of SAMD5 in prostate cancer is still unknown and needs further investigation.
Finally, some limitation of our research should be taken into account. Firstly, the differential expression of SAMD5 protein between prostate cancer and normal prostatic tissue and the exact molecular mechanism of SAMD5 in prostate cancer had not been investigated in our study. Secondly, the predictive effect of SAMD5 on prognosis after RP should be validated in another prospective clinical trial. Thirdly, the ethnic composition of the population in the TCGA database was mainly white and black; hence, whether our findings in the study could be extrapolated to other ethnicities remain unclear.
Conclusion
SAMD5 mRNA was strikingly overexpressed in prostate cancer and had powerful predictive ability on predicting post-operative BCR, independent of Gleason score and pathologic T stage. High expression of SAMD5 mRNA in prostate cancer was associated with poor prognosis after RP.
Abbreviations
- BCR:
-
Biochemical recurrence
- RP:
-
Radical prostatectomy
- TCGA:
-
The Cancer Genomic Atlas
- GEPIA:
-
Gene expression profiling interactive analysis
- TPM:
-
Transcripts per million
- AJCC:
-
American Joint Committee on Cancer
References
Bray F, Ferlay J, Soerjomataram I et al (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424. https://doi.org/10.3322/caac.21492
Cornford P, Bellmunt J, Bolla M et al (2017) EAU-ESTRO-SIOG guidelines on prostate cancer. Part II: treatment of relapsing, metastatic, and castration-resistant prostate cancer. Eur Urol 71:630–642. https://doi.org/10.1016/j.eururo.2016.08.002
Li D, Lv H, Hao X et al (2018) Prognostic value of serum alkaline phosphatase in the survival of prostate cancer: evidence from a meta-analysis. Cancer Manage Res 10:3125–3139. https://doi.org/10.2147/CMAR.S174237
Uhlen M, Zhang C, Lee S et al (2017) A pathology atlas of the human cancer transcriptome. Science. https://doi.org/10.1126/science.1260419
Wang Y, Shang Y, Li J et al (2018) Specific Eph receptor-cytoplasmic effector signaling mediated by SAM-SAM domain interactions. elife. https://doi.org/10.7554/eLife.35677
Matsuo T, Dat le T, Komatsu M et al (2014) Early growth response 4 is involved in cell proliferation of small cell lung cancer through transcriptional activation of its downstream genes. PLoS ONE 9:e113606. https://doi.org/10.1371/journal.pone.0113606
Watanabe T, Kobunai T, Akiyoshi T et al (2014) Prediction of response to preoperative chemoradiotherapy in rectal cancer by using reverse transcriptase polymerase chain reaction analysis of four genes. Dis Colon Rectum 57:23–31. https://doi.org/10.1097/01.dcr.0000437688.33795.9d
The Cancer Genome Atlas Research Network (2015) The molecular taxonomy of primary prostate cancer. Cell 163:1011–1025. https://doi.org/10.1016/j.cell.2015.10.025
Taylor BS, Schultz N, Hieronymus H et al (2010) Integrative genomic profiling of human prostate cancer. Cancer Cell 18:11–22. https://doi.org/10.1016/j.ccr.2010.05.026
Rhodes DR, Yu J, Shanker K et al (2004) ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia 6:1–6. https://doi.org/10.1016/S1476-5586(04)80047-2
Tang Z, Li C, Kang B et al (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45:W98–W102. https://doi.org/10.1093/nar/gkx247
Li B, Ruotti V, Stewart RM et al (2010) RNA-Seq gene expression estimation with read mapping uncertainty. Bioinformatics 26:493–500. https://doi.org/10.1093/bioinformatics/btp692
Thompson IM, Andriole GL, Blumenstein B et al (2003) AJCC cancer staging manual, 6th edn. Springer, New York
Cerami E, Gao J, Dogrusoz U et al (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2:401–404. https://doi.org/10.1158/2159-8290.CD-12-0095
Gao J, Aksoy BA, Dogrusoz U et al (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6:pl1. https://doi.org/10.1126/scisignal.2004088
Tripathi S, Pohl MO, Zhou Y et al (2015) Meta- and Orthogonal integration of influenza “OMICs” data defines a role for UBR4 in virus budding. Cell Host Microbe 18:723–735. https://doi.org/10.1016/j.chom.2015.11.002
Tomlins SA, Mehra R, Rhodes DR et al (2007) Integrative molecular concept modeling of prostate cancer progression. Nat Genet 39:41–51. https://doi.org/10.1038/ng1935
Varambally S, Yu J, Laxman B et al (2005) Integrative genomic and proteomic analysis of prostate cancer reveals signatures of metastatic progression. Cancer Cell 8:393–406. https://doi.org/10.1016/j.ccr.2005.10.001
Lapointe J, Li C, Higgins JP et al (2004) Gene expression profiling identifies clinically relevant subtypes of prostate cancer. Proc Natl Acad Sci USA 101:811–816. https://doi.org/10.1073/pnas.0304146101
Arredouani MS1, Lu B, Bhasin M et al (2009) Identification of the transcription factor single-minded homologue 2 as a potential biomarker and immunotherapy target in prostate cancer. Clin Cancer Res 15:5794–5802. https://doi.org/10.1158/1078-0432.CCR-09-0911
Grasso CS, Wu YM, Robinson DR et al (2012) The mutational landscape of lethal castration-resistant prostate cancer. Nature 487:239–243. https://doi.org/10.1038/nature11125
Luo JH, Yu YP, Cieply K et al (2002) Gene expression analysis of prostate cancers. Mol Carcinog 33:25–35. https://doi.org/10.1002/mc.10018
Yagai T, Matsui S, Harada K et al (2017) Expression and localization of sterile alpha motif domain containing 5 is associated with cell type and malignancy of biliary tree. PLoS ONE 12:e0175355. https://doi.org/10.1371/journal.pone.0175355
Lemmon MA, Schlessinger J (2010) Cell signaling by receptor tyrosine kinases. Cell 141:1117–1134. https://doi.org/10.1016/j.cell.2010.06.011
Acknowledgements
This research was based on public database: TCGA and GEPIA, and we are grateful for the extraordinary works of these project groups. We thank Bioinformatics Engineer Rang-Fei Zhu for his excellent pretreatment of TCGA-PCa data.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare that they have no conflict of interest.
Ethical approval
The patients’ information involved in our research were obtained from The Cancer Genome Atlas (TCGA) and Taylor Prostate dataset. All the patients and treatments complied with the principles laid down in the Declaration of Helsinki in 1964 and its later amendments or comparable ethical standards.
Informed consent
Informed consent was confirmed by all the patients participated in the TCGA-Prostate adenocarcinoma project and Taylor prostate project.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11255_2019_2096_MOESM1_ESM.tif
Supplementary material 1 Fig. S1 SAMD5 protein expression in Prostate cancer. a. The HPA research group detected SAMD5 protein expression in 10 prostate cancer samples via IHC technology, and found that 6 samples were stained with Low to medium strength. b. SAMD5 protein was shown to be expressed in prostate cancer cells (Brown dots) and located in cytoplasm or membrane. c. SAMD5 protein staining was negative in normal prostate tissue. (TIF 6927 KB)
11255_2019_2096_MOESM2_ESM.eps
Supplementary material 2 Fig. S2 Sensitive analysis of pooled HRs. a. The pooled HR calculated from univariable Cox regression model in two datasets was stable. After removing a specific study and calculating the pooled HR of the remaining study, both the point estimations dropped into the interval:1.25-3.09. b. The pooled HR calculated from multivariable Cox regression model in two datasets was stable. After removing a specific study and calculating the pooled HR of the remaining study, both the point estimations dropped into the interval:1.09-2.77. (EPS 1780 KB)
Rights and permissions
About this article
Cite this article
Li, F., Xu, Y. & Liu, RL. SAMD5 mRNA was overexpressed in prostate cancer and can predict biochemical recurrence after radical prostatectomy. Int Urol Nephrol 51, 443–451 (2019). https://doi.org/10.1007/s11255-019-02096-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11255-019-02096-3