Abstract
The present study describes a new regeneration system based on somatic embryogenesis from mature endosperm Passiflora cincinnata Mast. cultures. Moreover, the morpho-agronomic and phenological traits, as well as enzymatic activity of regenerated triploid emblings are compared to those of diploids. Mature endosperms were cultured on Murashige and Skoog medium supplemented with various concentrations (4.5–45.2 µM) of 2,4-dichlorophenoxyacetic acid (2,4-D) and 4.5 μM 6-benzylaminopurine (BA). No plant growth regulators were included in the control group. Embryogenic calli were observed only in treatments supplemented with 13.6 and 18.1 µM 2,4-D + 4.5 µM BA, with the highest number of somatic embryos per explant and regenerated plants (emblings) obtained with 18.1 µM 2,4-D. Most emblings (70%) were triploid (2n = 3x = 27), with a DNA amount (4.38 pg) similar to that of endosperm and 1.5 times greater than in diploid P. cincinnata seedlings (2n = 2x = 18), that contained 2.98 pg of DNA. While the number of organs and/or structures was akin to that in diploids, triploid emblings generally exhibited larger and longer vegetative and floral structures. The flower lifespan was also slightly altered by triploidy, nectar concentration was 27% higher, and the activity of plant defense enzymes β-1,3-glucanase and polyphenol oxidase was 29.8% and 22.1% higher. This study describes a new regeneration system for the production of phenotypic variants of this ornamental passion fruit species, opening new perspectives for future studies on genetic passion fruit breeding.
Key message
Mature endosperms of Passiflora cincinnata display high embryogenic competence when cultivated in an auxin-rich induction medium and the regenerated triploid emblings are morphologically distinct from their diploid counterparts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polyploidy is a strategic tool in plant breeding programs (Sattler et al. 2016; Touchell et al. 2020). Polyploid plants have more than two sets of chromosomes, and this condition often promotes desirable morpho-agronomic characteristics, favoring the development of more productive and/or more adaptable cultivars (Osborn et al. 2003). Polyploid organisms can arise in natural populations (Soltis and Burleigh 2009; Van de Peer et al. 2017) or be induced via somatic hybridization (Grosser and Gmitter 2011). The latter can occur from in vitro cultures of tissues with different ploidy levels (Thomas and Chaturvedi 2008; Wang et al. 2016; Abdullah et al. 2021) or from in vitro and ex vitro tissues treated with antimitotic agents (Dhooghe et al. 2011; Sumarji and Suparno 2017).
Genome duplication can cause significant genetic and epigenetic changes, possibly leading to phenological, phenotypic, and physiological alterations (Adams and Wendel 2005; Otto and Whitton 2007; Yang et al. 2011; Van de Peer et al. 2017). Triploid plants often have larger vegetative and reproductive organs, thicker, darker-colored leaves, and longer-lasting flowers. In addition, they display improved vigor and greater tolerance to biotic and abiotic factors, which confers superior adaptation to a wide range of environmental stresses; however, they are generally sterile owing to gamete imbalance (Comai 2005; Ranney 2006; Sun et al. 2011; Padoan et al. 2013; Santana-Vieira et al. 2016; Wang et al. 2016).
Given the remarkable features displayed by triploid individuals, plant regeneration systems based on in vitro endosperm culture have been established to obtain genotypes with larger vegetative and reproductive organs (ornamental plants) and seedless fruits (fruit crops) (Kumar et al. 2015; Wang et al. 2016; Iannicelli et al. 2020). The endosperm is a nutrient-rich reserve tissue found in angiosperm seeds, which is closely related to the growth and development of zygotic embryos. It forms following the fusion of a haploid spermatic nucleus with the two nuclei from the central cell of the embryo sac during fertilization. This step results in a single, naturally triploid tissue (Cailleau et al. 2010), explaining why endosperm in vitro culture offers a direct regeneration system for triploid plants (Thomas and Chaturvedi 2008; Wang et al. 2016).
Plant regeneration from the endosperm requires that parenchyma storage cells acquire competence to reprogram themselves and take on a new developmental route. Cellular competence is acquired by hormonal signals from auxins, such as 2,4-dichlorophenoxyacetic acid (2,4-D), 1-naphthaleneacetic acid, or indole-3-butyric acid, and cytokinins, such as 6-benzylaminopurine (BA) or kinetin, which are added to the culture medium (Hoshino et al. 2011). De novo shoot organogenesis (DNSO) constitutes the main regeneration pathway during in vitro endosperm culture (Hoshino et al. 2011). However, regeneration can proceed also from somatic embryos. Owing to the complexity of this morphogenetic process, whereby two meristematic poles (shoot and root apical meristems) need to be established synchronously, few studies have reported the induction of somatic embryogenesis from endosperm culture (Kosmiatin et al. 2014; Sukmara et al. 2014).
In Passiflora (Passifloraceae), a tropical genus valued for the production of passion fruit and its ornamental potential, regeneration of triploid plants by endosperm culture was reported in 1996 by Mohamed et al. However, it is only in recent years that this biotechnological tool has been tried in P. edulis Sims (Antoniazzi et al. 2018), the main commercial species of the genus, as well as P. cincinnata Mast. (Silva et al. 2020) and P. foetida L. (Mikovski et al. 2021), two ornamental species (Bernacci et al. 2005; Mikovski et al. 2019). In all these systems, regeneration occurred from adventitious buds in mature endosperms cultivated in cytokinin-rich induction medium.
Passion fruit somatic embryogenesis was induced successfully from in vitro cultures of zygotic embryos (Rocha et al. 2020). Based on the established regeneration system for P. cincinnata, in which somatic embryos were formed in induction medium supplemented with 18.1 µM 2,4-D and 4.5 µM BA (Silva et al. 2009), embryogenic cultures were optimized for various Passiflora spp. using 8.8–72.4 μM 2,4-D and 4.5 μM BA (Paim Pinto et al. 2011; Rosa et al. 2015; Prudente et al. 2017). P. cincinnata is a model species for passion fruit morphogenetic studies (Dias et al. 2009; Otoni et al. 2013; Vieira et al. 2018; Silva et al. 2020). Interestingly, the anthers of this species display high embryogenic competence (Silva et al. 2021) when cultivated in induction medium supplemented with 18.1 µM 2,4-D and 4.5 µM BA. Given the close ontogenetic, physiological, and functional relationship between zygotic embryos and endosperm, as well as the high embryogenic responsiveness of P. cincinnata tissues when cultured in medium containing a 4:1 ratio of auxin (2,4-D) to cytokinin (BA), we hypothesized that these culture conditions could induce the formation of somatic embryos from P. cincinnata endosperm storage cells.
In the present study, we describe a system for the regeneration of P. cincinnata triploid plants, based on new morphogenetic pathways. Moreover, to facilitate the use of specific genotypes in passion fruit breeding programs, we compare the morpho-agronomic, phenological, and physiological traits of diploid and triploid plants.
Materials and methods
Plant material
P. cincinnata seeds were isolated from ripe fruits and dried at room temperature. The seed coat was removed with the aid of a mini-vise. Then, the seeds were immersed in 70% (v/v) alcohol for 3 min and disinfected for 25 min with 100 mL of 2.5% (v/v) commercial NaClO containing three drops of Tween-20. The seeds were rinsed four times in distilled, autoclaved water and immersed for 10 h to facilitate endosperm isolation.
Induction of somatic embryogenesis
For the induction of somatic embryogenesis, endosperms were aseptically isolated from zygotic embryos and cultured in 90 × 15 mm polystyrene Petri dishes containing 30 mL of induction medium composed of MS basic salts (Calsson Labs, Smithfield, UT, USA) (Murashige and Skoog 1962), 0.01% (w/v) myo-inositol (Synth, Diadema, SP, Brazil), 3% (w/v) sucrose (Exodo Cientifica Ltda., Sumaré, SP, Brazil), and 8 g/L agar (Acumedia®; Neogen, Lansing, MI, USA). Culture medium was supplemented with different concentrations of 2,4-D (4.5, 9.0, 13.6, 18.1, 22.6, 27.1, 31.7, 36.2, and 45.2 µM) and 4.5 µM BA. No plant growth regulators (PGRs) were added to the control group. The pH was adjusted to 5.7 ± 0.1, and the culture medium was autoclaved at 121 °C and 1.1 atm for 20 min. Each treatment consisted of five Petri dishes with 10 endosperms each. The dishes were kept in a culture room in the absence of light and at 25 ± 2 °C for 30 days.
Embryogenic calli were transferred to Petri dishes containing maturation medium consisting of MS salts as described above, but without PGRs. The dishes were kept in a growth room under the same environmental conditions as described above, but with a 16-h photoperiod and 36 μmol/m2/s irradiance (Special Day Light, 20 W; Osram, Barueri, SP, Brazil) for 45 days.
Cotyledonary stage somatic embryos were isolated and subcultured in flasks containing MS-half strength medium, 0.01% (w/v) myo-inositol, 3% (w/v) sucrose, 8 g/L agar, and no PGRs. Emblings of roughly 10 cm in length, obtained after 90 days of cultivation, were transferred to plastic cups containing Plantmax HT® substrate (Eucatex Agro, Paulínia, SP, Brazil). The plastic cups were covered with transparent plastic bags and kept under a 16-h photoperiod and 25 ± 2 °C for 20 days.
Embling acclimatization
P. cincinnata emblings (n = 10), obtained from different embryogenic lines, measuring approximately 30 cm were transferred to 30-dm3 plastic pots filled with a 3:1 mixture of soil and substrate, and grown in a greenhouse. To compare the morpho-agronomic features of diploid and triploid plants, ten seeds of P. cincinnata were germinated in black polyethylene plastic bags of 28 × 14 cm containing Plantmax HT® substrate, kept in a greenhouse, and irrigated daily. After germination, three diploid plants were also cultivated under the same conditions and used as controls. The pots were spaced 1.5 m between rows and 4 m between plants. The conduction system was in a vertical spreader with a smooth wire at a height of 1.8 m, without elimination of terminal buds and lateral branches.
The foliar application of fertilizer was carried out monthly in the vegetative phase and every 15 days in the reproductive phase, at a dosage of 3–9 L/ha. The fertilizer consisted of N = 10%; P2O5 = 10%; K2O = 10%; Fe = 0.1%; Zn = 0.1%; Cu = 0.1%; S = 0.3%; and Mn = 0.5%.
Flow cytometry
For flow cytometry, approximately 20–30 mg of endosperm tissue, young and fresh leaves from diploid seedlings (control), and endosperm-derived plants regenerated by somatic embryogenesis (emblings) were chopped with a disposable steel razor blade in 1 mL WPB buffer to release the nuclei (Loureiro et al. 2007). Pisum sativum (2C DNA content = 1.96 pg) was used as an internal reference standard (Doležel 1997). Ten samples of each material were analyzed.
Previously macerated tissues were filtered through a 50-μm nylon mesh and collected in a polystyrene tube. The filtrate was stained with 25 μL propidium iodide solution (1 mg/mL; Sigma-Aldrich, St. Louis, MO, USA) supplemented with 2.5 µL RNase (20 mg/L). At least 10,000 nuclei per sample were analyzed in triplicate. Data were acquired using a CytoFLEX instrument (Beckman Coulter Life Sciences, Indianapolis, IN, USA) at the Institute of Biological Sciences of the Federal University of Juiz de Fora, and then plotted and analyzed using CytExpert 2.0.1 software. Nuclear DNA content (pg) was estimated using the following equation (Doležel and Bartos 2005): Sample (2C DNA) = G1 peak channel of sample × 9.09 (P. sativum DNA content) / G1 peak channel of P. sativum.
Cytogenetic analyses
Cytogenetic analyses were performed in seedlings (diploid control) and those emblings (n = 3) that, based on flow cytometry analysis, contained more DNA than the seedlings. Slides were prepared from young root tips pre-treated with 3 mM 8-hydroxyquinoline at 4 °C for 8 h and fixed in 3:1 (v/v) ethanol/acetic acid for 24 h at -20 °C. The root tips were submitted to enzymatic maceration in 4% (w/v) cellulase (Onozuka R-10; Serva, Heidelberg, Germany) and 40% (v/v) pectinase (Sigma-Aldrich) for 3 h at 37 °C. Cytological preparations were performed as described by Carvalho and Saraiva (1993), and stained with 4,6-diamidino-2-phenylindole. Slides were imaged using a BX51 fluorescence microscope with CellSens software (Olympus, Tokyo, Japan).
Morpho-agronomic characterization
Comparison of the morpho-agronomic characteristics of seed-derived diploid plants and endosperm-derived triploid P. cincinnata plants (n = 3) was carried out following the instructions for distinguishability, homogeneity, and stability assays proposed by the National Service for the Protection of Cultivars (Brasil 2016) for wild ornamental species and interspecific hybrids of the genus Passiflora, as well as the descriptors proposed by Fonseca et al. (2017) and Nobrega et al. (2017). The triploid plants were obtained from different explants. Comparisons were carried out only on plants that were already in the reproductive stage, and included 29 morpho-agronomic descriptors. The descriptors were evaluated based on the structures found in the middle third of each plant. Ten leaves and flowers from each plant were analyzed. Plant dimensions were measured with the aid of a digital caliper and an analog refractometer (Akrom, São Leopoldo, RS, Brazil) to quantify the Brix of the floral nectar. Values reported here represent the average of ten repetitions per genotype (diploid or triploid), and each repetition corresponds to the average of ten measurements of vegetative or reproductive structures per plant.
Phenological observations were also carried out within a period of 30 days to determine the average opening and closing times of the flowers. In total, three flowers per plant were evaluated, with 30 flowers per genotype.
Enzymatic evaluation
The activities of β-1,3-glucanases, polyphenol oxidase and peroxidase were quantified due to their involvement with disease resistance mechanisms in plants. To evaluate the enzymatic activity of diploid and triploid plants, we collected two leaves per plant (n = 3) from the middle third of the plant, placed them in labeled aluminum envelopes, snap-froze in liquid nitrogen, and stored at −80 °C.
The activity of β-1,3-glucanases was determined by colorimetric quantification of reducing sugars released from laminarin (Lever 1972). The protein extract was obtained by adding 2 mL of extraction medium containing 50 mM sodium phosphate buffer (pH 6.5), 1 mM phenylmethylsulfonyl fluoride, and 200 mg polyvinylpolypyrrolidone. For the reaction, 230 μL of 0.1 M sodium acetate buffer (pH 5.0), 250 μL laminarin substrate solution (4 mg/mL) and 20 μl plant extract were mixed and incubated at 45 °C for 30 min. Then, the reaction was stopped by the addition of 500 μL of 3,5-dinitrosalicylic acid, heated to 100 °C for 5 min, and cooled to 30 °C. Absorbance was read at 540 nm. The amount of reducing sugars formed was quantified using a standard curve of glucose concentrations. Results were expressed in mg glucose/min/mg protein.
Polyphenol oxidase activity was determined according to Duangmal and Apenten (1999), and the protein extract was obtained by macerating the leaf sample in 0.1 M sodium acetate buffer (pH 5.0). For the reaction, 100 μL enzymatic extract, 900 μL of 0.1 M sodium phosphate buffer (pH 6.8), and 20 mM catechol were mixed, readings were taken at 420 nm, and values were expressed as absorbance/min/mg fresh weight.
For the determination of peroxidase activity, the extract was obtained by maceration of leaf samples in liquid nitrogen followed by extraction in 0.1 M sodium acetate buffer (pH 5.2) containing 1 mM ethylenediaminetetraacetic acid. For the reaction, 2.9 mL of 50 mM potassium phosphate buffer (pH 6.0), 20.2 mM guaiacol, 90 mM H2O2, and 10 μL protein extract were mixed (Chance and Maehly, 1955). Optical density values at 470 nm were obtained using a spectrophotometer (FEMTO, São Paulo, SP, Brazil), and were expressed as absorbance/min/mg fresh weight.
Experimental design
The induction of somatic embryogenesis followed a completely randomized experimental design with five Petri dishes containing ten endosperms representing an experimental unit. The experiment included nine treatments (2,4-D concentrations) and a control (without PGRs). The evaluated parameters were explant responsiveness after 30 days of culture, mean number of somatic embryos per explant after 45 days in maturation medium, and mean number of regenerated plants per explant after 90 days of culture. Tukey’s test with a 5% significance level was used.
The morpho-agronomic descriptors and mean enzymatic activities of diploid (seed-derived) and triploid plants were submitted to a normality and homogeneity test (Bartlett), and evaluated by Student’s t-test to verify significant differences between the means of the two genotypes. A value of P < 0.05 was considered statistically significant. In case of an asymmetric pattern, data were subjected to logarithmic transformation and submitted to a non-parametric test using the Genes Program (Cruz, 2016). In addition, morpho-agronomic descriptors of diploid and triploid P. cincinnata plants were evaluated by principal component analysis (PCA) using the pcaMethods package in R (Stacklies et al. 2007), following z-score transformation of the data set (Worley and Powers 2013).
Results
Somatic embryogenesis is induced by specific auxin:cytokinin ratios
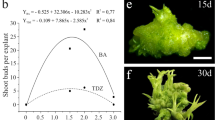
P. cincinnata endosperms were responsive to auxin-rich induction medium (Fig. 1a). However, yellowish pro-embryogenic masses were observed only in treatments supplemented with 13.6 and 18.1 µM 2,4-D + 4.5 µM BA (Fig. 1b). Differentiation of somatic embryos in the early stages of development could be observed after 15 days of in vitro culture (Fig. 1a). The highest number of somatic embryos (8.8 ± 1.3) was detected in induction medium supplemented with 18.1 µM 2,4-D + 4.5 µM BA (Fig. 1b). Control explants showed no response after 30 days of culture (data not shown).
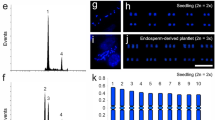
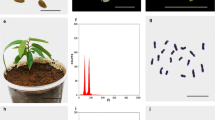
Somatic embryogenesis, nuclear DNA content, and chromosome assessment of Passiflora cincinnata emblings. a Overview of the regeneration process via somatic embryogenesis. From left to right: endosperm (en) (bar = 5 mm); embryonic callus showing a heart-shaped somatic embryo (h) (bar = 5 mm); torpedo (t) embryo stage with callus (ca) (bar = 2.5 mm); cotyledonary (co) embryo stage (bar = 5 mm); young embling (bar = 50 mm); and embling in the reproductive phase in the field (bar = 110 mm). b Number of somatic embryos per explant after 45 days in maturation medium. c Number of regenerated plants per explant. d Histograms comparing nuclear DNA content of P. cincinnata seedlings (upper panel), emblings (lower panel), and Pisum sativum (internal standard) determined using propidium iodide (PI) staining. e Metaphase plate showing 18 chromosomes of P. cincinnata seedlings. f Metaphase plate showing 27 chromosomes of P. cincinnata emblings. Bars (e, f) = 5 µm
Emblings were obtained from somatic embryos after 90 days of maturation in the absence of PGRs and germination in half-strength medium (Fig. 1a). The highest mean number of regenerated plants (7.0 ± 1.2) was obtained from explants previously cultivated in induction medium supplemented with 18.1 μM 2,4-D and 4.5 μM BA (Fig. 1c).
Even though the regenerated plants were acclimatized to greenhouse conditions, only 30% of them survived. Endosperm-derived emblings grown in the greenhouse entered the flowering phase 210 days after initiation of the somatic embryogenesis induction process (Fig. 1a).
Most endosperm-derived emblings are triploids
Flow cytometry analysis revealed differences in the amount of DNA between seedlings and emblings of P. cincinnata (Fig. 1d). P. cincinnata seedlings contained 2.98 pg of DNA, with a coefficient of variation of 2.6% (Fig. 1d). Instead, 70% of tested P. cincinnata emblings had on average 4.38 pg of DNA, or roughly 1.5 times more than diploid plants, and a coefficient of variation of 2.8% (Fig. 1d). The remaining 30% of emblings contained a similar amount of DNA as seedlings (data not shown).
P. cincinnata seedlings displayed 18 chromosomes (2n = 2x = 18) (Fig. 1e); whereas emblings with higher DNA content by flow cytometry exhibited 27 chromosomes (2n = 3x = 27) (Fig. 1f), confirming triploidy.
Morpho-agronomic characterization of P. cincinnata diploid and triploid plants
The greatest morpho-agronomic differences between diploid and triploid plants were observed with respect to the dimensions of vegetative and reproductive organs, while the number of organs or structures was the same (Table 1). Both genotypes displayed pentalobed leaves in the middle third of the plants (Fig. 2a). The leaves of triploid plants had larger petioles (Fig. 2b) and longer and wider leaf blades (Fig. 2c, d), but the number of extrafloral nectaries in the petiole and number of leaf blades were analogous (Table 1). The branches were cylindrical and predominantly dark green in diploid plants, but ranged in color from dark to purplish green (data not shown) and displayed a 17% larger diameter in triploid emblings (Fig. 2e).
Vegetative organs of diploid and triploid Passiflora cincinnata plants. a Leaf morphology. b Petiole length. c, d Leaf blade length (c) and width (d). e Stem width. Error bars denote the standard error. Asterisks refer to statistical differences between diploid and triploid plants within each parameter (t-test). Bar = 50 mm.
Morphological differences between diploid and triploid plants were even more evident in their reproductive organs (Fig. 3a). All quantitative descriptors, with the exception of style length, anther width, nectar chamber width, ovary length, ovary diameter, and peduncle length, were significantly higher in triploid plants (Fig. 3b–k; Table 1). Color intensity of floral organswas apparently greater in triploid plants (Fig. 3a), although it has not been quantified.
Flower features of diploid and triploid Passiflora cincinnata plants. a Flower organ morphology highlighting corona filaments (cf), androgynophore (a), ovary (ov), sepal (se), petal (pe), stamen filament (stf), and style (sty). b Hypanthium diameter. c Corona diameter. d Androgynophore length. e Ovary diameter. f Sepal length. g Sepal width. h Petal length. i Petal width. j Stamen filament length. k Style length. Error bars denote the standard error. Asterisks refer to statistical differences between diploid and triploid plants within each parameter (t-test). Bars = 20 mm
Flower lifespan and floral nectar concentration are altered by triploidy
Flowering of diploid and triploid plants occurred after 210 days of culture. It took 22–26 days for diploid plants and 17–21 days for triploids to move from the production of flower buds to anthesis. Anthesis of diploid flowers occurred at approximately 5:20 a.m. and they remained open for 14 h. In triploid emblings, anthesis occurred at 05:00 a.m. and the flowers remained open for up to 15 h. Nectar concentration was 27% higher in triploid emblings (39.26 ± 1.88% Brix) than in diploid plants (30.91 ± 1.4% Brix).
The activity of plant defense enzymes is affected by ploidy level
The activities of β-1,3-glucanase and polyphenol oxidase were 29.8% and 22.1% higher in triploid emblings, respectively, compared to diploid P. cincinnata (Fig. 4a, b). A similar trend was observed for peroxidase activity, although the difference was not significant (Fig. 4c).
Enzymatic activity of diploid and triploid Passiflora cincinnata plants. a β-1,3-glucanase. b Polyphenol oxidase. c Peroxidase. Abs, absorbance. FW, fresh weight. Glu, glucose. Error bars denote the standard error. Asterisks refer to statistical differences between diploid and triploid plants within each parameter (t-test)
Diploid and triploid P. cincinnata plants are morpho-physiologically distinct
Based on morphological characteristics, nectar concentration, and enzymatic activity, PCA analysis revealed that the first two principal components (PC1 and PC2) accounted for 84.2% of total variation (Fig. 5), with PC1 largely explaining the separation between triploids and diploids. Almost all evaluated traits displayed higher values in triploid plants.
Discussion
The DNSO-based regeneration of P. cincinnata triploid plants has been previously reported for endosperms cultivated in a cytokinin-rich medium (Silva et al. 2020). In the present study, we describe an alternative in vitro regeneration system capable of inducing somatic embryogenesis by altering the relationship between exogenously added PGRs and the light regimen. Whereas DNSO was previously induced by cultivating mature endosperms in medium supplemented with 9 µM BA under a 16:8 h photoperiod (Silva et al. 2020); here, somatic embryos were obtained through cultivation in medium supplemented with a higher auxin:cytokinin ratio (18.1 µM 2,4-D + 4.5 µM BA) in the absence of light (Fig. 6).
Schematic diagram summarizing the regeneration pathways established from embryonic tissues of Passiflora cincinnata. (1) Somatic embryogenesis from mature endosperm culture (present study). (2) De novo shoot organogenesis from mature endosperm culture (Silva et al. 2020). (3) Somatic embryogenesis from mature zygotic embryo culture (Silva et al. 2009)
The regeneration of endosperm-derived triploid emblings is uncommon, and existing reports point to the need to combine two or more PGRs to induce somatic embryogenesis. In most of these studies, auxins are required for the induction of embryogenic calli in in vitro endosperm cultures (Thomas and Chaturvedi 2008) and 2,4-D is the main PGR used for this purpose, regardless of explant type or plant species (Yang and Zhang, 2010; Nic-Can and Loyola-Vargas 2016). Interestingly, the highest number of somatic embryos and P. cincinnata emblings regenerated from endosperm culture was observed under the same growth conditions previously described for the induction of embryogenic cultures from zygotic embryos of this species (Silva et al. 2009; Paim Pinto et al. 2011; Vieira et al. 2018). Endosperms and zygotic embryos are embryonic structures with a shared origin linked to double fertilization and similar physiology. The endosperm plays an important role in supporting embryo growth by supplying nutrients and producing signals to regulate embryo development (Yan et al. 2014). These results suggest the conservation of hormonal signaling and similar endogenous conditions for the expression of totipotency during induction of somatic embryogenesis. Accordingly, Passiflora embryonic tissues could be used as simple and efficient systems for the regeneration of diploid and triploid plants (Fig. 6).
Determining the amount of nuclear DNA and the number of chromosomes is essential for the validation of polyploid production/regeneration systems. In the present study, 70% of endosperm-derived emblings had a 1.5 times higher DNA content (2n = 3x = 27) than their respective diploids (2n = 2x = 18). Compared to the DNSO pathway (Silva et al. 2020), the number of triploid plants obtained via somatic embryogenesis is slightly smaller, although still satisfactory, as no mixoploidy, aneuploidy, or ploidy above the 3 × level was observed among regenerated emblings. In Actinidia deliciosa, for example, most endosperm-derived plants were aneuploids (Góralski et al. 2005); whereas in Azadirachta indica, both diploid and triploid plants were obtained from endosperm-derived calli (Chaturvedi et al. 2003), as observed here.
Vegetative and reproductive organs of P. cincinnata were differently sized with respect to ploidy level. Triploid plants exhibited generally larger and longer vegetative and floral structures than their diploid counterparts, but an equal number of lateral organs. Higher ploidy induces an increase in cell volume to accommodate polyploid tissues (Malladi and Hirst 2010; Wu et al. 2013), leading to larger and more voluminous leaves, flowers, stems, and roots (Dhooghe et al. 2011; Kumar et al. 2015). These events can contribute to the development of new genotypes and phenotypes and, consequently, to greater intraspecific genetic diversity (Padoan et al. 2013; Serapiglia et al. 2015; Spoelhof et al. 2017).
In addition to greater floral size, triploidy changed also the phenology of P. cincinnata emblings. Floral phenology is frequently altered by polyploidy. According to Rezende et al. (2020), the flower traits most frequently affected by hybridization and polyploidy were floral morphology (83.8%), color (59.46%), and phenology (27.02%). In addition to morphological changes, the flower buds of triploid plants developed more quickly, showed a longer flower opening period, and higher Brix values than diploids. These modifications can affect the communication between flowers and pollinators, as well as accessibility of the pollinator to the floral reward, compromising the plant-pollinator match (Cortis et al. 2009; Van der Niet et al. 2014; Rezende et al. 2020). Future studies will determine if the floral changes observed in endosperm-derived plants have any specific effect on the P. cincinnata plant-pollinator interaction.
Triploid plants of P. cincinnata exhibited higher glucanase and polyphenol oxidase activities compared to diploid plants. These oxidative enzymes are important plant defense mechanisms directly associated with cell wall degradation by pathogens, disease susceptibility, and senescence (Thipyapong et al. 2004; Araji et al. 2014). At present, we cannot confirm that P. cincinnata triploid plants are more tolerant to pathogens compared to their respective diploids, as such experiments were beyond the scope of the present study. However, numerous studies have demonstrated that increased ploidy levels could strengthen enzyme activity, improving survival and resistance to drought (Padoan et al. 2013; del Pozo and Ramirez-Parra 2014), cold (Deng et al. 2012; Bertrand et al. 2015), salt stress (Chao et al. 2013; Elmaghrabi et al. 2013; Nezhad and Mansouri 2019), and even pathogens (Hias et al. 2018).
Stronger enzymatic activity in polyploid organisms is accompanied by upregulation of the respective genes (Caverzan et al. 2012; Wu et al. 2013), although it is not necessarily proportional to the DNA content of these organisms. In this study, the enzymatic activities of glucanase and polyphenol oxidase were 20% to 30% higher in triploid organisms. Chromosomal duplication may increase enzymatic activity, although it may not necessarily double it; some enzymes may not be affected at all and others may actually exhibit lower expression (Xing et al. 2011). Even though duplication does not introduce new genetic material and only generates additional copies of genes and chromosomes, genome alterations could disturb gene or enzyme patterns (Ranney 2006). The effects of neofunctionalization, mutations, and transposable elements in coding regions of regulatory genes, as well as epigenetic mechanisms, which are common in polyploids, may also contribute to altered gene expression profiles (Osborn et al. 2003).
In summary, we describe here an alternative regeneration system based on somatic embryogenesis that leads to triploid P. cincinnata plants from endosperm culture. We also show that the organs of these triploid emblings are larger and possess greater enzymatic activity than their dipoloid counterparts. We believe that this regeneration system will enable the production of phenotypic variants of this species, and stimulate future genetic studies on passion fruit breeding.
Data availability
Not applicable.
Code availability
Not applicable.
References
Abdullah M, Sliwinska E, Góralski G, Latocha P, Tuleja M, Widyna P, Popielarska-Konieczna M (2021) Effect of medium composition, genotype and age of explant on the regeneration of hexaploid plants from endosperm culture of tetraploidkiwiberry (Actinidia arguta). Plant Cell Tissue Organ Cult 147:569–582. https://doi.org/10.1007/s11240-021-02149-5
Adams KL, Wendel JF (2005) Polyploidy and genome evolution in plants. Curr Opin Plant Biol 8:135–41. https://doi.org/10.1016/j.pbi.2005.01.001
Antoniazzi CA, Faria RB, Carvalho PP, Mikovski AI, Carvalho IF, Matos EM, Otoni WC (2018) In vitro regeneration of triploid plants from mature endosperm culture of commercial passionfruit (Passiflora edulis Sims). Sci Hortic 238:408–415. https://doi.org/10.1016/j.scienta.2018.05.001
Araji S, Grammar TA, Gertzen R, Anderson SD, Mikulic-Petkovsek M, Veberic R, Phu ML, Solar A, Leslie CA, Dandekar AM, Escobar MA (2014) Novel roles for the polyphenol oxidase enzyme in secondary metabolism and the regulation of cell death in walnut. Plant Physiol 164:1191–1203. https://doi.org/10.1104/pp.113.228593
Bernacci LC, Melleti LMM, Soares-Scott MD, Passos IRS, Junqueira NTV (2005) Espécies de maracujá: caracterização e conservação da biodiversidade. In: Faleiro FG, Junqueira NTV, Braga MF (eds) Maracujá: germoplasma e melhoramento genético, 1st edn. EMBRAPA Planaltina, pp 559–586
Bertrand B, Bardil A, Baraulle H, Dusset S, Doulbeau S, Dubois E, Severac D, Dereeper A, Etienne H (2015) The greater phenotypic homeostasis of the allopolyploid Coffea arabica improved the transcriptional homeostasis over that of both diploid parents. Plant Cell Physiol 56:2035–2051. https://doi.org/10.1093/pcp/pcv117
Brasil (2016) Ministério da Agricultura, Pecuária e Abastecimento. Proteção de cultivares. Instruções para execução dos ensaios de distinguibilidade, homogeneidade e estabilidade de cultivares de maracujá (Passiflora L. e híbridos específicos) exceto Passiflora edulis Sims.
Cailleau A, Cheptou PO, Lenormand T (2010) Ploidy and the evolution of endosperm of flowering plants. Genetics 184:439–453. https://doi.org/10.1534/genetics.109.110833
Carvalho CRDE, Saraiva LS (1993) A new heterochromatin banding pattern revealed by modified HKG banding technique in maize chromosomes. Heredity 70:515–519. https://doi.org/10.1038/hdy.1993.74
Caverzan A, Passaia G, Rosa SB, Ribeiro CW, Lazzarotto F, Margis-Pinheiro M (2012) Plant responses to stresses: role of ascorbate peroxidase in the antioxidant protection. Genet Mol Biol 35:1011–1019. https://doi.org/10.1590/S1415-47572012000600016
Chance B, Maehly AC (1955) Assay of catalases and peroxidases. Methods Enzymol 2:764–775. https://doi.org/10.1016/S0076-6879(55)02300-8
Chao DY, Dilkes B, Luo H, Douglas A, Yakubova E, Lahner B, De S (2013) Polyploids exhibit higher potassium uptake and salinity tolerance in Arabidopsis. Science 341:658–659. https://doi.org/10.1126/science.1240561
Chaturvedi R, Razdan MK, Bhojwani SS (2003) Production of haploids of neem (Azadirachta indica A. Juss.) by anther culture. Plant Cell Rep 21:531–537. https://doi.org/10.1007/s00299-002-0565-6
Comai L (2005) The advantages and disadvantages of being polyploid. Nat Rev Genet 6:836–846. https://doi.org/10.1038/nrg1711
Cortis P, Vereecken NJ, Schiestl FP, Lumaga MRB, Scrugli A, Cozzolino S (2009) Pollinator convergence and the nature of species’ boundaries in sympatric Sardinian Ophrys (Orchidaceae). Ann Bot 104:497–506. https://doi.org/10.1093/aob/mcn219
Cruz CD (2016) Genes software—extended and integrated with the R, matlab and selegen. Acta Sci Agron 38:547–552. https://doi.org/10.4025/actasciagron.v38i3.32629
del Pozo JC, Ramirez-Parra E (2014) Deciphering the molecular bases for drought tolerance in Arabidopsis autotetraploids. Plant Cell Environ 37:2722–2737. https://doi.org/10.1111/pce.12344
Deng B, Du W, Liu C, Sun W, Tian S, Dong H (2012) Antioxidant response to drought, cold and nutrient stress in two ploidy levels of tobacco plants: low resource requirement confers polytolerance in polyploids. Plant Growth Regul 66:37–47. https://doi.org/10.1007/s10725-011-9626-6
Dhooghe E, Laere KV, Eeckhaut T, Leus L, Uylenbroeck JV (2011) Mitotic chromosome doubling of plant tissues in vitro. Plant Cell Tissue Organ Cult 104:359–373. https://doi.org/10.1007/s11240-010-9786-5
Dias LLC, Santa-Catarina C, Ribeiro DM, Barros RS, Floh ELS, Otoni WC (2009) Ethylene and polyamine production patterns during in vitro shoot organogenesis of two passion fruit species as affected by polyamines and their inhibitor. Plant Cell Tissue Organ Cult 99:199–208. https://doi.org/10.1007/s11240-009-9594-y
Doležel J (1997) Applications of flow cytometry for the study of plant genomes. J Appl Genet 38:285–302
Doležel LJ, Bartos J (2005) Plant DNA flow cytometry and estimation of nuclear genome size. Ann Bot 95:99–110. https://doi.org/10.1093/aob/mci005
Duangmal K, Apenten RKO (1999) A comparative study of polyphenoloxidases from taro (Colocasia esculenta) and potato (Solanum tuberosum var. Romano). Food Chem 64:351–359
Elmaghrabi AM, Ochatt S, Rogers HJ, Francis D (2013) Enhanced tolerance to salinity following cellular acclimation to increasing NaCl levels in Medicago truncatula. Plant Cell Tissue Organ Cult 114:61–70. https://doi.org/10.1007/s11240-013-0306-2
Fonseca KG, Faleiro FG, Junqueira NTV, Barth M, Feldberg NP (2017) Caracterização morfoagronômica e molecular de cultivares de maracujá ornamental. Pesqui Agropecu Bras 52:849–860. https://doi.org/10.1590/S0100-204X2017001000004
Góralski G, Popielarska M, Slesak H, Siwinska D, Batycka M (2005) Organogenesis in endosperm of Actinidia deliciosa cv. Hayward cultured in vitro. Acta Biol Crac Ser Bot 47:121–128
Grosser JW, Gmitter FG (2011) Protoplast fusion for production of tetraploids and triploids: applications for scion and rootstock breeding in citrus. Plant Cell Tissue Organ Cult 104:343–357. https://doi.org/10.1007/s11240-010-9823-4
Hias N, Svara A, Keulemans JW (2018) Effect of polyploidisation on the response of apple (Malus × domestica Borkh.) to Venturia inaequalis infection. Eur J Plant Pathol 151:515–526. https://doi.org/10.1007/s10658-017-1395-2
Hoshino Y, Miyashita T, Thomas TD (2011) In vitro culture of endosperm and its application in plant breeding: approaches to polyploidy breeding. Sci Hortic 130:1–8. https://doi.org/10.1016/j.scienta.2011.06.041
Iannicelli J, Guariniello J, Tossi VE, Regalado JJ, Di Ciaccio L, van Baren CM, Pitta Álvarez SI, Escandón AS (2020) The “polyploid effect” in the breeding of aromatic and medicinal species. Sci Hortic 260:108854. https://doi.org/10.1016/j.scienta.2019.108854
Kosmiatin M, Purwito A, Wattimena GA, Mariska I (2014) Induksi Embriogenesis Somatik dari Jaringan Endosperma Jeruk Siam (Citrus nobilis Lour.) cv Simadu. J Agron Indones 42:44–51
Kumar N, Singh AS, Kumari S, Reddy MP (2015) Biotechnological approaches for the genetic improvement of Jatropha curcas L.: a biodiesel plant. Ind Crops Prod 76:817–828
Lever M (1972) A new reaction for colorimetric determination of carboydrates. Anal Biochem 47:273–279
Loureiro J, Rodriguez E, Doležel J, Santos C (2007) Two new nuclear isolation buffers for plant DNA flow cytometry: a test with 37 species. Ann Bot 100:875–888. https://doi.org/10.1093/aob/mcm152
Malladi A, Hirst PM (2010) Increase in fruit size of a spontaneous mutant of “Gala” apple (Malusx domestica Borkh.) is facilitated by altered cell production and enhanced cell size. J Exp Bot 61:3003–3013. https://doi.org/10.1093/jxb/erq134
Mikovski AI, Silva NT, Souza CS, Machado MD, Otoni WC, Carvalho IF, Rocha DI, Silva ML (2019) Tissue culture and biotechnological techniques applied to passion fruit with ornamental potential: an overview. Ornam Hortic 25:189–199. https://doi.org/10.14295/oh.v25i2.2036
Mikovski AI, Silva NT, Silva LAS, Machado M, Barbosa LCS, Reis AC, Matos EM, Viccini LF, Souza CS, Machado MD, Otoni WC, Carvalho IF, Rocha DI, Silva ML (2021) From endosperm to triploid plants: a stepwise characterization of the de novo shoot organogenesis and morpho-agronomic aspects of an ornamental passion fruit (Passiflora foetida L.). Plant Cell Tissue Organ Cult 147:239–253. https://doi.org/10.1007/s11240-021-02120-4
Mohamed ME, Hicks RGT, Blakesley D (1996) Shoot regeneration from mature endosperm of Passiflora foetida. Plant Cell Tissue Organ Cult 46:161–164. https://doi.org/10.1007/BF00034851
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Nezhad FS, Mansouri H (2019) Effects of polyploidy on response of Dunaliella salina to salinity. J Mar Biolog Assoc UK 99:1041–1047. https://doi.org/10.1017/S0025315418001005
Nic-Can GI, Loyola-Vargas VM (2016) The role of the auxins during somatic embryogenesis. In: Loyola-Vargas V, Ochoa-Alejo N (eds) Somatic embryogenesis: fundamental aspects and applications. Springer, Cham, pp 171–182
Nobrega DS, Peixoto JR, Vilela MS, Faleiro FG, Gomes KPS, Sousa RMD, Nogueira I (2017) Agronomic descriptors and ornamental potential of passion fruit species. Ornam Hortic 23:357–362. https://doi.org/10.14295/oh.v23i3.1053
Osborn T, Pires C, Birchler JA, Auger DL, Chen ZJ, Li HS, Cornai L, Madlung A, Doerge RW, Colot V, Martienssen RA (2003) Understanding mechanisms of novel gene expression in polyploids. Trends Genet 19:141–147. https://doi.org/10.1016/S0168-9525(03)00015-5
Otoni WC, Pinto DPL, Rocha DI, Vieira LM, Dias LLC, Silva ML, Silva CVE, Lani RG, Silva LC, Tanaka FAO (2013) Organogênese e embriogênese somática em maracujá (Passiflora spp.). In: Aslam J, Srivastava OS, Sharma MP (eds) Somatic embryogenesis and gene expression, 1st edn. Narosa Publishing House, Nova Delhi, pp 1–17
Otto SP, Whitton J (2007) The evolutionary consequences of polyploidy. Cell 131:452–462. https://doi.org/10.1016/j.cell.2007.10.022
Padoan D, Mossad A, Chiancone B, Germana MA, Khan PSSV (2013) Ploidy levels in Citrus clementine affects leaf morphology, stomatal density and water content. Theor Exp Plant Physiol 25:283–290. https://doi.org/10.1590/S2197-00252013000400006
Paim Pinto DL, Almeida AMR, Rêgo MM, Silva ML, Oliveira EJ, Otoni WC (2011) Somatic embryogenesis from mature zygotic embryos of commercial passionfruit (Passiflora edulis Sims) genotypes. Plant Cell Tissue Organ Cult 107:521–530. https://doi.org/10.1007/s11240-011-0003-y
Prudente DO, Paiva R, Carpentier S, Swennen R, Nery FC, Silva LC, Panis B (2017) Characterization of the formation of somatic embryos from mature zygotic embryos of Passiflora ligularis Juss. Plant Cell Tissue Organ Cult 131:97–105. https://doi.org/10.1007/s11240-017-1266-8
Ranney TG (2006) Polyploidy: from evolution to new plant development. Combined Proceedings International Plant Propagators Society 56:137–142
Rezende L, Suzigan J, Amorin FW, Moraes AP (2020) Can plant hybridization and polyploidy lead to pollinator shift? Acta Bot Bras 34:229–242. https://doi.org/10.1590/0102-33062020abb0025
Rocha DI, Batista DS, Faleiro FG, Rogalski M, Ribeiro LM, Mercadante-Simões MO, Yockteng R, Silva ML, Soares WS, Pinheiro MVM, Pacheco TG, Lopes AS, Viccini LF, Otoni WC (2020) Maracujá: Passiflora spp. In: Litz RA, Pliego-Alfaro F, Hormaza JI (eds) Biotechnology of fruit and nut crop, 2nd edn. CABI, Wallingford, pp 381–408
Rosa YBCJ, Monte-Bello CC, Dornelas MC (2015) Species-dependent divergent responses to in vitro somatic embryo induction in Passiflora spp. Plant Cell Tissue Organ Cult 120:69–77. https://doi.org/10.1007/s11240-014-0580-7
Santana-Vieira DDS, Freschi L, Almeida LA, Moraes DHS, Neves DM, Santos LM (2016) Survival strategies of citrus rootstocks subjected to drought. Sci Rep 6:38775. https://doi.org/10.1038/srep38775
Sattler MC, Carvalho CR, Clarindo WR (2016) The polyploidy and its key role in plant breeding. Planta 243:281–296. https://doi.org/10.1007/s00425-015-2450-x
Serapiglia MJ, Gouker FE, Hart JF, Unda F, Mansfield SD, Stipanovic AJ, Smart LB (2015) Ploidy level affects important biomass traits of novel shrub willow (Salix) hybrids. Bioenergy Res 8:259–269. https://doi.org/10.1007/s12155-014-9521-x
Silva ML, Paim Pinto DL, Guerra MP, Floh EIS, Bruckner CH, Otoni WC (2009) A novel regeneration system for wild passion fruits species (Passiflora cincinnata Mast.) based on somatic embryogenesis from mature zygotic embryos. Plant Cell Tissue Organ Cult 99:47–54. https://doi.org/10.1007/s11240-009-9574-2
Silva NT, Silva LAS, Reis AC, Machado MD, Matos EM, Viccini LF, Otoni WC, Carvalho IF, Rocha DI, Silva ML (2020) Endosperm culture: a facile and efficient biotechnological tool to generate passion fruit (Passiflora cincinnata Mast.) triploid plants. Plant Cell Tissue Organ Cult 142:613–624. https://doi.org/10.1007/s11240-020-01887-2
Silva ML, Pinto DLP, Campos JMS, Carvalho IF, Rocha DI, Batista DS, Otoni WC (2021) Repetitive somatic embryogenesis from wild passion fruit (Passiflora cincinnata Mast.) anthers. Plant Cell Tissue Organ Cult 146:635–641. https://doi.org/10.1007/s11240-021-02083-6
Soltis DE, Burleigh JG (2009) Surviving the K-T mass extinction: new perspectives of polyploidization in angiosperms. Proc Natl Acad Sci USA 106:5455–5456. https://doi.org/10.1073/pnas.0901994106
Spoelhof JP, Soltis PS, Soltis DE (2017) Pure polyploidy: closing the gaps in autopolyploid research. J Syst Evol 55:340–352. https://doi.org/10.1111/jse.12253
Stacklies W, Redestig H, Scholz M, Walther D, Selbig J (2007) pcaMethods—a bioconductor package providing PCA methods for incomplete data. Bioinformatics 23:1164–1167. https://doi.org/10.1093/bioinformatics/btm069
Sukmara E, Sukamto LA, Bintang M (2014) Induksidankarakterpertumbuhankalus triploid dariendospermaavokad (Persea americana Mill.). Curr Biochem 1:20–28
Sumarji MP, Suparno MH (2017) The effectiveness of colchisin giving on watermelon ploidization (Citrullus vulgaris Schard). Int J Appl Environ 12:1951–1967
Sun DQ, Lu XH, Liang GL, Guo QG, Mo YW, Xie JH (2011) Production of triploid plants of papaya by endosperm culture. Plant Cell Tissue Organ Cult 104:23–29. https://doi.org/10.1007/s11240-010-9795-4
Thipyapong P, Hunt MD, Steffens JC (2004) Antisense downregulation of polyphenol oxidase results in enhanced disease susceptibility. Planta 220:105–117
Thomas TD, Chaturvedi R (2008) Endosperm culture: a novel method for triploid plant production. Plant Cell Tissue Organ Cult 93:1–14. https://doi.org/10.1007/s11240-008-9336-6
Touchell DH, Palmer IE, Ranney TG (2020) In vitro ploidy manipulation for crop improvement. Front Plant Sci 11:722. https://doi.org/10.3389/fpls.2020.00722
Van De Peer Y, Mizrachi E, Marchal K (2017) The evolutionary significance of polyploidy. Nat Rev Genet 18:411–424. https://doi.org/10.1038/nrg.2017.26
Van der Niet T, Peakall R, Johnson SD (2014) Pollinator-driven ecological speciation in plants: new evidence and future perspectives. Ann Bot 113:199–211. https://doi.org/10.1093/aob/mct290
Vieira LM, Silva PO, Fernandes AM, Rocha DI, Otoni WC (2018) Protocol for somatic embryogenesis in Passiflora cincinnata Mast (Passifloraceae). In: Jain MS, Gupta P (eds) Stepwise protocols for somatic embryogenesis of woody plants, 2nd edn. Springer International Publishing AG, Cham, pp 253–265
Wang X, Cheng ZM, Zhi S, Xu F (2016) Breeding triploid plants: a review. Czech J Genet Plant Breed 52:41–54. https://doi.org/10.17221/151/2015-CJGPB
Worley B, Powers R (2013) Multivariate analysis in metabolomics. Curr Metabolomics 1:92–107. https://doi.org/10.2174/2213235x11301010092
Wu L, Han Z, Wang S, Wang X, Sun A, Zu X (2013) Comparative proteomic analysis of the plant-virus interaction in resistant and susceptible ecotypes of maize infected with sugarcane mosaic virus. J Proteom 89:124–140. https://doi.org/10.1016/j.jprot.2013.06.005
Xing Y, Zhao X, Cai L (2011) Prediction of nucleosome occupancy in Saccharomyces cerevisiae using position-correlation scoring function. Genomics 98:359–366. https://doi.org/10.1016/j.ygeno.2011.07.008
Yan D, Duermeyer L, Leoveanu C, Nambara E (2014) The functions of the endosperm during seed germination. Plant Cell Physiol 55:1521–1533. https://doi.org/10.1093/pcp/pcu089
Yang X, Zhang X (2010) Regulation of somatic embryogenesis in higher plants. CRC Crit Rev Plant Sci 29:36–57. https://doi.org/10.1080/07352680903436291
Yang J, Dungrawala H, Hua H, Manukyan A, Abraham L, Lane W, Mead H, Wright J, Schneider BL (2011) Cell size and growth rate are major determinants of replicative lifespan. Cell Cycle 10:144–155. https://doi.org/10.4161/cc.10.1.14455
Acknowledgements
This work was supported by Fundação de Amparo à Pesquisa do Estado de Mato Grosso (FAPEMAT) (Cuiabá, MT); Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brazil (CAPES)—Finance Code 001; and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) (420913/2018-1). We also thank the CNPq for granting a scholarship to MM (DCR-314905/2018-9). We would like to thank Editage (www.editage.com) for English language editing.
Funding
This work was supported by Fundação de Amparo à Pesquisa do Estado de Mato Grosso (FAPEMAT) (Cuiabá, MT), Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG) (Belo Horizonte, MG; Grants APQ-00772–19 and APQ-02581-21), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) (Brasília, DF; Finance Code 001), and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) (Brasília, DF; Grant 420913/2018–1).
Author information
Authors and Affiliations
Contributions
MLS, DIR, and MDM designed the study; MDM and CSS established embryonic cultures; MM performed statistical analyses; ACR, EMM, SMS, and LFV performed cytogenetics and flow cytometry analyses; MDM, CSS, and DIR performed morphometric evaluations; IFC and MDM carried out enzymatic analyses; MDM, DIR, MLS, and WCO wrote the manuscript; MM, IFC, ACR, SMS, and LFV revised the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
All co-authors approved the final version of the manuscript.
Additional information
Communicated by Mohammad Reza Abdollahi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Machado, M.D., Souza, C.S., Machado, M. et al. Novel avenues for passion fruit in vitro regeneration from endosperm culture, and morpho-agronomic and physiological traits of triploid Passiflora cincinnata Mast. emblings. Plant Cell Tiss Organ Cult 150, 637–650 (2022). https://doi.org/10.1007/s11240-022-02318-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-022-02318-0