Abstract
Insight into control of proton transfer, a crucial attribute of cellular functions, can be gained from investigations of bacterial reaction centers. While the uptake of protons associated with the reduction of the quinone is well characterized, the release of protons associated with the oxidized bacteriochlorophyll dimer has been poorly understood. Optical spectroscopy and proton release/uptake measurements were used to examine the proton release characteristics of twelve mutant reaction centers, each containing a change in an amino acid residue near the bacteriochlorophyll dimer. The mutant reaction centers had optical spectra similar to wild-type and were capable of transferring electrons to the quinones after light excitation of the bacteriochlorophyll dimer. They exhibited a large range in the extent of proton release and in the slow recovery of the optical signal for the oxidized dimer upon continuous illumination. Key roles were indicated for six amino acid residues, Thr L130, Asp L155, Ser L244, Arg M164, Ser M190, and His M193. Analysis of the results points to a hydrogen-bond network that contains these residues, with several additional residues and bound water molecules, forming a proton transfer pathway. In addition to proton transfer, the properties of the pathway are proposed to be responsible for the very slow charge recombination kinetics observed after continuous illumination. The characteristics of this pathway are compared to proton transfer pathways near the secondary quinone as well as those found in photosystem II and cytochrome c oxidase.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Control of the kinetics and thermodynamics of proton transfer is key to numerous chemical processes in cells. The concept of hydrogen-bonds providing pathways for fast and efficient stepwise proton transfer in “proton wires” stems from the mechanism of charge migration in aqueous electrolysis introduced over 200 years ago (Agmon 1995). However, biological systems require explanation of the dependence of proton transfer on the detailed chemical nature of a proton transfer network extending from a proton donor to an acceptor in large protein-cofactor complexes. For example, catalytic active sites of proteins often achieve local control of protons through sequestration from bulk solvent followed by transfer of protons via hydrogen-bonded networks in multi-step transfers over 10–20 Å. The inhomogeneous distributions of electrostatic and hydrophobic interactions in proteins can tune the properties of the amino acid residues to favor proton transfer. Determination of how such interactions operate on specific amino acid residues remains a challenge.
Proton transfer in protein complexes is often coupled with electron transfer (Barbara et al. 1996; Wraight 2006; Migliore et al. 2014). Fundamental differences exist between their transfer mechanisms on a phenomenological level, with electrons being highly mobile and exhibiting quantum behavior, while protons are ~ 2000 times heavier and more semi-classical in their motion. Electrons tunnel over many length scales, and atomic and molecular orbital descriptions have been developed describing processes that facilitate electron transfer through well-defined pathways. These models are predictive of the electron transfer rates, which are primarily sensitive to the donor–acceptor separation distance. In contrast, proton tunneling is restricted to a hydrogen-bond length, preventing application of electron transfer models for proton transfer. The development of general principles of the coupling of protons to electrons during catalysis at an active site has progressed considerably (Hammes-Schiffer and Stuchebrukhov 2010; Weinberg et al. 2012; Gunner et al. 2013; Layfield and Hammes-Schiffer 2014). However, limitations arise in describing proton transfer over molecular distances in protein complexes and in the application of computer algorithms to the large membrane protein complexes that transfer protons in cells. The continued coordination of theoretical and experimental aspects will contribute to a better understanding of proton-coupled electron transfer in proteins (Hammes-Schiffer 2015; Gunner and Baker 2016).
Light-driven proton and electron transfer occurs in reaction centers from Rhodobacter sphaeroides, providing a system for probing the coupling of these processes in large biological complexes. These bacterial reaction centers are integral membrane protein complexes consisting of three protein subunits that surround two branches of cofactors. They perform the primary photochemistry in bacterial photosynthesis, where light energy is converted into a proton gradient across the cell membrane. After light is absorbed by the bacteriochlorophyll dimer, P, an electron is transferred through one of the branches of cofactors to the primary quinone QA, followed by transfer to the secondary quinone QB (Fig. 1). Proton transfer occurs when the transfer of a second electron to QB is accompanied by the uptake of two protons from the surrounding aqueous environment. The resulting quinol is released to the membrane and replaced with an exogenous quinone. These proton-coupled electron transfer reactions have been thoroughly studied with the proton transfer pathways from the solvent to QB being well mapped (Okamura et al. 2000; Wraight 2006; Gunner et al. 2013; Maróti 2019; Sugo et al. 2021; Wei et al. 2022).
Three-dimensional structure of the reaction center from R. sphaeroides. Shown are the amino acid residues that were tested for their contribution to proton release, including residues from the L subunit, Thr L130, Leu L131, Arg L135, Asp L155, Tyr L162, Ser L244, and Cys L247 (yellow) and from the M subunit, Arg M164, Asp M184, Thr M186, Ser M190, His M193, and Tyr M210 (blue). Also shown are the cofactors, including the bacteriochlorophyll dimer P (red), the primary quinone QA and secondary quinone QB (red), and the bacteriochlorophyll monomers, bacteriopheophytins, and non-heme iron (wheat)
On the other side of the protein, a protracted presence of the oxidized bacteriochlorophyll dimer, P+, is associated with the release of protons. For purified reaction centers, excitation using a short laser flash results in very limited proton release (Maróti and Wraight 1988; McPherson et al. 1988). However, continuous illumination produces a significant release of protons, up to six protons per reaction center for a 300 s light exposure (Kálmán and Maróti 1997). The recovery of this proton release in the dark occurs with a multi-phasic behavior. Complex kinetic behavior for charge recombination of the P+Q– state under continuous illumination has been shown by a variety of spectroscopic measurements, such as monitoring P+ using electron paramagnetic resonance spectroscopy (Andréasson and Andréasson 2003). Multiple pulses of saturating white light for up to 5 min results in the same outcome as observed for continuous illumination (van Mourik et al. 2001; Manzo et al. 2011; Serdenko et al. 2016). The multiphasic slow behavior depends on the presence of water, as continuous illumination (20 s) of dehydrated wild-type reaction centers samples results in only the normal fast recovery component (Malferrari et al. 2013). Combined measurement of the optical and proton release properties of reaction centers altered using mutagenesis show that both the proton release and altered charge recombination kinetics in wild-type upon continuous illumination are absent when the reaction centers contain the mutation Leu to His at L131 (Deshmukh et al. 2011a, b). Because they are both observed under continuous illumination, the link between the proton release and long-lived kinetics provides an interpretation that the light-adapted state differs in the protonation of amino acid residues near P.
If amino acid residues in the environment of P+ are participants in proton release, they should provide clues to the connection between proton and electron transfer. We surveyed protonatable and polar amino acid residues within 10 Å of P with the additional restriction that the residues could potentially form a proton release pathway from P to the periplasmic surface. In total, twelve prospective positions were chosen, six from the L subunit and six from the M subunit, for modification using site-directed mutagenesis to test which alterations had an impact on proton transfer (Fig. 1). Mutations were designed to remove polar and protonatable side chains, so that if the residues are involved in proton release, the altered reaction centers would show a decrease in the amount of proton release relative to wild-type. Structurally conservative substitutions at these residues were identified as Thr to Val at L130, Arg to Met at L135, Asp to Asn at L155, Tyr to Phe at L162, Ser to Ala at L244, Cys to Ala at L247, Arg to Met at M164, Asp to Asn at M184, Thr to Val at M186, Ser to Ala at M190, His to Leu at M193, and Tyr to Phe at M210. Using the reaction center structure, modeling showed that none of the substitutions produced any unfavorable steric interactions with the surrounding protein. The L and M subunits are symmetry-related, and two pairs of residues, Arg L135–Arg M164 and Asp L155–Asp M184, in the set are at comparable positions.
The known biochemistry and spectroscopic backgrounds of mutant reaction centers includes the electron transfer properties studied previously for changes of several of the residues in this set. Tyr M210 lies between P and the initial electron acceptors, and mutant reaction centers have altered rates of the initial electron transfer (Williams and Allen 2009; Jones 2009; Weaver et al. 2021). The rate and yield of electron transfer from cytochrome c2 to P+ are decreased in R. sphaeroides upon mutation of Tyr L162, which is located in a bridging position (reviewed in Axelrod and Okamura 2005). Ser M190 is also positioned in the region between the cytochrome and P. Charge-charge interactions with the dimer have been demonstrated when ionizable amino acid residues are placed at several positions, including Asn L170, Asp L155, Cys L247, and Asn M199 (Williams et al. 2001; Johnson and Parson 2002). In photosystem II, electron transfer from the Mn4CaO5 cluster to P680 is critically coupled with proton transfer involving YZ (D1-161) and His D1-190. The corresponding residues in the reaction center can be identified by the homology with photosystem II, and we have shown that Tyr can be oxidized at the corresponding positions in high potential reaction centers when substitutions create Tyr/His pairs at Arg M164, His M193, Arg L235, Tyr L144, and Tyr L164 (reviewed in Allen and Williams 2011). The previous studies provided a basis for the viability and characteristics of the modifications designed to probe proton transfer properties.
In this paper, we explore how changes in residues near P+ affect proton release and measure both light-induced pH changes and the kinetics of P+ reduction to examine the relationship between proton release and electron transfer. We investigate proton release/uptake in the reaction centers combined with optical spectroscopy to characterize the P+ state, using light to trigger the electron and proton transfer steps, which allows the contribution of the dark background to be removed. We report the use of these experimental approaches on a set of twelve reaction centers altered with site-specific mutations and compare these reaction centers to wild-type and to the previously described reaction center with the mutation Leu to His at L131. The overall spectroscopic properties of the reaction centers were characterized, and the long-lived recovery of P+ and the proton release were categorized. The results are discussed in terms of a proton release pathway and compared to proton transfer pathways in other biological complexes.
Methods
Mutagenesis and reaction center preparation
The twelve substitutions are designated as TV(L130) for Thr to Val at L130, RM(L135) for Arg to Met at L135, DN(L155) for Asp to Asn at L155, TF(L162) for Tyr to Phe at L162, SA(L244) for Ser to Ala at L244, CA(L247) for Cys to Ala at L247, RM(M164) for Arg to Met at M164, DN(M184) for Asp to Asn at M184, TV(M186) for Thr to Val at M186, SA(M190) for Ser to Ala at M190, HL(M193) His to Leu at M193, and YF(M210) for Tyr to Phe at M210 (Table S1). Reaction centers with the wild-type amino acid sequence and LH(L131) reaction centers with the mutation Leu to His at L131 have been described previously (Lin et al. 1994). Mutations in the L and M subunit genes were made by standard protocols and expressed in R. sphaeroides (Williams and Taguchi 1995; Thielges et al. 2005). Reaction centers were isolated using the detergent lauryl dimethylamine oxide to solubilize the protein from the cell membrane, purified using ion exchange chromatography, and then concentrated and exchanged into 15 mM Tris–Cl pH 8.0 and 0.05% Triton ×-100.
Optical spectroscopy
Steady-state optical spectra were measured from 700 to 1000 nm using a Varian Cary 6000i spectrophotometer (Agilent Technologies) using a scan rate of 15 nm/s. Light excitation was provided by continuous illumination from an Oriel tungsten lamp with a 860 nm interference filter under subsaturating conditions with a light fiber directed into the sample chamber. The lamp produced an intensity of 0.2 W/cm2 that was directed over a 1 cm2 area of the cuvette. This intensity of the light is approximately 30% of the minimum intensity required for the maximum absorption change at 865 nm for wild-type reaction centers, as previously described (Kálmán et al. 2005).
The kinetics of charge recombination from QA and QB were measured by monitoring the absorption at 865 nm after a 2–3 s illumination. Samples contained 1 µM reaction centers in 15 mM Tris–Cl pH 8.0 and 0.05% Triton ×-100. In some samples, electron transfer from QA to QB was blocked by the addition of terbutryn at a concentration of 100 µM prior to the measurement. The slow kinetics of P+ recovery were measured by monitoring the absorption at 865 nm for 30 min after a 5-min illumination. The average values and standard errors were determined from three independent measurements.
Proton release/uptake measurements
The release of protons from reaction centers in the light was measured as described previously (Kálmán et al. 2005; Deshmukh et al. 2011a, b). Samples with 60 µM reaction centers were prepared in 0.05% Triton ×-100, 100 mM NaCl and 400 µM terbutryn, in the absence of buffer, with the pH poised to 5.9 ± 0.1 by addition of diluted HCl and NaOH. The pH changes were measured with a Mettler micro pH electrode connected to an Orion SA720 pH meter. The yield of proton release was determined by measurement of the light-induced voltage change compared to the steady dark background, with an illumination time of 5 min. The average yields and standard errors were determined from three independent measurements.
Results
Overall spectroscopic properties
Reaction centers with the designed mutations could be purified and exhibited absorption spectra similar to wild-type reaction centers (Fig. S1). For wild-type reaction centers, the visible region has a Soret peak at 366 nm and smaller peaks at 531 nm and 598 nm from the bacteriopheophytins and bacteriochlorophylls, respectively. In the 700–1000 nm region, the spectra have three characteristic peaks, at 756 nm due to the bacteriopheophytins, 802 nm due to the bacteriochlorophyll monomers, and 865 nm due to P. For the reaction centers containing the site-directed mutations, the peaks were in similar positions for both the visible and near infrared regions. The peaks associated with the bacteriochlorophyll and bacteriopheophytin monomers were close to those of wild-type reaction centers, ranging from 799 to 805 nm and 755–756 nm, respectively. Some small differences were seen for the P band, which was positioned at 858–869 nm in the mutants. These shifts arise from small changes in the energy of the excited state of P due to the mutations. Shifts of similar extents have been reported for a range of alterations of amino acid residues in the vicinity of P. For example, such shifts are observed due to changes in electrostatic interactions between P and specific amino acid residues and result in only minor changes in the electron transfer properties (reviewed in Williams and Allen 2009; Jones 2009). The relative absorption values are similar for the mutants compared to wild-type, for example the 802 nm bands are approximately twice as large as the 865 nm bands. The close similarity of these spectra demonstrates that the altered reaction centers contain the cofactors found in wild-type and suggests that the altered amino acid residues have a minimal impact on the organization of the cofactors in the protein.
The ability of the modified reaction centers to perform electron transfer was measured by monitoring the changes to the absorption spectra after a short exposure to light. For wild-type reaction centers in the presence of terbutryn, the light-minus-dark difference spectrum shows spectral changes characteristic of the P+QA– state. In the 700–1000 nm region, illumination results in a decrease centered at 865 nm due to oxidation of P, and electrochromic shifts in the 760 and 800 nm bands due to interactions of the bacteriopheophytins and bacteriochlorophyll monomers with QA–. The experiments made use of non-saturating illumination, and the extent of absorption change reflected a number of competing reactions, including the yield of charge separation and the charge recombination rate. After illumination, the absorption changes decay due to recovery of the P+QA– state to the PQA state. For reaction centers containing the altered amino acid residues, illumination resulted in very similar spectral features (Fig. S2). Small shifts of the P+ band were evident for some mutant reaction centers matching those observed in the absorption spectra of those mutant reaction centers. The extent of the changes in the light differed in the mutant reaction centers, where the decrease at 865 nm ranged from 15 larger to 43% smaller than wild-type. Also evident were differences of the spectra immediately after illumination and after 5 min in the dark. For example, the TV(L130) reaction centers had larger absorption changes while the SA(M190) reaction centers had significantly smaller absorption changes remaining immediately after illumination compared to wild-type.
The rates of charge recombination from the P+QA– and P+QB– states were measured by monitoring the absorption recoveries at 865 nm after a short illumination (Figs. S3, S4, Table S2). In reaction centers, an electron is transferred from P to QA and followed by transfer to QB, producing the P+QB– charge-separated state. Once illumination is ended, the charge-separated state recovers to the ground state in the dark. Transfer between the two quinones can be blocked by the addition of terbutryn, allowing the charge recombination rate from the P+QA– state to be measured. For reaction centers in the presence of terbutryn, the rate of charge recombination from P+QA– was measured to be 9.7 s–1 for wild-type compared to a range of 6.4 to 11.9 s–1 for the mutants. For reaction centers without terbutryn, the observed rate of charge recombination was observed to be biphasic. One component had a rate corresponding to the charge recombination rate from P+QA– and the second component was slower, with rates in the range of 0.8 to 1.6 s–1 for the mutants compared to 1.6 s–1 for wild-type. The P+QA– charge recombination rate is sensitive to the energy difference between the charge-separated and ground states. Previous studies have shown that as the P+/P midpoint potential increases in mutants then the energy difference for charge recombination increases and correspondingly the rate becomes faster according to the Marcus relationship (Williams and Allen 2009). The changes in this rate in some mutant reaction centers may arise from such effects of the mutations on the P+/P redox potential. The changes observed for P+QB– charge recombination reflect the rate changes for the P+QA– charge recombination rates, as P+QB– charge recombination occurs through electron transfer from QB– to QA that is followed by charge recombination from the P+QA– charge-separated state. The variations in these overall properties were generally not correlated with the long-lived recovery of P+ or the proton release (see below and Table S2), in agreement with previous work (Deshmukh et al. 2011a, b).
Long-lived recovery of P+
Unlike the mono-exponential recoveries observed for charge recombination following a laser flash, complex behavior is observed for the absorption recovery of wild-type reaction centers after prolonged illumination (Kálmán and Maróti 1997; Deshmukh et al. 2011a, b). The recovery from the P+QA– state after a 5-min illumination was monitored at 865 nm for wild-type and the mutant reaction centers in the presence of terbutryn. In general, an initial, partial fast recovery was followed by a slow multiphasic recovery (Fig. 2). As previously reported, wild-type reaction centers showed a multi-phasic recovery of the charge-separated state after long illumination while the LH(L131) mutant reaction centers recovered quickly and showed only a negligible fraction of the slow absorption recovery (Deshmukh et al. 2011a, b). The overall recoveries for the set of mutant reaction centers showed a range of different relative contributions from the fast and slow recoveries, with the DN(L155), SA(L244), and HL(M193) mutant reaction centers having a predominant fast recovery with only a minor contribution from a slow recovery, and the TV(L130) and RM(M164) mutant reaction centers showing a long-lived P+ signal with incomplete recovery even on longer timescales (Fig. 2).
Kinetics of the recoveries of the absorbance changes at 865 nm after 5 min of illumination for reaction centers from wild-type (black), DN(M184) (brown), DN(L155) (red), and TV(L130) (blue). Traces are normalized using the absorbance at 865 nm measured at the end of the illumination. Conditions: 1 µM reaction centers in 15 mM Tris–Cl, pH 8.0, 0.05% Triton ×-100, 100 µM terbutryn
Over the large timescale of these absorption measurements, the patterns in three regions could be discerned. An initial fast exponential decay had a time constant of ~ 0.1 s and is assigned to P+QA– charge recombination, as the fitted rates matched the measured recombination rates described above. Another component observed in the 0.03 to 3 min time range could be fit with a time constant of ~ 1 min (Fig. S5). The amplitudes of this component were about 40% of the total with rates ranging from 0.5 to 1.2 min–1, and no correlation was evident with the amplitudes of the proton release (Table S2). After 5 min in the dark, the signal at 865 nm was generally still very slowly recovering. The extent of this long-lived P+ signal was characterized by calculating the average fraction of the signal remaining from 5 to 10 min after illumination. In the set of twelve mutant reaction centers, the absorption change attributed to the long-lived P+ ranged from 5 to 35% of the total change, compared to 22% for the wild-type reaction centers (Table S2).
Proton release
The light-induced release of protons was measured using reaction center samples without any buffer and poised at the pH value of 5.9. The prolonged illumination time of 5 min resulted in a saturation value for the release of protons. The extent of proton release was determined by the voltage change at the end of illumination for three independent samples of each mutant. The voltage change of 5.5 mV for wild-type corresponded to a significant release of six protons per reaction center, in agreement with previous measurements (Kálmán and Maróti 1997; Deshmukh et al. 2011a, b). The voltage changes for the mutant reaction centers were lower than wild-type, ranging from 0.3 to 4.7 mV (Table S2). A voltage change of less than 1 mV in the light was observed for DN(L155), SA(L244), SA(M190), and HL(M193), as well as LH(L131) reaction centers. After illumination, the samples generally recovered to the initial value in the dark, with the TV(L130) and RM(M164) reaction centers recovering more slowly than the others. Although the data did not permit analysis of the time course of the voltage change, the kinetics appeared to be multiphasic as previously published for wild-type and LH(L131) reaction centers (Deshmukh et al. 2011b).
Calculation of the relative proton release for the mutant reaction centers compared to wild-type yielded values ranging from 0.06 to 0.85 (Fig. 3). Similarly, the relative amount of the long-lived P+ signal can be expressed as having a range of 0.2 to 1.6 compared to the wild-type (Fig. 3). The proton release has a distinctive pH dependence with a maximum at pH 5.9, where the measurements were performed (Deshmukh et al. 2011a, b). The measurements of the electron transfer rates and absorption spectra were performed at pH 8.0, as these results show a very mild pH dependence, with no different features at pH 5.9 compared to pH 8.0, allowing comparison to previously reported data (Deshmukh et al. 2011a, b). The relative trends for the set of mutant reaction centers were thus comparable.
Comparison of results after five minutes of illumination for the set of twelve mutant reaction centers relative to wild-type, showing the classification into three groups (red, wheat, and blue). a The proton release for each type of reaction center was measured as a voltage change and normalized to the voltage change observed for wild-type (Table S2). b The extent of the long-lived P+ signal was measured by the change in absorbance at 865 nm remaining after 5 to 10 min in the dark and normalized to the value obtained for the wild-type (Fig. 2 and Table S2). c Structure of the reaction center showing the twelve amino acid residues colored according to their grouping and the bacteriochlorophyll dimer (green). View is down the two-fold symmetry axis of the protein
Discussion
This mutagenesis study examined the relationship between proton release and long-lived kinetics associated with the bacteriochlorophyll dimer P in the bacterial reaction center. The properties of a set of twelve reaction centers, each with a single amino acid substitution near P, were measured and compared to wild-type and the previously described reaction centers with the mutation Leu to His at L131 (Fig. 1). Experiments using optical spectroscopy demonstrated that the general spectral features for the mutant reaction centers were similar to wild-type. The peak positions and relative absorption values of the spectra of the mutant reaction centers were very close to those observed for wild-type, showing that the organization of cofactors remains unchanged. The mutant reaction centers were all capable of performing charge separation as evidenced by the light-minus-dark spectra that exhibited peaks typical of the P+QA– charge-separated state, and the P+QA– and P+QB– charge recombination times were within ~ 20% of the canonical times of 100 ms and 1 s, respectively. After prolonged illumination, the set of mutant reaction centers exhibited a range of recovery kinetics and significant differences in the extent of proton release (Figs. 2, 3). The observation that the residues could be grouped was used to propose their involvement in a proton release pathway as discussed below.
Classification of proton release mutants
The outcomes for the set of mutant reaction centers reveal a correlation between the extent of proton release and the characteristic features of the recovery kinetics. In response to prolonged illumination, the extent of proton release was measured to have relative values of 6 to 85% compared to wild-type. This broad range was not uniformly populated, but rather the mutant reaction centers could be classified into three distinct groupings based upon their proton release values (Fig. 3). The presence of a long-lived P+ signal in the recovery after prolonged illumination showed a large breadth of the extent that was 20 to 160% compared to wild-type, although the distinctions between the groups is less sharp. This connection between the proton release and recovery kinetics can be interpreted according to the concept that both reflect the transfer of protons along a pathway from the bacteriochlorophyll dimer P to the periplasmic surface in response to the oxidation state of P.
One distinct group contains mutations at Asp L155, Ser L244, Ser M190, and His M193. For these four mutant reaction centers, the extent of proton release is minimal, at ~ 10% that of wild-type. This lack of proton release assigns these amino acid residues as being essential for the light-induced proton transfer. Reaction centers with changes at Asp L155, Ser L244, and His M193 also showed the most dominant fast recovery kinetics. The Ser M190 mutation resulted in less change in the recovery kinetics compared to wild-type than the other mutations in this group. Similar characteristics of minimal proton release and fast recovery kinetics were found for reaction centers with the mutation Leu to His at L131.
The middle grouping is formed by reaction centers with changes at Arg L135, Tyr L162, Cys L247, Asp M184, Thr M186, and Tyr M210. None of these reaction centers had properties that were identical to wild-type, but the differences were limited and not at the extreme ends of the range of values for either proton release or slow kinetics observed in the set of mutants. The proton release of these six reaction centers was approximately 40 to 60% of the proton release of wild-type. They showed between 30 and 110% of the extent of the long-lived P+ signal compared to wild-type. While changes at these amino acid residues showed influences on proton release, the effects may be assigned as a general electrostatic response to the presence of P+ rather than resulting from participation as essential members of the proton transfer pathway.
The third group consists of reaction centers with alterations of Thr L130 and Arg M164. These two mutant reaction centers showed proton release that was ~ 80% that of wild-type. Because the proton release when these residues were changed was comparable to wild-type, Thr L130 and Arg M164 are assigned as having minimal roles in proton release during prolonged illumination. However, the recoveries of both the proton release and the long-lived recoveries of P+ of these two mutants were much slower than the other mutant reaction centers. Approximately 30% of the optical signal remained at long times, representing an increase of ~ 50% compared to wild-type. The presence of a significant amount of P+ remaining after long times in the dark shows that changing Thr L130 and Arg M164 has a distinctive impact during the period of the uptake of protons that occurs after illumination is stopped and the protonation states of the residues recover to their ground state values.
Identification of a proton release pathway from the bacteriochlorophyll dimer P
The effects of the mutations can be interpreted in terms of the extent of participation of each amino acid residue in a proton transfer pathway. A network of hydrogen-bonded amino acid residues forming a proton transfer pathway can include both protonatable residues that can act as proton donors and acceptors, such as His and Asp, as well as polar residues, such as Asn and Trp, which are not active participants but can anchor a pathway (Kaur et al. 2021). A key feature of a proton pathway is near complete blockage of proton release due to alteration of any of the residues as they act in concert and not independently. This dramatic impact differentiates participation in a proton pathway from a general pKa shift of an amino acid residue in the vicinity of P due to its light-driven oxidation. The partial proton release observed for the six mutant reaction centers in the middle group is likely due to a change in the pKa shift of the residues, with the extent of proton release related to the pKa shift and the distance to P+. However, the total loss of proton release observed for the four changes in the first group indicates that a single proton release pathway exists.
In contrast to wild-type, reaction centers with mutations at four amino acid residues, Asp L155, Ser L244, Ser M190, and His M193, showed very limited proton release and predominately fast kinetics. As a result, these residues are assigned as being critical participants in the transfer of protons from P+ to the periplasmic surface, suggesting their cooperative participation in a hydrogen-bonded pathway. In addition two residues, Thr L130 and Arg M164, are likely participating in proton uptake as P+ recovers to P in the dark. Three of the critical amino acid residues, Asp L155, Ser M190, and His M193, are located in the helices that lie perpendicular to the membrane on the loops between two transmembrane helices. The two residues whose mutations are associated with uptake of protons in the dark, Thr L130 and Arg M164, are positioned near the end of one of the transmembrane helices. Most of these residues are buried when cytochrome c2 is bound to the region of the helices in this loop, except for Arg M164 and His M193, which have solvent exposure (Axelrod and Okamura 2005).
The residues identified in the mutagenesis screening show the outline of a proton transfer pathway (Fig. 4). These residues are distributed throughout the periplasmic side of the reaction center, spanning a distance of ~ 25 Å, and are located between 5 and 10 Å from the bacteriochlorophyll dimer. Due to the long spanning distance of the pathway, the separations between the identified residues are too large to form the required extended hydrogen-bonding network. The incomplete pathway indicates the need to include amino acid residues that were not investigated by mutagenesis but are positioned at locations where they could contribute to the hydrogen-bond network. An examination of the three-dimensional structure shows the probable involvement of four additional amino acid residues, Trp L156, Ser L158, Asn M195, and Tyr M198. Trp L156 is located at the most deeply buried end of the pathway and is close to the bacteriochlorophyll dimer, while the other three are positioned midway along the pathway. Because water can serve as a participant in hydrogen-bonded networks, the pathway formed by the amino acid residues was supplemented by bound water molecules whose presence in the pathway provides the proper spacing for an extended hydrogen-bonded network (Xu et al. 2004). Along with the bound water molecules, we propose that the chain of nine amino acid residues, Thr L130–Trp L156–Asp L155–Ser L158–Tyr M198–Asn M195–Ser M190–His M193–Arg M164, participates in a hydrogen-bonded linear pathway that transfers protons between the environment of P+ and the periplasmic surface of the protein.
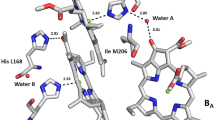
Structural model of the proposed proton pathway. A hydrogen-bond network was identified near the bacteriochlorophyll dimer (green) by the effects of changing three amino acid residues, Asp L155, Ser M190, and His M193 (purple), and including contributions from four additional amino acid residues, Trp L156, Ser L158, Asn M195, and Tyr M198 (cyan) and bound water molecules (red spheres). The pathway is anchored at each end by residues Thr L130 and Arg M164 (blue)
For the proposed proton transfer pathway, the release and uptake of protons are coupled with the formation and subsequent recovery of the P+QA– charge-separated state. Upon illumination, the P+QA– state is rapidly formed, while the release of protons occurs on a much slower timescale. The loss of the protons from the protein surrounding P+ would create a more negative environment that stabilizes the P+QA– state, resulting in the long-lived charge-separated state observed in the spectroscopic measurements after continuous illumination. Furthermore, the recovery back to the ground state requires the uptake of protons to restore the reaction centers back to the charge-neutral environment. Mutant reaction centers with less proton release in the light would have less of the long-lived recovery. The large P+ fractions remaining at long times for reaction centers with mutations at Thr L130 and Arg M164 represent an impaired ability to recover by proton uptake despite their significant proton release. These two residues are spatially separated. The mutation at M164 would disrupt the hydrogen-bond between residue Arg M164 and His M193, which is one of the key amino acid residues required for proton release. His M193 is close to the protein surface and probably serves as the initial site for proton uptake. Alteration of residue Thr L130 may inhibit the other terminus site through interaction with the nearby residue Trp L156 and a bound water molecule.
Nearly 20 bound water molecules are located in this region in high-resolution X-ray structures of the reaction center (Xu et al. 2004). The involvement of bound water molecules in the pathway is consistent with previous measurements of the dependence of the long-lived kinetics on the water content (Malferrari et al. 2011, 2013). After wild-type reaction center samples are prepared with a very low water content, exposure to prolonged illumination results in the recovery kinetics becoming significantly faster and being dominated by the P+QA– charge recombination. Thus, the long-lived kinetics appear to require the presence of bound water, supporting the involvement of water molecules in the pathway for proton release. In addition, a common feature of proton transfer pathways is the involvement of sites that hold protons prior to transfer. An examination of the high-resolution structures of the reaction centers suggests the source of the protons for the pathway as being a large cluster of bound water molecules found near Trp L156 at the beginning of the pathway.
While the residues identified as forming the proton transfer pathway are largely in close hydrogen-bonding contact with each other, residue Ser L244 is not near any of the other residues nor near any of the bound water molecules, despite its inclusion as one of the critical amino acid residues. This residue is located in immediate proximity to P and on the end of a transmembrane helix, and so mutation of L244 may have an indirect influence on the structural conformation of that region or the protonation states of nearby residues. The role of Leu L131, which is located near the beginning of the proposed pathway, in the proton transfer is also not clear. The mutation Leu L131 to His results in the blocking most of the proton release and a fast recovery (Deshmukh et al. 2011a, b), Because Leu is an aliphatic residue, a significant direct role in proton transfer is unlikely, so the mutation should result in an indirect influence on the pathway. The Leu to His at L131 mutation results in the formation of a hydrogen-bond to the keto-carbonyl of one of the bacteriochlorophylls in the dimer (reviewed in Williams and Allen 2009). The addition of this hydrogen-bond may result in structural rearrangements or rigidity changes that affect the pathway (Deshmukh et al. 2011a, b).
Our proposal for a proton release pathway refines previous models of the proton release and multiphasic kinetics in terms of molecular changes (Deshmukh et al. 2011b). Similar results were previously obtained for wild-type and LH(L131) reaction centers, with the multiphasic recovery kinetics including charge recombination and a component with a time of 10 to 60 s thought to be associated with changes associated with the primary quinone. The prolonged illumination was previously interpreted as producing a series of conformational states recovering with different time constants. No specific structural changes were identified for the different states, and no measurable changes near P were evident in studies examining light-induced structural changes using protein crystallography, although the light conditions and cryogenic temperatures would not have produced the long-lived states (Stowell et al. 1997; Katona et al. 2005). The current results suggest that the protonation states of residues in the protein environment of P+ contribute to the slow recovery kinetics. Determination of the relationship between the protonation states and conformational changes would require additional studies. In addition, the previous analysis produced the conjecture that Tyr M210 would have a critical role in proton release, based on its pKa and effect on the P+/P midpoint potential. Tyr M210 is located near P and has a role in tuning the primary electron transfer (Williams and Allen 2009; Jones 2009; Weaver et al. 2021). The protonation state of Tyr M210 could be vital for the function of the reaction center, but it does not appear to be essential for the major proton release pathway described here.
Proton uptake/release in reaction centers
Reaction centers convert light energy into a proton gradient by taking up protons on the inside of the cell through reduction and protonation of a quinone, a process that requires a fast turnover. In the presence of the exogenous donor cytochrome c2, light excitation of P leads to the PQB– state after reduction of P+ by the cytochrome. Since P is in the neutral state, it can be excited again, resulting in the transfer of a second electron, forming the QB–2 state, in a reaction coupled with the uptake of two protons to form a quinol (Okamura et al. 2000; Wraight 2006; Gunner et al. 2013; Maróti 2019; Sugo et al. 2021; Wei et al. 2022). These proton-coupled electron transfer reactions are facilitated by a group of ionizable and other polar residues that participate in a proton transfer pathway to the QB site and control the potential and pKa value of the quinone. Multiple pathways involving polar residues and bound water molecules participate in the transfer of protons over distances of 7 to 20 Å. The uptake of protons to the QB site and the release of protons near P both represent a response to electron transfer by a group of protonatable residues, polar residues, and bound water molecules. However, the release of protons near P reverses in the dark, reflecting that it does not contribute to the proton gradient in the cell. The rapid uptake of protons from the solvent to QB is made easier by the multiple pathways, while the release of protons from the P+ environment is slower. A conformational gating mechanism regulates the second electron transfer and consequent proton uptake for QB, suggesting the possibility that a gated rate-limiting step is responsible for the slow proton release in response to P+.
Consideration of the physiological relevance of the proton pathway requires addressing the overall electron and proton transfer cycle between the reaction center and cytochrome bc1 complex in R. sphaeroides (Lavergne et al. 2009). The quinol produced in the reaction center diffuses through the membrane to the cytochrome bc1 complex, where the stored energy is used to pump protons across the membrane. The resulting proton gradient drives the synthesis of ATP. In the meantime, cytochrome c2 shuttles electrons from the cytochrome bc1 complex to the reaction center. The process takes place in the membrane, which also contains light-harvesting complexes that capture light and direct the excitation energy to the reaction center. Given the numbers of the complexes and rates of the electron transfer and binding reactions, the turnover of the overall process is estimated to occur on the order of tens of ms, and the efficiency is very high under low light conditions (Geyer and Helms 2006; Sener et al. 2016; Singharoy et al. 2019). However, both experimental and modeling studies show that the overall efficiency in the cell decreases under high light conditions (Joliot et al. 2005; Sener et al. 2016; Singharoy et al. 2019). For example, quinol turnover in the cytochrome bc1 complex can become rate-limiting, with the result that the pool of cytochrome c2 becomes largely oxidized, although the cytochrome still binds to the reaction center (Moser and Dutton 1988; Geyer and Helms 2006; Pogorelov et al. 2007). We speculate that under such conditions, when the equilibrium is out of balance, a long-lived P+ is produced in the cell, with the consequent compensating release of protons. Because the cytochrome binding blocks access to the solution of a portion of the cytoplasmic surface of the reaction center, a relatively long proton transfer pathway is required.
Proton transfer in photosynthesis and respiration
Proton transfer plays an essential role in many biological systems, where the controlled transport of protons through the membrane is used to do work, such as the production of ATP by ATPase (Gunner et al. 2013). Bacteriorhodopsin and other proton pumps create a proton gradient across the membrane (Balashov 2000; Lanyi 2006). In this case, light absorption initiates structural changes that result in the removal of a proton from the cell interior, transport along proton pathways throughout the protein, and release on the opposite side. In an alternative photosynthetic route, a proton gradient can be achieved by coupling proton transfer with light-driven electron transfer. These processes require the involvement of proton-coupled electron transfer involving pathways in large membrane protein complexes, including photosystem II. Similar strategies are employed by cytochrome c oxidase in the reduction of molecular oxygen. The mechanisms employed in these well-studied systems can inform interpretation of the proton transfer events associated with oxidation of the bacteriochlorophyll dimer in reaction centers.
Photosystem II is evolutionarily related to bacterial reaction centers and also performs proton-coupled electron transfer (Wydrzynski and Satoh 2005; Vinyard and Brudvig 2017). Both proteins have a central core of cofactors arranged in two branches that facilitate the transfer of electrons from the primary donor to QB and the uptake of two protons after the second electron transfer. A major difference is that photosystem II performs the complex four-electron, four-proton process of water oxidation to dioxygen. Key questions remain open concerning the transfer of water molecules and protons associated with the catalytic oxidation steps at the active site, the CaMn4O5 metal cluster (Pantazis 2018; Ghosh et al. 2019; Hussein et al. 2021; Kaur et al. 2021). Upon light excitation, the oxidized primary donor of photosystem II is reduced by the redox-active tyrosine residue YZ, facilitated by proton transfer with the hydrogen-bonded His D1-190, and followed by electron transfer to YZ from the CaMn4O5 metal cluster. Multiple proton transfer paths from the CaMn4O5 metal cluster to the protein surface have been identified involving a large set of highly interacting residues, bound water molecules, as well as water channels through a distance of approximately 25 Å. Differences in the structures of the central core as well as the presence of additional subunits required for water oxidation make the overall region near the primary donor of photosystem II different than the bacterial reaction center, and so preclude recognition of comparable pathways. However, certain positions appear to retain a proclivity for interaction with the primary electron donor, including Arg M164 and His M193, which were identified in the proton transfer pathway. Mutagenesis of Arg M164 to Tyr in the reaction center produces a Tyr M164–His M193 pair that can participate in proton-coupled electron transfer to reduce highly oxidizing reaction centers suggesting this configuration with the primary donor facilitates proton-coupled electron transfer (Narváez et al. 2002).
Well-characterized proton transfer pathways are found in cytochrome c oxidase, which reduces molecular oxygen to water in a proton-coupled electron transfer reaction, with four protons delivered to the active site while four other protons are pumped through the protein (Wraight 2006; Kaila et al. 2010; Gunner et al. 2013; Cai et al. 2018). The active site contains heme a3, a Cu cofactor, and a redox-active tyrosine that sequentially receives four electrons from exogeneous cytochrome c, followed by binding of molecular oxygen and subsequent breaking of the covalent bond and formation of two water molecules. As the active site is reduced, protons are taken up from solution by two proton transfer pathways identified as the D and K channels. The K channel spans a relatively short distance and requires a Lys residue, while the D channel consists of several residues that are connected by a chain of bound water molecules to transfer multiple protons over a larger distance. The D channel pathway is linear, and mutation of either an Asp or Glu residue at the two ends results in loss of proton delivery to the active site. Dynamical changes have been proposed to regulate the pathways in cytochrome c oxidase, including the proposed fast gating of the D channel. The kindred pattern of a linear pathway of amino acid residues and waters that can be blocked by mutagenesis of specific residues implies that insights that have been gained from the cytochrome c oxidase pathways, such as the role of dynamics, can be applicable to better understand the proton release pathway in reaction centers.
Conclusions
The striking loss of proton release for specific mutations in our survey of residues near the bacteriochlorophyll dimer demonstrated their critical role as participants in a linear proton transfer pathway. The results indicate that, in wild-type reaction centers, a significant release of protons occurs through this pathway under prolonged illumination, accompanied by formation of a persistent charge-separated state. After cessation of light, the uptake of protons is concomitant with a slow recovery of this state. To varying degrees, alteration of each investigated residue had some effect on proton release, showing that transfer along the pathway in wild-type is delicately poised, involving multiple interactions with the protein environment surrounding the pathway. More precise establishment of the roles of the residues forming the proposed pathway will require additional experimental and computational investigations.
References
Agmon N (1995) The Grotthuss mechanism. Chem Phys Lett 244:456–462. https://doi.org/10.1016/0009-2614(95)00905-J
Allen JP, Williams JC (2011) The evolutionary pathway from anoxygenic to oxygenic photosynthesis examined by comparison of the properties of photosystem II and bacterial reaction centers. Photosynth Res 107:59–69. https://doi.org/10.1007/s11120-010-9552-x
Andréasson U, Andréasson LE (2003) Characterization of a semi-stable, charge-separated state in reaction centers from Rhodobacter sphaeroides. Photosynth Res 75:223–233. https://doi.org/10.1023/A:1023944605460
Axelrod HL, Okamura MY (2005) The structure and function of the cytochrome c2: reaction center electron transfer complex from Rhodobacter sphaeroides. Photosynth Res 85:101–114. https://doi.org/10.1007/s11120-005-1368-8
Balashov SP (2000) Protonation reactions and their coupling in bacteriorhodopsin. Biochim Biophys Acta 1460:75–94. https://doi.org/10.1016/S0005-2728(00)00131-6
Barbara PF, Meyer TJ, Ratner MA (1996) Contemporary issues in electron transfer research. J Phys Chem 100:13148–13168. https://doi.org/10.1021/jp9605663
Cai X, Haider K, Lu J, Radic S, Son CY, Cui Q, Gunner MR (2018) Network analysis of a proposed exit pathway for protons to the P-side of cytochrome c oxidase. Biochim Biophys Acta 1859:997–1005. https://doi.org/10.1016/j.bbabio.2018.05.010
Deshmukh SS, Williams JC, Allen JP, Kálmán L (2011a) Light-induced conformational changes in photosynthetic reaction centers: dielectric relaxation in the vicinity of the dimer. Biochemistry 50:340–348. https://doi.org/10.1021/bi101496c
Deshmukh SS, Williams JC, Allen JP, Kálmán L (2011b) Light-induced conformational changes in photosynthetic reaction centers: redox-regulated proton pathway near the dimer. Biochemistry 50:3321–3331. https://doi.org/10.1021/bi200169y
Geyer T, Helms V (2006) Reconstruction of a kinetic model of the chromatophore vesicles from Rhodobacter sphaeroides. Biophys J 91:927–937. https://doi.org/10.1529/biophysj.105.067561
Ghosh I, Khan S, Banerjee G, Dziarski A, Vinyard DJ, Debus RJ, Brudvig GW (2019) Insights into proton-transfer pathways during water oxidation in photosystem II. J Phys Chem B 123:8195–8202. https://doi.org/10.1021/acs.jpcb.9b06244
Gunner MR, Baker NA (2016) Continuum electrostatics approaches to calculating pKas and Ems in proteins. Methods Enzymol 578:1–20. https://doi.org/10.1016/bs.mie.2016.05.052
Gunner MR, Amin M, Zhu X, Lu J (2013) Molecular mechanisms for generating transmembrane proton gradients. Biochim Biophys Acta 1827:892–913. https://doi.org/10.1016/j.bbabio.2013.03.001
Hammes-Schiffer S (2015) Proton-coupled electron transfer: moving together and charging forward. J Am Chem Soc 137:8860–8871. https://doi.org/10.1021/jacs.5b04087
Hammes-Schiffer S, Stuchebrukhov AA (2010) Theory of coupled electron and proton transfer reactions. Chem Rev 110:6939–6960. https://doi.org/10.1021/cr1001436
Hussein R, Ibrahim M, Bhowmick A, Simon PS, Chatterjee R, Lassalle L, Doyle M, Bogacz I, Kim IS, Cheah MH, Gul S, de Lichtenberg C, Chernev P, Pham CC, Young ID, Carbajo S, Fuller FD, Alonso-Mori R, Batyuk A, Sutherlin KD, Brewster AS, Bolotovsky R, Mendez D, Holton JM, Moriarty NW, Adams PD, Bergmann U, Sauter NK, Dobbek H, Messinger J, Zouni A, Kern J, Yachandra VK, Yano J (2021) Structural dynamics in the water and proton channels of photosystem II during the S2 to S3 transition. Nat Commun 12:6531. https://doi.org/10.1038/s41467-021-26781-z
Johnson ET, Parson WW (2002) Electrostatic interactions in an integral membrane protein. Biochemistry 41:6483–6494. https://doi.org/10.1021/bi012131y
Joliot P, Joliot A, Verméglio A (2005) Fast oxidation of the primary electron acceptor under anaerobic conditions requires the organization of the photosynthetic chain of Rhodobacter sphaeroides in supercomplexes. Biochim Biophys Acta 1706:204–214. https://doi.org/10.1016/j.bbabio.2004.11.002
Jones MR (2009) Structural plasticity of reaction centers from purple bacteria. In: Hunter CN, Daldal F, Thurnauer MC, Beatty JT (eds) The purple phototrophic bacteria. Springer, Dordrecht, The Netherlands, pp 295–321. https://doi.org/10.1007/978-1-4020-8815-5_16
Kaila VRI, Verkhovsky MI, Wikström M (2010) Proton-coupled electron transfer in cytochrome oxidase. Chem Rev 110:7062–7081. https://doi.org/10.1021/cr1002003
Kálmán L, Maróti P (1997) Conformation-activated protonation in reaction centers of the photosynthetic bacterium Rhodobacter sphaeroides. Biochemistry 36:15269–15276. https://doi.org/10.1021/bi971882q
Kálmán L, Thielges MC, Williams JC, Allen JP (2005) Proton release due to manganese binding and oxidation in modified bacterial reaction centers. Biochemistry 44:13266–13273. https://doi.org/10.1021/bi051149w
Katona G, Snijder A, Gourdon P, Andréasson U, Hansson O, Andréasson LE, Neutze R (2005) Conformational regulation of charge recombination reactions in a photosynthetic bacterial reaction center. Nat Struct Mol Biol 12:630–631. https://doi.org/10.1038/nsmb948
Kaur D, Khaniya U, Zhang Y, Gunner MR (2021) Protein motifs for proton transfers that build the transmembrane proton gradient. Front Chem 9:660954. https://doi.org/10.3389/fchem.2021.660954
Lanyi JK (2006) Proton transfers in the bacteriorhodopsin photocycle. Biochim Biophys Acta 1757:1012–1018. https://doi.org/10.1016/j.bbabio.2005.11.003
Lavergne J, Verméglio A, Joliot P (2009) Functional coupling between reaction centers and cytochrome bc1 complexes. In: Hunter CN, Daldal F, Thurnauer MC, Beatty JT (eds) The purple phototrophic bacteria. Springer, Dordrecht, The Netherlands, pp 509–536. https://doi.org/10.1007/978-1-4020-8815-5_26
Layfield JP, Hammes-Schiffer S (2014) Hydrogen tunneling in enzymes and biomimetic models. Chem Rev 114:3466–3494. https://doi.org/10.1021/cr400400p
Lin X, Murchison HA, Nagarajan V, Parson WW, Allen JP, Williams JC (1994) Specific alteration of the oxidation potential of the electron donor in reaction centers from Rhodobacter sphaeroides. Proc Natl Acad Sci USA 91:10265–10269. https://doi.org/10.1073/pnas.91.22.10265
Malferrari M, Francia F, Venturoli G (2011) Coupling between electron transfer and protein–solvent dynamics: FTIR and laser-flash spectroscopy studies in photosynthetic reaction center films at different hydration levels. J Phys Chem B 115:14732–14750. https://doi.org/10.1021/jp2057767
Malferrari M, Mezzetti A, Francia F, Venturoli G (2013) Effects of dehydration on light-induced conformational changes in bacterial photosynthetic reaction centers probed by optical and differential FTIR spectroscopy. Biochim Biophys Acta 1827:328–339. https://doi.org/10.1016/j.bbabio.2012.10.009
Manzo AJ, Goushcha AO, Berezetska NM, Kharkyanen VN, Scott GW (2011) Charge recombination time distributions in photosynthetic reaction centers exposed to alternating intervals of photoexcitation and dark relaxation. J Phys Chem B 115:8534–8544. https://doi.org/10.1021/jp1115383
Maróti P (2019) Thermodynamic view of proton activated electron transfer in the reaction center of photosynthetic bacteria. J Phys Chem B 123:5463–5473. https://doi.org/10.1021/acs.jpcb.9b03506
Maróti P, Wraight CA (1988) Flash-induced H+ binding by bacterial photosynthetic reaction centers: influences of the redox states of the acceptor quinones and primary donor. Biochim Biophys Acta 934:329–347. https://doi.org/10.1016/0005-2728(88)90092-8
McPherson PH, Okamura MY, Feher G (1988) Light-induced proton uptake by photosynthetic reaction centers from Rhodobacter sphaeroides R-26. I. Protonation of the one-electron states D+QA–, DQA–, D+QAQB–, and DQAQB–. Biochim Biophys Acta 934:348–368. https://doi.org/10.1016/0005-2728(88)90093-X
Migliore A, Polizzi NF, Therien MJ, Beratan DN (2014) Biochemistry and theory of proton-coupled electron transfer. Chem Rev 114:3381–3465. https://doi.org/10.1021/cr4006654
Moser CC, Dutton PL (1988) Cytochrome c and c2 binding dynamics and electron transfer with photosynthetic reaction center protein and other integral membrane redox proteins. Biochemistry 27:2450–2461. https://doi.org/10.1021/bi00407a031
Narváez AJ, Kálmán L, LoBrutto R, Allen JP, Williams JC (2002) Influence of the protein environment on the properties of a tyrosyl radical in reaction centers from Rhodobacter sphaeroides. Biochemistry 41:15253–15258. https://doi.org/10.1021/bi0264566
Okamura MY, Paddock ML, Graige MS, Feher G (2000) Proton and electron transfer in bacterial reaction centers. Biochim Biophys Acta 1458:148–163. https://doi.org/10.1016/S0005-2728(00)00065-7
Pantazis DA (2018) Missing pieces in the puzzle of biological water oxidation. ACS Catal 8:9477–9507. https://doi.org/10.1021/acscatal.8b01928
Pogorelov TV, Autenrieth F, Roberts E, Luthey-Schulten ZA (2007) Cytochrome c2 exit strategy: dissociation studies and evolutionary implications. J Phys Chem B 111:618–634. https://doi.org/10.1021/jp064973i
Sener M, Strümpfer J, Singharoy A, Hunter CN, Schulten K (2016) Overall energy conversion efficiency of a photosynthetic vesicle. Elife 5:e09541. https://doi.org/10.7554/eLife.09541
Serdenko TV, Barabash YM, Knox PP, Seifullina NK (2016) The kinetic model for slow photoinduced electron transport in the reaction centers of purple bacteria. Nanoscale Res Lett 11:286. https://doi.org/10.1186/s11671-016-1502-x
Singharoy A, Maffeo C, Delgado-Magnero KH, Swainsbury DJK, Sener M, Kleinekathöfer U, Vant JW, Nguyen J, Hitchcock A, Isralewitz B, Teo I, Chandler DE, Stone JE, Phillips JC, Pogorelov TV, Mallus MI, Chipot C, Luthey- Schulten Z, Tieleman P, Hunter CN, Tajkhorshid E, Aksimentiev A, Schulten K (2019) Atoms to phenotypes: molecular design principles of cellular energy metabolism. Cell 179:1098–1111. https://doi.org/10.1016/j.cell.2019.10.021
Stowell MHB, McPhillips TM, Rees DC, Soltis SM, Abresch E, Feher G (1997) Light-induced structural changes in photosynthetic reaction center: implications for mechanism of electron-proton transfer. Science 276:812–816. https://doi.org/10.1126/science.276.5313.812
Sugo Y, Saito K, Ishikita H (2021) Mechanism of the formation of proton transfer pathways in photosynthetic reaction centers. Proc Natl Acad Sci USA 118:e2103203118. https://doi.org/10.1073/pnas.2103203118
Thielges M, Uyeda G, Cámara-Artigas A, Kálmán L, Williams JC, Allen JP (2005) Design of a redox-linked active metal site: manganese bound to bacterial reaction centers at a site resembling that of photosystem II. Biochemistry 44:7389–7394. https://doi.org/10.1021/bi050377n
van Mourik F, Reus M, Holzwarth AR (2001) Long-lived charge-separated states in bacterial reaction centers isolated from Rhodobacter sphaeroides. Biochim Biophys Acta 1504:311–318. https://doi.org/10.1016/S0005-2728(00)00259-0
Vinyard DJ, Brudvig GW (2017) Progress toward a molecular mechanism of water oxidation in photosystem II. Annu Rev Phys Chem 68:101–116. https://doi.org/10.1146/annurev-physchem-052516-044820
Weaver JB, Lin CY, Faries KM, Mathews II, Russi S, Holten D, Kirmaier C, Boxer SG (2021) Photosynthetic reaction center variants made via genetic code expansion show Tyr at M210 tunes the initial electron transfer mechanism. Proc Natl Acad Sci USA 118:e2116439118. https://doi.org/10.1073/pnas.2116439118
Wei RJ, Zhang Y, Mao J, Kaur D, Khaniya U, Gunner MR (2022) Comparison of proton transfer paths to the QA and QB sites of the Rb. sphaeroides photosynthetic reaction centers. Photosynth Res 152:153–165. https://doi.org/10.1007/s11120-022-00906-x
Weinberg DR, Gagliardi CJ, Hull JF, Murphy CF, Kent CA, Westlake BC, Paul A, Ess DH, McCafferty DG, Meyer TJ (2012) Proton-coupled electron transfer. Chem Rev 112:4016–4093. https://doi.org/10.1021/cr200177j
Williams JC, Allen JP (2009) Directed modification of reaction centers from purple bacteria. In: Hunter CN, Daldal F, Thurnauer MC, Beatty JT (eds) The purple phototrophic bacteria. Springer, Dordrecht, pp 337–353. https://doi.org/10.1007/978-1-4020-8815-5_18
Williams JC, Taguchi AKW (1995) Genetic manipulation of purple photosynthetic bacteria. In: Blankenship RE, Madigan MT, Bauer CE (eds) Anoxygenic photosynthetic bacteria. Kluwer, Dordrecht, pp 1029–1065. https://doi.org/10.1007/0-306-47954-0_48
Williams JC, Haffa ALM, McCulley JL, Woodbury NW, Allen JP (2001) Electrostatic interactions between charged amino acid residues and the bacteriochlorophyll dimer in reaction centers from Rhodobacter sphaeroides. Biochemistry 40:15403–15407. https://doi.org/10.1021/bi011574z
Wraight CA (2006) Chance and design—proton transfer in water, channels and bioenergetic proteins. Biochim Biophys Acta 1757:886–912. https://doi.org/10.1016/j.bbabio.2006.06.017
Wydrzynski TJ, Satoh K (2005) Photosystem II: the light-driven water:plastoquinone oxidoreductase. Springer, Cham. https://doi.org/10.1007/1-4020-4254-X
Xu Q, Axelrod HL, Abresch EC, Paddock ML, Okamura MY, Feher G (2004) X-ray structure determination of three mutants of the bacterial photosynthetic reaction centers from Rb. sphaeroides: altered proton transfer pathways. Structure 12:703–715. https://doi.org/10.1016/j.str.2004.03.001
Acknowledgements
This work was supported by the National Science Foundation CHE-1904860 (J.C.W. and J.P.A.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Allen, J.P., Chamberlain, K.D. & Williams, J.C. Identification of amino acid residues in a proton release pathway near the bacteriochlorophyll dimer in reaction centers from Rhodobacter sphaeroides. Photosynth Res 155, 23–34 (2023). https://doi.org/10.1007/s11120-022-00968-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-022-00968-x