Abstract
Two Ni(II) adamantane complexes, [Ni(bqad)Cl2] (1) and [Ni(bpad)(dmbp)(H2O)](ClO4)2·CH3OH H2O (2) (bqad = N,N′-bis(2-quinolinylmethyl) amantadine, bpad = N,N′-bis(2-pyridylmethyl)amantadine, dmbp = 5,5′-dimethyl-2,2′-bipyridine) have been synthesized and characterized by elemental analysis, infrared spectroscopy and single crystal X-ray diffraction. The nickel centers in complex 1 have a distorted tetragonal pyramidal geometry, while the coordination polyhedron of 2 can be described as a distorted octahedron. The reaction kinetics for reduction of p-nitrophenol to p-aminophenol catalyzed by these complexes has been investigated by UV–visible spectrophotometry. Complex 1 exhibits a higher turnover frequency of 1.4 min−1 for the reduction of p-nitrophenol.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nickel(II) complexes have attracted much attention in recent years because of their potential applications in molecule-based magnetic materials, water oxidation catalysis, asymmetrical catalysis, nuclease mimics and artificial superoxide dismutases [1–5]. Nickel complexes of quinoline-based ligands have been found to be effective catalysts for the coupling of aryl chlorides and opening/closing reactions of the tetrahydroquinazoline ring [6, 7]. Nickel(II) complexes of aromatic ligands can have potent catalytic activity toward C–N coupling reactions [8], methyl methacrylate polymerization [9], as superoxide dismutase mimics [10], and for the visible-light-driven production of hydrogen from water [11]. Meanwhile, p-nitrophenol (p-NP) is among the most notorious water pollutants with carcinogenic character and high toxicity because of its high solubility and stability in water [12, 13]. Moreover, p-aminophenol (p-AP) as the reduced product has commercial importance as an intermediate for the preparation of antipyretic and analgesic drugs [14, 15]. Therefore, the reduction of p-NP to p-AP is interesting for both the environmental protection and prodrug synthesis. The various reported catalysts for hydrogenation reactions of p-NP have been focused on gold, silver, platinum, palladium nanoparticles and other noble metal complexes [16, 17]; hence, their widespread application is limited by their high cost. Therefore, the design and synthesis of stable, noble-metal-free catalysts with high catalytic efficiency is desirable. Recently, several nickel-based nanoparticles and Ni(II) clusters have been reported to exhibit high catalytic performance for the reduction of p-NP [18, 19]. Therefore, the design and synthesis of nickel complexes as catalysts for p-nitrophenol reduction have received increasing attention.
In previous studies, we synthesized copper complexes of amantadine-based tripodal ligands and investigated their catalytic activities in dismutation of the superoxide anion [20, 21]. Adamantane is a highly symmetrical aliphatic hydrocarbon constructed from three fused symmetrical cyclohexane rings in the chair configuration, whose derivatives have been extensively applied in the field of clinic medicine [22, 23]. In this paper, we report the design and synthesis of two Ni(II) adamantane complexes, [Ni(bqad)Cl2] (1) and [Ni(bpad)(dmbp)(H2O)](ClO4)2·CH3OH H2O (2) (bqad = N,N′-bis(2-quinolinylmethyl) amantadine, bpad = N,N′-bis(2-pyridylmethyl)amantadine, dmbp = 5,5′-dimethyl-2,2′-bipyridine), which have been characterized by X-ray crystallography. Furthermore, their catalytic activities for the p-NP reduction have been investigated. The results show that nickel complex 1 can efficiently catalyze reduction of p-NP to p-AP in the presence of NaBH4 as the reducing agent.
Experimental
Materials and instrumentation
Organic reagents were of reagent grade, and solvents used in this research were purified by standard procedures. Water used in all physical measurements was double-distilled. The free ligands bqad and bpad were synthesized according to our previous work [20, 21] and were characterized by their elemental analysis and 1H NMR spectra. Elemental analyses for C, H and N were obtained on a Vario EL III instrument. As shown in appendix A of the supplementary material, infrared spectroscopy on KBr pellets was performed on a Shimadzu IR Prestige-21 infrared spectrophotometer from 4000 to 400 cm−1. The electronic spectra were recorded on a Shimadzu UV-2450 spectrophotometer.
Synthesis of complex 1
To a solution of NiCl2·6H2O (0.237 g, 1.0 mmol) in methanol (15 mL) was slowly added a solution of bqad (0.433 g, 1.0 mmol) in chloroform (5 mL) with stirring. After refluxing for 2 h, the solution was filtered and the filtrate was kept at room temperature for slow evaporation. After a week, dark blue block crystals suitable for X-ray analysis were obtained in 77% yield. Anal. Calcd. for C30H31Cl2N3Ni: C 63.98, N 7.46, H 5.55%; Found: C 63.62, N 7.33, H 5.82%. IR (KBr disc, cm−1): 3388(m), 2904(vs), 2852(s), 1599(vs), 1509(vs), 1425(s), 1367(m), 1199(w),1135(m), 947(m), 851(vs), 786(vs), 748(vs), 696(w), 631(w), 579(w).
Synthesis of complex 2
To a solution of Ni(ClO4)2·6H2O (0.365 g, 1.0 mmol) in methanol (20 mL) was slowly added a solution of bpad (0.333 g, 1 mmol) and bmpy (0.184 g, 1.0 mmol) in acetonitrile (10 mL) with stirring. After refluxing for 2 h, the solution was filtered and the filtrate was kept at room temperature for slow evaporation. After three days, blue hexagonal crystals suitable for X-ray analysis were obtained in 72% yield. Anal. Calcd. for C35H45Cl2N5NiO10: C 50.44, N 8.40, H 5.44%; Found: C 50.27, N 8.32, H 5.73%. IR (KBr disc, cm−1): 3443(s), 2903(vs), 2842(s), 1606(vs), 1481(vs), 1444(s), 1388(m), 1243(w),1093(vs), 923(w), 829(s), 767(s), 624(s).
X-ray crystallography
Single crystals of complexes 1 and 2 were used for X-ray diffraction analyses by mounting on the tip of a glass fiber in air. Data were recorded with a Bruker Smart Apex CCD diffractometer with graphite-monochromated Mo Kα radiation (λ = 0.71073 Å). Diffraction intensities were collected by the ω-scan technique. The reflections were corrected for Lorentz and polarization effects, and empirical absorption corrections were applied using the SADABS program [24]. Structure solutions were obtained readily using SHELXTL (XS) [25]. Hydrogen atoms were placed in idealized positions and were set as riding on the respective parent atoms. All non-hydrogen atoms were refined with anisotropic thermal parameters. The structures were refined (weighted least squares refinement on F 2) to convergence [26]. Olex2 was employed for the final data presentation. For complex 2, the anion ClO4 − and methanol solvent were refined with a disordered model. The occupancies of different parts were refined with free parameters. The ADP restrains (Rigu) and distances restrains (DFIX) were used to get better ADPs of the oxygen atoms. After treating the disorder, the SQUEEZE program in PLATON was used to exclude the highly disordered water molecule with a Q peak of 1.2 eÅ−3, which was judged to be negligible. Further crystallographic data and experimental details for structural analyses of both complexes are summarized in Table 1. Selected bond distances and angles are listed in Table 2. CCDC numbers are 1478260 for 1 and 1478261 for 2, respectively. These data can be obtained free of charge via http://www.ccdc.cam.ac.uk/conts/retrieving.html.
Catalytic reduction of p-NP
The reduction of p-nitrophenol (p-NP) to p-aminophenol (p-AP) with NaBH4 was investigated at ambient temperature. The UV–Vis absorption spectroscopy was used to evaluate the catalytic activity and stability, as the reactant p-NP has a strong absorption peak at 400 nm, while the product p-AP has a medium absorption peak at 300 nm. Stock solutions of p-NP (3 × 10−4 mol L−1), NaBH4 (3 × 10−2 mol L−1), complex 1 and 2 (3 × 10−5 mol L−1) were prepared. For the study of reaction kinetics, 1.0 mL of p-NP stock solution was transferred to a quartz optical cell and 1.0 mL of NaBH4 stock solution was added. Upon addition of the reducing agent, the band at λ = 400 nm remained unaltered until addition of the nickel catalyst stock solution (1 mL). Subsequently, the electronic spectrum of the reaction mixture was recorded every 60 s.
Results and discussion
Crystal structure analysis
The structures of both of these nickel(II) complexes have been confirmed by X-ray diffraction. ORTEP views of both complexes are shown in Fig. 1, together with the atomic numbering scheme. The crystallographic and refinement data are listed in Table 1. Selected bond lengths and bond angles for both complexes are listed in Table 2. Complexes 1 and 2 crystallize in the orthorhombic space group Pbca and tetragonal space group P42 /n, respectively. The nickel atom in each asymmetrical unit in 1 is penta-coordinated by a [N3Cl2] set, provided by three nitrogen atoms from a bqad ligand and two chloride anions. The asymmetrical unit in 2 consists of a Ni(II) center, bpad and dmbp ligands, one water ligand, two uncoordinated perchlorate anions and one methanol molecule. The Ni(II) in complex 2 is coordinated by two bipyridine N atoms, plus two pyridine N atoms of the bpad, together defining the equatorial plane. A water O atom and the tertiary amine N atom of the bpad ligand occupy the two axial positions, in an overall octahedral geometry. The axial bonds (Ni(1)–N(2) = 2.260 Å) are longer than the basal bonds (Ni(1)–N(1) = 2.060 Å).
For complex 1, the penta-coordinate geometry can be assessed by the trigonal index (τ = β − α)/60), where β and α are the largest coordination angles. The values of τ are 0 and 1 for a regular tetragonal pyramid and a perfect trigonal bipyramid, respectively [27]. The τ value of complex 1 is 0.47, indicating the distorted tetragonal pyramidal geometry. Three nitrogen donor atoms from the bqad ligand and one chloride ligand occupy the four corners of the square plane, while the remaining chloride ligand is located at the axial position. As expected, the axial Ni–Cl bond length in 1 is longer than other basal bonds in the coordination sphere [Ni(1)–Cl(1) = 2.3373(6) Å and Ni(1)–Cl(2) = 2.2860(6) Å], comparable to the values reported for other Ni(II) complexes with chloride co-ligands [28, 29]. The bonds lengths [Ni–N = 2.060(3) − 2.260(3) Å] in both 1 and 2 are similar to those reported for Ni(II) complexes of similar ligands [30, 31]. The length of Ni–Ow in 2 is 2.133(3) Å, which is in the normal range of Ni–O distances determined for other mononuclear Ni(II) complexes [32].
In order to explain the structural variations of the V-shaped ligand bqad and its Ni(II) complexes, the forming angle of three centroids from one N atom and two quinoline rings in bqad, called α V, is obtained from the crystal structure [33]. The V-shaped angle (α V) of free bqad is 101.63° [21], while the α V value for Ni(II) complex 1 is 110.48°. This shows that α V increases such that the two quinolone rings rotate around the σ type of C–C bond upon coordination. The dihedral angle between the two quinolone rings in the free ligand bqad is 74.615(33)° [21], whereas the value in 1 is 8.747(38)°. The change in α V and rotation of the aromatic rings can be ascribed to the electronic and steric effect as bqad distorts in order to construct Ni–N coordination bonds. As shown in Fig. 2, complex 1 is further assembled into a two-dimensional framework along the c-axis through offset face-to-face π–π stacking interactions of neighboring quinoline rings, with interplanar distances of 4.9524(16) and 5.6934(13) Å for 1.
UV–Vis and IR spectroscopy
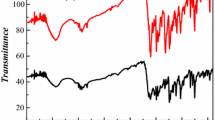
The electronic absorption spectra of both complexes in freshly prepared acetonitrile solution were obtained at room temperature. As shown in Fig. 3, the electronic spectra of the free ligand bqad displayed two sharp absorption peaks at 212 and 230 nm, which are assigned to absorptions of the phenanthroline chromophore. A broad band at 273 nm in the spectrum of bqad corresponds to an n–π* transition. New broad features at 304 and 310 nm for complex 1 and 2, respectively, are attributed to the LMCT transitions [34]. The FTIR spectra of complexes 1 and 2 (Fig. S1) showed sharp intense peaks at about 2903 and 2842 cm−1 due to symmetrical and asymmetrical CH stretches, respectively, of the adamantyl group. In order to study the binding state of the ligands, the IR spectra of the free ligands were compared with those of the complexes [35]. A peak at 1593 cm−1 for free ligand bqad is ascribed to the C=N breathing vibration of the quinoline group, which was shifted toward lower frequency at 1599 cm−1 in complex 1. A peak at 1588 cm−1 for free ligand bmpy is assigned to the stretching vibrations of the pyridine ring, which is shifted to 1606 cm−1 in complex 2. Sharp peaks at 1093 and 624 cm−1 in complex 2 are assigned to perchlorate anion [36], indicating that ClO4 − exists as the anion of equilibrium charge.
Catalytic degradation experiments
Catalytic reduction of p-NP to p-AP in the presence of NaBH4 has been extensively used to evaluate the activities of Ni-based catalysts in aqueous solution [37]. The catalytic reactions were monitored by UV–Vis spectroscopy at 1-min intervals. The reactant p-NP has a strong characteristic absorption peak at 400 nm, while the product p-AP has a medium absorption peak at 300 nm [38]. As shown in Fig. S2, almost no change in the peak at 400 nm in the absence of catalyst was observed after 24 h, indicating that the reduction does not proceed in the presence of NaBH4 alone. However, after addition of both complexes, as shown in Fig. S3, the intensity of the peak at 400 nm quickly decreased while the characteristic absorption of p-aminophenol at 300 nm appeared. The maximum conversions catalyzed by complexes 1 and 2 were up to 96.8% in 7 min and 89.7% in 11 min, respectively. Since the concentration of NaBH4 is 100-fold higher than that of p-NP, the rate of p-AP reduction can be evaluated by pseudo-first-order kinetics as follows:
here c t is the concentration of p-NP and k a is the rate constant. The p-NP concentration is proportional to the intensity of the peak at 400 nm. Thus, the above equation can be expressed as following:
A t and A 0 are the intensities of the p-NP absorption peak at t = t and t = 0, respectively. Fig. S4 shows plots of ln(A 0/A t) versus time for the reductions of p-NP catalyzed by complexes 1 and 2. The linear correlations indicate that the reaction follows first-order kinetics, and thus, the values of k a for complexes 1 and 2 are 7.1 × 10−3 and 2.7 × 10−3 s−1, respectively, as determined from the linear slopes. Furthermore, the turnover frequency (TOF) was calculated to determine the efficiencies of the catalysts. The TOF values of complexes 1 and 2 were 1.4 and 0.8 min−1, respectively, calculated as the moles of p-NP reduced per mole of nickel complex per unit time. As shown in Table 3, our nickel complexes showed higher catalytic efficiencies than most of other catalysts for the reduction of p-NP [39–43]. The higher catalytic activity of complex 1 for the reduction of p-NP compared with complex 2 could be attributed to the fact that complex 1 has an unsaturated coordination sphere and smaller steric hindrance compared with complex 2.
Conclusion
In summary, two Ni(II)-amantadine complexes have been synthesized and characterized by single crystal X-ray diffraction. The results of catalytic kinetics studies indicate that both complexes are active for the reduction of p-nitrophenol to p-aminophenol, and their catalytic performances vary with the coordination configuration and steric hindrance.
References
Zheng YZ, Zheng ZP, Chen XM (2014) Coord Chem Rev 258:1–15
Singh A, Spiccia L (2013) Coord Chem Rev 257:2607–2622
Ritleng V, Henrion M, Chetcuti MJ (2016) ACS Catal 6:890–906
Desbouis D, Troitsky IP, Belousoff MJ, Spiccia L, Graham B (2012) Coord Chem Rev 256:897–937
Chatterjee SK, Maji RC, Barman SK, Olmstead MM, Patra AK (2014) Angew Chem Int Ed 53:10184–10189
Zhang Q, Zhang XQ, Wang ZX (2012) Dalton Trans 41:10453–10464
Garcia-Deibe AM, Sanmartin-Matalobos J, Gonzalez-Bello C, Lence E, Portela-Garcia C, Martinez-Rodriguez L, Fondo M (2012) Inorg Chem 51:1278–1293
Shi J, Li FL, Li HX, Wang F, Yu H, Ren ZG, Zhang WH, Lang JP (2014) Inorg Chem Commun 46:159–162
Miao LL, Li HX, Yu M, Zhao W, Gong WJ, Gao J, Ren ZG, Wang HF, Lang JP (2012) Dalton Trans 41:3424–3430
Chai LQ, Zhang HS, Huang JJ, Zhang YL (2015) SpectroChim Acta A 137:661–669
Yuan YJ, Lu HW, Tu JR, Fang Y, Yu ZT, Fan XX, Zou ZG (2015) ChemPhysChem 16:2925–2930
Zhao S, Ma H, Wang M, Cao C, Xiong J, Xu Y, Yao S (2010) Photochem Photobiol Sci 9:710–715
Shilpa ML, Gayathri V (2016) Transit Metal Chem 41:393–401
Alexander S, Udayakumar V, Gayathri V (2011) Transit Metal Chem 37:1–6
Rode CV, Vaidya MJ, Jaganathan R, Chaudhari RV (2001) Chem Eng Sci 56:1299–1304
Jiang F, Li RM, Cai JH, Xu W, Cao AM, Chen DQ, Zhang X, Wang CR, Shu CY (2015) J Mater Chem A 3:19433–19438
Yang Y, Jin RX, Zhao S, Liu JH, Li YF, Yu XD, Shi Z, Xing Y (2016) Rsc Adv 6:32580–32585
Yang L, Kong J, Zhou D, Ang JM, Phua SL, Yee WA, Liu H, Huang Y, Lu X (2016) Chem Eur J 20:7776–7783
Yang Y, Ren Y, Sun C, Hao S (2014) Green Chem 16:2273–2280
Zhou YH, Liu XW, Chen LQ, Wang SQ, Cheng Y (2016) Polyhedron 117:788–794
Zhou YH, Tao J, Sun DL, Chen LQ, Jia WG, Cheng Y (2015) Polyhedron 85:849–856
Wang J, Ma C, Wang J, Jo H, Canturk B, Fiorin G, Pinto LH, Lamb RA, Klein ML, DeGrado WF (2013) J Med Chem 56:2804–2812
Davies WL, Grunert RR, Haff RF, McGahen JW, Neumayer EM, Paulshock M, Watts JC, Wood TR, Hermann EC, Hoffmann CE (1964) Science 144:862–863
Sheldrick GMS (1996) Program for Scaling and Correction of Area Detector Data. University of Göttingen, Göttingen
Sheldrick GM (2015) Acta Cryst C71:3–8
Dolomanov OV, Bourhis LJ, Gildea RJ, Howard JAK, Puschmann H (2009) J Appl Cryst 42:339–341
Addison AW, Rao TN, Reedijk J, van Rijn J, Verschoor GC (1984) Dalton Trans 1349–1356. doi:10.1039/DT9840001349
Batisai E, Lusi M, Jacobs T, Barbour LJ (2012) Chem Commun 48:12171–12173
Drahoš B, Herchel R, Trávníèek Z (2015) Inorg Chem 54:3352–3369
Jany T, Horstmann née Gruschka C, Bögge H, Stammler A, Glaser T (2015) Z Anorg Allg Chem 641:2157–2168
Powell-Jia DA, Pham MTN, Ziller JW, Borovik AS (2010) Inorg Chim Acta 363:2728–2733
Song X, Wen H, Ma C, Chen H, Chen C (2015) New J Chem 39:1734–1741
Wu HL, Wang H, Wang XL, Pan GL, Shi FR, Zhang YH, Bai YC, Kong J (2014) New J Chem 38:1052–1061
Wang Y, Xiao W, Mao JW, Zhou H, Pan ZQ (2013) J Mol Struct 1036:361–371
Patzer A, Schütz M, Möller T, Dopfer O (2012) Angew Chem Int Ed 51:4925–4929
Dobrzyñska D, Janczak J, Wojciechowska A, Helios K (2010) J Mol Struct 973:62–68
Zhu M, Zhou S, Yao C, Liao L, Wu Z (2014) Nanoscale 6:14195–14199
Kalarivalappil V, Divya CM, Wunderlich W, Pillai SC, Hinder SJ, Nageri M, Kumar V, Vijayan BK (2016) Catal Lett 146:474–482
Zhang P, Shao C, Zhang Z, Zhang M, Mu J, Guo Z, Liu Y (2011) Nanoscale 3:3357–3363
Sun T, Zhang Z, Xiao J, Chen C, Xiao F, Wang S, Liu Y (2013) Sci Rep 3:2527
Kandula S, Jeevanandam P (2016) Eur J Inorg Chem 2016:1548–1557
Demirci S, Sahiner N (2015) Water Air Soil Pollut 226:1–13
Wang Z, Fu H, Han D, Gu F (2014) J Mater Chem A 2:20374–20381
Acknowledgements
This work was supported by the Natural Science Foundation of Anhui Province (No. 1408085MKL21), Special and Excellent Research Fund of Anhui Normal University, Postgraduate Scientific Research and Innovation Project of Anhui Normal University (No. 2015cxsj142), and Undergraduate Innovative training program of Anhui Normal University (201610370486, 201610370490). Thanks to Dr. Fengfeng Wang (Institute of Materia Medica, Chinese Academy of Medical Science and Peking Union Medical College) for his expert crystallographic analysis.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhou, YH., Wang, SQ., Chen, LQ. et al. Effective reduction of p-nitrophenol catalyzed by nickel(II) adamantane complexes. Transit Met Chem 42, 175–180 (2017). https://doi.org/10.1007/s11243-017-0122-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11243-017-0122-3