Abstract
Hibiscus rosa-sinensis L. is a popular ornamental species valued for its large brightly coloured ephemeral flowers and has a range of health-promoting properties. The value of H. rosa-sinensis could be improved even further if there were ways to prolong the display life of its short-lived flowers, and to improve its frost tolerance. Development of an efficient plant transformation and regeneration procedure that allows introduction of genes into the plant will greatly facilitate this. Here we outline a transformation and regeneration procedure that is the first to produce transformed H. rosa-sinensis plants successfully. We first optimised callus induction and shoot regeneration efficiency. The highest shoot regeneration frequency of 66.7 % was achieved in the cultivar ‘Ruby’ when callus induced from axillary buds using a basal medium supplemented with 2.22 µM benzylaminopurine and 2.47 µM β-naphthoxyacetic acid was cultured on shoot regeneration medium. The frequency of shoot regeneration from callus was lower in ‘Ben James’ and absent in ‘Bright Light’, indicating genotypic differences. When axillary bud-derived callus of ‘Ruby’ was co-cultured with Agrobacterium tumefaciens harbouring a β-glucuronidase (GUS) reporter plasmid, 49 % of calli produced shoots on selection media. All tested plantlets were confirmed as transformed based on the presence of the GUS transgene in the genomic DNA and GUS activity measurements. Roots were induced on transgenic plantlets using half-strength basal medium supplemented with 2.85 µM indole-3-acetic acid. This simple protocol can be used to improve the ornamental, agronomic and health-promoting traits of H. rosa-sinensis hitherto recalcitrant to A. tumefaciens-mediated transformation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hibiscus rosa-sinensis L. (Malvaceae) is a popular glabrous shrub widely cultivated in the tropics as an ornamental plant. The plants produce prolific numbers of large attractive flowers of varied colours that survive for only a single day. In addition to being prized as an ornamental, the plant is also recognised as a rich source of health-promoting secondary metabolites. For example, the petal extracts of H. rosa-sinensis have been used as food colorants and for their antioxidant properties (Vankar and Srivastava 2008), and have been shown in a number of rat- and mouse-based studies to protect against the cancer-promoting effects of benzoyl peroxide and ultraviolet radiation (Sharma and Sultana 2004); to inhibit the diabetes promoting effects of streptozotocin and alloxan (Sachdewa and Khemani 2003; Venkatesh et al. 2008); and to confer cardioprotective effects (Gauthaman et al. 2006).
The development of an efficient transformation and regeneration system for H. rosa-sinensis will enable researchers to improve its quality in a number of ways. For example, by appropriate gene manipulation plants could be produced that have greater tolerance to frost or whose flowers have a much extended display life. The technology will also enable researchers to examine in detail the metabolic processes in H. rosa-sinensis that give rise to its health-promoting effects.
Research previously conducted on Hibiscus spp. has indicated that certain species are amenable to transformation and regeneration. Calli and other explants of some species of Hibiscus such as H. syriacus, H. sabdariffa, and H. cannabinus have been successfully transformed using Agrobacterium, and transgenic plants were regenerated (Banks et al. 1993; Yang et al. 1996; Gassama-Dia et al. 2004). On the other hand, for H. acetosella Welw. ex. Hiern, in vitro methods have been described only for micropropagation purposes (Sakhanokho 2008).
In vitro studies on H. rosa-sinensis have led to the development of protocols for micropropagation (Airò et al. 2009; Christensen et al. 2008) and to the production of virus-free plants, which is important because of the species’ widely reported high susceptibility to viruses (Mao et al. 2008; De Stradis et al. 2008; Huang et al. 2004). However, regeneration of whole plants from transformed tissue of H. rosa-sinensis has up until now been unsuccessful. Christensen et al. (2009) have more recently transformed H. rosa-sinensis calli using Agrobacterium rhizogenes but were unable to regenerate plants. Similarly, Mercuri et al. (2010) reported production of hairy roots of H. rosa-sinensis following transformation with A. rhizogenes strain ATCC 15384, and verified the transformation (T-DNA insertion) using polymerase chain reaction (PCR). They reported formation of adventitious bud-like structures although no plant regeneration was observed in culture within 1 year. Vazquez-Thello et al. (1996), in an attempt to transfer chilling and freezing tolerance from Lavatera thuringiaca to H. rosa-sinensis, successfully transformed cell suspensions of both species but were only able to regenerate plants of L. thuringiaca. Subsequently they fused the transgenic protoplasts from the two species and were able to select for hybrid calli successfully but the attempts to regenerate plants from these calli were unsuccessful.
The aim of this study was to develop an efficient reproducible protocol for the rapid transformation and regeneration of transgenic plants of H. rosa-sinensis. Here we report on the first successful regeneration of H. rosa-sinensis plants from axillary bud-derived calli transformed with Agrobacterium tumefaciens.
Materials and methods
Plant materials and establishment of cultures
The three cultivars of H. rosa-sinensis used in this study, ‘Ruby’, ‘Bright Lights’ and ‘Ben James’, were supplied by Lyndale Nurseries Auckland Ltd, Whenuapai, Auckland, New Zealand. Plants were grown under natural light in a temperature-controlled greenhouse at Plant & Food Research, Palmerston North, New Zealand. The glasshouse was heated at 15 °C and vented at 26 °C. Freshly harvested green shoots ca. 2–3 cm long were surface sterilised by immersion in 75 % (v/v) ethanol (Merck, Darmstadt, Germany) for 30 s, followed by washing in 3 % (v/v) sodium hypochlorite (Janola®, Auckland, New Zealand) containing 0.03 % (v/v) Tween® 20 (BDH Laboratory Supplies, Poole, England) for 20 min on a rotary shaker (50 rpm). The shoots were then washed three times with sterile water. Axillary bud, petiole and young leaf explants ranging in size from 4 mm to 6 mm were then excised for regeneration and transformation experiments. Unless otherwise stated, all chemicals used in this study were purchased from Sigma Chemical Company (St Louis, MO, USA).
Callus initiation, shoot regeneration, root induction and greenhouse acclimation
To determine the influence of genotype, explant, and plant growth regulators (PGR) on callus induction efficiency, three cultivars (‘Ruby’, ‘Bright Lights’ and ‘Ben James’), three different explants (leaf, petiole and axillary bud), and eight different combinations of auxins and 6-benzylaminopurine (BAP) were tested. To induce callus formation, leaf, axillary bud, and petiole explants were excised and placed on a basal H. rosa-sinensis medium (Christensen et al. 2008) supplemented with factorial combinations of BAP (2.22, 4.44 µM), β-naphthoxyacetic acid (NOA) (0, 2.47, 4.95 µM) and 2,4-dichlorophenoxyacetic acid (2,4-D) (0, 2.26, 4.52 µM) in 9 cm Petri plates (Table 1). Each Petri plate contained 25 mL of media and 10 explants. The plates were held in the dark at 24 ± 2 °C and callus induction frequency was estimated after 3 weeks of culture as the percentage of explants producing callus. The experiment was conducted in three replicates, each consisting of 10 explants.
To determine the ability of the initiated callus to induce shoots, calli derived from all three explant types in all three cultivars were transferred to a shoot regeneration medium (RE: basal media supplemented with 2.22 µM BAP) in 290 mL clear, wide-mouth tubs, with snap on lids containing 50 mL of media per tub. The cultures on RE medium were maintained at 24 ± 2 °C under light (30–40 µmol m−2 s−1, at shelf level provided by ‘‘Cool White’’ fluorescent tubes) with a 16 h photoperiod. The percentage of calli producing shoots was determined after 15 days. To induce roots, regenerated shoots (5–6 cm length) were separated from the callus and transferred individually to a Rooting Medium (RO) consisting of half-strength basal medium supplemented with 2.85 µM indole-3-acetic acid (IAA) in tubs as previously described by Airò et al. (2009). All the culture media used in this work were adjusted to pH 5.7 prior to autoclaving at 121 °C for 15 min and are listed in Table 1.
Plants with roots were deflasked to trays of a soil-free medium (bark:pumice 50:50) and placed in a fog tent within a containment greenhouse without supplementary lighting for 1 week before transfer to an open greenhouse bench, where they were misted every 10 min for 1 week. The bases of both the fog tent and the greenhouse bench were heated to 28 °C to facilitate root growth and plant establishment. After the first week, the plants were transferred to a greenhouse bench with capillary watering for another week before transfer to a greenhouse bench with overhead watering. The glasshouse was heated at 15 °C and vented at 26 °C during the period of acclimation.
Establishment of sensitivity of H. rosa-sinensis to kanamycin
To determine the concentration of kanamycin that inhibits shoot regeneration from non-transformed callus, a separate experiment was conducted using the best treatment combination for callus induction and plant regeneration (i.e. axillary bud-derived callus of ‘Ruby’ in CI1 media). Three-week-old axillary bud-derived callus of ‘Ruby’ induced on CI1 media were cultured on RE media supplemented with five concentrations of kanamycin from 0 to 165.12 µM at 41.28 µM intervals under the conditions described for plant regeneration above. The experiment was conducted in two replicates with 10 explants per replicate. Shoot regeneration was assessed after 15 days.
Plasmids and Agrobacterium strains
Transformation was carried out using A. tumefaciens strain GV3101. The strain harboured the binary vector pNWA37, which has an intron-containing bacterial uidA gene encoding β-glucuronidase (GUS) from pMOG410 (Hood et al. 1993) under the control of a double CaMV35S promoter in the pART27 binary vector (Gleave 1992). The binary vector contains the neomycin phosphotransferase (NPT II) gene, which confers resistance to kanamycin, driven by the nopaline synthase (NOS) promoter. The intron within the GUS coding sequence prevents Agrobacterium-derived GUS expression (Vancanneyt et al. 1990; Fig. 1).
A partial map of the T-DNA region of the binary vector used for H. rosa-sinensis plant transformation (not drawn to scale). The 1200 and 139 bp fragments that were amplified from genomic DNA for agarose gel PCR analysis and qPCR analysis respectively are indicated. LB left border, RB right border, GUS β-glucuronidase, NOS nopaline synthase terminator. For clarity, the NPTII (kanamycin resistance gene) driven by the nopaline synthase promoter present between the LB and double 35S cauliflower mosaic virus promoter (35S) is not indicated
Plant transformation
Single colonies of A. tumefaciens strain GV3101 harbouring pNWA37 were grown overnight at 28 °C on a shaker at 250 rpm in 3 mL of Luria Broth containing spectinomycin (301 µM), gentamycin (42 µM) and rifampicin (61 µM) as selective agents. Following overnight incubation the cells were harvested by centrifugation (6000×g, 5 min) and resuspended at a concentration of 104–105 cells mL−1 in Inoculation Medium [IM: basal liquid media supplemented with 100 µM acetosyringone in which sucrose was replaced with 2 % (w/v) glucose]. Three-week-old axillary bud-derived callus induced on CI1 media of H. rosa-sinensis ‘Ruby’ (best treatment from callus induction and plant regeneration step above) was immersed in IM for 10 min, blotted on sterile filter paper to remove excess bacteria and transferred onto new sterilised filter paper placed on Co-Cultivation Medium [CC: RE media in which sucrose was replaced with 2 % (w/v) glucose and supplemented with 100 µM acetosyringone]. After 3 days co-cultivation in the dark, the infected callus was transferred onto Selection Medium [SE: RE media supplemented with 82.56 µM kanamycin and 0.66 mM cefotaxime (AFT Pharmaceuticals, Auckland, NZ)]. The calli were transferred to fresh SE every 2 weeks. To induce roots, the putative transgenic shoots regenerated from inoculated callus were excised and transferred onto RO supplemented with 82.56 µM kanamycin. All steps involved in regeneration of transgenic shoots and their rooting, after the co-cultivation step, were carried out under the conditions described in media optimisation experiments above.
Molecular confirmation of transformation
PCR was used to confirm presence of the transgene in ten randomly selected independent shoots from separate calli. Leaves of shoots regenerated from un-inoculated calli served as the non-transformed control. Total genomic DNA (gDNA) was isolated by a standard phenol-based method from young leaves excised from kanamycin-resistant shoots and analysed for the presence of a 1200 bp 35S-GUS genomic fragment by performing PCR with forward primer 5′-ATGTGATATCTCCACTGACG-3′ and reverse primer 5′-AGTCCCGCTAGTGCCTTGTCCAG-3′. The PCR conditions were an initial denaturation at 94 °C for 3 min, followed by 35 cycles (at 94 °C for 1 min, 60 °C for 1 min, 72 °C for 1 min) and a final extension at 72 °C for 7 min. PCR products were separated on a 1 % w/v agarose gel stained with ethidium bromide and visualised under UV light.
To estimate transgene copy number equivalence and to confirm the transgenic nature of the regenerated plantlets, gDNA from the selected clones was further analysed by quantitative real time PCR (qPCR). Genomic DNA was quantified with a spectrophotometer (NanoDrop Technologies, DE, USA), diluted to 5 ng µL−1 in sterile distilled water, and used as a template in qPCR with GUS and 18S primer sets designed with the PrimerQuestSM software (Integrated DNA Technologies, Coralville, IA, USA). The primers for the GUS gene (forward, 5′-GATCACCTGCGTCAATGT-3′, and reverse, 5′-TTCCAGTACCTTCTCTGCC-3′) were selected to amplify a 139 bp fragment. For the reference gene, 18S, the primers (forward, 5′-CGCGCAAATTACCCAATCCTGACA-3′, and reverse, 5′-TTGCCCTCCAATGGATCCTCGTTA-3′) were selected to amplify a 122-bp fragment. Triplicate qPCR reactions were performed for each selected clonal line in a fluorimetric thermal cycler (Rotor Gene 6000; Corbett Research, Australia) using 25 μL PCR reaction volumes containing 10 μL of master mix (LightCycler® 480 SYBR Green I Master; Roche Diagnostics, Indianapolis, IN, USA), 1 μM of each primer and 20 ng of gDNA. qPCR conditions were an initial denaturation at 94 °C for 5 min, followed by 50 cycles of 94 °C for 30 s, 63 °C for 30 s, and a final extension at 72 °C for 45 s. Data acquisition and analysis were performed with Rotor Gene 6000 software ver. 1.7. A simple estimation of transgene copy number equivalence for H. rosa-sinensis was obtained using the comparative quantitation analysis software supplied with the Rotor Gene 6000 using the endogenous 18S gene as the reference gene.
Fluorimetric and histochemical assay of GUS activity
GUS protein from leaf tissue of the regenerated plants was extracted as described by Winichayakul et al. (2004). The GUS activity of the leaf tissue was quantified and localised histochemically as described by Hunter and Watson (2008). Total protein concentration in the extracts was measured using a Bio-Rad Protein Assay Kit (Bio-Rad Laboratories, Hercules, CA, USA).
Determination of total phenolic content
Phenolics were extracted from leaf tissues (30 mg) of H. Rosa-sinensis ‘Ruby’, Petunia hybrida and Solanum lycopersicum by adding 3 mL of methanol to the tissues after they had been ground in liquid nitrogen. Following the addition of methanol, a 2-mL aliquot was centrifuged at 10,000×g to produce a clarified solution (Ke and Saltveit 1989; Campos-Vargas and Saltveit 2002). The absorbance of the clarified methanol extract was read at 320 nm (Loaiza-Velarde et al. 1997) and expressed as absorbance per gram fresh weight. Data were the mean ± SD of 10 biological replicates.
Experimental design and statistical analysis
Each experiment had three replicates. A replicate was made up of three Petri plates with a minimum of ten explants. Data are presented as means and analysed by one-way ANOVA. For the callus induction experiment, a multifactor ANOVA was performed to assess the effects of cultivar, media and explant on callus induction frequency using StatGraph v2.0 (Statgraphics Centurion XV, StatPoint Technologies, Inc., Herndon, VA, USA). The significance of the results obtained for experiments on regeneration rate and molecular analysis were verified post hoc by Tukey’s Least Significant Difference test (P < 0.05).
Results
Optimisation of conditions for callus induction in H. rosa-sinensis
Callus was produced from all explant types of all cultivars and in all CI media described in Table 1. However, there were considerable differences in the frequency at which it occurred, ranging from 6.7 to 100 % (Table 2). The frequency of callus induction was highest for axillary bud explants and in these explants the frequency was only slightly affected by the type of CI media. In CI2 media, callus was produced from 100 % of the axillary bud explants from all three cultivars, whereas in the other CI media callus induction frequency from axillary buds ranged from 95.8 to 100 % depending on the cultivar. The frequency of callus induction was lowest for the leaf explants and was greatly affected by the type of cultivar and CI media used, ranging from 6.7 to 70 % (Table 2). The callus induction frequency from the leaf explants of ‘Bright Lights’ was particularly poor, with none of the CI media giving frequencies higher than 16.7 %. The type of CI media significantly affected callusing frequency for leaf explants of ‘Ruby’ and ‘Ben James’, with NOA giving two to threefold higher callusing frequencies than 2,4-D (Table 2). A similar enhancement in callusing frequency with NOA was observed for petiole explants of ‘Ben James’, but for petiole explants of ‘Ruby’ callusing frequency was high and independent of the type of auxin used (Table 2).
Efficiency of shoot regeneration from callus in H. rosa-sinensis
No shoots regenerated from calli derived from the leaf and petiole explants for any of the three cultivars (data not shown). Shoots were regenerated from calli derived from axillary buds of ‘Ruby’ and ‘Ben James’ (Fig. 2), but not from ‘Bright Lights’ (data not shown). Axillary bud-derived calli from ‘Ruby’ was the best tissue for shoot induction; the highest frequency of regeneration (66.7 %) being obtained from calli that had been induced in CI1 media (2.47 µM NOA and 2.22 µM BAP, Fig. 2). This combination of auxin and cytokinin was also the only combination that enabled shoot induction (15 %) from axillary bud-derived callus of ‘Ben James’ (Fig. 2).
Effect of plant growth regulator concentrations in callus induction media on shoot regeneration from axillary bud-derived callus transferred to regeneration media (basal + 2.22 μM 6-benzylaminopurine). Data represent mean regeneration percentage (n = 3) ± SD. Each replicate consisted of 10 calli and the number of shoots was counted after 15 days on regeneration media. NOA naphthoxyacetic acid, BAP 6-benzylaminopurine
Agrobacterium-mediated transformation and identification of the transgenic plantlets
To maximise the likelihood that the plants regenerated after transformation would be true transgenics, we determined the minimum concentration of kanamycin that would successfully kill 100 % of the non-transformed plants at the plant regeneration stage. The minimum concentration of kanamycin that completely inhibited regeneration of shoots from wild-type axillary bud-derived callus of ‘Ruby’ was 82.56 µM (Fig. 3) and this concentration was subsequently used in RE for selecting the transformed shoots after the co-cultivation step. Agrobacterium over-growth was successfully controlled by including cefotaxime at 0.66 mM in the kanamycin-supplemented RE (SE).
Effect of kanamycin concentration in media on shoot regeneration percentage from calli induced from axillary buds of H. rosa-sinensis ‘Ruby’. Data represent means (n = 10) ± SD. Regeneration %: (number of explants that produce adventitious shoots after 15 days on selection medium/total number of explants placed on the medium) × 100. The experiment was repeated twice
Putatively transformed adventitious shoots began to appear approximately 3 weeks after the calli had been transformed and placed on SE media. Elongated shoots from independent calli were aseptically separated and transferred to a modified RO Medium (half-strength basal medium supplemented with 82.56 µM kanamycin and 2.85 µM IAA—Table 1). Of the 70 axillary bud explants of ‘Ruby’ that were used, 67 produced callus and from these 33 (49.2 %) produced antibiotic-resistant shoots (Table 3).
Analysis of transformed plantlets
PCR performed on leaf gDNA with primers designed to amplify a 1200 bp region spanning regions of the CaMV35S promoter and GUS gene (Fig. 1) confirmed the presence of the transgene in leaves from all 10 putatively transformed shoots tested (Fig. 4). Melting curve analysis of the resultant products amplified by qPCR using primers designed to the GUS gene revealed that they all produced a single-peak profile, indicating that the fluorescence signals from all samples were generated from specific amplification of the GUS gene and not through spurious non-specific amplification. When the results from the qPCR were analysed using the comparative quantitation method, which compared GUS gDNA abundance in the lines with that of an endogenous reference gene (18 s rRNA), we were able to achieve an approximate estimation of whether the transformation procedure integrated high or low copy numbers of the transgene into the shoots. As shown in Fig. 5, the relative abundance of the GUS gDNA differed significantly in the 10 lines analysed when normalised to the abundance of the endogenous 18 S rRNA gene. Thus the lines T-9, T-12 and T-17 were significantly higher for GUS gDNA abundance than the lines T-1, T-3, T-6, T-8, T-15, T-16 and T-18 (Fig. 5).
Quantitative analysis of GUS genomic DNA isolated from leaves of 10 randomly selected H. rosa-sinensis ‘Ruby’ transgenic plantlets. The data shown are means from three independent assays. Means with different letters differ significantly (P < 0.05), Tukey’s least significant difference test. Error bars indicate SD. T- transgenic line number, C untransformed control plant
GUS activity in five of these lines was also quantified to further confirm the differences in copy number in the transformed shoots, using the assumption that higher copy number would result in higher activity of the GUS gene. The GUS activity of the leaf extract from the untransformed control was <2 pmol 4-MU min−1 mg protein−1, while the leaf extracts from kanamycin resistant shoots displayed a wide range of GUS activities (Fig. 6). The highest amounts of GUS activity, 1546.3 and 1644 pmol 4-MU min−1 mg protein−1, were found in leaf extracts from lines T9 and T17 respectively, which were also the ones that qPCR had shown to have the highest transgene copy numbers. Similarly, the lowest GUS activities of 66.1 and 83.3 pmol 4-MU min−1 mg protein−1 were found in leaf extracts from lines T3 and T15 (Fig. 6), which had the lowest transgene copy numbers according to GUS gDNA abundance (Fig. 5).
GUS specific activity in leaf extracts of H. rosa-sinensis ‘Ruby’ transgenic plants. The data shown are averages from three independent assays. Means with different letters differ significantly (P < 0.05), Tukey’s Least Significant Difference test. Error bars indicate SD. T- transgenic line number, C untransformed control plant
When histochemical analysis was performed, GUS activity could not be visualised in any of the transgenic leaves tested, with the exception of a microscopic blue spot on one leaf (data not shown). Interference in the visualisation of GUS staining in certain plants has been attributed to their high phenolic content (Serres et al. 1997). To determine whether this may be the cause of the lack of staining in H. rosa-sinensis, we examined the phenolic content of its tissue. For comparison, we also determined the total phenolic content in leaf extracts of petunia and tomato, two species that present difficulties in visualising GUS. The total phenolic content in H. rosa-sinensis leaf extracts was 0.73 OD320 g−1 FW which was 4.6- and 2.3-fold higher than those in petunia (0.16 OD320 g−1 FW) and tomato (0.32 OD320 g−1 FW) respectively (Fig. 7).
A scheme summarising the overall procedure and time needed for each step in the transformation and regeneration of H. rosa-sinensis is presented in Fig. 8. The stages from callus induction, shoot regeneration and rooting through to greenhouse-acclimated whole plants of ‘Ruby’ are illustrated in Fig. 9. The complete procedure from inducing callus to acclimating plants in the greenhouse took approximately 17 weeks.
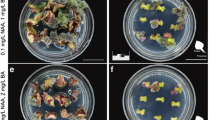
Plant regeneration from callus induced from axillary buds and Agrobacterium-mediated transformation of H. rosa-sinensis ‘Ruby’. a Callus induced from axillary bud explants on basal medium supplemented with 2.22 µM 6-benzylaminopurine and 2.47 µM β-naphthoxyacetic acid after 3 weeks. b Callus induced from an axillary bud on selective regeneration medium after 10 days of culture. c A plantlet regenerated on selective regeneration medium after 4 weeks. d A transgenic plantlet after 6 weeks on rooting medium. e Transgenic plants transferred to soil
Discussion
The genetic transformation of ornamental plants can considerably enhance the existing efforts of traditional and marker assisted breeding in generating new improved cultivars and allows testing of plant gene function in the context of whole plant development. Such biotechnological strategies until now have not been possible in H. rosa-sinensis because of the lack of an efficient transformation and regeneration procedure. Procedures to produce transformed plants are usually empirically determined as the type of explant, and the ratio and type of PGR needed for successful transformation and regeneration, can vary considerably depending upon plant genotype. This has clearly been illustrated for kenaf (Herath et al. 2005), Prunus avium (Feeney et al. 2007), chickpea (Tripathi et al. 2013) and Miscanthus (Hwang et al. 2014). In Actinidia eriantha for example, BAP in combination with zeatin was critical for successful plant regeneration, including those of transformed plants (Wang et al. 2006). In our study, ‘Ruby’ was more amenable to plant regeneration from callus induced in media supplemented with BAP and NOA. All the calli produced using 2,4-D as the auxin failed to regenerate plants.
Previous callus-based plant regeneration attempts on H. rosa-sinensis showed that callus could be induced from root explants grown on media supplemented with 2,4-D and BAP, but not on media supplemented with α-naphtheleneacetic acid (NAA) and BAP. N-(2-chloro-4-pyridyl)-N′-phenylurea and thidiazuron produced large yellow-green callus. However, none of these calli could be induced to produce shoots, even after 12 months in culture (Christensen et al. 2009). According to Jenderek and Olney (2001), the highest regeneration frequency in the Hibiscus genus has been achieved from callus induced from explants on NAA and BAP media. Mercuri et al. (2010) reported transformation of hairy roots of H. rosa-sinensis with the A. rhizogenes strain ATCC 15384 and they verified the transformation (T-DNA insertion) using PCR. They reported callus formation from one of many hairy roots derived from a cotyledon of a hybrid seed from the cross ‘Columbine’ × ‘Casanova Apricot’ and adventitious bud differentiation after one year of culture, but it was not clear if this bud was from a transformed hairy root.
We tested the ability of basal media supplemented with eight combinations of 2,4-D, NOA and BAP to induce callus on three types of explants (axillary bud, petiole and leaf) from three H. rosa-sinensis cultivars ‘Ruby’, ‘Bright Lights’ and ‘Ben James’. Our target was transformation of clonal H. rosa-sinensis, as it will be more practical for the improvement of selected elite genotypes. Like other researchers we found that callus induction, and subsequent shoot regeneration from the callus, was highly dependent upon genotype, explant type and plant growth regulator supplementation. Axillary buds are the explants of choice for callus initiation as callus was initiated at near 100 % in the axillary bud explants of all three cultivars independent of which auxin (NOA or 2,4-D) was used, or the ratio of auxin used in combination with BAP in the callus-inducing media. Callus induction from petiole and leaf explants was poor by comparison and showed greater dependency on the type of auxin present in the induction medium, with NOA being more effective at inducing callus than 2,4-D.
While callus induction is important, the differentiation of shoots from the callus is essential if transformed plants are to be produced. Many studies, including those on Dieffenbachia (Shen et al. 2007), Atropa (Song and Walworth 2013), Cicer (Tripathi et al. 2013) and Fraxinus (Palla and Pijut 2015) have shown that shoot organogenesis is induced in calli by the synergistic action of auxins with cytokinins. In our study, shoots could not be produced from callus derived from petiole or leaf explants from any of the cultivars. Shoots did regenerate from axillary bud-derived callus but the frequency was cultivar dependent. The highest shoot regeneration frequency (66.7 %) was obtained from axillary bud-derived callus of ‘Ruby’. The absolute amounts of NOA and BAP as well as their ratio appeared to have a large effect on the ability of the callus to regenerate shoots in that when both were present at lower concentrations, 2.47 and 2.22 µM respectively, the frequency of regeneration was 66.7 %, whereas when their concentrations were doubled, the shoot regeneration frequency dropped dramatically, to 5 %. Another factor for successful regeneration of shoots in our study could be the use of specialised media, previously optimised for H. rosa-sinensis by Christensen et al. (2008), in particular, the replacement of the traditionally used iron source ethylenediaminetetraacetic acid with ethylenediamine di-2-hydroxy-phenylacetate ferric, which provides higher iron availability in plant tissue culture media (Zawadzka and Orlikowska 2009). Furthermore, the pH in the media was balanced using 2-(N-morpholino)-ethanesulphonic acid as H. rosa-sinensis is known to be a pH-sensitive species (Valdez-Aguilar and Reed 2006). The regeneration protocol reported in this study is simple, involving the use of a single regeneration medium, on which visually distinct shoots were produced within 4 weeks. The shoots could be transferred to rooting medium after approximately 6 weeks of culture.
The regeneration efficiency reported here for ‘Ruby’ on selection media was 49 %, which is higher than for many other protocols developed for woody species such as Euonymus alatus (11.3 %; Chen et al. 2006) and Citrus paradisi (6.1 %; Costa et al. 2002). Kanamycin is the most widely used antibiotic in plant transformation when selecting for transformed plants, with concentrations typically ranging from 103.2 to 1.03 mM (Chawla 2002). Kanamycin sensitivity depends on the type of explant and species. A wide range of concentrations has been reported to inhibit organogenesis in Hibiscus spp., with H. syriacus (Yang et al. 1996) and H. cannabinus (Herath et al. 2004) requiring 103.2 µM whereas H. sabdariffa requiring twice as much kanamycin (Gassama-Dia et al. 2004), to inhibit shoot induction. In our study, shoot induction of H. rosa-sinensis ‘Ruby’ was completely inhibited at 82.56 µM kanamycin and 49 % of the calli produced shoots that grew on selective media. A randomly chosen subgroup of these putative transformed antibiotic-resistant plantlets, in total ten, were tested and shown using PCR amplification to have integrated the GUS transgene into their genomes.
Using primers specific to the GUS gene we further confirmed the transgenic nature of the plants by qPCR, which allowed an estimation of transgene copy number. Transgene copy number has traditionally been estimated by Southern blot analysis, and more recently other methods including fluorescence in situ hybridisation (Kallioniemi et al. 1996), multiplex amplifiable probe hybridisation (Armour et al. 2000) and microarray analysis (Lucito et al. 2000) have also been used. However, all these techniques are time consuming, require large amounts of gDNA, often involve the use of radioisotopes, and are laborious. qPCR is fast becoming the method of choice for transgenic plant analysis because it is a quick, robust, inexpensive and high-throughput alternative (Yuan et al. 2007). Using this technique, a reliable detection of specific PCR product was obtained and it was possible to estimate the transgene copy number through a high- to low-copy ordered series, using 18 S rRNA as the reference gene without a standard curve. The advantage of this approach is that it does not rely on the accuracy of DNA measurement for the standard curve and/or does not require a control plant sample with a known copy number (Bubner and Baldwin 2004; Li et al. 2004, Yuan et al. 2008). However, because the qPCR method gave only a rough estimation of transgene copy number, we further validated the transgenic plants by quantifying GUS activity on a subgroup of the transgenic plants.
Interestingly, none of the transformed plants showed GUS staining in the histochemical assay. Other authors have previously reported random, unpredictable or even no staining when in vivo histochemical staining of the GUS activity was performed on transformed plants (Serres et al. 1997; Chen et al. 2006; Batista et al. 2008). Plant secondary compounds are known to interfere with GUS staining in different tissues of plants (Jefferson et al. 1987). Phenolic compounds are one likely inhibitor group (Vainstein et al. 1993), which potentially react with GUS and other proteins through hydrogen bonding, oxidation and/or hydrophobic interactions immediately after being released from the cellular compartment (Loomis 1974). In our research, the total phenolic content of leaf extracts of H. rosa-sinensis ‘Ruby’ used in the transformation experiment was found to be several fold higher than those in petunia and tomato. These results may explain the lack of GUS staining observed in the transgenic lines obtained. However, GUS activity could be assayed in extracts from transgenic H. rosa-sinensis plants. Both histochemical and GUS activity assays result in the disruption of cellular membranes, leading to mixing of vacuolar-derived inhibitors with the cytoplasmic GUS. However, the disruption of cellular membranes in the GUS activity assay occurs by physical maceration of snap-frozen tissue, while no such macerations are involved in the histochemical assay. Thus, it is likely that the tissue maceration step performed at ultra-low temperature in liquid nitrogen during the GUS activity assay blocked the activation of such inhibitor/enzyme dynamics.
In our study the GUS activity assay was used to confirm the copy number estimations obtained using qPCR. A good correlation between GUS activity and qPCR-measured transgene abundance was not guaranteed because transgene expression can be affected both by site of transgene integration and by posttranscriptional gene silencing. However, our finding that high transgene copy number correlated well with higher rates of GUS activity provided confidence in the qPCR results and suggests that GUS activity assays can in some cases be useful for validating qPCR copy number estimations.
In conclusion, the results from the present study indicate that callus can be induced readily from axillary bud explants of H. rosa-sinensis using a combination of NOA and BAP and that the callus can be transformed with high efficiency using A. tumefaciens and induced to regenerate transformed shoots in a BAP-supplemented medium. All these processes however, are extremely dependent on the type of explant as well as on the cultivar. The qPCR results indicated that the transformed shoots carried different copy numbers of the transgene and the copy numbers corresponded with changes in measured GUS activity. Transgenic shoots could be rooted in an IAA-supplemented half-strength basal medium. The procedure described here may be used to transform H. rosa-sinensis, which has hitherto been recalcitrant to regeneration from transformed tissue.
References
Airò M, Giardina G, Farruggia G, Zizzo GV (2009) In vitro propagation of Hibiscus rosa-sinensis (L.). Acta Hortic 812:107–112
Armour JA, Sismani C, Patsalis PC, Cross G (2000) Measurement of locus copy number by hybridisation with amplifiable probes. Nucleic Acids Res 28:605–609
Banks SW, Gossett DR, Lucas MC, Millhollon EP, LaCelle M (1993) Agrobacterium-mediated transformation of kenaf (Hibiscus cannabinus L.) with the β-glucuronidase (GUS) gene. Plant Mol Biol Report 11:101–104
Batista D, Fonseca S, Serrazina S, Figueiredo A, Pais MS (2008) Efficient and stable transformation of hop (Humulus lupulus L.) var. Eroica by particle bombardment. Plant Cell Rep 27:1185–1196
Bubner B, Baldwin IT (2004) Use of real-time PCR for determining copy number and zygosity in transgenic plants. Plant Cell Rep 23:263–271
Campos-Vargas R, Saltveit ME (2002) Involvement of putative chemical wound signals in the induction of phenolic metabolism in wounded lettuce. Physiol Plant 114:73–84
Chawla HS (2002) Introduction to plant biotechnology, 2nd edn. Science Publishers, Plymouth
Chen Y, Lu L, Deng W, Yang W, McAvoy R, Zhao D, Pei Y, Luo K, Duan H, Smith W, Thammina C, Zheng X, Ellis D, Li Y (2006) In vitro regeneration and Agrobacterium-mediated genetic transformation of Euonymus alatus. Plant Cell Rep 25:1043–1051
Christensen B, Sriskandarajah S, Serek M, Müller R (2008) In vitro culture of Hibiscus rosa-sinensis L.: influence of iron, calcium and BAP on establishment and multiplication. Plant Cell Tissue Organ Cult 93:151–161
Christensen B, Sriskandarajah S, Müller R (2009) Transformation of Hibiscus rosa-sinensis L. by Agrobacterium rhizogenes. J Hortic Sci Biotechnol 84:204–208
Costa MGC, Otonoi WC, Moore GA (2002) An evaluation of factors affecting the efficiency of Agrobacterium-mediated transformation of Citrus paradisi (Macf) and production of transgenic plants containing carotenoid biosynthetic genes. Plant Cell Rep 21:365–373
De Stradis A, Parrella G, Vovlas C, Ragozzino A (2008) Vein yellowing of Hibiscus rosa-sinensis caused by eggplant mottled dwarf virus in southern Italy. J Plant Pathol 90:359–361
Feeney M, Bhagwat B, Mitchell JS, Lane WD (2007) Shoot regeneration from organogenic callus of sweet cherry (Prunus avium L.). Plant Cell Tissue Organ Cult 90:201–214
Gamborg O, Miller RA, Ojima K (1968) Nutrient requirements of suspension cultures of soybean root cells. Exp Cell Res 50:151–158
Gassama-Dia YK, Sanè D, Ndoye M (2004) Direct genetic transformation of Hibiscus sabdariffa L. Afr J Biotechnol 3:226–228
Gauthaman KK, Saleem MTS, Thanislas PT, Prabhu V, Krishnamoorthy K, Devaraj NS, Somasundara JS (2006) Cardioprotective effect of the Hibiscus rosa-sinensis flowers in an oxidative stress model of myocardial ischemic reperfusion injury in rat. BMC Complement Altern Med 6:3
Gleave AP (1992) A versatile binary vector with a T-DNA organizational structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol Biol 20:1203–1207
Herath SP, Suzuki T, Hattori K (2004) Multiple shoot regeneration from young shoots of kenaf (Hibiscus cannabinus). Plant Cell Tissue Organ Cult 77:49–53
Herath SP, Suzuki T, Hattori K (2005) Factors influencing Agrobacterium mediated genetic transformation of kenaf. Plant Cell Tissue Organ Cult 82:201–206
Hood EE, Gelvin SB, Melchers LS, Hoekema A (1993) New Agrobacterium helper plasmids for gene transfer to plants. Transgenic Res 2:208–218
Huang JG, Fan ZF, Li HF, Tian GZ, Hu JS (2004) First report of Tomato mosaic virus on Hibiscus rosa-sinensis in China. Plant Dis 88:683
Hunter DA, Watson LM (2008) The harvest-responsive region of the Asparagus officinalis asparagine synthetase promoter reveals complexity in the regulation of the harvest response. Funct Plant Biol 35:1212–1223
Hwang O, Cho M, Han YJ, Kim SH, Kim DS, Hwang I, Kim JI (2014) Agrobacterium-mediated genetic transformation of Miscanthus sinensis. Plant Cell Tissue Organ Cult 117:51–63
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Jenderek MM, Olney AJ (2001) Hibiscus syriacus plant regeneration from callus. Int Plant Propag Soc Prod 50:565–568
Kallioniemi A, Visakorpi T, Karhu T, Pinkel D, Kallioniemi OP (1996) Gene copy number analysis by fluorescence in situ hybridization and comparative genomic hybridization. Methods 9:113–121
Ke D, Saltveit ME (1989) Wound-induced ethylene production, phenolic metabolism and susceptibility to russet spotting in iceberg lettuce. Physiol Plant 76:412–418
Li ZW, Hansen JL, Liu Y, Zemetra RS, Berger PH (2004) Using real-time PCR to determine transgene copy number in wheat. Plant Mol Biol Report 22:179–188
Loaiza-Velarde JG, Tomás-Barberá FA, Saltveit ME (1997) Effect of intensity and duration of heat-shock treatments on wound-induced phenolic metabolism in Iceberg lettuce. J Am Soc Hortic Sci 122:873–877
Loomis WD (1974) Overcoming problems of phenolics and quinines in the isolation of plant enzymes and organelles. Methods Enzymol 31:528–544
Lucito R, West J, Reiner A, Alexander J, Esposito D, Mishra B, Powers S, Norton L, Wigler M (2000) Detecting gene copy number fluctuation in tumor cells by microarray analysis of genomic representations. Genome Res 10:1726–1736
Mao MJ, He ZF, Yu H, Li HP (2008) Molecular characterization of cotton leaf curl multan virus and its satellite DNA that infects Hibiscus rosa-sinensis. Chin J Virol 24:64–68
Mercuri A, Braglia L, Benedetti Ld, Ballardini M, Nicoletti F, Bianchini C (2010) New genotypes of Hibiscus rosa-sinensis through classical breeding and genetic transformation. Acta Hortic 855:201–208
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Palla KJ, Pijut PM (2015) Agrobacterium-mediated genetic transformation of Fraxinus americana hypocotyls. Plant Cell Tissue Organ Cult 120:631–641
Sachdewa A, Khemani LD (2003) Effect of Hibiscus rosa sinensis Linn. ethanol flower extract on blood glucose and lipid profile in streptozotocin induced diabetes in rats. J Ethnopharmacol 89:61–66
Sakhanokho HF (2008) In vitro multiple shoot induction and plant regeneration from shoot apices of Hibiscus acetosella Welw. Ex. Hiern. J Crop Improv 21:201–208
Serres R, McCown B, Zeldin E (1997) Detectable β-glucuronidase activity in transgenic cranberry is affected by endogenous inhibitors and plant development. Plant Cell Rep 16:641–646
Sharma S, Sultana S (2004) Effect of Hibiscus rosa-sinensis extract on hyperproliferation and oxidative damage caused by benzoyl peroxide and ultraviolet radiations in mouse skin. Basic Clin Pharmacol Toxicol 95:220–225
Shen X, Chen J, Kane ME (2007) Indirect shoot organogenesis from leaves of Dieffenbachia cv. Camouflage. Plant Cell Tissue Organ Cult 89:83–90
Song G, Walworth A (2013) Agrobacterium tumefaciens-mediated transformation of Atropa belladonna. Plant Cell Tissue Organ Cult 115:107–113
Tripathi L, Singh AK, Singh S, Singh R, Chaudhary S, Sanyal I, Amla DV (2013) Optimization of regeneration and Agrobacterium-mediated transformation of immature cotyledons of chickpea (Cicer arietinum L.). Plant Cell Tissue Organ Cult 113:513–527
Vainstein A, Fisher M, Ziv M (1993) Application of reporter genes to carnation transformation. Hortic Sci 28:1122–1124
Valdez-Aguilar LA, Reed DW (2006) Comparison of growth and alkalinity-induced responses in two cultivars of Hibiscus (Hibiscus rosa-sinensis L.). Hortic Sci 41:1704–1708
Vancanneyt G, Schmidt R, O’Connor-Sanchez A, Willmitzer L (1990) Construction of an intron-containing marker gene: splicing of the intron in transgenic plants and its use in monitoring early events in Agrobacterium-mediated plant transformation. Mol Gen Genet 220:245–250
Vankar PS, Srivastava J (2008) Comparative study of total phenol, flavonoid contents and antioxidant activity in Canna indica and Hibiscus rosa-sinensis: prospective natural food dyes. Int J Food Eng 4:14–20
Vazquez-Thello A, Yang LJ, Hidaka M, Uozomi T (1996) Inherited chilling tolerance in somatic hybrids of transgenic Hibiscus rosa-sinensis × transgenic Lavatera thurigiaca selected by double-antibiotic resistance. Plant Cell Rep 15:506–511
Venkatesh S, Thilagavathi J, Sundar SD (2008) Anti-diabetic activity of flowers of Hibiscus rosa-sinensis. Fitoterapia 79:79–81
Wang T, Ran Y, Atkinson RG, Gleave AP, Cohen D (2006) Transformation of Actinidia eriantha: a potential species for functional genomics studies in Actinidia. Plant Cell Rep 25:425–431
Winichayakul S, Moyle RL, Coupe SA, Davies KM, Farnden KJF (2004) Analysis of the asparagus (Asparagus officinalis) asparagine synthetase gene promoter identifies evolutionarily conserved cis-regulatory elements that mediate Suc-repression. Funct Plant Biol 31:63–72
Yang LJ, Vazquez-Tello A, Hidaka M, Masaki H, Uozomi T (1996) Agrobacterium-mediated transformation of Hibiscus syriacus and regeneration of transgenic plants. Plant Tissue Cult Lett 13:161–167
Yuan JS, Burris J, Stewart NR, Mentwab A, Stewart CN Jr (2007) Statistical tools for transgene copy number estimation based on real-time PCR. BMC Bioinformatics 8:1–12
Yuan JS, Wang D, Stewart NR, Mentwab A, Stewart CN Jr (2008) Statistical methods for efficiency adjusted real-time PCR analysis. Biotechnol J 3:112–123
Zawadzka M, Orlikowska T (2009) Influence of FeEDDHA on in vitro rooting and acclimatisation of red raspberry (Rubus idaeus L.) in peat and vermiculite. J Hortic Sci Biotechnol 84:599–603
Acknowledgments
We thank Sriya Pathirana and Andrew Mullan for tissue culture media preparation and Ian King for maintaining the plants in the greenhouse. We also thank Nick Albert for supplying the binary vector pNWA37, and Mary Christey and Murray Boase for critical reading of the manuscript. The first author is grateful to Prof. Vernieri Paolo and Prof. Serra Giovanni for a scholarship granted by University of Pisa (Italy) and Scuola Superiore Sant’Anna (Italy) for a year and half of work at Plant and Food Research, New Zealand.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Trivellini, A., Ferrante, A., Hunter, D.A. et al. Agrobacterium tumefaciens-mediated transformation of axillary bud callus of Hibiscus rosa-sinensis L. ‘Ruby’ and regeneration of transgenic plants. Plant Cell Tiss Organ Cult 121, 681–692 (2015). https://doi.org/10.1007/s11240-015-0738-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-015-0738-y