Abstract
In this study, we identified two Mn2+ sites in apo-Photosystem II (PSII) using the pulsed electron–electron double resonance (PELDOR). A Mn2+ ion was bound to apo-PSII on the deactivation of the oxygen-evolving complex. The electron–electron magnetic dipole interaction of the Mn2+ to YD· was estimated to be 2.4 MHz. The site was assigned at the position between His332 and Glu189 in the D1 polypeptide, which is close to the Mn1 site in mature PS II. Using recent structures observed under electron microscopes (EM), the location of the Mn2+ site on photoactivation was reevaluated. The position between Asp170 and Glu189 in the D1 polypeptide is a good candidate for the initial high-affinity site for photoactivation. Based on a comparison with the PELDOR results, the two EM structures were evaluated.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
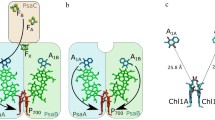
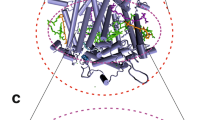
Photosynthesis is an essential function of energy conversion in organism on earth. Oxygen evolution occurs in the Mn cluster of photosystem II (PSII), a protein complex consisting of D1/D2 subunits with pseudo C2 symmetry, surrounded by proteins (Lubitz et al. 2019; Shen 2015; McEvoy and Brudvig 2006). Studies on the reactions have been developed by the highly resolved X-ray crystal structures of PSII (Fig. 1) (Umena et al. 2011; Kern et al. 2018; Suga et al. 2019). Recent electron microscopy (EM) studies provide detailed information on the location of composed proteins to understand chemical reactions (Wei et al. 2016; Huang et al. 2021; Gisriel et al. 2020; Zabret et al. 2021).
Photoactivation is the formation of Mn cluster induced by light reactions, which is one of the main subjects for PSII studies(Bao and Burnap 2016; Burnap 2004; Ono 2001; Dismukes et al. 2005; Becker et al. 2011). Oxygen evolution is deactivated by NH2OH or Tris treatments with a reduction in OEC and removal of extrinsic proteins. For photoactivation, a two-quantum mechanism has been proposed (Tamura and Cheniae 1987; Radmer and Cheniae 1971; Ono and Inoue 1987). In this model, a Mn2+ ion is initially bound to a specific high-affinity site in the apo-PSII, and a second Mn2+ ion is bound to another affinity site after oxidizing the initial Mn2+ (Campbell et al. 2000; Ono and Mino 1999). During the sequential process, a Mn–Mn dimer was formed as an intermediate (Barra et al. 2006). The extrinsic proteins assist D1 assembly for photoactivation (Bondarava et al. 2005; Zabret et al. 2021; Avramov et al. 2020; Huang et al. 2021; Tokano et al. 2020).
We have previously reported the location of Mn2+ in apo-PSII (Asada and Mino 2015), where the Mn cluster was removed by NH2OH-treatment with EDTA after the removal of the extrinsic PsbP/PsbQ proteins (NH2OH/NaCl/EDTA treatment). The magnetic dipole interaction between Mn2+ and YD· was estimated to be 1.7 MHz (Asada and Mino 2015). Based on the mature PSII structure, the Mn2+ affinity site was assigned to Asp170 and Glu333 in the D1 protein (Asada and Mino 2015). Zhang et al. reported the 2.55 Å resolved X-ray crystal structure of Mn depleted PSII, similar to that of the matured PSII (Zhang et al. 2017). One or two Mn ions were detected around the location of Mn1 and Mn2 in the matured PSII (Zhang et al. 2017). However, recent EM studies have shown some conformational fluctuation of D1 and D2 C-terminus, and D1 D–E loop in PSII (Gisriel et al. 2020; Zabret et al. 2021; Huang et al. 2021), where the assembly of the PSII complex was supported by the Psb27 polypeptide. In addition, an AFM study has shown the conformational fluctuation of CP43 E-loop of the extrinsic proteins depleted PSII (Tokano et al. 2020).
It is important to match the active sites on the molecular structure and chemical functional sites to understand the photoactivation process. In the deactivation process, the strong binding of Mn2+ has also been found in apo-PSII (Tamura and Cheniae 1987; Zhang et al. 2017). The Mn2+ binding sites on the deactivation process and the high-affinity Mn2+ site on the photoactivation process should be identified.
Materials and methods
PS II membranes were prepared as described previously (Berthold et al. 1981) with some modification (Asada and Mino 2015). For the study of the leftover Mn2+ site on the deactivation process, the membranes of 0.5 mgChl/ml were incubated for 1 min at 4 °C in a buffer containing 1 mM NH2OH, 400 mM sucrose, 20 mM NaCl, and 20 mM Mes/NaOH (pH 6.5), and centrifuged at 35,000 × g for 20 min. Low concentrations (< 2 mM) of NH2OH treatment maintain PsbP/Q for higher plants (Tamura and Cheniae 1985). To study of the affinity site on the deactivation, the membranes were washed thrice with the same buffer (0.5 mgChl/ml) without EDTA· Na2 (NH2OH treatment). The membranes were centrifuged and inserted into 5 mm diameter EPR tubes. For the study of the initial high-affinity Mn2+ site for photoactivation, the extrinsic (PsbP/Q) proteins were removed by 1 M NaCl treatments and NH2OH treated with EDTA· Na2 (NH2OH/NaCl/EDTA treatment)(Asada and Mino 2015). The membranes were finally washed with a buffer containing 20 μM MnCl2, 400 mM sucrose, 20 mM NaCl, and 20 mM Mes/NaOH (pH 6.5) at 1 mgChl/ml, then centrifuged (Ono and Mino 1999). PELDOR signals are detected only the Mn2+ at high affinity site even in the absent of Ca2+ (Asada and Mino 2015). Tris-treated PSII was prepared as previously described(Ono and Mino 1999), and washed with a buffer containing 20 μM MnCl2, 400 mM sucrose, 20 mM NaCl, and 20 mM Mes/NaOH (pH 6.5) at 1 mgChl/ml, then centrifuged.
CW-EPR measurements were performed using a Bruker ESP-300E EPR spectrometer with a standard cavity (TE102). PELDOR (DEER) was performed using Bruker E580 with an Oxford Instruments liquid helium cryostat (CF935P, Oxford GB) as described previously (Asada and Mino 2015). A π/2-t-π-t-echo sequence was used to measure the ESE field-swept spectra. A π/2-t1-π-t1-t2-π-t2-π-echo sequence was used for the PELDOR, where the different signals in t1 of 200–256 ns at the time interval of 8 ns and t2 of 1400 ns were summed. The pulse length were 24 ns. The signals were obtained at 10 K and the repetition time, 1000 μs.
The PELDOR spectra were simulated as previously described (Asada and Mino 2015). The atomic coordinates of PDB 4UB6 (Suga et al. 2019), PDB 6WJ6 (Gisriel et al. 2020) and PDB 7NHO (Zabret et al. 2021) were used for simulations. The g-factors for two electron spins were assumed to be 2.0046 for YD· and 2.0093 for Mn2+.
To quantify the Mn ions in PS II, HNO3 of 10% volume was added to 2 mgChl/ml of PSII and centrifuged. The supernatant was transferred to calibrated 50 µl capillary tubes. Figure 2A shows the EPR spectra of (a) untreated and (b–f) NH2OH-treated PS II. Figure 2B shows the amount of Mn2+ in HNO3 treated PSII after 1 mM NH2OH-treatment. The amount of 4 Mn2+ was evaluated using the untreated PS II. Three Mn2+ ions were easily removed after 1 mM NH2OH treatment for 1 min. One Mn ion was detected in NH2OH-treated (no EDTA) PSII. The leftover Mn2+ ion is tightly bound to PSII, which is difficult to remove without EDTA. After three washes after NaCl treatment in the presence of 50 mM EDTA, the Mn2+ ions were removed (> 99%).
A EPR spectra of the supernatant of the HNO3 treated PSII. (a) untreated, (b–f) 1 mM NH2OH treated PSII with (b) no EDTA, (c) 1 mM EDTA, (d) 10 mM EDTA, (e) 50 mM EDTA and (f) 50 mM EDTA after NaCl treatment and B The amount of Mn2+ in HNO3 treated PSII after 1 mM NH2OH-treatment. The number of Mn2+ was normalized by the untreated PSII. The membranes were washed for 3 times. Measurement conditions: m.w. freq., 9.78 GHz; m.w. power, 10 mW; field modulation, 10 G; temperature, RT
Results and discussion
The spin quantization shows that one Mn ion was bound to PSII and three Mn ions are easily removed after reduction of manganese cluster (Fig. 2B), which indicates that there is one Mn binding site in the deactivation process. It is notable that the leftover Mn-binding is too strong to remove in the presence of the extrinsic proteins even with EDTA (Fig. 2c, d). We labelled the leftover Mn2+ site in the deactivation process as ‘site1’. Figure 3 shows the ESE-field swept EPR spectrum in the NH2OH-treated PSII, denoted as “site1”. The PELDOR signal was observed at YD· pumped with Mn2+. Figure 4 shows the PELDOR signal of the Mn2+ (site 1) with YD· in the NH2OH-treated PSII. The membranes were washed thrice without EDTA buffer after 1 mM NH2OH treatment (NH2OH treatment). The electron–electron magnetic dipole interaction between Mn2+ and YD· was estimated to be 2.4 MHz. The interaction with the Mn2+ site in the deactivation process (site1) is larger than that with the initial high-affinity site for photoactivation, denoted as site2 (Fig. 4B). During the deactivation process of the Mn cluster in the presence of Mn2+ binding, the PSII structure might be similar to the mature PSII (Zhang et al. 2017). Therefore, we simulated the PELDOR signal using the structure of mature PSII. Figure 4A(b–f) show the simulated PELDOR signals, where the Mn2+ is located at positions (b) Mn1, (c) Mn4, (d) Ca, (e) Mn3, and (f) Mn2 in the mature PSII (PDB 4UB6), respectively. The comparisons showed that the location of Mn2+ is closer to YD· than that of the native Mn1. Using the point dipole approximation, the distance between Mn2+ and the center of the ring in YD· is estimated as 28.1 Å, whereas the position of Mn1 and Mn2 were estimated as 29.3 Å and 32 Å, respectively. Figure 5A shows the possible location of the Mn2+ (gray dot surface) overlapped with the mature PSII structure. The site is located between nitrogen (NE) in His332 and oxygen (OE) in Glu189. The location is slightly different from the X-ray structure, which is 0.6–0.7 Å close to YD from the midpoint of the atoms in His332 and Glu189. The Mn ion at the site is possible to connect with His332 and Glu189 linearly. Therefore, the main difference is ascribed to the loss of the octahedral coordination of the Mn cluster and the formation of axial bidentate binding. When Mn2–4 and O1–5 are removed, the octahedral coordinates of Mn1 would be easily deactivated. On the other hand, such a structural modification would be required the replacements of the ligands, such as water molecules. Therefore, the stability of the coordinates should be evaluated by other methods, such as chemical calculations.
ESE field swept spectra of the 1 mM NH2OH treated PS II washed with (a) no EDTA and (b) 50 mM EDTA for 4 times. Experimental conditions: microwave frequency, 9.49 GHz; pulse length, 16 ns for π/2 pulses and 24 ns for π pulses; interval τ between the mw pulses, 200 ns; repetition time, 1000 μs; temperature, 10 K. The arrows show the resonant magnetic fields for the observation and pumping microwave frequencies in the PELDOR measurement
A PELDOR signals of (a) the experimental NH2OH treated PSII and (b–f) simulated signals. The simulated traces are for (b) Mn1, (c) Mn4, (d) Ca, (e) Mn3 and (f) Mn2 in the crystal structure (PDB 4UB6), assuming that Mn2+ is located among the cofactors. B PELDOR signals of (a) the NH2OH/NaCl/EDTA treated PSII and (b–e) simulated signals. The locations of Mn2+ for traces (b–d) are assumed from the EM structures (PDB 6WJ6). The traces are for (b) position 1: the centers of the four oxygen atoms of Asp170 and Glu189, (c) position 2: the triangle of two oxygen atoms of Asp170 and Glu189 and one nitrogen in H332, (d) position 3: the triangle of three oxygen atoms of Asp170, Glu189, and Tyr161 in the PDB 6WJ6. These locations are indicated in Fig. 6B. The locations of Mn2+ for trace (e) is assumed from the EM structures (PDB 7NHO). The baseline signals for the experimental PELDOR have been subtracted. The simulations were performed as previously described (Asada and Mino 2015). The gray markers are used for the calculations in Figs. 5, 6, and 7 as the peak positions
The possible position for the leftover Mn2+ high-affinity site in the NH2OH treated PSII. Curved grey-surface (dots) represent the coordinates corresponding to the PELDOR simulation. The positions show the first positive peak at t′ = 410–440 ns of the PELDOR signal on A the crystal structure (PDB 4UB6) and B EM structure (PDB 6WJ6)
Recent EM analysis has reported some conformational fluctuation in PSII (Zabret et al. 2021; Huang et al. 2021). Therefore, we simulated the PELDOR spectrum using the EM structures. Figure 5B shows the possible location of Mn2+ in the EM structure (gray dot surface) (PDB 6WJ6). The binding site (site1) is located on the line between nitrogen (NE) in His332 and oxygen (OE) in Glu189. As for distance estimation, Glu333 in the EM structure may be close to the site1 surface, however, it is not a candidate for the leftover Mn2+ (Zhang et al. 2017). These results show that the local structure of the Mn2+ site (site1) fits the conformationally fluctuated PSII rather than the matured PSII. After the removal of other Mn and oxygen ions in the presence of Mn1, His332, and Glu189 shifted to thermally stable locations. In both the X-ray and EM structures, the leftover Mn in the NH2OH treated PSII was ascribed to Mn1 in the mature PSII (Fig. 1).
We have previously assigned the location of the reconstituted Mn2+ site (NH2OH/EDTA/NaCl treatment) between Asp170 and Glu333 in the D1 polypeptide, based on the matured PSII structure (Asada and Mino 2015). However, EM studies have shown that apo-PSII has a conformationally fluctuated structure. Figure 4B shows the previous PELDOR signal between reconstituted Mn2+ and YD· in apo-PSII (Asada and Mino 2015), where the extrinsic proteins PsbP and PsbQ were depleted by NaCl treatments before NH2OH treatment. Gisriel et al. suggested that the location of Mn2+ is between Asp170 and Glu189 based on the conformationally fluctuated structure (Gisriel et al. 2020). Zabret et al. have shown the location of an attaching metal ion in the EM structure (Zabret et al. 2021). Therefore, we reevaluated the previous PELDOR analysis (Asada and Mino 2015) based on these EM structures (Zabret et al. 2021; Gisriel et al. 2020).
First, we compared the PELDOR signal with the EM structure obtained by Gisriel et al. (PDB 6WJ6). Figure 6A illustrates the coordinates and possible surface (gray) for the reconstituted Mn2+ site (site2, the high-affinity site) based on the PELDOR results. As the EM structure has no Mn ion (Gisriel et al. 2020), we tested three positions of the Mn2+ binding in the structure: position 1, the centers of the four oxygen atoms of Asp170 and Glu189; position 2, the triangle of two oxygen atoms of Asp170 and Glu189 and one nitrogen in H332, and position 3, the triangle of three oxygen atoms of Asp170, Glu189 and Tyr161. These positions are shown in Fig. 6B. Traces b–d in Fig. 4B show the simulated signals for positions 1–3 in Fig. 6B, respectively. The results show that the location of Mn2+ in the conformationally fluctuated structure fits well with position 1 between the four oxygen atoms of Asp170 and Glu189.
A the possible position of the Mn2+ high-affinity site for the NH2OH/NaCl treated PSII. Curved grey-surface (dots) represent the coordinates corresponding to the PELDOR simulation. The positions, where PELDOR signals have the first negative peak at t′ = 290–320 ns at first positive peak at t′ = 575–595 ns, were collected on the EM structure (PDB 6WJ6). B Assumed Mn locations on EM structure. The label 1–3 corresponds to traces (b)–(d) in Fig. 4B, respectively
The EM structure by Gisriel et al. does not have a binding metal around the candidate sites, whereas that by Zabret et al. has a binding metal (Zabret et al. 2021), which may be a Mn ion. Figure 7 shows the PELDOR results overlapping on EM structure by Zabret et al. (PDB 7NHO), where Psb27 is released from the complex. Panels A and B show the possible locations for the Mn sites at site1 and site2, respectively. The locations of the sites in the EM structure do not fit the PELDOR results for both Mn sites. For site1, the location given by PELDOR overlaps with the side chain of the amino acids. As for site2, the location of the metal coordinate is far from the PELDOR result. To fit the location of the Mn2+ relative to the YD position for the PELDOR results, the correction of 1.9 Å should be required for the local structure around Mn2+ in the EM structure. The inconsistencies are ascribed to the differences in both the backbone structure and the local structure. We evaluated the difference of the backbone as the distance between the ring centers of YD and YZ. The distance is estimated as 30.1 Å in the EM structure by Gisriel et al. (6WJ6), which is the same as that in X-ray structure (4UB6), but 30.9 Å in the EM structure by Zabret et al. (PDB 7NHO). Zabret et al. have reported that the structural difference with Cβ carbons is averaged 1.3 Å in the EM structure (Zabret et al. 2021).
The possible position of the Mn2+ high-affinity site for A the leftover one in the deactivation process and B the high-affinity site for photoactivation. Curved gray surface (dots) represent the coordinates corresponding to the PELDOR simulation, overlapped on the EM structure (PDB 7NHO). The positions, where PELDOR signals have A the first negative peak at t′ = 290–320 ns and first positive peak at t′ = 575–595 ns or B the first positive peak at t′ = 410–440 ns were collected on the EM structure
Zabret et al. have reported the other two types of EM intermediates, labelled PDB 7NHP and PDB7NHQ, which include the assembly factors Psb27/Psb28/Psb34 and Psb28/Psb34, respectively. We checked the distances between the ring centers of YD and Mn to evaluate the differences of the structures. The distances are 32.2 Å for PDB7NHO, 31.8 Å for PDB7NHP and 32.4 Å for PDB7NHQ, respectively. These structural modifications were so small that the difference by the assembly factors do not influence to the conclusion. On the other hand, it cannot be excluded the possibility that dimerization of PSII derives structural modification, because these EM structures are monomer.
PsbO is an essential polypeptide for oxygen evolution. We have shown the PELDOR results using PsbO-containing PSII (NaCl/NH2OH/EDTA treatment) for the site2. In contract, the EM-PSII structures had no PsbO. Figure 8 shows the difference in the PELDOR signals in NaCl/NH2OH treated and Tris-treated PSII for the site2. The PELDOR signals in both treatments for site2 are the same oscillation frequencies, indicating that PsbO binding does not result in structural modification of the D1/D2 structure.
Three-pulsed PELDOR signals of (a) NaCl/NH2OH-treated and (b) Tris-treated apo-PS II samples in the present of 20 μM MnCl2. Experimental conditions: microwave frequencies, 9.52 GHz for observation and 9.80 GHz for pumping pulses; pulse length, 16 and 24 ns for π/2 and π pulses; magnetic field, 348.6 mT; repetition time, 1000 μs; temperature, 8 K. The different signals in τ of 1400–1480 ns at the time interval of 40 ns were summed
This study showed the locations of two different Mn2+ sites (sites 1 and 2) in deactivated PSII. After the NH2OH treatment of 1 mM without EDTA (NH2OH treatment), three Mn ions were easily removed from the OEC. However, one Mn2+ ion was not easily removed in the presence of extrinsic proteins (Fig. 2). The extremely high affinity of Mn2+ can be ascribed to the binding of His332 and Glu189, where the Mn1 coordinates change from octahedral to bidentate. To deplete the leftover Mn, highly concentrated EDTA is required with the removal of extrinsic proteins. The results show that the deactivation and reassembly for manganese cluster are the different processes. In the deactivation process, the site1 is reserved in the presence of the extrinsic proteins. However, in the reassembly process, the site2 is the highest Mn-binding site in the absence of the extrinsic proteins. In the two quantum mechanism for photoactivation, two Mn-binding sites are assumed (Tamura and Cheniae 1987; Radmer and Cheniae 1971; Ono and Inoue 1987; Avramov et al. 2020). As the relative structure around His332 and Glu189 did not change in the reported structures, the position between His332 and Glu189 could be a good candidate as an affinity site for the photoactivation process. However, the site (site1) is no longer the highest affinity site in the initial state of photoactivation, when the highest affinity Mn2+ site (site2) would be located between Asp170 and Glu189. The very high affinity of the site1 would be related to the binding of the extrinsic proteins. The two-quantum mechanism has proposed the replacement of oxidized Mn2+ with another site (Bao and Burnap 2016; Burnap 2004; Ono 2001; Dismukes et al. 2005; Becker et al. 2011; Zhang et al. 2017), and finally converted to the complete Mn cluster (Sato et al. 2021; Avramov et al. 2020). The PELDOR results show that site1 is a candidate for another Mn site. The location of Mn1 in the matured PSII is a substantial candidate for the second binding site (Sato et al. 2021). The native Mn1 in the matured PSII is a ligand with His332 and Glu189 at a right angle. As site1 is an axial ligand with His332 and Glu189, a large structural rearrangement would be included during the sequential reactions.
Under physiological conditions (in vivo) of cyanobacterial PSII, the degraded D1 polypeptide is replaced by another D1 protein. Moreover the Mn cluster is reassembled with the support of Psb27 (Zabret et al. 2021; Huang et al. 2021). Under experimental conditions (in vitro) without the replacement of the D1 protein, photoactivation would be possible under the occupation of the strongly binding Mn2+ in the second affinity site (site1), which might assist the reassembly of the Mn cluster.
This study shows the location of Mn affinity sites using pulsed EPR by comparing with the crystal and EM structures. Although the photoactivation mechanism is still unclear, advanced EPR is a key technique for tracing the location of Mn ions during the reactions.
References
Asada M, Mino H (2015) Location of the high-affinity Mn2+ site in photosystem II detected by PELDOR. J Phys Chem B 119(32):10139–10144. https://doi.org/10.1021/acs.jpcb.5b03994
Avramov AP, Hwang HJ, Burnap RL (2020) The role of Ca2+ and protein scaffolding in the formation of nature’s water oxidizing complex. Proc Natl Acad Sci USA 117(45):28036–28045. https://doi.org/10.1073/pnas.2011315117
Bao H, Burnap RL (2016) Photoactivation: the light-driven assembly of the water oxidation complex of photosystem II. Front Plant Sci. https://doi.org/10.3389/fpls.2016.00578
Barra M, Haumann M, Loja P, Krivanek R, Grundmeier A, Dau H (2006) Intermediates in assembly by photoactivation after thermally accelerated disassembly of the manganese complex of photosynthetic water oxidation. Biochemistry 45(48):14523–14532. https://doi.org/10.1021/bi061842z
Becker K, Cormann KU, Nowaczyk MM (2011) Assembly of the water-oxidizing complex in photosystem II. J Photochem Photobiol B 104(1–2):204–211. https://doi.org/10.1016/j.jphotobiol.2011.02.005
Berthold DA, Babcock GT, Yocum CF (1981) A highly resolved, oxygen-evolving photosystem-II preparation from spinach thylakoid membranes—electron-paramagnetic-res and electron-transport properties. FEBS Lett 134(2):231–234. https://doi.org/10.1016/0014-5793(81)80608-4
Bondarava N, Beyer P, Krieger-Liszkay A (2005) Function of the 23 kDa extrinsic protein of photosystem II as a manganese binding protein and its role in photoactivation. Biochim Biophys Acta 1708(1):63–70. https://doi.org/10.1016/j.bbabio.2005.01.005
Burnap RL (2004) D1 protein processing and Mn cluster assembly in light of the emerging Photosystem II structure. Phys Chem Chem Phys 6(20):4803–4809. https://doi.org/10.1039/b407094a
Campbell KA, Force DA, Nixon PJ, Dole F, Diner BA, Britt RD (2000) Dual-mode EPR detects the initial intermediate in photoassembly of the photosystem II mn cluster: the influence of amino acid residue 170 of the D1 polypeptide on Mn coordination. J Am Chem Soc 122(15):3754–3761. https://doi.org/10.1021/ja000142t
Dismukes GC, Ananyev GM, Watt R (2005) Photo-assembly of the catalytic manganese cluster. Photosystem II, vol 22. Advances in photosynthesis and respiration. Springer, Berlin, pp 609–626
Gisriel CJ, Zhou KF, Huang HL, Debus RJ, Xiong Y, Brudvig GW (2020) Cryo-EM structure of monomeric photosystem II from Synechocystis sp. PCC 6803 lacking the water–oxidation complex. Joule 4(10):2131–2148. https://doi.org/10.1016/j.joule.2020.07.016
Huang G, Xiao Y, Pi X, Zhao L, Zhu Q, Wang W, Kuang T, Han G, Sui S-F, Shen J-R (2021) Structural insights into a dimeric Psb27-photosystem II complex from a cyanobacterium Thermosynechococcus vulcanus. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.2018053118
Kern J, Chatterjee R, Young ID, Fuller FD, Lassalle L, Ibrahim M, Gul S, Fransson T, Brewster AS, Alonso-Mori R, Hussein R, Zhang M, Douthit L, de Lichtenberg C, Cheah MH, Shevela D, Wersig J, Seuffert I, Sokaras D, Pastor E, Weninger C, Kroll T, Sierra RG, Aller P, Butryn A, Orville AM, Liang MN, Batyuk A, Koglin JE, Carbajo S, Boutet S, Moriarty NW, Holton JM, Dobbek H, Adams PD, Bergmann U, Sauter NK, Zouni A, Messinger J, Yano J, Yachandra VK (2018) Structures of the intermediates of Kok’s photosynthetic water oxidation clock. Nature 563(7731):421–425. https://doi.org/10.1038/s41586-018-0681-2
Lubitz W, Chrysina M, Cox N (2019) Water oxidation in photosystem II. Photosynth Res 142:105–125. https://doi.org/10.1007/s11120-019-00648-3
McEvoy JP, Brudvig GW (2006) Water-splitting chemistry of photosystem II. Chem Rev 106(11):4455–4483. https://doi.org/10.1021/cr0204294
Ono T (2001) Metallo-radical hypothesis for photoassembly of (Mn)4-cluster of photosynthetic oxygen evolving complex. Biochim Biophys Acta 1503(1–2):40–51. https://doi.org/10.1016/s0005-2728(00)00226-7
Ono TA, Inoue Y (1987) Reductant-sensitive intermediates involved in multi-quantum process of photoactivation of latent O2-evolving system. Plant Cell Physiol 28(7):1293–1299
Ono T, Mino H (1999) Unique binding site for Mn2+ ion responsible for reducing an oxidized YZ tyrosine in manganese-depleted photosystem II membranes. Biochemistry 38(27):8778–8785. https://doi.org/10.1021/bi982949s
Radmer R, Cheniae GM (1971) Photoactivation of manganese catalyst of O2 evolution. II. A two-quantum mechanism. Biochim Biophys Acta 253(1):182–186. https://doi.org/10.1016/0005-2728(71)90243-x
Sato A, Nakano Y, Nakamura S, Noguchi T (2021) Rapid-scan time-resolved ATR-FTIR study on the photoassembly of the water-oxidizing Mn4CaO5 cluster in photosystem II. J Phys Chem B 125(16):4031–4045. https://doi.org/10.1021/acs.jpcb.1c01624
Shen JR (2015) The structure of photosystem II and the mechanism of water oxidation in photosynthesis. Annu Rev Plant Biol 66:23–48. https://doi.org/10.1146/annurev-arplant-050312-120129
Suga M, Akita F, Yamashita K, Nakajima Y, Ueno G, Li H, Yamane T, Hirata K, Umena Y, Yonekura S, Yu L-J, Murakami H, Nomura T, Kimura T, Kubo M, Baba S, Kumasaka T, Tono K, Yabashi M, Isobe H, Yamaguchi K, Yamamoto M, Ago H, Shen J-R (2019) An oxyl/oxo mechanism for oxygen-oxygen coupling in PSII revealed by an x-ray free-electron laser. Science 366(6463):334–338. https://doi.org/10.1126/science.aax6998
Tamura N, Cheniae G (1985) Effects of photosystem II extrinsic proteins on microstructure of the oxygen-evolving complex and its reactivity to water analogs. Biochim Biophys Acta 809(2):245–259. https://doi.org/10.1016/0005-2728(85)90068-4
Tamura N, Cheniae G (1987) Photoactivation of the water-oxidizing complex in photosystem II membranes depleted of Mn and extrinsic proteins. I. Biochemical and kinetic characterization. Biochim Biophys Acta 890(2):179–194. https://doi.org/10.1016/0005-2728(87)90019-3
Tokano T, Kato Y, Sugiyama S, Uchihashi T, Noguchi T (2020) Structural dynamics of a protein domain relevant to the water-oxidizing complex in photosystem II as visualized by high-speed atomic force microscopy. J Phys Chem B 124(28):5847–5857. https://doi.org/10.1021/acs.jpcb.0c03892
Umena Y, Kawakami K, Shen JR, Kamiya N (2011) Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature 473(7345):55–60. https://doi.org/10.1038/nature09913
Wei XP, Su XD, Cao P, Liu XY, Chang WR, Li M, Zhang XZ, Liu ZF (2016) Structure of spinach photosystem II-LHCII supercomplex at 3.2 Å resolution. Nature 534(7605):69–74. https://doi.org/10.1038/nature18020
Zabret J, Bohn S, Schuller SK, Arnolds O, Moller M, Meier-Credo J, Liauw P, Chan A, Tajkhorshid E, Langer JD, Stoll R, Krieger-Liszkay A, Engel BD, Rudack T, Schuller JM, Nowaczyk MM (2021) Structural insights into photosystem II assembly. Nat Plants 7:524–538. https://doi.org/10.1038/s41477-021-00895-0
Zhang M, Bommer M, Chatterjee R, Hussein R, Yano J, Dau H, Kern J, Dobbek H, Zouni A (2017) Structural insights into the light-driven auto-assembly process of the water-oxidizing Mn4CaO5-cluster in photosystem II. Elife 6:e26933. https://doi.org/10.7554/eLife.26933
Funding
This work was partly supported by a Nanotechnology Platform Program < Molecule and Material Synthesis > (JPMXP0S21MS1007) of the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan (to H.M), and JSPS KAKENHI Grant Number JP20H05096 (to H.M.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mino, H., Asada, M. Location of two Mn2+ affinity sites in photosystem II detected by pulsed electron–electron double resonance. Photosynth Res 152, 289–295 (2022). https://doi.org/10.1007/s11120-021-00885-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-021-00885-5