Abstract
Soybean is a difficult crop to manipulate through Agrobacterium tumefaciens-mediated genetic transformation. Plant–bacterium interaction plays an important role in the transformation process. Being rich with isoflavones, soybean may have an adverse effect on the A. tumefaciens-mediated genetic transformation. To investigate whether high content of endogenous isoflavones in soybean is a serious obstacle in achieving high efficient Agrobacterium-mediated transformation in soybean, a series of experiments on inhibition of isoflavone biosynthesis were conducted to improve upon soybean transformation efficiency. Results indicated that soybean isoflavones inhibited A. tumefaciens growth and respiration, the transformation efficiency [β-glucuronidase (GUS) transient expression] was negatively correlated with the phenylalanine ammonia-lyase activity and isoflavones content. The biosynthesis of soybean isoflavones was partially inhibited by sonication treatment and applying antagonists in co-culture medium and thereby decreased the adverse effects of isoflavones on Agrobacterium infection. A discernible improvement in transformation efficiency was achieved when sonication at 40 kHz for 3 min was applied along with Agro-infection and the explants were cultured on co-culture medium containing 20 μM α-aminooxyacetic acid (AOA), with the percentage of GUS transient expression as 41.4 %, being 3.6 times higher than that not sonicated and co-cultured on medium without AOA. Sonication was found not only to simply make micro-wounds for Agrobacterium to penetrate or releasing phenolic compounds for induced Agrobacterium vir gene expression; it disturbed the biosynthesis of isoflavones at the transcription level and decreased the adverse effects of isoflavones on soybean transformation, and thereby improving soybean transformation efficiency.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soybean [Glycine max (L.) Merrill.] is a dicotyledonous plant widely used for human and animal food and considered to be a compatible host for Agrobacterium (Pedersen et al. 1983), but difficult to transform via Agrobacterium tumefaciens mediated methods (Larkin 2001), although quite a few studies has been reported (Hinchee et al. 1988; Parrott et al. 1989; Meurer et al. 1998; Yan et al. 2000; Paz et al. 2006; Hong et al. 2007; Muhammad et al. 2010; Liu et al. 2014). In order to enhance soybean transformation via Agrobacterium various parameters have been evaluated. These parameters include the use of the super-virulent A. tumefaciens strain (Meurer et al. 1998; Ko et al. 2003, 2004; Yukawa et al. 2007, 2008), employment of mutant bacteria that constitutively express their vir genes (Hansen et al. 1994; Benzle et al. 2014), adding the phenolic compound like acetosyringone during inoculation and/or co-culture medium to induce expression of vir genes (Godwin et al. 1991), adding antioxidant substances, such as l-cysteine and thiol compounds in co-culture medium, to prevent tissue browning and necrosis (Olhoft et al. 2001, 2003, 2004, 2007; Zeng et al. 2004; Paz et al. 2004, 2006), and screening soybean genotypes that were competent to Agrobacterium infection and regeneration (Meurer et al. 1998; Paz et al. 2004; Song et al. 2013a, b). Sonication-assisted Agrobacterium-mediated transformation (SAAT) has also been used for transforming various tissue types and plant species to increase the number of infection sites (Trick and Finer 1997; Santarém et al. 1998; Meurer et al. 1998; Jiang et al. 2004; Zaragoza et al. 2004; Dutta et al. 2013; Teixeira da Silva and Dobránszki 2014). Those research works improved soybean transformation efficiency substantially either by strengthening the ability of Agrobacterium infection or by reducing tissue browning and necrosis to some extent. Although a few reports claimed high efficiency ranging from 9.4 to 26.2 % (Olhoft et al. 2001, 2003; Dang and Wei 2007), these milepost works in soybean genetic transformation were scarcely successfully replicated by other laboratory. Currently available transformation protocols for soybean transformation remained inefficient in most studies, with efficiency ranging from 0.2 to 2 %, very rarely up to 5–6 % (Atif et al. 2013) and limited to a few cultivars (Trick and Finer 1997). Soybean transformation efficiency was affected by many factors, genotypes, tissue types, A. tumefaciens strain, medium component and culture condition, transformant selection, etc., even the skills of operator. These may be the main reason for the poor repeatability. To date it has not been fully understand why soybean transformation via Agrobacterium is inefficient in most studies. One of the limitations for efficient plant transformation is a lack of understanding of gene expression during the selection and regeneration processes, plant–bacterium interactions also may prohibit or negatively affect the transformation process (Mello-Farias and Chaves 2008).
The defense response is considered to be one of the most important governing plant–microbe interaction during the transformation process and may strongly influence transformation efficiency as well (Mello-Farias and Chaves 2008; Anndrea Pitzschke 2013). Plants produce signal molecules that induce resistance responses following wounding or pathogen invasion (Mello-Farias and Chaves 2008). Soybean isoflavones play diverse roles in plant–microbe interactions (Subramanian et al. 2004, 2005). A variety of genes involved in regulating plant defense responses have already been identified in soybean (Larkin 2001), including many genes in isoflavone kind of secondary metabolic pathway. Wingender et al. (1989) detected enhanced CHS expression in the roots of sterile-grown soybean seedlings infected with Agrobacterium C58. Silencing genes of isoflavone synthase (IFS) or chalcone reductase (CHR) in soybean roots led to a nearly complete (95 %) suppression of all isoflavone metabolites in roots (Graham et al. 2007) and enhanced susceptibility to Phytophthora sojae (Subramanian et al. 2005; Graham et al. 2007).
Exogenous chemicals can be used in the regulation of soybean isoflavones synthesis. Phenylalanine ammonia-lyase (PAL) is the first enzyme in the plant secondary metabolism pathway. The isoflavonoid biosynthesis in cell suspension cultures of kudzu [Pueraria Lobata (WILLD) Ohwi] was found to be significantly inhibited by α-aminooxyacetic acid (AOA), a specific inhibitor of phenylalanine ammonia lyase (Li et al. 2009). Actinomycin D and cycloheximide inhibited the increase of PAL activity in sweet potato [Ipomoea batatas (Lam.) L.] (Wang and Xue 1981).
SAAT method was used in the transformation of various tissue types and plant species (genotypes), especially, that are considered to be difficult to transform (Bakshi et al. 2011; Subramanyam et al. 2011; King et al. 2014). It was believed that the micro-wounds produced by sonication treatment, allowing Agrobacterium to infect deeper within the plant tissue, were the main reason for the improvement in transformation efficiency by SAAT method (Trick and Finer 1997; Meurer et al. 1998). It was speculated that the micro-wound in plant tissues due to SAAT treatment released compounds that facilitated the growth and accumulation of bacteria under aerobic conditions so enhances transformation efficiency (Finer and Finer 2000). The candidate compounds were focused on phenolic compounds (Song et al. 2013a, b). However, decreased CHS expression following sonication treatment was detected (Larkin 2001), indicating that sonication may disturb the synthesis of isoflavones at the transcription level, rather than simply provide micro-wounds for Agrobacterium to penetrate into or release phenolic compounds for induced Agrobacterium vir gene expression.
From the above review of literature, we speculate that the low efficiency of soybean transformation is partially due to the inhibitory effects of isoflavones, a type of secondary metabolites, available in plenty in soybean, upon Agrobacterium infection. To our knowledge, literature on blocking isoflavone synthesis to improve soybean transformation has not been reported. The objectives of this study were to investigate whether endogenous soybean isoflavones were the obstacles to high efficient genetic transformation of soybean via the A. tumefaciens-mediated method and whether the inhibition of isoflavone biosynthesis by physical and chemical measures could decrease or eliminate its adverse effect on soybean genetic transformation and thereby improve upon transformation efficiency.
Materials and methods
Soybean cultivar Jihuang 13 was used as plant material. A. tumefaciens strain EHA105 (Hood et al. 1993) harboring a binary vector pCAMBIA2201 (GenBank: AF234314.1), containing both a GUS-intron gene and an NPTIIselectable marker, was used as the tested bacterium.
Effects of isoflavone on Agrobacterium tumefaciens growth
Agrobacterium tumefaciens strain EHA105::pCAMBIA2201 from a glycerol stock stored at −70 °C was incubated for 2–3 d at 28 °C on solid YEP medium (yeast extract 10 g l−1, peptone 10 g l−1, NaCl 5 g l−1, pH 7.2) containing 50 mg l−1 kanamycin and 25 mg l−1 rifampicin until colonies appeared. A single colony was inoculated in 30 ml of liquid YEP medium containing 50 mg l−1 kanamycin and 25 mg l−1 rifampicin for 4 h with 200 rpm shaking at 28 °C. These A. tumefaciens cultures were used as the initial solution to evaluate the effects of isoflavones on Agrobacterium growth. Treatments and their preparation were as listed in Table 1. The Agrobacterium solutions with different concentration of daidzein were cultured with 200 rpm shaking at 28 °C and their absorbance at 600 nm were detected using a spectrophotometer (UV-3200, Mapada Instruments, China) for 38 h at 2 h intervals. YEP medium containing different concentrations of daidzein was used as blank in the OD600nm measurements to eliminate the influence of daidzein on luminosity. Logistic equation of Agrobacterium growth was obtained by nonlinear regression analysis with SPSS11.0 software.
Respiration assay: Agrobacterium cultured overnight in YEP medium with approximately 0.8 OD600nm was centrifuged at 5,000 rpm and 4 °C for 10 min. The pellet was washed with 0.9 % NaCl solution once and then re-suspended in physiological saline solution to reach a final concentration of 10 g l−1. Agrobacterium respiration was assayed using an Oxygraph (Hansatech Oxygraph, Hansatech Instruments Ltd, Pentney, Norfolk, United Kingdom). The reaction systems include 1.8 ml of 0.1 M phosphate buffer, pH 7.2, 0.2 ml of 10 g l−1 glucose, and 0.5 ml of 10 g l−1 Agrobacterium suspension. Different concentration of daidzein was added to evaluate the effect of isoflavones on Agrobacterium respiration.
Plant explants preparation
Seeds were sterilized in a sealed desiccator with chlorine gas generated by the reaction between 100 ml of chlorine bleach and 4 ml 12 N HCl for 24 h and subsequently germinated on B5 medium (Gamborg et al. 1968) at 25 °C under 16 h/8 h light/dark photoperiod for 4 days. The cotyledonary node explants were prepared by removing the majority of the hypocotyl tissue approximately 0.5 cm below the cotyledonary node and removing the epicotyl and axillary buds about 1 mm above the cotyledonary node as described by Hong et al. (2007).
Soybean cotyledonary node transformation
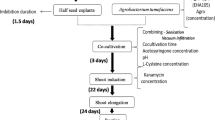
Recovery from −70 °C glycerol stock and propagation of A. tumefaciens strain EHA105::pCAMBIA2201 in liquid YEP medium were described above. When the OD600nm had reached to 0.8, the A. tumefaciens cultures were centrifuged for 5 min at 5,000 rpm and re-suspended in infection medium, i.e., 1/10 liquid MS salt with B5 vitamins medium containing 2 g l−1 2-(N-morpholine)-ethane sulphonic acid (MES), 0.05 mg l−1 benzylaminopurine (BA), 0.1 mg l−1 2,4-dichlorophenoxyacetic acid (2,4-D), 30 g l−1 sucrose, and 40 mg l−1 acetosyringone (AS), adjusting OD600nm to approx. 0.2–0.3. Soybean cotyledonary node transformations were prepared in groups: 15–20 explants were placed in a 50 ml glass tube containing 25 ml of infection medium and sonicated for 0 or 3 min in a bath sonicator KH2200B, then incubated for another 15 min at ambient temperature. Parallel experiments without Agrobacterium infection were conducted for evaluating the effects of antagonists of isoflavone synthesis and sonication on PAL activity and isoflavone accumulation. The explants were blotted dry on sterile Whatman paper to remove excess water before incubating for 3 days in darkness at 25 °C on solid co-culture medium, which had the same ingredients as the infection medium plus 1 mM dithiothreitol (DTT), 1 mM sodium thiosulfate, 3.3 mM cysteine, and 5.0 g l−1 agar, with or without 20 μM AOA. Fifteen explants were cultured on one plate per treatment with a minimum of three replicates. After co-culture, the explants were rinsed twice with sterile water and maintained on FNL medium (Samoylov et al. 1998) supplemented with 660 mg asparagine, 2 g l−1 MES, 0.2 mg l−1 BA, 0.1 mg l−1 Thidiazuron (TDZ), 0.2 mg l−1 2,4-D, 30 g l−1 sucrose, and 4.5 g l−1 agar, and 500 mg l−1 cefotaxime pH 5.8 for 7 days at 25 °C under 16 h/8 h light/dark photoperiod. Shoot induction and rooting followed the methods described by Hong et al. (2007). 100 mg l−1 kanamycin were used to select the transformant. GUS activity in explants 3 days after transformation and newly regenerated plantlets were assayed histochemically as described by Jefferson (1987). Percentage response was determined as the number of cotyledonary nodes of staining blue with X-gluc at the regenerable area divided by the number of cotyledonary nodes assayed.
Effects of antagonists of isoflavone synthesis and sonication on PAL activity and isoflavone accumulation
Explants at different time points after infection (24 h for Agrobacterium infected explants, and multiple time points between 0 and 30 h for those not infected by Agrobacterium) were sampled for the assay of PAL activity and determination of isoflavones content. The PAL activity assay followed the method described in “Guide for modern plant physiology experiments” [eds. Institute of Shanghai plant physiology, Chinese academy of sciences, Shanghai plant physiology society (China) 1999]. Ten explants per treatment was ground into homogenate in 10 ml 0.1 M Boric acid buffer solution (pH 8.8) on ice, the homogenate was filtrated through double-layer gauze and centrifuged for 30 min at 13,000 rpm and 4 °C condition, the supernatant was used as raw enzyme extracts. The reaction was performed in a 5 ml volume with 0.01 M Boric acid buffer (pH 8.8), 0.1 M Phenylalanine and 100 μl enzyme extracts at 37 °C for 1 h. Parallel control without 0.1 M phenylalanine was included for each treatments. Their absorbance at 290 nm was determined with a spectrophotometer (UV-3200, Mapada Instruments, China). The enzyme unit was defined as the difference between A290 nm in test tube and that in control tube divided by 1,000, the PAL activity was calculated as enzyme unit per hour per mg protein. Isoflavones were extracted with 80 % ethanol and sonicated at 40 kHz for 30 min at 50 °C in a bath sonicator (KH2200B, Kunshan Hechuang, China). Its absorbance at 260 nm was detected with a spectrophotometer (UV-3200, Mapada Instruments, China) to calculate the content of isoflavones according to the standard curve made from genistin as the standard isoflavone.
Expression of genes important for isoflavone biosynthesis
Quantitative RT-PCR assays for PAL, CHS, IFS1 and IFS2 were performed involving Agrobacterium-infected cotyledonary node explants incubated for 5 h on co-culture medium. Five explants were frozen immediately, ground in liquid nitrogen, then total RNA was extracted using 2 ml RNArose LS Reagent (Watson Biotechnologies, China) as described by Liu et al. (2011). The RNA was digested with DNase I (RNase Free) for 30 min to remove the contamination of genomic DNA. The cDNA was converted from RNA by performing reverse transcription PCR (RT-PCR) with RNA PCR Kit (AMV) Ver. 3.0 according to the manufactures introduction (Takara, Dalian, China) and was used as template in real-time quantitative PCR. Two-step real-time quantitative PCR was performed using the comparative Ct method on the 7500/7500 Fast Real-Time PCR System (Applied Biosystems, USA) according to the manufacturer’s instructions. The reactions were performed in a 20 µl volume with 0.25 µM primers. The nucleotides dNTP, Taq DNA polymerase, Mg2+, SYBR Green fluorescent dye, ROX reference dye and PCR buffer were included in the 2× RealStar Power SYBR Mixture (GenStar Biosolutions, Beijing, China). The cycle condition was preheating at 95 °C for 30 s; 40 cycles of 95 °C for 15 s and 60 °C for 60 s. The final dissociation stage was automatically run by the equipment to generate a melting curve and consequently verify the specificity of the amplification products. The relative quantification of gene expression was calculated as 2−ΔΔCт. The primer sequences were as follows: PAL forward: 5′-ATTATGGATTCAAGGGAGCT-3′, PAL reverse: 5′-AATGAGGAAAGTGGAGGACA-3′; IFS1 forward: 5′-GGCCACCTTCACCTCTTAAA-3′, IFS1 reverse: 5′-AGCCGAAGGAGAGAGAGAATA-3′; IFS2 forward: 5′-CCCTTCATAGGACACCTTCATC-3′, IFS2 reverse: 5′-CATGGAGCCAAAGTAGAGAGAG-3′; CHS forward: 5′-GGTCAACCCAAGTCCAAGAT-3′, CHS reverse: 5′-GGCGAAGGCCTAATAGTTTAGT-3′; Actin forward: 5′-GTGTCAGCCATACTGTCCCCATTT-3′, Actin reverse: 5′-GTTTCAAGCTCTTGCTCGTAATCA-3′. Actin was used as the endogenous control gene.
Results
Isoflavones inhibited Agrobacterium growth
The maximum OD600nm after 36 h of culture was decreased by 68.5 and 82.2 % for 1.0 and 2.0 mM daidzein treatments, respectively, compared to 0 mM daidzein treatment; lower concentration (0.17–0.34 % (v/v)) of solvent DMSO in YEP medium had negligible influence on Agrobacterium growth (Fig. 1a). The maximum velocity (Vmax) of Agrobacterium growth was decreased by 81.8 and 92.6 %, compared to 0 mM daidzein treatment, for 1.0 and 2.0 mM daidzein treatments, respectively; the appearance time of Vmax (Tmax) was delayed by about 5 h (Table 2). The respiration rate of Agrobacterium was inhibited by 29.7 and 36.9 % for 1 and 2 mM daidzein treatments, respectively, relative to their solvent control (Table 3).
The time course of Agrobacterium growth in YEP medium supplemented with different concentration of daidzein. Left demonstrate change of OD600nm with time course; right demonstrate change of Agrobacterium growth rate with time course. Daidzein was dissolved in DMSO to form a 600 mM stock and then added to YEP medium at low dosages. The Agrobacterium solution with different concentrations of daidzein were cultured with 200 rpm shaking at 28 °C. YEP medium containing different concentrations of daidzein was used as blank in OD600nm measurement. The data of OD600nm were collected from three replications and the whiskers represent the standard deviation about the mean. The Agrobacterium growth rate was obtained by the first order derivative of logistic regression equation
Antagonists of isoflavone synthesis and sonication interfered with isoflavone biosynthesis
Agrobacterium infection stimulated the expression of genes important for isoflavone biosynthesis (Fig. 2). The relative quantification (RQ) of gene expression after 5 h of infection, averaged between treatments with and without AOA addition, was increased by 56.8, 148.1, 70.6 and 118.3 %, for genes PAL, IFS1, IFS2 and CHS, respectively, when compared with that not transformed control. Antagonists and sonication decreased the expression of genes important for isoflavone biosynthesis (Figs. 2, 3). Antagonists of 20 μM AOA treatments decreased the relative expression of the PAL gene by 31.1 %, the IFS1 gene by 43.8 %, the IFS2 gene by 14.3 %, and the CHS gene by 47.0 %, relative to that of on medium without AOA. Sonication at 40 kHz for 3 min along with Agrobacterium infection decreased the relative gene expression by 32.6, 35.3, 56.5, and 21.8 %, for the genes PAL, IFS1, IFS2 and CHS, respectively, relative to that of Agrobacterium infection only.
Effects of α-aminooxyacetic acid (AOA) on expression of genes important for isoflavone biosynthesis. Samples were collected from three biological replications after Agrobacterium infection and incubated for 5 h on co-culture medium. Real-time quantitative PCR was performed using the comparative Ct method on the 7500/7500 Fast Real-Time PCR System (Applied Biosystems, USA) according to the manufacturer’s instructions. Actin was used as the endogenous gene control. Bars represent the mean data of relative quantification and whiskers represent the SD
Effects of sonication on expression of genes important for isoflavone biosynthesis. Samples were collected from three biological replications after Agrobacterium infection and Incubated for 5 h on co-culture medium. Real-time quantitative PCR was performed using the comparative Ct method on the 7500/7500 Fast Real-Time PCR System (Applied Biosystems, USA) according to the manufacturer’s instructions. Actin was used as the endogenous gene control. Bars represent the mean data of relative quantification and whiskers represent the SD
Antagonists of isoflavone synthesis AOA and sonication treatment decreased PAL activity relative to the control (Table 4). The highest PAL activity of control plants was at 12 h after treatment and was 44.1 % higher than that of at 0 h. The peak of PAL activity was not obvious for 20 μM AOA treatments, with only a 5.8 % increase at 6 h after treatment, relative to that of at 0 h; then the PAL activity decreased gradually to 83.1 % (at 24 h) and 73.9 % (at 30 h) of that at 0 h, respectively. The PAL activity of sonication treatment remained lower level and no obvious peak appeared throughout the tested period. Agrobacterium infection stimulated the increment of PAL activity by 62.6 % and increased isoflavone content by 4.7 % after 24 h of treatment, averaged among treatments of with (without) AOA and sonication, compared to that not infected with Agrobacterium (Table 5). Antagonists 20 μM AOA treatment decreased the PAL activity by 25.6 % and isoflavone content by 5.9 % on average compared to that without AOA treatment. Sonication at 40 kHz for 3 min decreased the PAL activity by 4.0 % and isoflavone content by 22.8 % on average compared to that without sonication treatment.
Inhibition of isoflavone biosynthesis improved soybean transformation
The inhibition of isoflavone biosynthesis by either antagonists or sonication increased the T-DNA delivery efficiency as evidenced by transient GUS expression (Table 6). When the cotyledonary nodes were infected with Agro strain EHA105::pCAMBIA2201 without sonication and co-cultured on AOA deficient medium, an average of 9.0 % explants displayed GUS activity. The percentage of GUS transient expression for 20 μM AOA treatment was 2.1-fold that of without AOA treatment, averaged between treatments with and without sonication; the data for sonication treatment was averagely 1.9-fold that of no sonication treatment. A spectacular enhancement of GUS expression was observed when sonication was applied along with Agrobacterium infection and the explants were cultured on co-culture medium containing 20 μM AOA, with the percentage of GUS transient expression as 41.41 %, being 3.6 times higher than that without sonication and AOA treatment. Different effects on following tissue proliferation and transformant selection were observed for sonication and antagonists AOA treatment, respectively. Sonication treatment at 40 kHz for 3 min resulted in about 30 % explants loss of regenerative ability due to sonication, while, 20 μM AOA treatment increased the percentage of explants with regenerated shoot buds by about ten percentage points (Table 6). Results of GUS staining indicated that GUS gene was not only transiently expressed in explants but also stably expressed in regenerated shoot buds (Fig. 4). The effects of antagonists and sonication on stable transformation need to be further evaluated.
Transformation of soybean cotyledonary nodes with Agrobacterium strain EHA105::pCAMBIA2201. a Explant preparation, arrow indicates explant. b–d Shoot induction on selection medium. e, f Transient expression of GUS after 3 days co-culture. g, h and l Stable expression of GUS in regenerated new buds (the right one in l was from not transformed control). Bars represent 3 mm
Discussion
Transgenic soybean plants were produced via both Agrobacterium-mediated genetic transformation (Hinchee et al. 1988) and particle bombardment (McCabe et al. 1988). Nevertheless, works aimed to improve upon soybean transformation efficiency did not bring major breakthroughs during the following two decades (Yamada et al. 2010, 2012; Wiebke et al. 2011; Mariashibu et al. 2012). This status forced us to think about the mechanism underline the recalcitrant transformation of soybean. It was speculated that plant–bacterium interactions might prohibit or negatively affect the transformation process (Mello-Farias and Chaves 2008), and better understanding the ongoing molecular battle between agrobacteria and attacked hosts provide a breakthrough toward developing transformation protocols for recalcitrant plant species (Andrea Pitzschke 2013).
Many genes in isoflavone secondary metabolism are involved in plant defense responses (Larkin 2001), the best-characterized plant defense genes are members of the CHS family. Wounding and Agrobacterium infection could enhance the isoflavone metabolic pathway. Wingender et al. (1989) once detected enhanced CHS expression in sterile soybean roots infected with Agrobacterium C58 with maximum expression at 4–8 h after infection. In the present study, we detected enhanced expression of PAL, IFS1, IFS2, and CHS gene important for isoflavone biosynthesis due to wounding and Agrobacterium infection. The enhanced expression of defensive genes may well explain the low efficiency in soybean transformation. Soybean isoflavones play diverse roles in plant–microbe interactions (Subramanian et al. 2005). Isoflavones inhibited A. tumefaciens growth and respiration, the Vmax of Agrobacterium growth was decreased by more than 95 % and Agrobacterium respiration was inhibited by 35 % above with 2 mM daidzein treatment. The Tmax was delayed about 5 h by 1–2 mM daidzein treatments, implying that extending the co-culture time to three more days may further improve the efficiency of transformation. Whether the inhibition of isoflavone biosynthesis and by what could decrease or eliminate its adverse effect on soybean transformation? Silencing the isoflavone synthase (IFS) gene or chalcone reductase (CHR) in soybean roots led to a nearly complete (95 %) suppression of all isoflavone metabolites in roots (Graham et al. 2007), and enhanced susceptibility to Phytophthora sojae (Subramanian et al. 2005; Graham et al. 2007). Both physical (sonication) and chemical (antagonists) measures were adopted in this study to inhibit soybean isoflavone biosynthesis. The results showed that sonication and antagonists of isoflavone synthesis AOA were all found to be effective in decreasing the expression of genes governing isoflavone biosynthesis and PAL activity, also significantly decreased isoflavones accumulation. The isoflavone content was decreased by 5.9 % with 20 μM AOA treatment and 22.8 % with sonication at 40 kHz for 3 min treatment. Significant negative correlations were observed between the percentage of GUS transient expression and the relative quantification of gene expression as well as the PAL activity (Table 7), the isoflavone content was also negatively correlated with the transformation efficiency but didn’t achieved significant level (r = −0.8819, n = 4). The results implied that decreasing the intensity of defense reaction during Agrobacterium infection is more important than lowering isoflavone content itself.
Sonication has been used to enhance Agrobacterium-mediated transformation in a few different plant species (Trick and Finer 1997; Meurer et al. 1998; Bakshi et al. 2011; Subramanyam et al. 2011; Dutta et al. 2013; Teixeira da Silva and Dobránszki 2014; King et al. 2014). It was generally agreed that enhanced transformation from using SAAT most likely results from the micro-wounds both on the surface of and deep within the target tissue, which permit Agrobacterium to penetrate deep within the plant tissue and infect a large number of plant cells (Santarém et al. 1998; Finer and Finer 2000). The micro-wounds may aid in releasing compounds to facilitate growth and the accumulation of bacteria (Finer and Finer 2000). The candidate metabolites were focused on phenolic compounds (Song et al. 2013a, b). SAAT treatment was not effective at post co-cultivation period with decreased shoot proliferation from cotyledonary node of some soybean genotypes (Meurer et al. 1998). This might attribute to a not appropriate ultrasound intensity and duration. The appropriate intensity and duration depend on species and tissue types. Sonication at 50 kHz for more than 10 s resulted in tissue turned white and death a few days after sonication in immature pods of soybean (Santarém et al. 1998). Increased GUS-stained spots were observed, but few explants survived to 10 days when the cotyledonary nodes of mature soybean seeds were sonicated for 600 s (Meurer et al. 1998). The standardized sonication time was determined as 2 s for immature cotyledons (Santarém et al. 1998) and 6 s for mature cotyledons (Meurer et al. 1998) based on the transient GUS expression and continued tissue proliferation. The sonication treatment was implemented at 40 kHz for 3 min in this study and resulted in a 1.9-fold increase of transient GUS expression and about 30 % explants loss of regenerative ability due to sonication at the same time (Table 6), a rather longer sonication duration and lighter tissue damage than that previously reported (Santarém et al. 1998). This may be attributed to the different intensity and direction of the ultrasound wave. The waterbath sonicator used in this study was 40 kHz with only vertical wave from the bottom; its ultrasound is weaker than that of sonicator (Model PC5) which was 55 kHz with both vertical and horizontal wave output (Santarém et al. 1998; Meurer et al. 1998; Subramanyam et al. 2011). The tissue type may also affect the sonication effects and need further experimentation. Sonication treatment decreased the enhanced expression of the PAL, IFS1, IFS2 and CHS genes, PAL activity, and isoflavones accumulation owing to wounding and Agrobacterium infection and promoted the transient GUS expression, indicating that sonication may not simply provide micro-wounds for Agrobacterium to penetrate or release phenolic compounds to induce A. vir gene expression, but it may disturb the synthesis of isoflavones at the transcription level, which decreases the adverse effects of isoflavones on soybean transformation. Expression profiling by RNA-Seq (Quantification) analysis indicated that 3,524 (1,538) genes were down-regulated (up-regulated) by inhibition of isoflavone biosynthesis. These differentially expressed genes involved into three GO categories: cellular component, molecular function, and biological process. KEGG Pathway enrichment analysis showed that the DEGs were enriched in fatty acid biosynthesis, flavone and flavonol biosynthesis, flavonoid biosynthesis, phenylpropanoid biosynthesis, propanoate metabolism, pentose and glucuronate interconversion, DNA replication, etc. (unpublished data). These results were generally in consistent with the view points of the present study that sonication and antagonists interfered with isoflavone biosynthesis. Sonication may also involve the regulation of circadian rhythm-plant (unpublished data).
In summary, higher content of isoflavones in soybean was the major obstacles to achieve high efficient Agrobacterium-mediated soybean transformation and negatively correlated with the percentage of GUS transient expression. Blocking the pathway of isoflavone synthesis decreased its adverse effects on soybean transformation. The most effective measure was sonication at 40 kHz for 3 min along with Agrobacterium infection and 20 μM AOA antagonists was added in the infection and co-culture medium. A synergistic effect was observed by the combined use of sonication and antagonists AOA, with the percentage of GUS transient expression as 41.4 %, being 3.6 times higher than that not sonicated and co-cultured on AOA deficient medium. Sonication was found not simply created micro-wounds for Agrobacterium to penetrate or releasing phenolic compounds to induce A. vir expression, but rather it disrupted the synthesis of isoflavones at transcription level and decreased the adverse effects of isoflavones on soybean transformation, and thereby improved soybean transformation efficiency.
Abbreviations
- PAL:
-
Phenylalanine ammonia-lyase
- AOA:
-
α-Aminooxyacetic acid
- AS:
-
Acetosyringone
- SAAT:
-
Sonication-assisted Agrobacterium-mediated transformation
References
Atif RM, Patat-Ochatt EM, Svabova L, Ondrej V, Klenoticova H, Jacas L, Griga M, Ochatt SJ (2013) Gene transfer in legumes. Prog Bot 74:37–100
Bakshi S, Sadhukhan A, Mishra S, Sahoo L (2011) Improved Agrobacterium-mediated transformation of cowpea via sonication and vacuum infiltration. Plant Cell Rep 30:2281–2292
Benzle KA, Finer KR, Marty DM, McHale LK, Goodner BW, Taylor CG, Finer JJ (2014) Isolation and characterization of novel Agrobacterium strains for soybean and sunflower transformation. Plant Cell Tissue Organ Cult. doi:10.1007/s11240-014-0679-x
Dang W, Wei ZM (2007) An optimized Agrobacterium-mediated transformation for soybean for expression of binary insect resistance genes. Plant Sci 173:381–389
Dutta I, Kottackal M, Tumimbang E, Tajima H, Zaid A, Blumwald E (2013) Sonication-assisted efficient Agrobacterium-mediated genetic transformation of the multipurpose woody desert scrub Leptadenia pyrotechnica. Plant Cell Tissue Organ Cult 112:289–301
Finer KR, Finer JJ (2000) Use of Agrobacterium expressing green fluorescent protein to evaluate colonization of sonicated-assisted Agrobacterium-mediated transformation treated soybean cotyledons. Lett Appl Microbiol 30:406–410
Gamborg OL, Miller RA, Ojima K (1968) Nutrient requirements of suspension cultures of soybean root cells. Exp Cell Res 50:151–158
Godwin I, Todd G, Ford-Loyd B, Newbury HI (1991) The effects of acetosyringone and pH on Agrobacterium-mediated transformation vary according to plant species. Plant Cell Rep 9:671–675
Graham TL, Graham MY, Subramanian S, Yu O (2007) RNAi silencing of genes for elicitation or biosynthesis of 5-deoxyisoflavonoids suppresses race-specific resistance and hypersensitive cell death in Phytophthora sojae infected tissues. Plant Physiol 144:728–740
Hansen G, Das A, Chilton MD (1994) Constitutive expression of the virulence genes improves the efficiency of plant transformation by Agrobacterium. Proc Natl Acad Sci USA 91:7603–7607
Hinchee MAW, Connor-Ward DV, Newell CA, McDonnell RE, Sato SJ, Gasser CS, Fischhoff DA, Re DB, Fraley RT, Horsch RB (1988) Production of transgenic soybean plants using Agrobacterium-mediated DNA transfer. Bio/Technology 6:915–922
Hong HP, Zhang H, Olhoft P, Hill S, Wiley H, Toren E, Hillebrand H, Jones T, Cheng M (2007) Organogenic callus as the target for plant regeneration and transformation via Agrobacterium in soybean (Glycine max (L.) Merr.). In Vitro Cell Dev Biology-Plant 43:558–568
Hood EE, Gelvin SB, Melcher LS, Hoekema A (1993) New Agrobacterium helper plasmids for gene transfer to plants. Transgenic Res 2:208–218
Institute of Shanghai Plant Physiology, Chinese Academy of Sciences, Shanghai Plant Physiology Society (China), (ed) (1999) Guide for modern plant physiology experiments. Science press, Beijing
Jefferson RA (1987) Assaying chimeric genes in plants: the GUS gene fusion system. Plant Mol Biol Rep 5:387–405
Jiang L, Maoka T, Komori S, Fukamachi H, Kato H, Ogawa K (2004) An efficient method for sonication assisted Agrobacterium-mediated transformation of coat protein (CP) coding genes into papaya (Carica papaya L.). J Exp Biol 3:189–198
King JL, Finer JJ, McHale LK (2014) Development and optimization of agroinfiltration for soybean. Plant Cell Rep. doi:10.1007/s00299-01-1694-4
Ko TS, Korban SS (2004) Enhancing the frequency of somatic embryogenesis following Agrobacterium-mediated transformation of immature cotyledons of soybean [Glycine max (L.) Merrill]. In Vitro Cell Dev Biology-Plant 40:552–558
Ko TS, Lee S, Krasnyanski S, Kothan SS (2003) Two critical factors are required for efficient transformation of multiple soybean cultivar: Agrobaeterium strain and orientation of immature cotyledonary explants. Theor Appl Genet 107:439–447
Larkin KM (2001) Optimization of soybean transformation using SAAT and GFP. Wooster: OARDC/OSU p. 126. (Thesis—Master)
Li HQ, Yang H, Zhang JJ, Wan XC, Fang CB (2009) The effect of specific inhibitors of phenylalanine ammonia-lyase and 4-coumarate-CoA ligase on isoflavone biosynthesis in Kudzu cell suspension culture. Chin J Trop Crops 30(1):47–52
Liu ZH, Zhang HM, Li GL, Guo XL, Chen SY, Liu GB, Zhang YM (2011) Enhancement of salt tolerance in alfalfa transformed with the gene encoding for betaine aldehyde dehydrogenase. Euphytica 178:363–372
Liu SC, Zhang GC, Yang LF, Mii M, Gai JY, Zhu YL (2014) Bialaphos-resistant transgenic soybeans produced by the Agrobacterium-mediated cotyledonary-node method. J Agr Sci Technol 16:175–190
Mariashibu TS, Subramanyam K, Arun M, Mayavan S, Rajesh M, Theboral J, Manickavasagam M, Ganapathi A (2012) Vacuum infiltration enhances the Agrobacterium-mediated genetic transformation in Indian soybean cultivars. Acta Physiol Plan. doi:10.1007/s11738-012-1046-3
McCabe DE, Swain WF, Martinell BJ, Christou P (1988) Stable transformation of soybean (Glycine max) by particle acceleration. Biotechnology 6:923–926
Mello-Farias PC, Chaves ALS (2008) Advances in Agrobacterium-mediated plant transformation with emphases on soybean. Sci Agric 65:95–106
Meurer CA, Dinkin RD, Collins GB (1998) Factors affecting soybean cotyledonary node transformation. Plant Cell Rep 18:180–186
Muhammad Z, Bushra M, Salman AM, Muhammad FC (2010) Expression of genes in transgenic soybean (L.) leads to changes in plant phenotype, leaf morphology, and flowering time. Plant Cell Tissue Organ Cult 103:227–236
Olhoft PM, Lin K, Galbraith J, Nielsen NC (2001) The role of thiol compounds in increasing Agrobacterium-mediated transformation of soybean cotyledonary-node cells. Plant Cell Rep 20:731–737
Olhoft PM, Flagel E, Donovan CM, Somers DA (2003) Efficient soybean transformation using hygromycin B selection in the cotyledonary-node method. Planta 216:723–735
Olhoft PM, Flagel LE, Somers DA (2004) T-DNA locus structure in a large population of soybean plant transformed using the Agrobacterium-mediated cotyledonary-node method. Plant Biotechnol J 2(4):289–300
Olhoft PM, Bernal LM, Grist LB, Hill DS, Mankin SL, Shen Y, Kalogerakis M, Wiley H, Toren E, Song HS, Hillebrand H, Jones T (2007) A novel Agrobacterium rhizogenes-mediated transformation method of soybean [Glycine max (L.) Merrill] using primary-node explants from seedlings. In Vitro Cell Dev Biology-Plant 43:536–549
Parrott WA, Hoffman LM, Hildebrand DF, Williams EG, Collins GB (1989) Recovery of primary transformants of soybean. Plant Cell Rep 7:615–617
Paz MM, Shou H, Guo Z, Zhang Z, Banerjee A, Wang K (2004) Assessment of conditions affecting Agrobacterium-mediated soybean transformation using the cotyledonary node explant. Euphytica 136:167–179
Paz MM, Martinez JC, Kalvig AB, Fonger TM, Wang K (2006) Improved cotyledonary node method using an alternative explant derived from mature seed for efficient Agrobacterium-mediated soybean transformation. Plant Cell Rep 25:206–213
Pedersen HC, Christiansen J, Wyndaele R (1983) Induction and in vitro culture of soybean crown gall tumors. Plant Cell Rep 2:201–204
Pitzschke A (2013) Agrobacterium infection and plant defense—transformation success hangs by a thread. Front Plant Sci. doi:10.3389/fpls.2013.00519
Samoylov VW, Tucker DM, Thibaud-Nissen F, Parrott WAA (1998) liquid-medium-based protocol for rapid regeneration from embryogenic soybean cultures. Plant Cell Rep 18:49–54
Santarém ER, Trick HN, Essig JS, Finer JJ (1998) Sonication-assisted Agrobacterium-mediated transformation of soybean immature cotyledons: optimization of transient expression. Plant Cell Rep 17:752–759
Song KT, WC Yim, Jung GH, Kim SL, Kwon YU, Lee BM (2013a) Relationship of transformation efficiency and metabolites induced in Korean soybean cotyledons treated with sonication. Korean J Crop Sci Hanguk Jakmul Hakhoe Chi 58(2):119–127
Song ZY, Tian JL, Fu WZ, Li L, Lu LH, Zhou L, Shan ZH, Tang GX, Shou HX (2013b) Screening Chinese soybean genotypes for Agrobacterium-mediated genetic transformation suitability. J Zhejiang Univ Sci B 14(4):289–298
Subramanian S, Hu X, Lu G, Odell JT, Yu O (2004) The promoters of two isoflavone synthase genes respond differentially to nodulation and defense signals in transgenic soybean roots. Plant Mol Biol 54:623–639
Subramanian S, Graham MY, Yu O, Graham TL (2005) RNA interference of soybean isoflavone synthase genes leads to silencing in tissues distal to the transformation site and to enhanced susceptibility to Phytophthora sojae. Plant Physiol 137:1345–1353
Subramanyam K, Subramanyam K, Sailaja KV, Srinivasulu M, Lakshmidevi K (2011) Highly efficient Agrobacterium-mediated transformation of banana cv. Rasthali (AAB) via sonication and vacuum infiltration. Plant Cell Rep 30:425–436
Teixeira da Silva JA, Dobránszki J (2014) Sonication and ultrasound: impact on plant growth and development. Plant Cell Tissue Organ Cult 117:131–143
Trick HN, Finer JJ (1997) SAAT: sonication-assisted Agrobacterium mediated transformation. Transgenic Res 6:329–336
Wang JW, Xue YL (1981) Studies on plant phenylalanine ammonia-lyase I. The effect of phytohormone on the increase in phenylalanine ammonia-lyase (PAL) and cinnamic acid 4-hydroxylase (CA4H) activity and the sequence of concomitant changes of enzyme activity in sweet potato root tuber discs. Acta Phytophysiol Sin 7(4):373–380
Wiebke SB, Droste A, Pasquali G, Osorio MB, Bucker NL, Passaglia LMP, Bencke M, Homrich MS, Margis PM, Bodanese ZM (2011) Transgenic fertile soybean plants derived from somatic embryos transformed via the combined DNA-free particle bombardment and Agrobacterium system. Euphytica 177:343–354
Wingender R, Rohrig H, Horicke C, Wing D, Schell J (1989) Differential regulation of soybean chalcone synthase genes in plant defense, symbiosis, and upon environmental stimuli. Mol Gen Genet 218:315–322
Yamada T, Watanabe S, Arai M, Harada K, Kitamura K (2010) Cotyledonary node pre-wounding with a micro-brush increased frequency of Agrobacterium-mediated transformation in soybean. Plant Biotechnol 27:217–220
Yamada T, Takagi K, Ishimoto M (2012) Recent advances in soybean transformation and their application to molecular breeding and genomic analysis. Breed Sci 61:480–494
Yan B, Srinivas RM, Collins GB, Dinkins RB (2000) Agrobacterium tumefaciens -mediated transformation of soybean [Glycine max (L.) Merrill] using immature zygotic cotyledon explants. Plant Cell Rep 19:1090–1097
Yukawa K, Kaku H, Tanaka H, Koga-Ban Y, Fukuda M (2007) Enhanced soybean infection by the legume “supervirulent” Agrobacterium tumefaciens strain KAT23. Biosci Biotechnol Biochem 71:1676–1682
Yukawa K, Kaku H, Tanaka H, Koga-Ban Y, Fukuda M (2008) Enhanced soybean infection by the legume “Super-Virulent” Agrobacterium tumefaciens strain KAT23. Biosci Biotechnol Biochem 72:1809–1816
Zaragoza C, Munoz-Bertomeu J, Arrillaga I (2004) Regeneration of herbicide-tolerant black locust transgenic plants by SAAT. Plant Cell Rep 11:832–838
Zeng P, Vadnais DA, Zhang Z, Polacco JC (2004) Refined glufosinate selection in Agrobacterium-mediated transformation of soybean [Glycine max (L.) Merrill]. Plant Cell Rep 22:478–482
Acknowledgments
This work was supported by National Major Project for Transgenic Crops of Chinese Agriculture Ministry (Grant No. 2014ZX0800402B); Natural Science Foundation of Hebei Province, China (Grant No. C2013301033); Key project for fundamental research of Hebei Province, China (Grant No. 14962903D).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, YM., Zhang, HM., Liu, ZH. et al. Inhibition of isoflavone biosynthesis enhanced T-DNA delivery in soybean by improving plant–Agrobacterium tumefaciens interaction. Plant Cell Tiss Organ Cult 121, 183–193 (2015). https://doi.org/10.1007/s11240-014-0693-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-014-0693-z