Abstract
Steviol glycosides (SGs) and gibberellins (GAs) share the same molecular basis. However, the coordination of their respective biosynthetic pathways is very intriguing. Thus, the present study aimed to investigate the role of plant growth regulators, gibberellic acid (GA3), chlorocholine chloride (CCC), and paclobutrazol (PBZ), on the metabolism of Stevia rebaudiana and identify possible ameliorates of the evaluated parameters when CCC and PBZ-treated plants were subsequently treated with GA3. For this, explants were cultured in the absence or presence of 2 mg L−1 GA3, CCC, or PBZ (Step 1). After 20 days, half explants incubated with CCC and PBZ were treated with 2 mg L−1 GA3 and the other half, as well as the rest of the explants, were sub-cultured in their respective initial conditions for 20 days (Step 2). GA3-treated plants showed increased stevioside and phenolic compounds content, as well as a downregulation of most of the SGs/GAs biosynthesis-related genes, with a more pronounced effect upstream of steviol. Following this trend, CCC downregulated some MEP pathway genes, including SrDXS, SrDXR, SrCDPS, and SrKS, and upregulated SrUGT6G1. PBZ also upregulated SrUGT76G1 and inhibited five genes of the MEP pathway and all genes coding for kaurenoid pathway enzymes. The obtained results highlight the capability of GA3 to reverse the negative effects of PBZ on the pattern of many transcripts and to additionally increase the stevioside content to levels comparable to those found in field-grown plants.
Graphic abstract

Key message
A feedback loop in in vitro cultured stevia plants in response to GA3, CCC and PBZ was demonstrated. GA3-treated plants showed increases in stevioside and phenolic compound contents.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stevia rebaudiana Bertoni, commonly referred as sweet leaf, is a herbaceous perennial plant of the Asteraceae family native to the South American region (Paraguay and Brazil) and has a great importance due to its sweet glycosides, called steviol glycosides (SGs) (Geuns 2003). Steviol glycosides are a group of tetracyclic diterpenes, having 100–300 times the sweetness of sucrose (Karimi et al. 2019) and a wide spectrum for many therapeutic applications (Ullah et al. 2019). Additionally, evidence has shown that stevia is safe for human consumption and does not show side effects or cause health risks (Chughtai et al. 2020).
Steviol glycosides and gibberellins (GAs) share the same biosynthetic 2-C-methyl-D-erythritol-4-phosphate (MEP) pathway in plastids that culminates with the formation of the IPP and DMAPP (Totté et al. 2003). After the formation of these two basic isoprene units, geranylgeranyl diphosphate (GGDP) is synthesised in a reaction catalysed by geranylgeranyl diphosphate synthase (GGDPS). In sequence, GGDP is cyclised and rearranged into (−)-kaurene by the consecutive action of two enzymes namely copalyl diphosphate synthase (CPPS) and kaurene synthase (KS). Downstream of this, (−)-kaurene is oxidised in the next step by kaurene oxidase (KO) to form kaurenoic acid. From this point, divergence of the steviol and gibberellin biosynthesis pathways occurs (Humphrey et al. 2006). 13α-Hydroxylation of kaurenoic acid catalysed by kaurenoic acid 13-hydroxylase (KAH) leads to the formation of steviol, the ‘backbone’ of SGs, whereas the oxidation of kaurenoic acid by kaurenoic acid oxidase (KAO) forms gibberellic acid12 (GA12), which, in turn, is the precursor for the synthesis of all kinds of GAs (Richman et al. 2005; Brandle and Telmer 2007). Finally, glycosylation of steviol governed by cytosolic UDP-glucosyltransferases (UGTs), SrUGT74G1, SrUGT76G1, SrUGT85C2 (Richman et al. 2005), SrUGT91D2 (Wang et al. 2016), and SrUGT93E1 (Li et al. 2018) results in the diversity of SGs depending on the bond type and number of added glucose units (Richman et al. 2005).
Gibberellins are plant hormones that control growth and development of the plant (Hernández-García et al. 2020). Different steps of the GAs biosynthesis pathway can be inhibited by plant growth retardants (Karimi et al. 2019). Chlorocholine chloride (CCC), which has the trade name Cycocel®, and paclobutrazol (PBZ) are two growth retardants known for their anti-GA effects. In addition, CCC and PBZ are molecules with a recognised role in protecting against stress and by their ability to promote secondary metabolism changes. Some experiments have been performed to evaluate the effect of PBZ (Karimi et al. 2014; Lucho et al. 2018a; Hajihashemi 2018) and CCC (Dey et al. 2013a, b; Karimi et al. 2014; Kundu et al. 2014) on the phytochemicals of stevia plants with encouraging results. As far as we know, information concerning the effects of CCC on a molecular level in stevia and others plants is very limited, although this compound is considered the growth retardant most widely used on a global scale (Rademacher 2016). Another question that remains open is about the effect of PBZ on both the synthesis of SGs and the expression of SG-related genes (in the three stages of the SGs biosynthesis pathway) in in vitro cultured stevia plants. Unlike CCC, with only one study that evaluated its effect on the expression of the SrKAH gene (Kundu et al. 2014), at least three studies have been conducted with PBZ in stevia plants (Hajihashemi et al. 2013; Hajihashemi and Geuns 2017; Lucho et al. 2018a).
The effect of GA3 on SG-related gene expression in stevia plants has also been studied, showing increases in the expression of key genes (Kumar et al. 2012; Hajihashemi et al. 2013), although decreased expression would be expected given the possible negative feedback processes. As GAs and SGs compete for the same substrates, a possible downregulation of the SrKAO gene (coding for the first specific GAs synthesis enzyme) would trigger an accumulation of kaurenoic acid (KA). With increased KA availability, the enzyme upstream from steviol biosynthesis (SrKAH) would be ready to catalyse the first step of the SG-specific pathway, increasing the metabolic flux towards the formation of SGs. Thus, starting from the point where GA3, CCC, and PBZ have the potential to manipulate part of the diterpenoids pathway, including SGs formation, we hypothesised that these compounds could improve specialised metabolites accumulation in stevia plants. Therefore, the objective of the present study was to investigate the physiological, biochemical, and molecular responses to GA3, CCC, and PBZ and identify possible ameliorates on the evaluated parameters when CCC- and PBZ-treated plants were subsequently treated with GA3.
Material and methods
Plant material and in vitro culture of Stevia rebaudiana
Stevia rebaudiana Bertoni plants were collected in the Pelotas region, south of Brazil (31°42′43.2″S 52°10′28.7″W). This specie was previously identified, and then, a voucher was deposited in the Federal University of Pelotas herbarium under number #26,649, as well as registration (A3AF671), in the National System of Management of Genetic Heritage and Associated Traditional Knowledge (SisGen) was carried out. Plants were cultivated for four months in a greenhouse (28 ± 2 °C temperature and 40–60% relative humidity). For in vitro establishment, aerial branches were rinsed several times under running tap water for 10 min (rustic washing), and then they were taken to the laminar air flow chamber, where they were shaken for 15 min in a calcium hypochlorite 1.5% (v/v) solution containing three drops of Tween® 20 surfactant per litre. The stem portions were rinsed thrice with sterilised distilled water, and then, the explants were pre-treated with a modified Murashige and Skoog medium (MS; Murashige and Skoog 1962) with macronutrients at half-strength (MS/2) containing 4000 µL L−1 Plant Preservative Mixture® for 90 min. After disinfection, nodal explants were excised and placed in test tubes containing growth regulator-free MS/2 medium, supplemented with 250 mg L−1 casein hydrolysate, 30 g L−1 sucrose, 1 g L−1 charcoal, and 7.3 g L−1 agar and the pH adjusted to 5.8 before autoclaving (1 atm, 120 °C, 20 min). Cultures were incubated in a growth room at 25 ± 2 °C under a photon flux density of 48 μmol m−2 s 1 (16 h light / 8 h dark). After in vitro establishment, plants were in vitro propagated using glass flasks under the same conditions as described above. Aseptic and healthy in vitro plantlets were used for further studies.
Treatments and growth determinations
Node explants (2.0 cm long) were cultured in MS/2 nutrient medium in the absence (control, CON) or presence of 2 mg L−1 GA3, CCC, or PBZ (Step 1). After 20 days, half explants cultured in MS/2 media containing CCC and PBZ were sub-cultured in media containing 2 mg L−1 GA3 and the rest of explants, including those from GA3, were sub-cultured in the same conditions as step 1 (Step 2). The experiment was therefore comprised of the following treatments (Step 1 medium + Step 2 medium): T1- Control (CON + CON); T2- CCC (CCC + CCC); T3- PBZ (PBZ + PBZ); T4- GA3 (GA3 + GA3); T5- CCC + GA3, and T6- PBZ + GA3 (Supplementary Fig. 1). Flasks covered with aluminium foil were incubated under the same light and temperature conditions described above. The number of nodal segments, shoots, leaves, and roots, as well as root length and shoot height, from every explant were assessed in a non-destructive way at the end of each of the two steps, 20 days (Step 1) and 40 days (Step 2) after the beginning of the experiment. The experiment was repeated twice.
RNA extraction, cDNA synthesis, and RT-qPCR reactions
Leaf samples (located at the 3rd and 4th nodes from the apex) were harvested after 40 days (Step 2) and immediately frozen in liquid nitrogen. Total RNA was isolated and quantified and its quality/integrity checked. cDNA synthesis was carried out as described by Lucho et al. (2018a). Cycling conditions were optimised and are listed in Supplementary Table 1, together with a detailed description of the primers used. Some of the primer sequences were obtained from Modi et al. (2014), and others were designed based on sequences from the Genbank database on the NCBI website. The specificity of the primers used for RT-qPCR reactions was checked (Supplementary Figs. 2 and 3. The housekeeping gene Ubiquitin (UBQ) was selected as an internal standard to normalise the data (Lucho et al. 2018b). Relative gene expression was calculated with the 2−△△Ct method (Livak and Schmittgen 2001). Three technical repetitions were performed for each biological replicate, including samples for the control treatment as a template-free control.
Determination of specialised metabolites
Sample preparation and quantification of SGs via HPLC
The extraction and identification of stevioside was performed according to Lucho et al. (2019) with minor modifications. Steviol glycosides content was determined using an HPLC–DAD Jasco LC-NetII/ADC. Separations were carried out at 40 °C as described by JECFA (2010) on a Luna C-18 reverse-phase column (250 mm × 4 mm, 5 μm) supplied by Phenomenex. A mixture 10 mM phosphate buffer pH 2.6:acetonitrile (68:32, v/v) was used as a mobile phase and applied at a flow rate of 1 mL min−1. Stevioside content was expressed as mg equivalents of rebaudioside A per gram dry weight.
Quantification of total soluble phenol and flavonoid content via spectrophotometry
The determination of total soluble phenol content (TPC) was carried out according to López-Orenes et al. (2013) using gallic acid (0–2000 µM) as a standard, while the flavonoid content (FLA) was determined according to Kim et al. (2003). A standard curve of rutin (quercetin-3-rutinoside) was used for calibration (0–2000 µM). Absorbance readings for TPC and FLA were determined at 765 nm and 510 nm, respectively, using a Ultrospec® 7000/7000PC UV–Visible spectrophotometer. TPC content was expressed as mg of gallic acid equivalents and FLA content as mg of rutin equivalents per gram dry weight.
Statistical analyses
The experiment was set up in a completely randomised factorial design with 6 treatments, and each treatment consisted of five flasks with four explants per flask. The results correspond to the mean ± standard deviation. Analysis of variance (ANOVA) and Tukey’s Test at a 5% probability level (P < 0.05) were performed to calculate the significant differences among treatments using R software. Pearson correlation analysis was performed to examine whether there was a correlation between SGs content, gene transcription, and morphological characteristics. In addition, multivariate principal component analysis (PCA) was applied to detect possible groupings among all variables.
Results
Effects of GA3 and its biosynthesis inhibitors, CCC and PBZ, on Stevia growth parameters
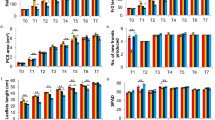
After 20 days of in vitro culture, CCC treatment significantly increased the number of shoots more than two-fold when compared to the control plants (Table 1). Following this trend, the number of both leaves and roots was also significantly higher in CCC-treated plants (10.33 ± 0.11 and 2.89 ± 0.50, respectively) than control plants (6.45 ± 0.23 and 0.56 ± 0.23, respectively). Regarding the number of nodal segments, the highest mean value was observed in PBZ-treated plants (4.44 ± 0.28). Both inhibitors showed a positive effect on the root system with CCC promoting the greatest number of roots and PBZ promoting their growth (4.15 ± 0.39 cm). Overall, at the end of the step 1, no reductions in growth parameters were observed when explants were treated with CCC and PBZ.
At the end of the step 2, the only growth parameter that was not significantly affected by treatment was the number of nodes (Table 2 and Supplementary Fig. 4). PBZ reduced plant height by about 25%, and the application of GA3 was not able to reverse this effect. Notably, PBZ also had negative effects on the root system, presenting less (1.78 ± 0.12) and shorter roots (1.35 ± 0.15 cm) than control plants (5.22 ± 0.23 and 2.95 ± 0.29 cm, respectively). A similar trend was observed in plants treated with GA3 both alone and in combination with CCC (CCC + GA3). The latter being the treatment that provoked a significant reduction of almost all analysed growth parameters. Surprisingly, the CCC treatment did not have a negative effect on plant height, resulting in an average value of 4.48 ± 0.02 cm, which was not significantly different from the value obtained in control plants (4.27 ± 0.19 cm).
Effects of GA3 and its biosynthesis inhibitors, CCC and PBZ, on the transcription of SG-related genes
The evaluation of transcription patterns in the regulatory SG-related genes under the effect of GA3 and its biosynthesis inhibitors CCC and PBZ are shown in Fig. 1 and Supplementary Table 2. The application of CCC caused downregulation of the first two genes in the MEP pathway, SrDXS and SrDXR and the last one, SrHDR, besides upregulation of the SrCMS and SrMCS genes. The negative effect of CCC on gene expression could not be reversed by the presence of GA3. Stevia plants treated only with GA3 showed downregulation of all genes in this pathway, exceto to SrMCS. Although PBZ provoked a noticeable increase in SrMCS gene relative expression, five of the seven genes that make up the MEP pathway were downregulated, including SrDXS, SrDXR, SrCMK, SrHDS, and SrHDR. Interestingly, treatment with GA3, after exposure to PBZ during step 1 reversed this effect, except for the last gene (SrHDR).
Genes encoding kaurenoid enzymes, namely SrGGDPS, SrCDPS, SrKS, and SrKO, were downregulated by both PBZ and GA3 treatments. Although, PBZ downregulated the expression of the four genes, treatment with GA3 in step 2 was able to reverse the negative effects on the expression of SrKS and SrKO. Notably, SrKO encodes the target for the inhibitory action of PBZ. Regarding the effect of CCC on the expression of these four genes, there was a significant decrease in the SrCDPS and SrKS genes’ relative expression (both genes coding for the targets of CCC inhibition). In this case, the presence of GA3 in step 2 did not reverse the negative effect on gene expression.
Concerning results of the relative expression of the six genes involved in the last stage of SGs biosynthesis, none of the treatments positively regulated SrKAH gene expression relative to the control plants. In relation to the expression levels of UGT genes, GA3 alone treatment downregulated SrUGT85C2, SrUGT76G1, SrUGT73E1, and SrUGT91D2 and upregulated SrUGT74G1 gene expression, whereas CCC and PBZ upregulated SrUGT76G1 and SrUGT91D2 and PBZ downregulated SrUGT85C2 and SrUGT73E1 gene expression. Regarding sequential treatments, CCC + GA3 resulted in the lowest levels of expression for all UGT genes. Interestingly, PBZ + GA3 treatment always reversed the negative effects of PBZ and even increased the UGT expression levels when the inhibitor had no apparent effect relative to the control.
Content of specialised metabolites
Effect of GA3, CCC, and PBZ on SGs content
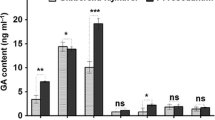
Stevioside content was highest (32.18 ± 0.76 mg rebaudioside A eq. g−1 DW) in plants treated with GA3 alone (Fig. 2). When GA3 was used in step 2, after the application of CCC or PBZ, stevioside content was lower than that of the control (15.56 ± 0.16, 11.03 ± 0.51, and 18.95 ± 0.59 mg rebaudioside A eq. g−1 DW, respectively). This same trend was observed in stevia plants that were only treated with CCC (13.67 ± 0.06 mg rebaudioside A eq. g−1 DW), whereas PBZ had no influence on stevioside content, presenting values very close to those obtained in control plants (18.33 ± 0.39 mg rebaudioside A eq. g−1 DW). It was not possible to detect the presence of rebaudioside A in any sample analysed by HPLC.
Stevioside contents in extracts from Stevia rebaudiana under the effect of GA3 and its inhibitors PBZ and CCC after 40 days of in vitro growth. Columns with different capital letters indicate significant differences between treatments based on ANOVA followed by Tukey test P ≤ 0.05. Error bars indicate the standard deviation around the mean (n = 5)
Effect of GA3, CCC, and PBZ on phenolic compound content
The results showed significant differences for both TPC and FLA content (Fig. 3 and Supplementary Table 3). Plants that were treated with only GA3 showed an increase in TPC (18.40 ± 0.34 mg gallic acid eq. g−1 DW), while plants treated with PBZ showed a decrease (2.38 ± 0.18 mg gallic acid eq. g−1 DW) when compared to control plants (9.12 ± 0.67 mg gallic acid eq. g−1 DW) (Fig. 3a). Regarding FLA, the results showed that none of the treatments caused a significant increase in their content. In contrast, PBZ alone or in combination with GA3 caused a decrease in FLA content (6.96 ± 0.20 and 6.98 ± 0.43 mg rutin eq. g−1 DW, respectively). Furthermore, CCC alone or in combination with GA3 did not provoke significant changes (Fig. 3b).
Total soluble phenolic (a) and flavonoids (b) contents in extracts from Stevia rebaudiana under the effect of GA3 and its inhibitors PBZ and CCC after 40 days of in vitro growth. Columns with different uppercase letters indicate significant differences between treatments based on ANOVA followed by Tukey test P ≤ 0.05. Error bars indicate the standard deviation around the mean (n = 5)
Trait correlations
Pearson’s test was carried out to establish the possible correlations between metabolite content (stevioside and phenolic compounds), gene transcription levels, and morphological characteristics (Fig. 4). No positive correlation was observed for any of the SG-related genes and stevioside content. However, there was a weak positive correlation between the SrUGT74G1 gene and stevioside accumulation (r2 = 0.178). Additionally, there was a positive correlation between phenolic compounds and stevioside. This correlation could be found for both FLA (r2 = 0.789) and TPC (r2 = 0.248). Correlations were also found between the number of shoots and both TPC and FLA content, and in the case of TPC content, also between this parameter and those related with root system development. Overall, specialised metabolite production showed a positive correlation with the analysed growth parameters.
Pearson correlations between SG-related genes expression, stevioside content and morphologic parameters in Stevia rebaudiana under the effect of GA3 and its inhibitors PBZ and CCC after 40 days of in vitro growth. Color intensity and the size of the circle are proportional to the correlation coefficients. (Color figure online)
Multivariate principal component analysis (PCA)
PCA was performed with all evaluated parameters, accounting for 83.3% of the total variance in the data set, with 67.2% in the first principal component (PC1) and 16.1% in the second one (PC2). Based on these results, a biplot graph was constructed (Fig. 5). The variables that most contributed to the distinction among treatments in PC1 were the genes encoding SrCMK and SrKO with 0.06 and SrHDS, SrUGT85C2, SrKAH, SrUGT73E1, SrHDR, SrGGDPS, SrCMS, SrCDPS, SrKS, and SrDXR with 0.05. For PC2, the most important variables were TPC (0.18), the gene encoding SrMCS (0.13), and some growth parameters, such as number of leaves and shoots, both with 0.12. In addition, the specialised metabolites, FLA (0.12) and stevioside (0.07), also contributed. Regarding the treatments in PCA analysis, CON and CCC were in the same quadrant, which could reflect the similarities in the responses to these treatments, mainly with regard to the growth of plants in vitro and the content of phenolic compounds. Conversely, GA3 remained alone in the first quadrant, due in part to its unique role in inducing specialised metabolites in stevia plants.
Principal component analysis (PCA) based on all parameters evaluated in response to the six treatments applied to Stevia rebaudiana: Control (CON); chlorocholine chloride (CCC); paclobutrazol (PBZ); gibberellic acid (GA3); chlorocholine chloride + gibberellic acid (CCC + GA3) and paclobutrazol + gibberellic acid (PBZ + GA3). The cos2 values are used to estimate the quality of the representation
Discussion
The goal of this study was not just to optimise the production of specialised metabolites in stevia using PGRs but also to evaluate the possible negative side effects associated with their application. Thus, the first part of this study was aimed at assessing the effect of these molecules on growth parameters and to determine if the presence of GA3 was able to reverse the possible deleterious effects of CCC and PBZ on stevia. Surprisingly, CCC had no significant reduction effect on the growth parameters assessed in stevia plants; instead, it promoted the largest number of shoots, leaves, and roots during step 1. Following this same trend, PBZ resulted in the highest mean values for nodal segment number and root length. According to Rogach et al. (2020), GAs biosynthesis inhibitors can inhibit and even remove apical dominance, thereby favouring the growth of lateral buds. This may partly explain the improvement in the number of shoots and nodal segments in plants treated with CCC and PBZ, respectively.
The positive effects of CCC and PBZ on some growth parameters during step 1 were not observed at the end of step 2. Conversely, PBZ was harmful, especially regarding root development (formation and growth). This result apparently disagrees with previous studies, in which application of PBZ resulted in an increased root number in in vitro cultured plants (Wang and Yao 2020). The response of plant materials to in vitro conditions can change with successive subcultures (Lajara et al. 2015; Wang and Yao 2020), which could account for the observed differences. PBZ can exert its inhibitory effect on root development through the increase of endogenous cytokinin levels that, in turn, downregulate the expression of genes involved in GAs synthesis (Fonouni-Farde et al. 2018).
Since SGs are mainly produced and accumulated in stevia leaves, the number of these organs generated in response to the different treatments applied is a growth parameter that deserves attention. Apart from CCC, no other PGR treatment resulted in clear-cut variations in leaf number at the end of step 1. Chlorocholine chloride increased the number of leaves, which is in line with results previously obtained by Dey et al. (2013b), who reported increased production of leaf biomass in in vitro-cultured stevia, and with those reported by Karimi et al. (2019) in stevia plants grown in greenhouse conditions. However, the positive effect of CCC on the number of leaves did not remain in step 2, highlighting the importance of assessing the evolution of plant material behaviour upon successive subcultures in in vitro conditions.
Biosynthesis of diterpenes, including SGs, can be initiated through two pathways: plastidial MEP and cytosolic MVA (Brandle and Telmer 2007). However, studies carried out by Wölwer-Rieck (2012) in the leaves of stevia showed that the C5 basic building blocks, IPP and DMAPP, are predominately synthesised through the MEP pathway. Treatment with GA3 alone downregulated almost all MEP genes (six out of seven), and all genes in stage 2 of the GAs/SGs biosynthesis pathways. Moreover, expression of the SrKAH gene and some UDP-glucosyltransferases (SrUGT73E1, SrUGT85C2, and SrUGT76G1) were downregulated, whereas the expression of SrUGT74G1 was upregulated. This expression pattern could be the result of negative feedback due to an increase in the GAs pool within leaf tissues. The bioactive forms of GAs stimulate the expression of genes involved in GAs catabolism and repress the expression of biosynthetic genes (Hernández-García et al. 2020). Overall, literature is scarce regarding the effects of GA3 on the expression profile of SG-related genes and other terpenoids, and further work that considers the expression of all GAs biosynthesis (e.g., KAO; GA20ox, Ga3ox) and catabolic (GA2ox) genes together with qualitative and quantitative analysis of endogenous GAs is required.
Chlorocholine chloride and PBZ are two growth retardants with sites of action in the 2nd stage of the GAs/SGs biosynthesis pathways. According to our results, CCC had a strong influence on the MEP pathway (three out of seven genes downregulated), a particular interference on genes encoding kaurenoid enzymes (downregulation of SrCDPS and SrKS), and little impact on the genes directly involved in the synthesis of SGs. According to Rademacher (2016), CCC inhibits CDPS at a lower degree than it inhibits KS. However, the results of gene expression did not allow us to forecast these differences in enzymatic activity since their relative expression was similar (0.15 ± 0.05 and 0.24 ± 0.04 for SrCDPS and SrKS, respectively).
Concerning the PBZ treatment, its negative effect on the expression of MEP genes was more pronounced than that of CCC (five out of the seven genes that make up the pathway were downregulated). These results contrasted with those of a previously published study by our group, in which we observed that PBZ had only a small effect on MEP pathway gene expression in stevia plants grown in hydroponic systems (Lucho et al. 2018a). Additionally, all genes encoding the enzymes responsible for the condensation steps leading to the formation of kaurenoic acid were downregulated, which is partially in accordance with other studies (Hajihashemi et al. 2013; Hajihashemi and Geuns 2017; Lucho et al. 2018a). Treatment of stevia with GA3 in step 2, after PBZ treatment during step 1, was able to revert the expression of the SrKS gene, and even enhanced the expression of SrKO (the gene encoding the enzyme that constitutes the target of PBZ inhibitory action) relative to the control plants. These results contrasted with those reported by Hajihashemi et al. (2013), who found that GA3 treatment was unable to reverse the negative effect of PBZ on SrKO gene expression. However, differences in experimental conditions (e.g., simultaneous vs sequential application of PGRs), as well as plant material origins, among other factors, may help to explain these opposing results. Interestingly, the ability of GA3 treatment to reverse the negative effects of PBZ on gene expression was also observed in the two other stages of the SGs biosynthesis pathway (Fig. 6).
Schematic representation of biosynthetic steps involved in phenolic compound biosynthesis and SGs/GAs in Stevia rebaudiana and points of inhibition by plant growth retardants (CCC and PBZ). Differential expression pattern of genes encoding enzymes of the 1st stage (green box), 2nd stage (cream box), and 3rd stage (white) are shown, as well as a simplified pathway of GAs biosynthesis (blue). Enzymatic flow is depicted as arrows from substrates to products. Enzymes catalyzing each step are shown into each arrow. Solid upward arrows in red (GA3), light gray (CCC), and dark gray (PBZ) represent upregulation by the respective PGR, whereas those solid downward arrows represent downregulation and the hollow red arrows represent reversal of negative effect by GA3 treatment. A single arrow indicates a one-step reaction, and double arrows represent multiple-step pathway
Apart from their inhibitory effect on gene expression, CCC and PBZ also upregulated the expression of some genes at the 1st and 3rd stages of the GAs/SGs biosynthesis pathways. Both inhibitors promoted increased expression of the SrUGT76G1 gene, which is a very interesting result, since the UGT76G1 enzyme is crucial for the formation of the sweetest SGs, including tetraglucosylated rebaudioside-A (Reb-A), pentaglucosylated rebaudioside-D (Reb-D), and hexaglucosylated rebaudioside-M (Reb-M) making them a production target as high-potency natural sweeteners (Olsson et al. 2016).
This paper reports for the first time, the transcription pattern of SrUGT73E1 and SrUGT91D2 in stevia plants and interestingly, expression of these two genes was positively correlated with key genes of the SGs biosynthesis pathway, including SrKAH, UGT85C2, and UGT76G1. This aspect is remarkable because it has been demonstrated that SrKA13H and SrUGT85C2 are regulatory genes influencing carbon flux between SGs/GAs biosynthetic routes (Guleria and Yadav 2013). Previous studies reported a correlation between the transcription of SrUGT85C2 and SGs accumulation (Hajihashemi et al. 2013; Yoneda et al. 2018). Moreover, the SrKAH gene showed a positive correlation with 12 of the 17 genes in the pathway. The KAH enzyme is upstream from steviol and is of great interest in biotechnology (Brandle and Telmer 2007; Kundu et al. 2014).
Regarding STEV quantification in GA3-treated plants, the results showed twice STEV content as that in control plants. Interestingly, GA3-treated plants showed upregulation of SrUGT74G1 (the gene encoding the enzyme that glycosylates steviolbioside at the C-19 position, forming STEV). Kumar et al. (2012) also showed that GA3 upregulated the SrUGT74G1 gene in stevia plants, however they did not report the content of SGs. Other authors have reported that STEV content increased (Hajihashemi and Geuns 2017; Yoneda et al. 2018), decreased (Karimi et al. 2015), or did not show significant changes (Pazuki et al. 2019) in response to GA3 treatment. Overall, our investigations found a higher STEV amount in GA3-treated, in vitro-cultured stevia plants than that reported by Bondarev et al. (2001) and Ladygin et al. (2008) (less than 5 mg g−1 DW in in vitro leaves and about 25 mg g−1 DW in leaves from intact plants, respectively).
It seems paradoxical that the highest STEV levels were obtained when the expression of almost all GAs/SGs biosynthesis genes were downregulated in response to GA3. Relatively high levels of exogenous GA3 were expected to repress the GAs/SGs biosynthesis pathways by negative feedback, thus decreasing product accumulation. However, the levels of SGs in stevia leaves were several orders of magnitude higher than those of GAs (Geuns 2003), which suggests that although biosynthesis intermediaries are common for both pathways, regulatory mechanisms might differ between biosynthesis routes as some authors have previously proposed (Karimi et al. 2015).
Additionally, PBZ-treated plants showed STEV levels very similar to those obtained in control plants (about 18 mg g−1 DW). Based on the statement that the SrKO gene has two functional copies (Humphrey et al. 2006), Karimi et al. (2019) recently suggested that one copy may be more related to SGs biosynthesis, while the other would be related to GAs biosynthesis. This might be why treatment with PBZ, even when blocking the SrKO gene, showed a STEV content similar to that of the control plants. Conversely, CCC-treated plants showed a decrease in STEV content. Karimi et al. (2014, 2019) observed similar trends in relation to the negative effects of CCC on the accumulation of SGs. A possible reason for this STEV decrease in CCC-treated plants may be related to its inhibitory action on two enzymes (CDPS and KS), instead of on only one (KO), as that seen for PBZ.
In the search for differences and similarities between treatments, PCA analysis was performed (Fig. 5). Based on the obtained results, the factors that had the most influence on the separation of samples were those related to gene expression in PC1 and growth parameters in PC2. Overall, the GA3 alone treatment influenced all the evaluated parameters, unlike its inhibitors. In addition, GA3 reversed the negative effects of PBZ, mainly at the molecular level, partially confirming our hypothesis. Collectively, ours results suggest that the application of GA3 may be a viable way to improve the production of stevioside and phenolic compounds in stevia plants and that this compound can be considered a pivotal regulator of SGs/GAs biosynthesis-related genes. Besides, this study provides the first model for regulation of the expression of SG-related genes in stevia leaves and explores other factors that may influence its biosynthesis (Fig. 7). A better understanding of these pathways and of how external factors, including PGRs (GA3, CCC, and PBZ), influence them will allow us to take full advantage of the richness that stevia plants offer.
Model depicting the metabolic flux in Stevia rebaudiana in response to GA3, CCC and PBZ. Feedback loop (a), simple illustration of the biosynthetic steps of SGs (b) and factors that can influence their synthesis (c). Dashed arrows represent multistep pathways, whereas solid arrows represent only one step (X, x = major and minor inhibitory activity, respectively)
Conclusion
Chlorocholine chloride and PBZ showed different effects in relation to the morphological characteristics and specialised metabolite synthesis in stevia plants. However, at the molecular level, both inhibitors upregulated the SrUGT76G1 gene. Interestingly, among the three PGRs evaluated, GA3 had the most negative impact on gene expression both when used alone and especially after treatment with CCC. Despite GA3 not having a positive impact on transcription patterns, the obtained results demonstrated that STEV and TPC contents were much higher than those observed in the control plants. Thus, our experiment confirmed that exogenous supplementation of GA3 caused negative feedback on SGs/GAs biosynthesis genes and the negative modulator role of CCC and PBZ on the kaurenoid genes acid condensation step. Besides, we demonstrated for the first time a feedback loop in stevia plants cultured in vitro in response to GA3 and its inhibitors CCC and PBZ.
Abbreviations
- ANOVA:
-
Analysis of variance
- CCC:
-
Chlorocholine chloride
- CDPS:
-
Copalyl diphosphate synthase
- CMK:
-
4-Diphosphocytidyl-2-C-methyl-D-erythritol kinase
- CMS:
-
4-Diphosphocytidyl-2-C-methyl-D-erythritol synthase
- DMAPP:
-
Dimethylallyl diphosphate
- DXR:
-
Deoxyxylulose-5-phosphate reductase
- DXS:
-
Deoxyxylulose-5-phosphate synthase
- FLA:
-
Total soluble flavonoids
- GAs:
-
Gibberellins
- GA3 :
-
Gibberellic acid
- GA2ox:
-
Gibberellin 2-oxidase
- GA20ox:
-
Gibberellin 20-oxidase
- GA3ox:
-
Gibberellin 3-oxidase
- GGDP:
-
Geranylgeranyl diphosphate
- GGDPS:
-
Geranylgeranyl diphosphate synthase
- HDR:
-
1-Hydroxy-2-methyl-2(E)-butenyl-4-diphosphate reductase
- HDS:
-
1-Hydroxy-2-methyl-2(E)–butenyl-4-diphosphate synthase
- IPP:
-
Isopentenyl diphosphate
- KAH:
-
Kaurenoic acid 13-hydroxylase
- KAO:
-
Kaurenoic acid oxidase
- KO:
-
Kaurene oxidase
- KS:
-
Kaurene synthase
- MCS:
-
4-Diphosphocytidyl-2-C-methyl-D-erythritol-2,4-cyclodiphosphate synthase
- MEP:
-
2-C-methyl-D-erythritol-4 phosphate
- MVA:
-
Mevalonate
- PBZ:
-
Paclobutrazol
- PGRs:
-
Plant growth regulators
- Reb-A:
-
Rebaudioside-A
- Reb-D:
-
Rebaudioside-D
- Reb-M:
-
Rebaudioside-M
- SGs:
-
Steviol glycosides
- STEV:
-
Stevioside
- TPC:
-
Total soluble phenol content
- UGT73E1:
-
UDP glucosyltransferase-73E1
- UGT74G1:
-
UDP glucosyltransferase-74G1
- UGT76G1:
-
UDP glucosyltransferase-76G1
- UGT85C2:
-
UDP glucosyltransferase-85C2
- UGT91D2:
-
UDP glucosyltransferase-91D2
References
Bondarev N, Reshetnyak O, Nosov A (2001) Peculiarities of diterpenoid steviol glycoside production in in vitro cultures of Stevia rebaudiana Bertoni. Plant Sci. https://doi.org/10.1016/S0168-9452(01)00400-9
Brandle JE, Telmer PG (2007) Steviol glycoside biosynthesis. Phytochemistry. https://doi.org/10.1016/j.phytochem.2007.02.010
Chughtai MFJ, Imran P, Zahoor T et al (2020) Nutritional and therapeutic perspectives of Stevia rebaudiana as emerging sweetener; a way forward for sweetener industry. CYTA J Food . https://doi.org/10.1080/19476337.2020.1721562
Dey A, Paul S, Kundu S et al (2013a) Elevated antioxidant potential of chlorocholine chloride-treated in vitro grown Stevia rebaudiana Bertoni. Acta Physiol Plant. https://doi.org/10.1007/s11738-013-1214-0
Dey A, Kundu S, Bandyopadhyay A et al (2013b) Efficient micropropagation and chlorocholine chloride induced stevioside production of Stevia rebaudiana Bertoni. CR Biol. https://doi.org/10.1016/j.crvi.2012.11.007
Fonouni-Farde C, McAdam E, Nichols D et al (2018) Cytokinins and the CRE1 receptor influence endogenous gibberellin levels in Medicago truncatula. Plant Signal Behav. https://doi.org/10.1080/15592324.2018.1428513
Geuns JMC (2003) Stevioside. Phytochemistry. https://doi.org/10.1016/s0031-9422(03)00426-6
Guleria P, Yadav SK (2013) Agrobacterium mediated transient gene silencing (AMTS) in Stevia rebaudiana: insights into steviol glycoside biosynthesis pathway. PLoS One. https://doi.org/10.1371/journal.pone.0074731
Hajihashemi S (2018) Physiological, biochemical, antioxidant and growth characterizations of gibberellin and paclobutrazol-treated sweet leaf (Stevia rebaudiana B) herb. J Plant Biochem Biotechnol. https://doi.org/10.1007/s13562-017-0428-4
Hajihashemi S, Geuns JMC (2017) Steviol glycosides correlation to genes transcription revealed in gibberellin and paclobutrazol-treated Stevia rebaudiana. J Plant Biochem Biotechnol. https://doi.org/10.1007/s13562-017-0399-5
Hajihashemi S, Geuns JMC, Ehsanpour AA (2013) Gene transcription of steviol glycoside biosynthesis in Stevia rebaudiana Bertoni under polyethylene glycol, paclobutrazol and gibberellic acid treatments in vitro. Acta Physiol Plant. https://doi.org/10.1007/s11738-013-1226-9
Hernández-García J, Briones-Moreno A, Blázquez MA (2020) Origin and evolution of gibberellin signaling and metabolism in plants. Semin Cell Dev Biol. https://doi.org/10.1016/j.semcdb.2020.04.009
Humphrey TV, Richman AS, Menassa R et al (2006) Spatial organization of four enzymes from Stevia rebaudiana that are involved in steviol glycoside synthesis. Plant Mol Biol. https://doi.org/10.1007/s11103-005-5966-9
JECFA (2010) Steviol glycosides. In: Compendium of Food Additive Specifications. FAO JECFA Monographs 10:17-21
Karimi M, Ahmadi A, Hashemi J et al (2014) Effect of two plant growth retardants on steviol glycosides content and antioxidant capacity in Stevia (Stevia rebaudiana Bertoni). Acta Physiol Plant. https://doi.org/10.1007/s11738-014-1498-8
Karimi M, Hashemi J, Ahmadi A et al (2015) Opposing effects of external gibberellin and daminozide on Stevia growth and metabolites. Appl Biochem. https://doi.org/10.1007/s12010-014-1310-7
Karimi M, Ahmadi A, Hashemi J et al (2019) Plant growth retardants (PGRs) affect growth and secondary metabolite biosynthesis in Stevia rebaudiana Bertoni under drought stress. S Afr J Bot. https://doi.org/10.1016/j.sajb.2018.11.028
Kim D, Jeong SW, Lee CY (2003) Antioxidant capacity of phenolic phytochemicals from various cultivars of plums. Food Chem. https://doi.org/10.1016/S0308-8146(02)00423-5
Kumar H, Kaul K, Bajpai-Gupta S et al (2012) A comprehensive analysis of fifteen genes of steviol glycosides biosynthesis pathway in Stevia rebaudiana (Bertoni). Gene. https://doi.org/10.1016/j.gene.2011.10.015
Kundu S, Dey A, Bandyopadhyay A (2014) Stimulation of in vitro morphogenesis, antioxidant activity and over expression of kaurenoic acid 13-hydroxylase gene in Stevia rebaudiana Bertoni by chlorocholine chloride. Acta Physiol Plant. https://doi.org/10.1007/s11738-014-1639-0
Ladygin VG, Bondarev NI, Semenova GA et al (2008) Chloroplast ultrastructure, photosynthetic apparatus activities and production of steviol glycosides in Stevia rebaudiana in vivo and in vitro. Biol Plant. https://doi.org/10.1007/s10535-008-0002-y
Lajara MA, López-Orenes A, Ferrer MA et al (2015) Long-term exposure treatments revert the initial SA-induced alterations of phenolic metabolism in grapevine cell cultures. Plant Cell Tissue Organ Cult. https://doi.org/10.1007/s11240-015-0800-9
Li W, Zhou Y, You W et al (2018) Development of Photoaffinity Probe for the Discovery of Steviol Glycosides Biosynthesis Pathway in Stevia rebaudiana and Rapid Substrate Screening. ACS Chem Biol. https://doi.org/10.1021/acschembio.8b00285
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. https://doi.org/10.1006/meth.2001.1262
López-Orenes A, Ros-Marín AF, Ferrer MA et al (2013) Antioxidant capacity as a marker for assessing the in vitro performance of the endangered Cistus heterophyllus. Sci World. https://doi.org/10.1155/2013/176295
Lucho SR, do Amaral MN, Milech C et al (2018a) Elicitor-induced transcriptional changes of genes of the steviol glycosides biosynthesis pathway in Stevia rebaudiana Bertoni. J Plant Growth Regul. https://doi.org/10.1007/s00344-018-9795-x
Lucho SR, do Amaral MN, Benitez LC et al (2018b) Validation of reference genes for RT-qPCR studies in Stevia rebaudiana in response to elicitor agents. Physiol Mol Biol Plants. https://doi.org/10.1007/s12298-018-0583-7
Lucho SR, do Amaral MN, Auller PA et al (2019) Salt stress-induced changes in in vitro cultured stevia rebaudiana bertoni: effect on metabolite contents antioxidant capacity and expression of steviol glycosides-related biosynthetic genes. J Plant Growth Regul. https://doi.org/10.1007/s00344-019-09937-6
Modi A, Litoriya N, Prajapati V et al (2014) Transcriptional profiling of genes involved in steviol glycoside biosynthesis in Stevia rebaudiana Bertoni during plant hardening. Dyn Dev. https://doi.org/10.1002/dvdy.24157
Murashige T, Skoog F (1962) A Revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Olsson K, Carlsen S, Semmler A et al (2016) Microbial production of next-generation stevia sweeteners. Microb Cell Fact. https://doi.org/10.1186/s12934-016-0609-1
Pazuki A, Aflaki F, Yücesan B et al (2019) Effects of cytokinins, gibberellic acid 3, and gibberellic acid 4/7 on in vitro growth, morphological traits, and content of steviol glycosides in Stevia rebaudiana. Plant Physiol Biochem. https://doi.org/10.1016/j.plaphy.2019.02.009
Rademacher W (2016) Chemical regulators of gibberellin status and their application in plant production. Ann Plant Rev. https://doi.org/10.1002/9781119312994.apr0541
Richman A, Swanson A, Humphrey T et al (2005) Functional genomics uncovers three glucosyltransferases involved in the synthesis of the major sweet glucosides of Stevia rebaudiana. Plant J. https://doi.org/10.1111/j.1365-313X.2004.02275.x
Rogach VV, Voytenko LV, Shcherbatiuk MM et al (2020) Morphogenesis, pigment content, phytohormones and productivity of eggplants under the action of gibberellin and tebuconazole. Regul Mech Biosyst. https://doi.org/https://doi.org/10.15421/022017
Totté N, Van den Ende W, Van Damme EJM et al (2003) Cloning and heterologous expression of early genes in gibberellin and steviol biosynthesis via the methylerythritol phosphate pathway in Stevia rebaudiana. Can J Bot. https://doi.org/10.1139/b03-032
Ullah A, Munir S, Mabkhot Y et al (2019) Bioactivity profile of the diterpene isosteviol and its derivatives. Molecules. https://doi.org/10.3390/molecules24040678
Wang Y, Yao R (2020) Increased endogenous gibberellin level inhibits root growth of Pinus massoniana Lamb plantlets during long-term subculture. Vitro Cell Dev-Pl. https://doi.org/10.1007/s11627-020-10067-y
Wang J, Li S, Xiong Z, Wang Y (2016) Pathway mining-based integration of critical enzyme parts for de novo biosynthesis of steviol glycosides sweetener in Escherichia coli. Cell Res. https://doi.org/10.1038/cr.2015.111
Wölwer-Rieck U (2012) The Leaves of Stevia rebaudiana (Bertoni), Their Constituents and the Analyses Thereof: A Review. J Agric Food Chem. https://doi.org/10.1021/jf2044907
Yoneda Y, Shimizu H, Nakashima H et al (2018) Effect of treatment with gibberellin, gibberellin biosynthesis inhibitor, and auxin on steviol glycoside content in Stevia rebaudiana Bertoni. Sugar Tech. https://doi.org/10.1007/s12355-017-0561-3
Acknowledgements
The authors gratefully acknowledge the CNPq (Conselho Nacional de Desenvolvimento Cientifíco e Tecnológico) for their financial support and research fellowship EJBB and VJB, as well as the FAPERGS (Fundação de Amparo à Pesquisa do Estado do Rio Grande do Sul). This study was financed in part by CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brasil—Finance Code 001. This work was partially carried out at IBV (Instituto de Biotecnología Vegetal), UPCT (Spain).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: SRL and EJBB. Performed the experiments: SRL, CM, LA and MNA. Analysed the data: SRL, MNA and EJBB. Wrote the paper: SRL, AAC and EJBB. Corrected the manuscript: MAF, AAC, VJB and EJBB.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Mohammad Faisal.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11240_2021_2059_MOESM1_ESM.pdf
Supplementary Figure 1 Schematic representation of the experimental conditions. Stevia mother plants grown in greenhouse (A), in vitro establishment (B and C) and multiplication (D) before applying treatments (E) (PDF 108 kb)
11240_2021_2059_MOESM2_ESM.pdf
Supplementary Figure 2 Agarose gel electrophoresis (1%) of PCR products amplified from Stevia rebaudiana under the effect of GA3 and its inhibitors PBZ and CCC after 40 days of in vitro growth. (A and B) Specificity of primer pairs used for RT-qPCR reactions. Lane 1 is 100pb DNA ladder; 2: DXS; 3: DXR; 4: CMS; 5: CMK; 6: MCS; 7: HDS; 8: HDR; 9: GGDPS; 10: CDPS; 11: KS; 12: KO; 13: KAH; 14: UGT85C2; 15: UGT74G1; 16: UGT76G1; 17: UGT73E1; 18: UGT91D2; 19: UBQ and C: Negative control (PDF 413 kb)
11240_2021_2059_MOESM3_ESM.pdf
Supplementary Figure 3 Specificity of the primers used for the RT-qPCR reactions in Stevia rebaudiana under the effect of GA3 and its inhibitors PBZ and CCC after 40 days of in vitro growth (PDF 261 kb)
11240_2021_2059_MOESM4_ESM.pdf
Supplementary Figure 4 Effect of GA3 and its inhibitors PBZ and CCC on growth and development of Stevia rebaudiana after 40 days of in vitro growth (PDF 857 kb)
Rights and permissions
About this article
Cite this article
Lucho, S.R., do Amaral, M.N., Milech, C. et al. Gibberellin reverses the negative effect of paclobutrazol but not of chlorocholine chloride on the expression of SGs/GAs biosynthesis-related genes and increases the levels of relevant metabolites in Stevia rebaudiana. Plant Cell Tiss Organ Cult 146, 171–184 (2021). https://doi.org/10.1007/s11240-021-02059-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-021-02059-6