Abstract
In plants, somatic embryo development is regulated by a complex group or network of transcription factors (TFs). The LEAFY COTYLEDON (LEC) TFs are significant key regulators that promote the initiation of somatic embryo formation and biological processes of the embryo maturation phase. The LEC gene has been implicated to act as unique regulators in plant embryogenesis, growth and development via diverse signaling pathways. In the present review, we summarize the current advances in our understanding of the LEC TFs in plant biology including embryogenesis. Recent discoveries would be advantageous to unlock the mysteries of LEC TF genes of different molecular mechanisms in plant cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Embryogenesis is an essential stage which represents developmental plasticity in higher plant species (Yang and Zhang 2010). Somatic embryogenesis (SE) is an important in vitro regeneration method in modern crop breeding which describes the single cell differentiation into a mature embryo through different development periods (Kumar and Van Staden 2017). In the SE pathway, haploid or diploid somatic cells develop into differentiated plants through different embryological stages (globular, heart, torpedo and cotyledonary-shaped) without fusion of gametes (Williams and Maheswaran 1986; Kumar and Van Staden 2017). In general, two distinct phases are involved in the whole process of plant embryogenesis; early morphogenesis phase which includes the formation of embryogenic cells and tissues, and maturation phase that permit the embryo to enter into a desiccated state (West and Harada 1993; Goldberg et al. 1994; Méndez-Hernández et al. 2019). Studies on the genetic mechanisms confirmed that a number of transcription factors (TFs) have been identified which are responsible for inducing somatic embryogenesis when ectopically expressed. A predominant number of SE-inducing genes encode TFs including SOMATIC EMBRYOGENESIS RECEPTOR-LIKE KINASE (SERK) (Schmidt et al. 1997; Kumar and Van Staden 2019), BABY BOOM (BBM) (Boutilier et al. 2002; Jha and Kumar 2018), LEAFY COTYLEDON (LEC) (Stone et al. 2001; Gaj et al. 2005), AGAMOUS-LIKE 15 (AGL15) (Harding et al. 2003), WUSCHEL (WUS) (Zuo et al. 2002), and EMBRYO MAKER (Tsuwamoto et al. 2010) have been identified, which are responsible for the induction of differentiated somatic cells and somatic embryo formation. Among the TF genes, the LEC genes are reported to have an important role in controlling several aspects of embryogenesis including embryo development (Gaj et al. 2005; Braybrook and Harada 2008 (Table 1)). The LEC TFs (LEC1, LEC2 and FUSCA3 (FUS3)) are unique regulators of embryogenesis in that they are required to explain the molecular mechanisms for controlling embryo development, morphogenesis and embryo maturation (Harada 2001). The LEC TFs establish environments that encourage the initiation of somatic embryo formation and cellular processes of the maturation phase. The cessation of embryo morphogenesis, synthesis and storage of accumulated macromolecules, acquirement of desiccation tolerance and desiccation of the seed are the major features of the maturation phase (Harada 1997; Vicente-Carbajosa and Carbonaro 2005; Braybrook and Harada 2008). In early embryogenesis, LEC TFs are needed to specify suspensor cell fate and cotyledon identity (Lotan et al. 1998; Meinke et al. 1994; Keith et al. 1994; West et al. 1994; Stone et al. 2001), whereas during late embryogenesis LEC TFs are required for maturation phase for the expression of maturation-specific genes (West et al. 1994; Baumlein et al. 1994). Thus, LEC TFs are candidate gene regulators, which play a key role in controlling many aspects of embryogenesis including morphogenesis and maturation. The LEC1, LEC2 and FUS3 encode two distinct classes of TFs. The LEC1 gene encodes an extensive sequence similarity to the HEMEACTIVATED PROTEIN 3 (HAP3) subunit of the CCAAT-binding TF, an isoform found in seed plants (Lotan et al. 1998; Stone et al. 2001; Kwong et al. 2003; Braybrook and Harada 2008). In Arabidopsis thaliana, based on sequence similarity HAP3 subunits can be divided into two different classes such as LEC1-type and the non-LEC1-type (Lee et al. 2003). Both the LEC1 types are essential for embryogenesis and embryo development (Kwong et al. 2003). The LEC2 and FUS3 genes encodes B3 domain TFs, a DNA-binding motif, which acts in developing seeds (Luerssen et al. 1998; Stone et al. 2001). Ectopic expression of the LEC genes, LEC1 and LEC2 activate SE in vegetative cells and were found to be adequate in embryo development (Lotan et al. 1998; Stone et al. 2001). It was hypothesized that Arabidopsis PICKLE (PKL), which encodes a CHD3-chromatin-remodeling factor, is responsible for repression of the LEC genes during seed germination (Ogas et al. 1999; Rider et al. 2003). Consistent with expression of the LEC genes, PKL mutants accumulate storage products and promote embryonic identity in culture. These embryonic features are repressed by exogenous GA and enhanced by GA synthesis inhibitors (Ogas et al. 1997). The LEC2 and FUS3 TFs activates the genes involved in the accumulation of storage macromolecules in the embryo during maturation. In addition, LEC2 and FUS3 TFs are implicated in repression of GA biosynthesis during seed development (Gazzarrini et al. 2004; Curaba et al. 2004).
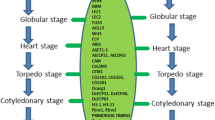
As shown in Fig. 1, LEC1 gene activates YUC10, whereas LEC2 induces YUC2 and YUC4, an auxin biosynthesis enzyme (Stone et al. 2008). The LEC2 gene also induces the IAA30 (negative regulator of auxin signaling) (Braybrook et al. 2006; Kumar and Van Staden 2017; Jha and Kumar 2018). The activation of YUC genes increase the endogenous auxin levels which obviates the necessity for exogenous auxin and provides a critical insight into LEC-mediated SE.
Schematic model which explain the role of LEC TFs in somatic embryogenesis. Ectopic expression of LEC genes induces embryogenesis without exogenous auxin. Based on the articles reviewed we suggest that the endogenous auxin level was increased by LEC1-mediated activation of YUC10 gene and LEC2-mediated activation of YUC2 and YUC4 gene that encodes auxin biosynthesis and IAA30 (negative regular of auxin signaling), which modulate the auxin-mediated signaling during embryogenesis. The FUS3 repressed biosynthesis of GA. Arrows with dotted line indicate transcriptional regulation that molecular mechanisms are not clear and arrows with solid line indicates direct transcriptional regulation by molecular evidence
The role of LEC genes in embryogenesis have been reported in many plant species such as A. thaliana (Lotan et al. 1998; Stone et al. 2001; Gaj et al. 2005; Wójcikowska et al. 2013), Zea mays (Zhang et al. 2002), Daucus carota hypocotyl (Yazawa et al. 2004), Coffea canephora seedlings (Nic-Can et al. 2013), Medicago sativa protoplasts (Domoki et al. 2006), Medicago truncatula leaves (Orlowska et al. 2017), Theobroma cacao leaf tissue (Alemanno et al. 2007; Fister et al. 2018) and c axillary buds (Brand et al. 2019). Additionally, few studies also revealed diverse biological processes of LECs such as regulation of gene sets, involved in seed development (Pelletier et al. 2017) and for enhancing oil yield in Camelina and Arabidopsis seeds (Zhu et al. 2018). The main purpose of this review is to provide brief insights on the recent discoveries and current advances of the LEC TFs in the area of plant embryogenesis.
Ectopic expression of LEC genes induces embryogenesis without exogenous auxin. Based on the articles reviewed we suggest that the endogenous auxin level was increased by LEC1-mediated activation of YUC10 gene and LEC2-mediated activation of YUC2 and YUC4 gene that encodes auxin biosynthesis and IAA30 (negative regular of auxin signaling), which modulate the auxin-mediated signaling during embryogenesis.
LEC TFs genes are crucial during somatic embryogenesis
SE developmental pathway comprises complex network of cellular processes and expression of several signaling pathways. A considerable number of studies related to molecular genetics confirmed that induction of spontaneous embryogenesis is due to ectopic expression of TF genes (Salvo et al. 2014; Horstman et al. 2017a; Jha and Kumar 2018). LECs TFs are central regulators of plant cell totipotency (Gaj et al. 2005), are responsible for initiation and control of maturation phase during embryogenesis and cause formation of somatic embryos when expressed ectopically (Braybrook and Harada 2008). During embryogenesis the zygote undergoes two different developmental stages i.e. morphogenesis and maturation. In morphogenesis, the basic plant body is established and the embryo is expressed as a shoot (plumule) and root (radicle) axis. (Harada 2001; Laux and Jurgens 1997). During maturation phase several metabolic activities allow the embryo to germinate (Harada 1997). Storage proteins and lipids are stored in protein and lipid bodies throughout the embryo and these are utilized by the growing seedling as a nutrient source.
In addition, at this stage the embryo acquires the ability to withstand desiccation; seeds desiccate at the late development of embryogenesis (Harada 2001).
It is so that the LEC1/2 and FUS3 participate in an important role in controlling several aspects of embryogenesis including early morphogenesis and late maturation phase (Harada 2001; Parcy et al. 1997; Nambara et al. 2000; Kroj et al. 2003; Gaj et al. 2005). The LEC genes are essential to maintain suspensor cell identity during morphogenesis phase in embryogenesis (Meinke and Yeung 1993). A further function during morphogenesis of the LEC genes is the specification of cotyledon identity (Meinke 1992; Meinke et al. 1994; West et al. 1994). Cotyledons are reverting partially to a leaf-like organ and incompletely specified in the absence of LEC gene activity.
During maturation phase, the LEC genes regulate and are responsible for the storage of macromolecule synthesis and accumulation. The LEC genes ectopically expressed in plants accumulates lipids and proteins characteristic of seeds in reproductive and vegetative tissues (Stone et al. 2001, 2008; Mendoza et al. 2005; Baud et al. 2007; Wang et al. 2007a, b). Storage protein synthesis and lipid accumulation is defective in loss-of-function LEC mutants, however, these LEC mutants are involved in the accumulation of starch grains and protein in the distal tips and in basal regions respectively (Meinke et al. 1994; Ikeda et al. 2006). Likewise, a number of researchers have identified that the expression of genes usually active during maturation phase, including genes related to storage lipid and protein accumulation, is defective in LEC mutants (Harada 2001; Bäumlein et al. 1994; Parcy et al. 1997).
The LEC TFs directly target the genes involved in the synthesis of storage macromolecule. The LEC2 and FUS3 TF genes bind with the RY sequence repeats, which is conserved in the 5′ flanking regions of seed protein genes and involved in the regulation of transcription of these genes (Dickinson et al. 1988; Kroj et al. 2003; Reidt et al. 2000; Monke et al. 2004; Braybrook et al. 2006). During maturation phase, ABSCISIC ACID INSENSITIVE3 (ABI3) interacts with LEC genes to regulate the seed protein genes (Kroj et al. 2003; To et al. 2006). Activation of LEC1 seed protein gene is dependent on the ABI3 and other LEC genes. Ectopic expression of LEC1 activates the ABI, LEC2 and FUS3 genes (Kagaya et al. 2005a).
A complex relationship among the LEC TFs for the regulation of embryo maturation has been shown in Fig. 2. During embryo maturation phase LEC TFs exhibit complex regulatory interrelationships to activate several genes to induce maturation. The LEC2 activates both LEC1 and FUS3 (Stone et al. 2008), whereas, LEC1 activates LEC2 and FUS3 (Kagaya et al. 2005b; To et al. 2006). LEC TFs interact with GA; LEC2 directly induces and activates AGL15 (Braybrook et al. 2006). The AGL15 positively activates GA degrading enzyme GA3ox2 (Wang et al. 2004; Kumar and Van Staden 2017). The FUS3 represses the GA3ox1 and GA3ox2 (GA biosynthesis genes) (Gazzarrini et al. 2004; Curaba et al. 2004). PKL represses LEC genes in seedlings (Ogas et al. 1997; Rider Jr. et al. 2003). PKL mutants express LEC genes ectopically and the overexpression of LEC genes is enhanced by GA synthesis inhibitors. However, PKL-mediated repression of the maturation process is still unclear. These characteristics revealed that the LEC TF genes play an essential role in controlling embryogenesis processes in plants. The LEC genes have distinct differences, although they share similar mutant phenotypes. Finally, the LEC genes are candidate markers that coordinate embryogenesis being involved in both morphogenesis and maturation stages.
Schematic overview of regulation of the embryo maturation phase by LEC TFs. During embryo maturation phase LEC TFs exhibit complex regulatory interrelationship to activate several genes to induce maturation. The LEC2 activates both LEC1 and FUS3 (Stone et al. 2008), whereas, LEC1 activates LEC2 and FUS3 (Kagaya et al. 2005b; To et al. 2006). LEC TFs interact with GA; LEC2 directly induces and activates AGL15 (Braybrook et al. 2006). The AGL15 positively activates GA degrading enzyme GA3ox2 (Wang et al. 2004; Kumar and Van Staden 2017). The FUS3 represses the GA3ox1 and GA3ox2 (GA biosynthesis genes) (Gazzarrini et al. 2004; Curaba et al. 2004). PKL represses LEC genes in seedlings (Ogas et al. 1997; Rider et al. 2003). pkl mutants express LEC genes ectopically and the overexpression of LEC genes is enhanced by GA synthesis inhibitors. However, PKL-mediated repression of maturation process is still unclear
Other transcription factor genes controlling plant embryogenesis
SE consists of various developmental phases which initiates with embryonic induction. While the cells are in induction phase, several genes are functional, possibly due to biotic or abiotic stresses or extrinsic hormones. The induction phase can further be classified into three sub stages: embryogenic dedifferentiation, totipotency expression and embryogenic commitment. Cell dedifferentiation is a cellular regression process in which mature cells are converted into transient stages.
As discussed above, LEC genes have been found to be key regulators for embryogenesis when ectopically expressed. However, other TF genes have also been recognized to promote embryogenesis which includes SERK (Schmidt et al. 1997; Hecht et al. 2001), BBM (Boutilier et al. 2002), AGL15 (Harding et al. 2002), WUS (Zuo et al. 2002), and EMBRYO MAKER (Tsuwamoto et al. 2010).
SERK role is identified during embryogenesis in several plant species such as A. thaliana (Hecht et al. 2001), Z. mays (Zhang et al. 2011), M. truncatula (Nolan et al. 2009), T. cacao (Santos et al. 2005) and T. nigrescens (Pilarska et al. 2016). In a study on Arabidopsis, by Hecht et al. (2001), AtSERK1 was found to be highly expressed during early embryogenesis. The study suggested that AtSERK1 gene is initially expressed during megasporogenesis in the megaspore and in cells of embryo sac till fertilization stage. However, least expression of the same gene has been found in matured vascular tissues. ZmSERK1 and ZmSERK2 genes isolated from Z. mays, has been found to express during embryogenesis (Zhang et al. 2011). Interestingly, ZmSERKs genes expression are associated to embryo development and hormone signaling. These studies indicates that SERK gene is involved in cell to embryonic transition in plant cells.
WUS encodes the homoeodomain TF, reported to play an important role in plant embryogenesis. It has been observed that WUS is positively up-regulated during SE in various plant species (Zuo et al. 2002; Zheng et al. 2014; Tvorogova et al. 2019). Ectopic expression of WUS gene was shown to be involved in vegetative-to-embryonic transition in all tissues (leaf petiole, leaves, stem and root), without adding exogenous growth hormones in A. thaliana (Zuo et al. 2002). In C. canephora, overexpression of WUS significantly enhanced the embryo development up to 400%, and also increased the SE in a heterologous system, however exogenous PGRs were essential for the initiation of SE (Arroyo-Herrera et al. 2008).
A. thaliana WUS (AtWUS) significantly increased embryogenic callus formation (47.75%) in G. hirsutum (cotton), when ectopically expressed (Zheng et al. 2014), and also positively upregulated LEC1, LEC2 and FUS3 in the embryogenic callus. Similarly, Bouchabké-Coussa et al. (2013) also revealed that WUS overexpression significantly promoted (×3) embryogenic capacity and triggered in vitro regeneration competence in cotton when WUS was expressed ectopically. However, these researchers also examined that WUS overexpression resulted in the initiation of embryo-like structures (abnormal) and that leaf-like structures developed on the somatic embryos (Bouchabké-Coussa et al. 2013).
In tobacco, Zhou et al. (2018) uncovered a novel function of WOXs in regulating embryo patterning, and confirmed by expression pattern analysis that WOX2 and WOX9 are essential for early embryo patterning. In a recent report with M. truncatula, it was showed that the WOX9 homolog, MtWOX9-1, participates in embryogenesis and its overexpression enhances embryogenic capacity by changing the expression levels of various SE-associated genes (Tvorogova et al. 2019). These findings confirmed that WUS and WOX family members have an important impact on improving SE competence in plant cells.
BBM TF is a master regulator, which induces embryo development without any exogenous PGRs (Boutilier et al. 2002; Jha and Kumar 2018). In a breakthrough report, it was observed that BBM transcriptionally regulates LEC1/2, ABI3 and FUS3 network during plant embryogenesis (Horstman et al. 2017b). This observation indicates that LEC1 and FUS3 are crucial for embryo development, where as ABI3 and LEC2 positively regulates BBM-mediated SE. However, it is a context and dose-dependent mechanism. In a breakthrough report by Boutilier et al. (2002), it was found that in Arabidopsis an ortholog gene (AtBBM) and in B. napus two ortholog genes (BnBBM1 and BnBBM2) were recognized and it was revealed that overexpression of these ortholog genes encourage embryo development. A transgene constructs 35S::BBM and UBI::BBM were used for transformation in Arabidopsis and B. napus respectively and responsible for cotyledon-shaped embryo development on post-germination organs. Interestingly, in P. tomentosa, overexpression of BBM-mediated embryogenesis significantly improved regeneration pathway (Deng et al. 2009). BBM induces embryo development from P. tomentosa calli, when expressed ectopically. Approximately 12 embryo were developed from 6 calli after 28 days, however, among 12 only 6 embryo survived and developed into complete plantlets (Deng et al. 2009). Similarly, in T. cacao, an ortholog gene (TcBBM) has been identified, which is found to promote the vegetative to embryonic transition of T. cacao somatic cells (Florez et al. 2015).
Expression level of TcBBM gene was found throughout the embryogenesis process including several stages such as globular-stage, heart-stage, early and late torpedo stage and cotyledonary stages. These expression levels led to phenotype in T. cacao, without any exogenous PGRs for direct embryogenesis, however, TcBBM overexpression enhanced embryonic potential significantly. Overall, these findings showed that TcBBM transcriptional level plays a vital role in embryogenesis and it could use as marker gene in T. cacao tissue for embryonic growth (Florez et al. 2015).
AGL15 encodes a MADS domain TF that is expressed during embryogenesis, although not exclusively (Heck et al. 1995; Rounsley et al. 1995; Perry et al. 1999; Wang et al. 2004; Zheng et al. 2016). In Arabidopsis, AGL15 can stimulate SE and lead to extended periods (over 12–19 years to date), when expressed ectopically (Harding et al. 2003; Thakare et al.2008; Zheng et al. 2016). A transgene (35S promoter:AGL15), promotes SE from apical region of shoots which is germinated in the medium supplemented with 2,4-D (Harding et al. 2003; Thakare et al.2008). In addition, overexpression of AGL15-like TF gene is responsible for early embryogenesis in Zea mays (Salvo et al. 2014). Moreover, gene encoding putative ortholog, GmAGL15 (isolated from Glycine max) can enhance embryo development in Arabidopsis (Thakare et al. 2008). However, loss-of-function alleles of agl15 showed significant reduction in SE (Thakare et al. 2008). Finally, the different TF genes are master regulators that coordinate SE being involved in both early and late embryo development.
LEC TFs gene mediated oil content accumulation
Oilseed crop improvement is one of the major objectives to fulfil the ever-increasing oil needs by humans and for biodiesel production. A number of plants accumulate oils in the seeds with several beneficial effects. Mainly plant oil is synthesized as triacylglycerols (TAGs) from fatty acyl-CoA and glycerol-3-phosphate (Ohlrogge and Browse 1995; Shen et al. 2010).
Seed oil content in plants is controlled by several phases in the oil biosynthetic pathway. Oilseed accumulation and biosynthesis are influenced by various genes which are involved directly or indirectly in embryo or seed development (Wang et al. 2007a, b; Shen et al. 2010; Tan et al. 2011; Zhu et al. 2018). In A. thaliana, two important TFs LEC1 and WRINKLED1 (WRI1) have been found which are involved in the regulation of oil accumulation (Lotan et al. 1998; Cernac and Benning 2004). Several studies by pioneer scientists have been documented that overexpression of TFs enhance the oil production in plants when compared to the overexpression of pathway enzymes (Broun 2004; Grotewold 2008; Van Erp et al. 2014). The LEC TFs are key regulators of embryogenesis and are also involved in fatty acid biosynthesis by increasing the expression of genes.
In Zea mays (maize), ZmLEC1 (maize LEC1) is overexpressed as a key regulator and increases the seed oil production (Shen et al. 2010). The ZmLEC1 gene homolog exhibited 41% identity to Arabidopsis LEC1 in amino acid sequence. Overexpression of ZmLEC1 enhanced the oil content by 48.7% in transgenic maize, however, seed germination and leaf growth reduced significantly (Shen et al. 2010). The transgenic leaves were 40–50% shorter, and were narrow and dark green in colour. Transgenic ZmLEC1 seedlings shoot and root growth were slower, resulting in reduced height of the plant in the field.
In a recent promising report, LEC gene was shown to increase oil production in Arabidopsis and Camelina seeds (Zhu et al. 2018). By using Agrobacterium-mediated floral dip method ZmLEC1 binary vector were constructed, driven by seed-specific serine carboxypeptidase-like (SCPL17) and acyl carrier protein (ACP5) promoters and introduced into Arabidopsis and Camelina for expression. The overexpression of ZmLEC1 enhanced the total oil content by < 20% in Arabidopsis and < 26% in Camelina mature seeds (Zhu et al. 2018). Interestingly, there was no phenotypic variation or abnormal growth identified throughout the life cycle of both the plants. These results suggest that ZmLEC1, a master regulator, trigger and increases the oil content in Arabidopsis and Camelina seeds and might be useful for the enhancement of oil production in different crops or oilseed crop improvement. Similarly, in Brassica napus, overexpression of BnLEC1 and Bn LEC1-like TFs significantly increases the total seed oil content by 2–20% in transgenic seeds without any abnormal effects on agronomic traits (Tan et al. 2011). In a report by Angeles-Núñez and Tiessen (2011), they proposed that overexpression of LEC2 TF reduced the seed oil content by 30% while maintaining high levels of sucrose (140%) and starch (> fivefold more) in transgenic Arabidopsis seeds. Future research with more extensive analysis may help to understand the molecular mechanisms on how LEC genes are involved in the expression of genes for fatty acid biosynthesis.
LEC crucial for seed development
Seed development (SD) is a critical and complex phase of the higher plant life cycle. A seed comprises three different regions (filial embryo, filial endosperm and maternal seed coat) with distinct variation on a common genotype (Jo et al. 2019). Furthermore, each region contains distinct subregions, cell and tissues. SD process starts with a double fertilization event that generate the zygote and endosperm (Goldberg et al. 1994; Harada 2001). Many TFs have been shown to express and regulate diverse processes during SD (Pradhan et al. 2014; Jia et al. 2014; Devic and Roscoe 2016; Jo et al. 2019). Among the TFs involved in SD, LEC1 has been considered to be a central regulator of SD (Harada 2001; To et al. 2006; Braybrook and Harada 2008; Pelletier et al. 2017; Jo et al. 2019). LEC1 TF acts sequentially and controls diverse processes at several stages of SD (Pelletier et al. 2017). During SD, LEC1 acts indirectly to regulate diverse processes by activating TFs controlling structural genes, however, LEC1 also regulate directly by establishing a feed-forward loop (FFL) network with association of other TFs (Mangan and Alon 2003). In addition, LEC1 also interacts with several other TFs and activates a particular set of genes during SD (Huang et al. 2015b).
LEC1 is also positively involved in chloroplast biogenesis and photosynthesis during SD (Pelletier et al. 2017; Jo et al. 2019). In Arabidposis and Glycine max (soybean) embryos, LEC1 TF transcriptionally activates and expressed a genes encoding the light-reaction components of photosystems I and II and other set of genes involved in photosynthesis and chloroplast biogenesis (Pelletier et al. 2017). It was also identified that LEC1 TF also regulates and control endosperm development (Lotan et al. 1998). In rice, LEC1 control endosperm development through its interaction with AP2 TFs (Zhang and Xue 2013; Xu et al. 2016).
It was found that LEC1 may regulate directly or indirectly different TFs to regulate gene sets involved in early and late stages of SD (Junker et al. 2012; Pelletier et al. 2017; Jo et al. 2019). The LEC1 directly regulates the LEC2, ABI3 and FUS3 TFs, which all are master regulators of seed maturation (Santos-Mendoza et al. 2008; Braybrook and Harada 2008; Boulard et al. 2017, 2018). It confirms that LEC1 transcriptionally regulates ABI3 and FUS3 and together they form a feed-forward loop (FFL) network, a three-gene pattern and regulates a target gene (Mangan and Alon 2003). Similarly, LEC1 directly regulates WRINKLED1 (WRI1) (TF which plays a key role in seed maturation), and make a FFL network and directly regulate genes involved in the fatty acid accumulation during SD in Arabidopsis (Baud et al. 2007; To et al. 2012; Jo et al. 2019). Moreover, LEC1 TF also control SD indirectly by regulating the expression of TFs that independently control SD.
Few studies suggested that LEC1 may interact with other TFs to regulate diverse development processes during SD (Parcy et al. 1997; To et al. 2006; Pelletier et al. 2017; Jo et al. 2019). In a recent report published in PNAS, they propose that LEC1 acts sequentially and interacts with different TFs and respond to different developmental signals during seed development (Pelletier et al. 2017). LEC1 interacts with LEC2, ABI3 and FUS3 TFs and control gene expression in seeds and are involved in the regulation of diverse processes during seed maturation (Devic and Roscoe 2016; Boulard et al. 2018; Lepiniec et al. 2018).
LEC1 control the maturation phase by interacting with B3 and bZIP (basic leucine zipper TF) TFs which accumulate during SD (Mendes et al. 2013; Baud et al. 2016).
In addition, LEC1 interacts with PHYTOCHROME INTERACTING FACTOR4 (PIF4) (a transcriptional modulator), which is responsible for the expression of hypocotyl elongation related genes through G box element (Huang et al. 2015b). LEC1 also interacts with TCL2 to repress trichome formation during embryogenesis (Huang et al. 2015a). Finally, the interaction of LEC1 with many other TFs provides an outline to define how LEC1 regulate and express distinct gene sets during different phases of SD. In future, LEC1 interaction with all TFs and their impact on LEC1 could provide novel insights into the multitasking of LEC1 during SD. In A. thaliana, FUS3 phosphorylation at SnRK1 (conserved eukaryotic kinase complex) sites positively regulates seed yield and plant growth at heat stress (Chan et al. 2017). They concluded that FUS3 phosphorylation plays an important role for SD and plant growth at high temperature. However, the molecular mechanism by which FUS3 regulates is still elusive. Finally, LEC1 TF has been identified as a central regulator of SD, however, very little is known about the mechanisms by which LEC1 controls and regulates diverse biological processes of SD.
Multi-functionality of LEC TFs during plant development
The LEC TFs acts as a master regulator and are involved in diverse functions including plant embryogenesis, growth and development. In A. thaliana, Junker and Baumlein 2012 and Junker et al. 2012 identified that LEC1 TF affects light and brassinosteroid (BR) signaling during embryogenesis. In addition, LEC1 expression has also been detected in etiolated seedlings (Warpeha et al. 2007; Siefers et al. 2009; Junker et al. 2012).
Lateral root development is critical for higher plants and is responsible for the uptake of water and nutrient acquisition for the growth and development of plants (Charlton 1996; Tang et al. 2016). Embryonic master regulators LEC2 and FUS3 are involved in the lateral root formation by regulating YUC functions (Tang et al. 2016). In A. thaliana, a FUS3 and LEC2 complex function synergistically and activates auxin biosynthesis and YUC gene during lateral root formation. However, expression of FUS3 during lateral root formation is activated by LEC2 (Tang et al. 2016). In future, more extensive studies will hopefully provide novel findings for the better understanding of the molecular mechanisms of lateral root formation.
A number of studies have been reported that during post-embryonic development cell fate determination is controlled by TFs (Peris et al. 2010; Perianez-Rodriguez et al. 2014). Few researchers documented a significant role of LEC1 in post-embryonic cell differentiation, including formation of trichomes, mesophyll cells and vascular tissue (Junker and Baumlein 2012; Junker et al. 2012). The regulation of trichome formation by different TFs has been well studied by several researchers (Marks and Feldmann 1989; Oppenheimer et al. 1991; Wang and Chen 2014; Zhao et al. 2008; Zhou et al. 2014). Huang et al. (2015a) documented that LEC1 is positively involved in cell fate determination during post-embryonic development in A. thaliana. They found that LEC1 interacts in vitro with transcription repressors such as TRICHOMELESS1/2 (TCL1/2), CAPPICE (CPC) and ENHANCER OF TRY AND CPC1 (ETC1) to repress trichome formation. It was identified that TCL1 was highly expressed in developing seeds, whereas TCL2 in cotyledons (Wang et al. 2007a, b; Gan et al. 2011). The interaction with these repressors provides a mechanism by which LEC1 regulates cell fate determination.
In a recent breakthrough report by Tao et al. (2019), they discovered that LEC2 and FUS3 TFs are involved in expression of key flowering gene and embryonic resetting in Arabidopsis. The LEC2 and FUS3 TFs compete against VAL1 and VAL2 (epigenome readers) to disrupt the Polycomb silencing during early embryogenesis. Furthermore, LEC2 and FUS3 recruit the FRIGIDA (scaffold protein) in order to establish an active chromatin state, resulting in the activation of FLC (FLOWERING LOCUC C, a floral repressor) and erasing the parental memory in early somatic embryos during winter cold. However, LEC2 and FUS3 were silenced during post-embryonic phase (Tao et al. 2019). Reprogramming of epigenetic mechanisms during embryogenesis by LEC TF is well reported (Tao et al. 2017).
Further research will help to unlock the different biological and molecular mechanism underlying these processes. These results suggest that over-expression of LEC gene has been used for multifunction in different plant species. Increasing current advances and better understanding of the mechanism of LEC genes will lead to new opportunities and development of different biological applications.
Conclusions and future perspectives
The findings presented reveal that the LEC transcription factor genes have emerged as a master regulator that controls diverse aspects of somatic embryogenesis and has potential application in the plant biology. The LEC TFs are used as candidate markers to define the molecular mechanisms that control the initiation and maturation phase of SE. The expression of the LEC gene provides clear evidence of its role in embryogenesis and diverse developmental signaling pathways including oil content accumulation, cell fate determination, lateral root development and chloroplast biogenesis and photosynthesis during seed development. The LEC gene acts as a master regulator to participate in initiation and maturation of somatic embryos but how the LEC-mediated cellular process initiates the maturation phase is still unclear. In addition, how the LEC TFs control signaling transmission specificity to regulate initiation and maturation of somatic embryo at the molecular level remains unclear. In addition, LEC also acts as a pioneer TF gene, which activates different sets of genes and controls diverse biological processes during SD. However, we are only at the beginning to understand the potential insight and molecular mechanism by which LEC1 regulates diverse functions of SD. Recent discoveries have explored the multiple roles of LEC TFs in diverse aspects of plant growth and development. However, a few future challenges still need to be clarified such as LEC1 and LEC2 are involved in the diverse signaling pathways related to embryogenesis including embryo morphogenesis and maturation, but how these pathways are regulated remains unclear. In addition, how is the specificity of these LEC TFs obtained? Apart from known processes, what additional physiological and biological processes are regulated by LEC TFs? The underlying molecular mechanism by which LEC regulates diverse biological processes of SD is still unclear. Therefore, research should shed some light on how these LEC TFs control embryogenesis and several aspects of plant dynamics. In order to decode these regulatory networks, a single-molecule imaging technology will be required to understand the diverse functions of individual LECs in different signaling pathways. Together, structural studies of different LECs may open new roadmaps for better understanding their signaling specificity and developmental plasticity. It would also help to find new insights into the molecular mechanisms and unexplored signaling pathways for the better understanding of the functions of LEC TFs in plant cells.
Abbreviations
- ABI3 :
-
ABSCISIC ACID INSENSITIVE3
- BBM :
-
BABY BOOM
- FUS3 :
-
FUSCA3
- AGL15 :
-
Agamous-Like 15
- IAA30:
-
Indole acetic acid inducible 30
- LEC :
-
LEAFY COTYLEDON
- PGRs:
-
Plant growth regulators
- PKL :
-
PICKLE
- SD:
-
Seed development
- SE:
-
Somatic embryogenesis
- SERK :
-
SOMATIC EMBRYOGENESIS RECEPTOR LIKE KINASE
- TCL:
-
TRICHOMELESS
- TFs:
-
Transcription factors
- WUS :
-
WUSCHEL
References
Alemanno L, Devic M, Niemenak N, Sanier C, Guilleminot J, Rio M, Verdeil JL, Montoro P (2007) Characterization of leafy cotyledon1-like during embryogenesis in Theobroma cacao L. Planta 227:853–866
Angeles-Núñez JG, Tiessen A (2011) Mutation of the transcription factor LEAFY COTYLEDON 2 alters the chemical composition of Arabidopsis seeds, decreasing oil and protein content, while maintaining high levels of starch and sucrose in mature seeds. J Plant Physiol 168:1891–1900
Arroyo-Herrera A, Gonzalez AK, Moo RC, Quiroz-Figueroa F, Loyola-Vargas V, Rodriguez-Zapata L, Burgeff D’Hondt C, Suárez-Solís VM, Castano E (2008) Expression of WUSCHEL in Coffea canephora causes ectopic morphogenesis and increases somatic embryogenesis. Plant Cell Tissue Organ Cult 94:171–180
Baud S, Mendoza MS, To A, Harscoët E, Lepiniec L, Dubreucq B (2007) WRINKLED1 specifies the regulatory action of LEAFY COTYLEDON2 towards fatty acid metabolism during seed maturation in Arabidopsis. Plant J 50:825–838
Baud S, Kelemen Z, Thevenin J, Boulard C, Blanchet S, To A, Payre M, Berger N, Effroy-Cuzzi D, Franco-Zorrilla JM, Godoy M, Solano R, Thevenon E, Parcy F, Lepiniec L, Dubreucq B (2016) Deciphering the molecular mechanisms underpinning the transcriptional control of gene expression by master transcriptional regulators in Arabidopsis seed. Plant Physiol 171:1099–1112
Bäumlein H, Miséra S, Leurben H, Kölle K, Horstman C, Wobus U, Müller AJ (1994) The FUS3 gene of Arabidopsis thaliana is a regulator of gene expression during late embryogenesis. Plant J 6:379–387
Bouchabké-Coussa O, Obellianne M, Linderme D, Montes E, Maia-Grondard A, Vilaine F, Pannetier C (2013) Wuschel overexpression promotes somatic embryogenesis and induces organogenesis in cotton (Gossypium hirsutum L.) tissues cultured in vitro. Plant Cell Rep 32:675–686
Boulard C, Fatihi A, Lepiniec L, Dubreucq B (2017) Regulation and evolution of the interaction of the seed B3 transcription factors with NF-Y subunits. Biochim Biophys Acta Gene Regul Mech 1860:1069–1078
Boulard C, Thevenin J, Tranquet O, Laporte V, Lepiniec L, Dubreucq B (2018) LEC1 (NF-YB9) directly interacts with LEC2 to control gene expression in seed. Biochim Biophys Acta 1861:443–450
Boutilier K, Offringa R, Sharma VK, Kieft H, Ouellet T, Zhang L, Hattori J, Liu C, van Lammeren AAM, Miki BLA, Custers JBM, van Lookeren Campagne MM (2002) Ectopic Expression of BABY BOOM Triggers a Conversion from Vegetative to Embryonic Growth. Plant Cell 14:1737–1749
Brand A, Quimbaya M, Tohme J, Chavariagga-Aguirre P (2019) Arabidopsis LEC1 and LEC2 orthologous genes are key regulators of somatic embryogenesis in Cassava. Front Plant Sci 10:673
Braybrook SA, Harada JJ (2008) LECs go crazy in embryo development. Trends Plant Sci 13:624–630
Braybrook SA, Stone SL, Park S, Bui AQ, Le BH, Fischer RL, Goldberg RB, Harada JJ (2006) Genes directly regulated by LEAFY COTYLEDON2 provide insight into the control of embryo maturation and somatic embryogenesis. Proc Natl Acad Sci USA 103:3468–3473
Broun P (2004) Transcription factors as tools for metabolic engineering in plants. Curr Opin Plant Biol 7:202–209
Cernac A, Benning C (2004) WRINKLED1 encodes an AP2/EREB domain protein involved in the control of storage compound biosynthesis in Arabidopsis. Plant J 40:575–585
Chan A, Carianopol C, Tsai AY, Varatharajah K, Chiu RS, Gazzarrini S (2017) SnRK1 phosphorylation of FUSCA3 positively regulates embryogenesis, seed yield, and plant growth at high temperature in Arabidopsis. J Exp Bot 68:4219–4231
Charlton WA (1996) Lateral root initiation. In: Waisel Y, Eshel A, Kafkafi U (eds) Plant roots: the hidden half. Marcel Dekker, New York, pp 149–174
Curaba J, Moritz T, Blervaque R, Parcy F, Raz V, Herzog M, Vachon G (2004) AtGA3ox2, a key gene responsible for bioactive gibberellin biosynthesis, is regulated during embryogenesis by LEAFY COTYLEDON2 and FUSCA3 in Arabidopsis. Plant Physiol 136:3660–3669
Deng W, Luo KM, Li ZG, Yang YW (2009) A novel method for induction of plant regeneration via somatic embryogenesis. Plant Sci 177:43–48
Devic M, Roscoe T (2016) Seed maturation: simplification of control networks in plants. Plant Sci 252:335–346
Dickinson CD, Evans RP, Nielsen NC (1988) RY repeats are conserved in the 5'-flanking regions of legume seed-protein genes. Nucleic Acids Res 16:371
Domoki M, Györgyey J, Bíró J, Pasternak TP, Zvara Á, Bottka S, Puskás LG, Dudits D, Fehér A (2006) Identification and characterization of genes associated with the induction of embryonic competence in leaf-protoplast-derived alfalfa cells. Biochimet Biophys Acta 1759:543–551
Feeney M, Frigerio L, Cui Y, Menassa R (2013) Following vegetative to embryonic cellular changes in leaves of Arabidopsis overexpressing LEAFY COTYLEDON2. Plant Physiol 162:1881–1896
Fister AS, Landherr L, Perryman M, Zhang Y, Guiltinan MJ, Maximova SN (2018) Glucocorticoid receptor-regulated TcLEC2 expression triggers somatic embryogenesis in Theobroma cacao leaf tissue. PLoS ONE 13:e0207666
Florez SL, Erwin RL, Maximova SN, Guiltinan MJ, Curtis WR (2015) Enhanced somatic embryogenesis in Theobroma cacao using the homologous BABY BOOM transcription factor. BMC Plant Biol 15:121
Gaj MD, Zhang S, Harada JJ, Lemaux PG (2005) LEAFY COTYLEDON genes are essential for induction of somatic embryogenesis of Arabidopsis. Planta 222:977–988
Gan LJ, Xia K, Chen JG, Wang SC (2011) Functional characterization of TRICHOMELESS2, a new single-repeat R3 MYB transcription factor in the regulation of trichome patterning in Arabidopsis. BMC Plant Biol 11:176
Gazzarrini S, Tsuchiya Y, Lumba S, Okamoto M, McCourt P (2004) The transcription factor FUSCA3 controls developmental timing in Arabidopsis through the hormones gibberellin and abscisic acid. Dev Cell 7:373–385
Goldberg RB, De Paiva G, Yadegari R (1994) Plant embryogenesis: zygote to seed. Science 266:605–614
Grotewold E (2008) Transcription factors for predictive plant metabolic engineering: are we there yet? Curr Opin Biotechnol 19:138–144
Harada JJ (1997) Seed maturation and control of germination. In: Larkins BA, Vasi IK (eds) Advances in cellular and molecular biology of plants cellular and molecular biology of seed development, vol 4. Kluwer Academic Publishers, Dordrecht, pp 545–592
Harada JJ (2001) Role of Arabidopsis LEAFY COTYLEDON genes in seed development. J. Plant Physiol 158:405–409
Harding EW, Tang W, Nichols KW, Fernandez DE, Perry SE (2003) Expression and maintenance of embryogenic potential is enhanced through constitutive expression of AGAMOUSLIKE15. Plant Physiol 133:653–663
Hecht V, Vielle-Calzada JP, Hartog MV, Schmidt ED, Boutilier K, Grossniklaus U, de Vries SC (2001) The Arabidopsis SOMATIC EMBRYOGENESIS RECEPTOR KINASE 1 gene is ex- pressed in developing ovules and embryos and enhances embryogenic competence in culture. Plant Physiol 127:803–816
Heck GR, Perry SE, Nichols KW, Fernandez DE (1995) AGL15, a MADS domain protein expressed in developing embryos. Plant Cell 7:1271–1282
Horstman A, Bemer M, Boutilier K (2017a) A transcriptional view on somatic embryogenesis. Regeneration 4:201–216
Horstman A, Li M, Heidmann I, Weemen M, Chen B, Muino JM, Angenent GC, Boutilier K (2017b) The BABY BOOM transcription factor activates the LEC1-ABI3-FUS3-LEC2 network to induce somatic embryogenesis. Plant Physiol 175:848–857
Huang M, Hu Y, Liu X, Li Y, Hou X (2015a) ArabidopsisLEAFY COTYLEDON1 controls cell fate determination during post-embryonic development. Front Plant Sci 6:955
Huang M, Hu Y, Liu X, Li Y, Hou X (2015b) Arabidopsis LEAFY COTYLEDON1 mediates postembryonic development via interacting with PHYTOCHROME-INTERACTING FACTOR4. Plant Cell 27:3099–3111
Ikeda M, Umehara M, Kamada H (2006) Embryogenesis-related genes; its expression and roles during somatic and zygotic embryogenesis in carrot and Arabidopsis. Plant Biotechnol 23:153–161
Irikova T, Grozeva S, Denev I (2012) Identification of BABY BOOM and LEAFY COTYLEDON genes in sweet pepper (Capsicum annuum L.) genome by their partialgene sequences. Plant Grow Regul 67:191–198
Jha P, Kumar V (2018) BABY BOOM (BBM): a candidate transcription factor gene in plant biotechnology. Biotechnol Lett 40:1467–1475
Jia H, Suzuki M, McCarty DR (2014) Regulation of the seed to seedling developmental phase transition by the LAFL and VAL transcription factor networks. Wiley Interdiscip Rev Dev Biol 3:135–145
Jo L, Pelletier JM, Harada JJ (2019) Central role of the LEAFY COTYLEDON1 transcription factor in seed development. J Integ Plant Biol 61:564–580
Jr Rider SD, Henderson JT, Jerome RE, Edenberg HJ, Romero- Severson J, Ogas J (2003) Coordinate repression of regulators of embryogenic identity by PICKLE during germination in Arabidopsis. Plant J 35:33–43
Junker A, Bäumlein H (2012) Multifunctionality of the LEC1 transcription factor during plant development. Plant Signal Behav 7:1718–1720
Junker A, Mönke G, Rutten T, Keilwagen J, Seifert M, Thi TMN, Renou JP, Balzergue S, Viehöver P, Hähnel U, Ludwig-Müller J, Altschmied L, Conrad U, Weisshaar B, Bäumlein H (2012) Elongation-related functions of LEAFY COTYLEDON1 during the development of Arabidopsis thaliana. Plant J 71:427–442
Kagaya Y, Okuda R, Ban A, Toyoshima R, Tsutsumida K, Usui H, Yamamoto A, Hattori T (2005a) Indirect ABA-dependent regulation of seed storage protein genes by FUSCA3 transcription factor in Arabidopsis. Plant Cell Physiol 46:300–311
Kagaya Y, Toyoshima R, Okuda R, Usui H, Yamamoto A, Hattori T (2005b) LEAFY COTYLEDON1 controls seed storage protein genes through its regulation of FUSCA3 and ABSCISIC ACID INSENSITIVE3. Plant Cell Physiol 46:399–406
Keith K, Kraml M, Dengler NG, McCourt P (1994) fusca3: a heterochronic mutation affecting late embryo development in Arabidopsis. Plant Cell 6:589–600
Kim HU, Jung SJ, Lee KR, Kim EH, Lee SM, Roh KH, Kim JB (2014) Ectopic overexpression of castor bean LEAFY COTYLEDON2 (LEC2) in Arabidopsis triggers the expression of genes that encode regulators of seed maturation and oil body proteins in vegetative tissues. FEBS Open Bio 4:25–32
Kirkbride RC, Fischer RL, Harada JJ (2013) LEAFY COTYLEDON1, a key regulator of seed development, is expressed in vegetative and sexual propagules of Selaginella moellendorffii. PLoS ONE 8:e67971
Kroj T, Savino G, Valon C, Giraudat J, Parcy F (2003) Regulation of storage protein gene expression in Arabidopsis. Development 130:6065–6073
Kumar V, Van Staden J (2017) New insights into plant somatic embryogenesis: an epigenetic view. Acta Physiol Plant 39:194
Kumar V, Van Staden J (2019) Multi-tasking of SERK-like kinases in plant embryogenesis, growth and development: current advances and biotechnological applications. Acta Physiol Plant 41:31
Kwong RW, Bui AQ, Lee H, Kwong LW, Fischer RL, Goldberg RB, Harada JJ (2003) LEAFY COTYLEDON1-LIKE defines a class of regulators essential for embryo development. Plant Cell 15:5–18
Laux T, Jurgens G (1997) Embryogenesis: a new start in life. Plant Cell 9:989–1000
Ledwoń A, Gaj MD (2011) LEAFY COTYLEDON1, FUSCA3 expression and auxin treatment in relation to somatic embryogenesis induction in Arabidopsis. Plant Grow Regul 65:157–167
Lee H, Fischer RL, Goldberg RB, Harada JJ (2003) Arabidopsis LEAFY COTYLEDON1 represents a functionally specialized subunit of the CCAAT binding transcription factor. Proc Natl Acad Sci USA 100:2152–2156
Lepiniec L, Devic M, Roscoe TJ, Bouyer D, Zhou DX, Boulard C, Baud S, Dubreucq B (2018) Molecular and epigenetic regulations and functions of the LAFL transcriptional regulators that control seed development. Plant Reprod 31:291–307
Liu Z, Ge XX, Qiu WM, Long JM, Jia HH, Yang W, Dutt M, Wu XM, Guo WW (2018) Overexpression of the CsFUS3 gene encoding a B3 transcription factor promotes somatic embryogenesis in Citrus. Plant Sci 277:121–131
Lotan T, Ohto M, Yee KM, West MAL, Lo R, Kwong RW, Tamagishi K, Fisher RL, Goldberg RB, Harada JJ (1998) Arabidopsis LEAFY COTYLEDON1 is sufficient to induce embryo development in vegetative cells. Cell 93:1195–1205
Luerssen H, Kirik V, Herrmann P, Miséra S (1998) FUSCA3 encodes a protein with a conserved VP1/AB13-like B3 domain which is of functional importance for the regulation of seed maturation in Arabidopsis thaliana. Plant J 15:755–764
Manan S, Ahmad MZ, Zhang G, Chen B, Haq BU, Yang J, Zhao J (2017) Soybean LEC2 regulates subsets of genes involved in controlling the biosynthesis and catabolism of seed storage substances and seed development. Front Plant Sci 8:1604
Mangan S, Alon U (2003) Structure and function of the feed-forward loop network motif. Proc Natl Acad Sci USA 100:11980–11985
Marks MD, Feldmann KA (1989) Trichome development in Arabidopsis-thaliana.1. T-DNA tagging of the Glabrous1 gene. Plant Cell 1:1043–1050
Meinke DW (1992) A homoeotic mutant of Arabidopsis thaliana with leafy cotyledons. Science 258:1647–1650
Meinke DW, Yeung EC (1993) Embryogenesis in angiosperms: development of the suspensor. Plant Cell 5:1371–1381
Meinke DW, Franzmann LH, Yeung EC (1994) Leafy Cotyledon mutants of Arabidopsis. Plant Cell 6:1049–1064
Mendes A, Kelly AA, van Erp H, Shaw E, Powers SJ, Kurup S, Eastmond PJ (2013) bZIP67 regulates the omega-3 fatty acid content of Arabidopsis seed oil by activating fatty acid desaturase3. Plant Cell 25:3104–3116
Méndez-Hernández HA, Ledezma-Rodríguez M, Avilez-Montalvo RN, Juárez-Gómez YL, Skeete A, Avilez-Montalvo J, De-la-Peña C, Loyola-Vargas VM (2019) Signaling overview of plant somatic embryogenesis. Front Plant Sci 10:77
Mendoza MS, Dubreucq B, Miquel M, Caboche M, lepineic L, (2005) LEAFY COTYLEDON 2 activation is sufficient to trigger the accumulation of oil and seed specific mRNAs in Arabidopsis leaves. FEBS Lett 579:4666–4670
Min L, Hu Q, Li Y, Xu J, Ma Y, Zhu L, Yang X, Zhang X (2015) LEAFY COTYLEDON1-CASEIN KINASE I-TCP15-PHYTOCHROME INTERACTING FACTOR4 network regulates somatic embryogenesis by regulating auxin homeostasis. Plant Physiol 169:2805–2821
Mönke G, Altschmied L, Tewes A, Reidt W, Mock HP, Bäumlein H, Conrad U (2004) Seed-specific transcription factors ABI3 and FUS3: molecular interaction with DNA. Planta 219:158–166
Nambara E, Hayama R, Tsuchiya Y, Nishimura M, Kawaide H, Kamiya Y, Naito S (2000) The role of ABI3 and FUS3 loci in Arabidopsis thaliana on phase transition from late embryo development to germination. Dev Biol 220:412–413
Nic-Can GI, López-Torres A, Barredo-Poll F, Wrobel K, LoyolaVargas VM, Rojas-Herrera R, De-la-Peña C (2013) New Insights into somatic embryogenesis: LEAFY COTYLEDON1, BABY BOOM1 and WUSCHEL-RELATED HOMEOBOX4 are epigenetically regulated in Coffea canephora. PLoS ONE 8:e72160
Nolan KE, Kurdyukov S, Rose RJ (2009) Expression of the SOMATIC EMBRYOGENESIS RECEPTOR-LIKE LINASE1 (SERK1) gene is associated with developmental change in the life cycle of the model legume Medicago truncatulata. J Exp Bot 60:1759–1771
Ogas J, Cheng JC, Sung ZR, Somerville C (1997) Cellular differentiation regulated by gibberellin in the Arabidopsis thaliana pickle mutant. Science 277:91–94
Ogas J, Kaufmann S, Henderson J, Somerville C (1999) PICKLE is a CHD3 chromatin-remodeling factor that regulates the transition from embryonic to vegetative development in Arabidopsis. Proc Natl Acad Sci USA 96:13839–13844
Ohlrogge J, Browse J (1995) Lipid biosynthesis. Plant Cell 7:957–970
Oppenheimer DG, Herman PL, Sivakumaran S, Esch J, Marks MD (1991) A myb gene required for leaf trichome differentiation in Arabidopsis is expressed in stipules. Cell 67:483–493
Orlowska A, Igielska R, Lagowska K, Kepczynska E (2017) Identification of LEC1, L1Land Polycomb Repressive Complex 2 genes and their expression during the induction phase of Medicago truncatula Gaertn. somatic embryogenesis. Plant Cell Tiss Organ Cult 129:119–132
Parcy F, Valon C, Kohara A, Misera S, Giraudat J (1997) The ABSCISIC ACID-INSENSITIVE3, FUSCA3, and LEAFY COTYLEDON1 loci act in concert to control multiple aspects of Arabidopsis seed development. Plant Cell 9:1265–1277
Pelletier JM, Kwong RW, Park S, Le BH, Baden R, Cagliari A, Hashimoto M, Munoz MD, Fischer RL, Goldberg RB, Harada JJ (2017) LEC1 sequentially regulates the transcription of genes involved in diverse developmental processes during seed development. Proc Natl Acad Sci USA 114:6710–6719
Perianez-Rodriguez J, Manzano C, Moreno-Risueno MA (2014) Post-embryonic organogenesis and plant regeneration from tissues: two sides of the same coin? Front Plant Sci 5:219
Peris CIL, Rademacher EH, Weijers D (2010) Green beginnings—pattern formation in the early plant embryo. Plant Dev 91:1–27
Perry SE, Lehti MD, Fernandez DE (1999) The MADS-domain protein AGAMOUS-like 15 accumulates in embryonic tissues with diverse origins. Plant Physiol 120:121–129
Pilarska M, Malec P, Salaj J, Bartnicki F, Konieczny R (2016) High expression of SOMATIC EMBRYOGENESIS RECEPTOR-LIKE KINASE coincides with initiation of various developmental pathways in in vitro culture of Trifolium nigrescens. Protoplasma 253:345–355
Pradhan S, Bandhiwal N, Shah N, Kant C, Gaur R, Bhatia S (2014) Global transcriptome analysis of developing chickpea (Cicer arietinumL.) seeds. Front Plant Sci 5:698
Reidt W, Wohlfarth T, Ellerstrom M, Czihal A, Tewes A, Ezcurra I, Rask L, Bäumlein H (2000) Gene regulation during late embryogenesis: the RY motif of maturation-specific gene promoters is a direct target of the FUS3 gene product. Plant J 21:401–408
Roscoe TJ, Vaissayre V, Paszkiewicz G, Clavijo F, Kelemen Z, Michaud C, Lepiniec L, Dubreucq B, Zhou DX, Devic M (2019) Regulation of FUSCA3 expression during seed development in Arabidopsis. Plant Cell Physiol 60:476–487
Rounsley SD, Ditta GS, Yanofsky MF (1995) Diverse roles for MADS box genes in Arabidopsis development. Plant Cell 7:1259–1269
Salvo SAGD, Hirsch CN, Buell CR, Kaeppler SM, Kaeppler HF (2014) Whole transcriptome profiling of maize during early somatic embryogenesis reveals altered expression of stress factors and embryogenesis-related genes. PLoS ONE 29:e111407
Santos MO, Romanoa E, Yotoko KSC, Tinoco MLP, Dias BBA, Aragao FJL (2005) Characterization of the cacao somatic embryogenesis receptor-like kinase (SERK) gene expressed during somatic embryogenesis. Plant Sci 168:723–729
Santos-Mendoza M, Dubreucq B, Baud S, Parcy F, Caboche M, Lepiniec L (2008) Deciphering gene regulatory networks that control seed development and maturation in Arabidopsis. Plant J 54:608–620
Schmidt ED, Guzzo F, Toonen MA, de Vries SC (1997) A leucine rich repeat containing receptor-like kinase marks somatic plant cells competent to form embryos. Development 124:2049–2062
Shen B, Allen WB, Zheng P, Li C, Glassman K, Ranch J, Nubel D, Tarczynski MC (2010) Expression of ZmLEC1 and ZmWRI1 increases seed oil production in maize. Plant Physiol 153:980–987
Shires ME, Florez SL, Lai TS, Curtis WR (2017) Inducible somatic embryogenesis in Theobroma cacao achieved using the DEX-activatable transcription factor-glucocorticoid receptor fusion. Biotechnol Lett 39:1747–1755
Siefers N, Dang KK, Kumimoto RW, Bynum WE, Tayrose G, Holt BF (2009) Tissue-specific expression patterns of Arabidopsis NF-Y transcription factors suggest potential for extensive combinatorial complexity. Plant Physiol 149:625–641
Stone SL, Kwong LW, Yee KM, Pelletier J, Lepiniec L, Fischer RL, Goldberg RB, Harada JJ (2001) LEAFY COTYLEDON2 encodes B3 domain transcription factor that induces embryo development. Proc Natl Acad Sci USA 98:11806–11811
Stone SL, Braybrook SA, Paula SL, Kwong LW, Meuser J, Pelletier J, Hsieh T, Fischer RL, Goldberg RB, Harada JJ (2008) Arabidopsis LEAFY COTYLEDON2 induces maturation traits and auxin activity: Implications for somatic embryogenesis. Proc Natl Acad Sci USA 105:3151–3156
Suzuki M, Wang HHY, McCarty DR (2007) Repression of the LEAFY COTYLEDON 1/B3 Regulatory Network in Plant Embryo Development by VP1/ABSCISIC ACID INSENSITIVE 3-LIKE B3 Genes. Plant Physiol 143:902–911
Tan H, Yang X, Zhang F, Zheng X, Qu C, Mu J, Fu F, Li J, Guan R, Zhang H, Wang G, Zuo J (2011) Enhanced seed oil production in canola by conditional expression of Brassica napus LEAFY COTYLEDON1and LEC1-LIKE in developing seeds. Plant Physiol 156:1577–1588
Tang LP, Zhou C, Wang SS, Yuan J, Zhang XS, Su YH (2016) FUSCA3 interacting with LEAFY COTYLEDON2 controls lateral root formation through regulating YUCCA4 gene expression in Arabidopsis thaliana. New Phyt 213:1740–1754
Tao Z, Shen L, Gu X, Wang Y, Yu H, He Y (2017) Embryonic epigenetic reprogramming by a pioneer transcription factor in plants. Nat 551:124–128
Tao Z, Hu H, Luo X, Jia B, Du J, He Y (2019) Embryonic resetting of the parental vernalized state by two B3 domain transcription factors in Arabidopsis. Nat Plant 5:424–435
Thakare D, Tang W, Hill K, Perry SE (2008) The MADS-domain transcriptional regulator AGAMOUS-Like 15 promotes somatic embryo development in Arabidopsis and soybean. Plant Physiol 146:1663–1672
To A, Valon C, Savino G, Guilleminot J, Devic M, Giraudat J, Parcy F (2006) A network of local and redundant gene regulation governs Arabidopsis seed maturation. Plant Cell 18:1642–1651
To A, Joubes J, Barthole G, Lecureuil A, Scagnelli A, Jasinski S, Lepiniec L, Baud S (2012) WRINKLED transcription factors orchestrate tissue-specific regulation of fatty acid biosynthesis in Arabidopsis. Plant Cell 24:5007–5023
Tsuwamoto R, Yokoi S, Takahata Y (2010) Arabidopsis EMBRYOMAKER encoding an AP2 domain transcription factor plays a key role in developmental change from vegetative to embryonic phase. Plant Mol Biol 73:481–492
Tvorogova VE, Fedorova YA, Potsenkovskaya EA, Kudriashov AA, Efremova EP, Kvitkovskaya VA, Wolabu TW, Zhang F, Tadege M, Lutova LA (2019) The WUSCHEL-related homeobox transcription factor MtWOX9-1 stimulates somatic embryogenesis in Medicago truncatula. Plant Cell Tissue Organ Cult 138:517–527
Van Erp H, Kelly AA, Menard G, Eastmond PJ (2014) Multigene engineering of triacylglycerol metabolism boosts seed oil content in Arabidopsis. Plant Physiol 165:30–36
Vicente-Carbajosa J, Carbonaro P (2005) Seed maturation: developing an intrusive phase to accomplish a quiescent state. Int J Plant Sci 49:645–651
Wang SC, Chen JG (2014) Regulation of cell fate determination by single-repeat R3 MYB transcription factors in Arabidopsis. Front Plant Sci 5:133
Wang F, Perry SE (2013) Identification of direct targets of FUSCA3, a key regulator of Arabidopsis seed development. Plant Physiol 161:1251–1264
Wang H, Caruso LV, Downie AB, Perry SE (2004) The embryo MADS domain protein AGAMOUS-Like 15 directly regulates expression of a gene encoding an enzyme involved in gibberellins metabolism. Plant Cell 16:1206–1219
Wang H, Guo J, Lambert KN, Lin Y (2007a) Developmental control of Arabidopsis seed oil biosynthesis. Planta 226:773–783
Wang SC, Kwak SH, Zeng QN, Ellis BE, Chen XY, Schiefelbein J, Chen JG (2007b) TRICHOMELESS1 regulates trichome patterning by suppressing GLABRA1 in Arabidopsis. Development 134:3873–3882
Warpeha KM, Upadhyay S, Yeh J, Adamiak J, Hawkins SI, Lapik YR, Anderson MB, Kaufman LS (2007) The GCR1, GPA1, PRN1, NF-Y signal chain mediates both blue light and abscisic acid responses in Arabidopsis. Plant Physiol 143:1590–1600
West MAL, Harada JJ (1993) Embryogenesis in higher plants: an overview. Plant Cell 5:1361–1369
West MAL, Yee KM, Danao J, Zimmerman JL, Fischer RL, Goldberg RB, Harada JJ (1994) LEAFY COTYLEDON1 is an essential regulator of late embryogenesis and cotyledon ldentity in Arabidopsis. Plant Cell 6:1731–1745
Williams EG, Maheswaran G (1986) Somatic embryogenesis: factors influencing coordinated behaviour of cells as an embryogenic group. Ann Bot 57:443–462
Wójcikowska B, Jaskóla K, Gasiorek P, Meus M, Nowak K, Gaj MD (2013) LEAFY COTYLEDON2 (LEC2) promotes embryogenic induction in somatic tissues of Arabidopsis, via YUCCA-mediated auxin biosynthesis. Planta 238:425–440
Xu JJ, Zhang XF, Xue HW (2016) Rice aleurone layer specific OsNF-YB1 regulates grain filling and endosperm development by interacting with an ERF transcription factor. J Exp Bot 67:6399–6411
Yang X, Zhang X (2010) Regulation of somatic embryogenesis in higher plants. Crit Rev Plant Sci 29:36–57
Yazawa K, Takahata K, Kamada H (2004) Isolation of the gene encoding Carrot leafy cotyledon 1 and expression analysis during somatic and zygotic embryogenesis. Plant Physiol Biochem 42:215–223
Zhang JJ, Xue HW (2013) OsLEC1/OsHAP3E participates in the determination of meristem identity in both vegetative and reproductive developments of rice. J Integr Plant Biol 55:232–249
Zhang SB, Wong L, Meng L, Lemaux PG (2002) Similarity expression patterns of knotted1 and ZmLEC1 during somatic and zygotic embryogenesis in Maize (Zea mays L.). Planta 215:191–194
Zhang S, Liu X, Lin Y, Xie G, Fu F, Liu H, Wang J, Gao S, Lan H, Rong T (2011) Characterization of a ZmSERK gene and its relationship to somatic embryogenesis in a maize culture. Plant Cell Tiss Org Cult 105:29–37
Zhao M, Morohashi K, Hatlestad G, Grotewold E, Lloyd A (2008) The TTG1-bHLH-MYB complex controls trichome cell fate and patterning through direct targeting of regulatory loci. Development 135:1991–1999
Zheng W, Zhang X, Yang Z, Wu J, Li F, Duan L, Liu C, Lu L, Zhang C, Li F (2014) AtWuschel promotes formation of the embryogenic callus in Gossypium hirsutum. PLoS ONE 9:e87502
Zheng Q, Zheng Y, Ji H, Burnie W, Perry SE (2016) Gene regulation by the AGL15 transcription factor reveals hormone interactions in somatic embryogenesis. Plant Physiol 172:2374–2387
Zhou LM, Zheng KJ, Wang XY, Tian HN, Wang XL, Wang SC (2014) Control of trichome formation in Arabidopsis by poplar single-repeat R3 MYB transcription factors. Front Plant Sci 5:262
Zhou X, Guo Y, Zhao P, Sun M (2018) Comparative analysis of WUSCHEL-related homeobox genes revealed their parent-of-origin and cell type-specific expression pattern during early embryogenesis in Tobacco. Front Plant Sci 9:311
Zhu S, Wang J, Ye J, Zhu A, Guo W, Deng X (2014) Isolation and characterization of LEAFY COTYLEDON 1-LIKE gene related to embryogenic competence in Citrus sinensis. Plant Cell Tiss Organ Cult 119:1–13
Zhu Y, Xie L, Chen GQ, Lee MY, Loque D, Scheller HV (2018) A transgene design for enhancing oil content in Arabidopsis and Camelina seeds. Biotechnol Biofuels 11:46
Zuo J, Niu QW, Frugis G, Chua NH (2002) The WUSCHEL gene promotes vegetative-to-embryonic transition in Arabidopsis. Plant J 30:349–359
Acknowledgements
The authors acknowledge the University of KwaZulu-Natal (UKZN), South Africa for providing financial support. All the authors read and approved the final version of the manuscript. We apologize to all colleagues whose work has not been cited in this manuscript due to space constraints.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Additional information
Communicated by Konstantin V. Kiselev.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kumar, V., Jha, P. & Van Staden, J. LEAFY COTYLEDONs (LECs): master regulators in plant embryo development. Plant Cell Tiss Organ Cult 140, 475–487 (2020). https://doi.org/10.1007/s11240-019-01752-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-019-01752-x