Abstract
Somatic embryogenesis (SE) is an intricate in vitro multi-step biotechnological tool used to develop embryos/plants from a single or a group of somatic cells. It is a model technique for understanding various plant developmental pathways. A lot of research is going on to elucidate the mechanism underlying the process of SE. This study was aimed at the identification of SE related proteins in a medicinally important plant, Catharanthus roseus via label free liquid chromatography–mass spectroscopy (LC–MS). LC–MS is a sensitive and reliable technique than the gel based techniques, using LC–MSMS in tandem for separation and identification of proteins. Here, we are reporting for the first time SE related proteins in C. roseus by using gel free shotgun proteomic approach. The non embryogenic and embryogenic calli of C. roseus were used for comparative proteome analysis. A total of 3573 proteins were identified in both embryogenic and non embryogenic calli of which 1511 proteins were found to be common in both the calli. In non embryogenic callus 982 proteins while in embryogenic callus 1079 proteins were exclusively identified, which were associated with varied cellular functions. The most of these proteins function in different metabolic processes and stress responses. More than 72 stress responsive proteins and isoforms were observed exclusively in embryogenic callus including glutathione S transferase, ascorbate peroxidase, catalase, superoxide dismutase, alkylhydro peroxidase, SOD Fe N domain containing protein, pyridine nucleotide disulphide oxidoreductase, thioredoxin reductase. The role of plant growth regulators (PGRs) in inducing stress cause switching on/off of several genes has been discussed, led biochemical and molecular alterations in acquiring somatic embryogenic competence.
Key message
Proteomic map of Catharanthus roseus was prepared. A total of 3573 proteins were identified, of which 1079 were embryogenic. These proteins have role in metabolic and stress responses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nature has bestowed on mankind a vast and diversified plant community, which unfortunately is currently exploited at an alarming rate (Chen et al. 2016). A lot of measures are being taken up nationally and internationally to conserve plant diversity and sustainably use plant resources for the welfare of mankind. One of the strategies for plant diversity conservation is the in vitro production of plants or plant parts to meet both the goals of plant propagation and their commercial utilization. Tissue culture technology has been very promising in fulfilling the goal of in vitro propagation and conservation of elite plant germplasm. This has only been possible after the discovery of totipotency of plant cells and associated process such as somatic embryogenesis. SE is an intricate in vitro multi-step biotechnological tool used to develop embryos/plants from a single or a group of somatic cells. The process has been utilized to address various problems and helps shortening the breeding period, synthetic seed development, wide hybridization via embryo rescue, scale up plant propagation etc (Lu et al. 2017).
Plant growth regulators (PGRs) play a very significant role in the azygotic embryogenesis process. Different PGRs are used depending on the nature of plant and the explants used. Among PGRs, auxin or cytokinin or both may be added to the culture medium (Imin et al. 2005; Nolan et al. 2003). Auxin is essentially used for the induction of SE and its concentration is subsequently lowered or completely omitted for further development of embryos. However, in C. roseus, auxin is critical for callus induction while the combinations of BAP and NAA are effective in inducing embryos (Mujib et al. 2014).
Somatic embryogenesis has been reported in a vast array of plants of diverse plant groups including ferns. Out of such vast plant diversity, an immensely important medicinal plant, C. roseus was selected for present study. The plant produces different alkaloids, the most important being vincristine and vinblastine, both having anti-cancerous properties. In SE transition of cells from non embryogenic to embryogenic state undergo an array of morphogical, biochemical and molecular changes, often triggered by stressful in vitro micro-environment (Schmidt et al. 1997; Feher 2015). Study of these morphological, biochemical and molecular changes is important for elucidating and understanding the molecular mechanism underlying the SE (Mordhorst et al. 1997; Mahdavi-Darvari et al. 2014; Heringer et al. 2015). SE is extensively studied at molecular level (Rupps et al. 2016; Orłowska et al. 2017) and several genes and proteins have been reported to take part in the process (Lu et al. 2017) but, still it is the least understood process (Feher 2015). Various tools and techniques are used to identify embryo specific proteins. One of the most reliable and sensitive tool that is currently used for the identification and quantification of proteins is the gel free based proteome analysis via liquid chromatography–mass spectroscopy (Karpievitch et al. 2010). Gel free proteomic approach is much more reliable and could identify those proteins that have low abundance and cannot be identified by gel-based techniques (Washburn et al. 2001). LC–MS is the bottom up approach in which the intricate protein mixtures are digested enzymatically, generating fragments of peptides which are ionized by different methods, identified on the basis of mass/charge (m/z) ratio, subsequently rolled up to proteins and further identification and quantification is made by comparing the ionized fragments with already identified proteins using various softwares like SEQUEST (Eng et al. 1994; Karpievitch et al. 2010). The present study was conducted to make a proteomic map of SE and to differentiate embryogenic callus from non-embryogenic tissue based on protein profiles in C. roseus. The study identifies a number of proteins that are exclusively present in the embryogenic calli most of which are stress related proteins. The present study shows stress as key player in the acquisition of embryogenic competence by non embryogenic/somatic cells.
Materials and methods
Seed germination and establishment of culture
The seeds of Catharanthus roseus (L.) G. Don were collected from the herbal garden of JamiaHamdard, New Delhi. The seeds were surface sterilized and placed in conical flask containing 50 ml solidified MS (Murashige and Skoog 1962), medium without any plant growth regulator (Junaid et al. 2008; Mujib et al. 2014). Seedlings were allowed to grow and various explants (leaf, hypocotyl and stem) were taken and inoculated in test tubes containing 8.0 ml solidified MS supplemented with 0.5, 1.0, 1.5,2.0, 2.5, and 3.0 mg/l 2,4-dichlorophenoxyacetic acid (2,4-D) for callus. After a few weeks, the induced callus of each explant was separately sub cultured in the same medium for proliferation. The calli obtained from different explants were transferred separately to MS, supplemented with various concentrations and combinations of N6-benzyladenine (0.5, 1.0, 1.5 and 2.0 mg/l) + naphthalene acetic acid (0.5, 1.0, 1.5 and 2.0 mg/l) Within 2–3 weeks, hypocotyl-induced embryogenic callus produced somatic embryos only while callus from other explants (leaf and stem) were observed to be non-embryogenic (no embryo formed) therefore was discontinued for protein study. The combination of 2.0 mg/l BAP and 1.5 mg/l NAA was noted to be the best treatment for somatic embryo induction. Repeated sub culturing of embryo bearing callus was conducted for subsequent embryo proliferation process. Some of the embryogenic calli with developed embryos were transferred to the MS medium added with 2.0 mg/l GA3 (gibberellic acid) for embryo maturation.
Protein extraction from tissues
The working samples of embryogenic and non embryogenic calli from hypocotyl were drawn separately for the extraction of total protein. The Isaacson et al. (2006) protocol (phenol method) was followed for the extraction of protein. Two and a half (2.5) grams of callus (pooled from three to five test tubes) of each (embryogenic and non embryogenic) was crushed to fine powder in liquid nitrogen using pre-chilled mortar pestle. The callus powder was suspended in 10 ml solution of phenylmethane sulfonylfluoride (PMSF), 4-(2-hydroxyethyl)-l-piperazine ethane sulfonic acid (HEPES), β-mercaptoethanol and sucrose containing extraction buffer in the Oak Ridge tubes. The solution was vortexed for 1.5 min and 15 ml of phenol was added to the solution and vortexed for another 1.5 min. Oak Ridge tubes containing the solution were sealed and covered with ice in tray for incubation on orbital shaker for 30 min in order to mix the solution. The solution was centrifuged at 5500 rpm at 4 °C for 10 min. The upper phenolic phase was carefully transferred to the separate tubes; 15 ml of ice-cold ammonium-acetate 0.1 M solution was added and incubated overnight at − 20 °C for protein precipitation. The proteins were pelleted by centrifuging at 10,000 rpm for 15 min at 4 °C. The pellet was washed thrice, first by ice cold methanol, incubated for 1 h. After centrifugation at 10,000 rpm for 10 min at 4 °C the pellet was resuspended in ice cold acetone and incubated for 1 h. The protein was again pelleted at 10,000 rpm for 10 min at 4 °C and the same step was repeated. Pellet obtained was carefully transferred to fresh 2 ml Eppendorf tubes and then solubilized in 6 M Guanidium chloride, dissolved in 0.1 M Tris–HClwith pH 8.5. The Guanidium chloride solution was boiled at 100 °C for 10 min later 200 µl solution was added to each 2 ml Eppendorf tube containing pellet and was vortexed for 2 min. The sample (extracted proteins) was boiled for 5 min and was again vortexed for complete solubilization and then centrifuged at 10,000 rpm for 10 min for pelleting out the debris. The supernatant (upper liquid phase) was transferred to fresh centrifuge tube and stored at − 80 °C.
Sample preparation for LC–MS
Sample was heated at 90 °C for 10 min for better lysis and then precipitated by trichloroacetic acid or TCA acetone. Precipitate was obtained, washed 3 times with cold acetone and was directly solubilized in 0.1% Rapigest (water) in 50 mM ammonium bicarbonate (ABC) (pH ~ 7). Reduction of samples was done using 5 mM Tris (2-carboxyethyl) phosphine [TCEP] (sigma) in 25 mM ABC (pH ~ 7) at 37 °C for 30 min and were alkylated using 55 mM iodoacetamide (IAA) (Sigma) in 25 mM ABC (pH ~ 7) at room temperature in dark for 30 min. Trypsin (Promega) was added in the ratio of 1:50 for overnight digestion at 37 °C. Rapigest was removed by spining at 13,400 rpm for 10 min (pellet was discarded); supernatant was collected in another tube for desalting column. Desalting of samples was done according to manufacturer’s protocol (https://assets.thermofisher.com/TFSAssets/LSG/manuals/MAN0011495_Pierce_C18_SpinCol_UG.pdf). The samples were vacuum dried by speedvac and finally reconstituted in 0.1% Formic Acid, FA. About ~ 3 µg amount of peptide was loaded on the column [PepMap RSLC C18 2 µm × 50 cm (Thermo scientific)]. The column temperature was set at 40 °C. The general reverse phase nano-Liquid Chromatography (Thermo Scientific Easy-nLC 1200) setting (Table 1) was used and the optimized LC program is given below:
Mobile phases for two solvent gradient elution were: A (water + 0.1% Formic acid) and B (acetonitrile + 0.1% formic acid). All solvents were of LC–MS grade (Fischer Scientific), Lock mass of 445.12003 Da was used for internal calibration. The Mass Spectroscopy, MS (Thermo Scientific Q Exactive Orbitrap), run time was 0–123 min, polarity was positive, default charge state was 2 with MS (MS1) setting as microscan 1, resolution was 70,000, automatic gain control (AGC) 3e6, maximum IT (ion transfer) time was 60 mS, number of scan range was 1, scan ranges were 350–2000 m/z and spectrum was obtained in profile mode. MS MS (MS2) setting like microscan was 1, resolution 17,500, AGC 1e5, maximum IT (ion transfer) 120 mS, loop count 10, maximum number of precursor to be plexed in single event was 1, isolation window 1.5 m/z, isolation offset was 0.0 m/z, fixed first mass 100 m/z, normalized collision energy 27, and the data dependent settings as minimum automatic gain control (AGC) target was 1.00 e2, charge exclusion was unassigned and dynamic exclusion time was 50.0 S.
About ~ 3 µg of sample was loaded on C18 reverse phase column, obtained raw data was finally analyzed in Proteome Discoverer 2.2 (Thermo Scientific).The databases were directly downloaded from http://www.uniprot.org in FASTA format. The maximum allowed missed cleavage was 3, minimum peptide length for search was 4, maximum peptide length was 144, precursor mass tolerance 10 ppm, fragment mass tolerance 0.05 Da, dynamic modification—oxidation of methionine and acetylation at N terminus of proteins, static modification was carbamido methylation, target FDR (false discovery rate) 0.01 (for decoy database search) and validation was based on q-value.
Results and discussion
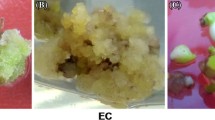
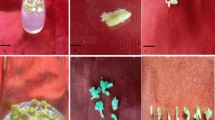
On inoculation in solid MS medium added with various 2,4-D concentrations (0.5, 1.0, 1.5, 2.0, 2.5, and 3.0 mg/l) more than 90% of the explants responded in culture and produced whitish callus within 4–5 weeks. Of all the tested concentrations, 2.0 mg/l was observed to be the most effective dose for callus induction. The induced callus from different explants was separately sub cultured and placed into test tubes containing 8–10 ml MS medium for callus proliferation. After 3–4 weeks, significant amount of new callus was observed with increased (more than 2.0 g of callus) biomass (Fig. 1a, c). Callus from each source was separately transferred into two solid MS media, one supplemented with same PGR (2.0 mg/l 2,4-D) for the maintenance of callus and the other was amended with 2.0 mg/l BAP and 1.5 mg/l NAA for induction of somatic embryos. Hypocotyl-induced callus turned into embryogenic calli while calli obtained from leaf and stem became non embryogenic in nature. Within 3–4 weeks, globular embryos were formed (Fig. 1b); later other developmental stages of embryos were induced from hypocotyl embryogenic calli. Some of the embryogenic calli with developing embryos were transferred to solid MS, amended with 2.0 mg/l GA3 for embryo maturation (Fig. 1d).
Total protein was extracted separately from embryogenic callus designated henceforth as F1 and non embryogenic callus designated as F2, were subject to LC–MS analysis. Proteins were identified and quantified using Protein Discoverer software. A total of 3573 proteins were identified among which 1511 proteins were present or common in both F1 and F2. LC-MS analysis revealed that 1079 proteins were exclusively found in embryogenic callus, F1 and 982 proteins were present only in F2 i.e. non embryogenic callus (Fig. 2). The overall abundance of proteins was found high in F1 (Fig. 3).
The embryogenic cells are metabolically highly active as the somatic cells while attaining embryogenic competence undergo thousands of physiological and biochemical alterations (Fehér et al. 2003; Jiménez 2005). In our study, we observed that among the core exclusive proteins in F1, 732 proteins are associated with metabolic processes while in F2 the number of proteins was only 685 indicating that the embryogenic cells are intensely active. This observation is very consistent with previous reports that suggest differentially expressed proteins are mostly related to metabolic processes in embryogenic cells (Heringer et al. 2015, 2018). The presence of dense cytoplasm, prominent nucleus, fragmented vacuole, highly active nucleus with large nucleolus, high nucleus to cytoplasm ratio and low level of hetrochromatin are some of those features confirming active embryogenic nature in other studies (Verdeil et al. 2007; Campos et al. 2017). Among the exclusive proteins found in F1, a large number was noted to regulate carbohydrate, protein and lipid metabolism. A significant number of proteins seem to be associated with transcription and translation, DNA and RNA binding, ribosome, transport and other related physiological processes. A classification of F1 exclusive protein list based on molecular function is given in Fig. 4, and also a classification on the basis of the biological processes is given in Fig. 5. The house keeping proteins of both F1 and F2 were almost the same and are not discussed here. The proteins exclusively present or expressed differentially were the main focus of this study. Out of core 1079 proteins found exclusively in the F1, 210 proteins are present in cytoplasm, 128 associated with cellular and organelle membranes and 55 proteins were found to be present in nucleus (Fig. 6). Five isoforms of ATP synthase subunit alpha (fragment), three ATP synthase subunit beta, one ATP synthase subunit d (mitochondrial), ATPase ASNA1 homolog, ATPase subunit 4, ATPase (AAA-type, CDC48 protein), and ATP/GTP-binding family proteins were identified exclusively in F1 suggesting the intense energy metabolism or very high requirement of ATP synthesis in order to fulfill the demand of energy in highly active cells. This observation was further supported by finding of translation related proteins in significant number in F1. In this study, seven 40S ribosomal proteins/isoforms, three 60S acidic ribosomal protein, two 60S ribosomal protein L6, one 60S ribosomal L6-like protein, one 60S ribosomal protein L26-1, and six other ribosomal proteins/isoforms were identified. Different translation factors like eukaryotic translation initiation factor 2c, six other proteins, indicating intense protein synthesis activity and high energy demand in F1. High protein synthesis may necessitate the need of higher number of different Heat shock Proteins (HSP) that assist in proper folding of the newly synthesized proteins. Nearly 12 isoforms of HSP70 and HSP90, six other chaparones and chaparonins were also found. Intense protein synthesis was accompanied with a large number of transport proteins suggesting high transportation of different micro and macro-molecules within and outside cells. One of these transporter proteins was a nuclear transport protein, importin, that has been earlier reported to be differentially expressed in embryogenic cells (Zhao et al. 2015). Here, in this study, two AP-2 complex subunit alpha were exclusively present in F1; AP-2 subunit alpha is a subunit of adaptor protein complex 2 (AP-2). Adaptor protein complexes are associated with vesicular transport of proteins in different membrane traffic pathways, play important role in cargo selection and vesicle formation in clathrin- dependent endocytosis (uniprot.org). Three clathrin heavy chains were also identified exclusively in embryogenic callus suggesting the cells are actively engaged in protein trafficking. As proteins are intensely synthesized in highly active cells, the older proteins need to be degraded and constituents recycled (Heringer et al. 2018), which is carried out by proteasome and ubiquitin (Guerra and Callis 2012). In this present investigation, 18 recycling and degradation proteins or isoforms of proteasome (16) and ubiquitin (2) were present exclusively in F1 (Table 2).
Molecular function wise classification of 1079 proteins exclusively found in embryogenic callus (F1). Y-axis shows the number of proteins and X-axis the molecular function. It may be noted that some proteins have more than one molecular function and calculated multiple times and hence the sum of all categories would not be equal to 1079
Biological process-wise classification of 1079 proteins found exclusively in embryogenic callus. Y-axis represents the number of proteins and X-axis represents biological processes. Some of the proteins are involved in two or more biological processes, calculated multiple times and hence the sum would not add up to total 1079
Stress related protein accumulation
As described earlier, stress has a very important role in the acquisition of embryogenic competence by non embryogenic or somatic cells in plants. Various stress responsive proteins have been reported to accumulate during SE. Glutathione S-transferase (GST) for example, is induced in embryogenic cells of several investigated plants like Cyclamen persicume (Hoenemann et al. 2012), Picea asperata (Jing et al. 2016), Zea mays (Ge et al. 2017) etc. GST plays significant role in various developmental processes including detoxification of xenobiotics, abiotic, disease and environmental stress tolerance. In C. roseus F1, more than 72 stress responsive proteins/isoforms were observed exclusively e.g.ascorbate peroxidase, catalase, glutathione S transferase, alkylhydro peroxidase, superoxide dismutase, SOD Fe N domain containing protein, pyridine nucleotide disulphideoxidoreductase, thioredoxinreductase (Table 2) etc. The presence of vast array of stress related proteins in F1 and fewer in F2 suggests that rigorous changes in molecular or biochemical environment may be the reason for the paradigm shift in morphogenetic route. This observation is getting support from previous reports, suggesting stress as a main cause to switch on/off of thousands of genes (Feher 2015; Horstman et al. 2017), cause a wide array of morphological, biochemical and molecular changes, resulting in acquisition of embryogenic competence of somatic cells (Pulianmackal et al. 2014).
In Catharanthus, we identified several isoforms of 14-3-3 protein in both embryogenic (F1) and non embryogenic (F2) tissues, many of them were exclusively found in embryogenic callus (F1) (Table 2) and only two isoforms in the non embryogenic callus. This shows its possible role in SE. Therefore, our results are consistent with the reports of the presence of 14-3-3 proteins in embryogenic cells in different plants like Saccharum (Heringer et al. 2015, 2017), Larix principisrupprechtii (Zhao et al. 2015), Zea mays (Ge et al. 2017) Araucaria angustifolia (Fraga et al. 2016). Earlier reports indicated 14-3-3 proteins’ role in plant immunity, stress responses, metabolism, transcription, signal transduction, cell cycle control, programmed cell death and protein trafficking, beside SE (Oh 2010). These are regulatory, acidic proteins, binding in phosphorylation dependent manner to target proteins like phosphothreonine and phosphoserine and thus have significant role in plant growth and development.
Proteomic analysis of Catharanthus revealed the presence of Heat Shock protein 70 (HSP70), Heat Shock protein 90 (HSP 90) and their several isoforms (more than 23) in both the types of calli i.e. F1 and F2. Fourteen HSPs were found exclusively in F1 (Table 2), and Supplementary Table 1 enlists accession numbers and other details of proteins found exclusively in the F1 callus. The significant number of HSPs in F1 suggests their possible role in induction of SE. Rest of the HSPs were expressed either in both or exclusively in F2 callus but the number was less than that of F1. A variety of heat shock proteins and their isoforms were previously reported to be abundantly accumulated in embryogenic cells of different plant genera e.g. Saccharum (Heringer et al. 2015), Larix principisrupprechtii (Zhao et al. 2015), Araucaria angustifolia (Fraga et al. 2016). These proteins accumulate abundantly under different stress conditions and their role is essentially in proper folding of other proteins (Park and Seo 2015).
In this Catharanthus study, two Late embryogenesis abundant (LEA) proteins, Late embryogenesis abundant protein accession number A0A072UGZ8, and Dehydrin accession number A0A1U9Y619; and other related proteins like Ca2+ binding, DNA binding and repair etc. were identified exclusively in F1; confirming that these proteins play a very important role in embryo induction and development during SE process. Interestingly, these proteins were observed to be accumulated early i.e. in globular stage of embryogenic callus. LEA, a large class of hydrophilic proteins however, was previously reported to abundantly accumulate at late stages of zygotic embryo development (Ikeda et al. 2006). Although the exact function of such a large class of proteins is still unknown, it is now known that these proteins protect the cellular damages during abiotic stress (Tolleter et al. 2010; Hincha and Thalhammer 2012), accumulate late during embryo development and are also induced during in vitro embryogenesis (Ikeda et al. 2006).
Proteomic study of Catharanthus tissue showed pathogenesis related proteins (PR),PR-4B accession number P29063, two chitinases, accession numbers G7IDV3 and Q9FS45, osmotin-like protein (accession number Q2MJK2) were present only in F1 (Table 2, Supplementary Table 1). These results are in agreements with previous report of the presence or accumulation of pathogen related proteins present in Cichorium embryogenic cells in high abundance as compared to non embryogenic callus (Helleboid et al. 2000). Presence of several PRs in embryogenic callus indicate that the cells are under stress and need various strategies for stress tolerance and protection from cellular damage, for which cells may have recruited the PR proteins as one way to get endurance under such conditions. PR accumulation occurs under various stresses like salicylic acid, UV light, heat, cold, wounding, heavy metal, hormone e.g. ethylene (Helleboid et al. 2000). Hence PR proteins appear to be involved in equipping the cells with endurance under stress condition.
Out of 1079 proteins exclusively found in F1 embryogenic callus, 336 proteins were uncharacterized and their function is unknown. Presence in high number of these proteins suggests their possible role in SE, but unfortunately, more information regarding these proteins is still to come. More than 334 proteins have abundance twice or more in F1 than in F2. A list of these protein accession numbers, abundance ratio of F1/F2 in decreasing order and other parameters is attached (Supplementary Table 2). Increased abundances of the 334 proteins in embryogenic callus twice or more as found in non embryogenic callus, shows their possible role the attainment of embryogenic competence. The maximum proteins among the 334 enlisted are related to energy metabolism, protein synthesis and stress related which further confirms the active nature of embryogenic cells, high ATP/energy demand and stress as the key player in attainment of embryogenic potential.
It is important to mention here that all culture conditions like nutrient concentration, light, temperature, humidity etc. were the same for both embryogenic and non embryogenic callus except the PGRs used. So we may infer from the study that the PGRs may induce stress, cause gross morphological, physical, molecular and biochemical changes, enrouting the cells to SE developmental pathway.
Conclusion and future prospects
Catharanthus roseus shows the potentiality of SE at high frequency in MS medium supplemented with BAP and NAA. SE was only induced in hypocotyl derived calli, calli from other explant sources were non embryogenic. A sizable 1079 proteins were identified in embryogenic callus exclusively having diverse functions. These proteins have functional homology with a number of proteins identified earlier from embryogenic cells in many other investigated plants. Some of the newly identified proteins were discussed with reference to previous others. The study suggests that a large number of stress related proteins accumulate in embryogenic cells of C. roseus, may activate an array of different signaling cascades in acquiring embryogenic competence. The abundance of different proteins, regulating metabolic processes indicates that the embryogenic cells are highly active compared to non embryogenic cells. The identification of SE related proteins in C. roseus is a step to understand the process of SE and further need to be consolidated at genomic and transcriptomic level. The SE related proteins isolated from diverse plant groups, could be used as important markers for the identification of embryogenic cells. Further research is however; necessary to unveil the mechanism underlying the process of SE as the process of SE has often been considered as a model to understand plant developmental pathways. The elucidation of SE proteomic maps may broaden its scope and applicability to other economically important and endangered plants that need rapid propagation and conservation.
References
Campos NA, Panis B, Carpentier SC (2017) Somatic embryogenesis in coffee: the evolution of biotechnology and the integration of omics technologies offer great opportunities. Front Plant Sci 8:1460. https://doi.org/10.3389/fpls.2017.01460
Chen SL, Yu H, Luo HM, Wu Q, Li CF, Steinmetz A (2016) Conservation and sustainable use of medicinal plants: problems, progress, and prospects. Chin Med 11:37. https://doi.org/10.1186/s13020-016-0108-7
Eng JK, McCormack AL, Yates JR (1994) An approach to correlate MS/MS data to amino acid sequences in a protein database. J Am Soc Mass Spectrum 5:976–989
Feher A (2015) Somatic embryogenesis—stress-induced remodeling of plant cell fate. Biochem Biophys Acta 1849:385–402
Fehér A, Pasternak TP, Dudits D (2003) Transition of somatic plant cells to an embryogenic state. Plant Cell Tissue Org Cult 74:201–228
Fraga HP, Vieria LN, Heringer AS, Puttkammer CC, Silveira V, Guerra MP (2016) DNA methylation and proteome profiles of Araucaria angustfolia (Bertol) Kuntzeembryogenic cultures as affected by plant growth regulators supplementation. Plant Cell Tissue Org Cult 125(2):353–374
Ge F, Hu H, Huang X, Zhang Y, Wang Y, Li Z, Zou C, Peng H, Li L, Gao S, Pan G, Shen Y (2017) Metabolomic and proteomic analysis of maize embryonic callus induced from immature embryo. Sci Rep 7(1):1004. https://doi.org/10.1038/s41598-017-01280-8
Guerra DD, Callis J (2012) Ubiquitin on the move: the ubiquitin modification system plays diverse roles in the regulation of endoplasmic reticulum- and plasma membrane-localized proteins. Plant Physiol 160(1):56–64. https://doi.org/10.1104/pp.112.199869
Helleboid S, Hendriks T, Bauw G, Inze D, Vasseur J, Hilbert JL (2000) Three major somatic embryogenesis related proteins in Cichorium identified as PR proteins. J Exp Bot 51:1189–1200
Heringer AS, Barroso T, Macedo AF, Santa-Catarina C, Souza GHMF, Floh EIS, Souza-Filho GA, Silveira V (2015) Label-free quantitative proteomics of embryogenic and non-embryogenic callus during sugarcane somatic embryogenesis. PLoS ONE. https://doi.org/10.1371/journal.pone.0127803
Heringer AS, Reis RS, Passaman LZ, de Souza-Filho GA, Santa-Catarina C, Silveira V (2017) Comparative proteomics analysis of the effect of combined red and blue lights on sugarcane somatic embryogenesis. Acta Physiol Plant 39:52. https://doi.org/10.1007/s11738-017-2349-1
Heringer AS, Santa-Catarina C, Silveira V (2018) Insights from proteomic studies into plant somatic embryogenesis. Proteomics. https://doi.org/10.1002/pmic.201700265
Hincha DK, Thalhammer A (2012) LEA proteins: IDPs with versatile functions in cellular dehydration tolerance. Biochem Soc T 40:1000–1003
Hoenemann C, Ambold J, Hohe A (2012) Gene expression of a putative glutathione S-transferase is responsive to abiotic stress in embryogenic cell cultures of Cyclamen persicum.. Electronic J Biotechnol 15:1–6
Horstman A, Li M, Heidmann I, Weemen M, Chen B, Muino JM, Angenent GC, Boutilier K (2017) The Baby Boom transcription factor activates the LEC1-ABI3-FUS3-LEC2 network to induce somatic embryogenesis. Plant Physiol 175:848–857
https://assets.thermofisher.com/TFSAssets/LSG/manuals/MAN0011495_Pierce_C18_SpinCol_UG.pdf
Ikeda M, Umehara M, Kamada H (2006) Embryogenesis-related genes; its expression and roles during somatic and zygotic embryogenesis in carrot and Arabidopsis. Plant Biotechnol 23:153–161
Imin N, Nizamidin M, Daniher D, Nolan KE, Rose RJ, Rolfe BG (2005) Proteomic analysis of somatic embryogenesis in Medicago truncatula. Explant cultures grown under 6-benzylaminopurine and 1-naphthaleneacetic acid treatments. Plant Physiol 137:1250–1260. https://doi.org/10.1104/pp.104.055277
Isaacson T, Damasceno CM, Saravanan RS, He Y, Catala C, Saladie M, Rose JK (2006) Sample extraction techniques for enhanced proteomic analysis of plant tissues. Nat Protoc 1(2):769–774
Jiménez VM (2005) Involvement of plant hormones and plant growth regulators on in vitro somatic embryogenesis. Plant Growth Regul 47:91–110
Jing D, Zhang J, Xia Y, Kong L, OuYang F, Zhang S, Zhang H, Wang J (2016) Proteomic analysis of stress-related proteins and metabolic pathways in Picea asperata somatic embryos during partial desiccation. Plant Biotechnol J. https://doi.org/10.1111/pbi.12588
Junaid A, Mujib A, Fatima S, Sharma MP (2008) Cultural conditions affect somatic embryogenesis in Catharanthus roseus L. (G.) Don. Plant Biotechnol Rep 2:179–189
Karpievitch YV, Ashoka DP, Gordon AA, Richard DS, Alan RD (2010) Liquid chromatography mass spectrometry-based proteomics: biological and technological aspects. Ann Appl Stat 4(4):1797–1823. https://doi.org/10.1214/10-AOAS341
Lu D, Wei W, Zhou W, Linda D, Xiao J, Yu L (2017) Establishment of a somatic embryo regeneration system and expression analysis of somatic embryogenesis-related genes in Chinese chestnut (Castanea mollissima Blume). Plant Cell Tissue Org Cult 130(3):601–616
Mahdavi-Darvari F, Noor NM, Ismanizan I (2014) Epigenetic regulation and gene markers as signals of early somatic embryogenesis. Plant Cell Tiss Org Cult 120(2):407–422
Mordhorst AP, Toonen MAJ, deVries SC (1997) Plant embryogenesis. Crit Rev Plant Sci 16(6):535–576
Mujib A, Ali M, Isah T, Dipti T (2014) Somatic embryo mediated mass production of Catharanthus roseus in culture vessel (bioreactor)—a comparative study. Saudi J Biol Sci 21(5):442–449
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15(3):473–497
Nolan KE, Irwanto RR, Rose RJ (2003) Auxin up-regulates MtSERK1 expression in both Medicago truncatula root-forming and embryogenic cultures. Plant Physiol 133:218–230. https://doi.org/10.1104/pp.103.020917
Oh CS (2010) Characteristics of 14-3-3 proteins and their role in plant immunity. Plant Pathol J 26(1):1–7. https://doi.org/10.1186/s13020-016-0108-7
Orłowska A, Igielski R, Łagowska K, Pczyska EK (2017) Identification of LEC1, L1L and polycomb repressive Complex2 genes and their expression during the induction phase of Medicago truncatula Gaertn. somatic embryogenesis. Plant Cell Tissue Organ Cult 129:119–132. https://doi.org/10.1007/s11240-016-1161-8
Park CJ, Seo YS (2015) A review of the molecular chaperones for plant immunity. Plant Pathol J 31(4):323–333. https://doi.org/10.5423/PPJ.RW.08.2015.0150
Pulianmackal AJ, Kareem AV, Durgaprasad K, Trivedi ZB, Prasad K (2014) Competence and regulatory interactions during regeneration in plants. Front Plant Sci 5:142
Rupps A, Raschke J, Rümmler M, Linke B, Zoglauer K (2016) Identification of putative homologs of Larix decidua to Babyboom (BBM), leafy cotyledon1 (LEC1), Wuschel-related Homeobox2 (WOX2) and somatic embryogenesis receptor-like kinase (SERK) during somatic embryogenesis. Planta 243:473–488
Schmidt ED, Guzzo F, Toonen MA, de Vries SC (1997) A leucine-rich repeat containing receptor-like kinase marks somatic plant cells competent to form embryos. Development 124:2049–2062
Tolleter D, Hincha DK, Macherel D (2010) A mitochondrial late embryogenesis abundant protein stabilizes model membranes in the dry state. BBA Gen Sub 1798:1926–1933
Verdeil JL, Alemanno L, Niemenak N, Tranbarger TJ (2007) Pluripotent versus totipotent plant stem cells: dependence versus autonomy? Trends Plant Sci 12:245–252
Washburn MP, Wolters D, Yates JR (2001) Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat Biotechnol 19(3):242–247. https://doi.org/10.1038/85686
Zhao J, Li H, Fu S, Chen B, Sun W, Zhang J (2015) An iTRAQ-based proteomics approach to clarify the molecular physiology of somatic embryo development in Prince Rupprecht’s larch (Larix principis-rupprechtii Mayr). PLoS ONE. https://doi.org/10.1371/journal.pone.0119987
Acknowledgements
The first author is thankful to University Grant Commission (UGC) for providing financial assistance with (Grant No. 2061530497). The authors are also thankful to the Department of Botany, Jamia Hamdard, and University of Delhi for providing laboratory and other research facilities for this research.
Author information
Authors and Affiliations
Contributions
BG, AF, NZ conducted most of the experiments. BG also made the draft of article. AM and MVR edited the article.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest in this research.
Additional information
Communicated by Francisco de Assis Alves Mourão Filho.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11240_2019_1563_MOESM1_ESM.xlsx
Supplementary Table 1. Proteins/isoforms (accession numbers) exclusively found in F1 i.e. embryogenic tissue of Catharanthus roseus—Supplementary material 1 (XLSX 102 KB)
11240_2019_1563_MOESM2_ESM.xlsx
Supplementary Table 2. Accession numbers of the proteins found in embrogenic callus (F1) having high abundance (twice or more) than that of non embryogenic callus (F2). The ratio (F1/F2) protein abundances are listed in decreasing order—Supplementary material 2 (XLSX 45 KB)
Rights and permissions
About this article
Cite this article
Gulzar, B., Mujib, A., Rajam, M.V. et al. Identification of somatic embryogenesis (SE) related proteins through label-free shotgun proteomic method and cellular role in Catharanthus roseus (L.) G. Don. Plant Cell Tiss Organ Cult 137, 225–237 (2019). https://doi.org/10.1007/s11240-019-01563-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-019-01563-0