Abstract
Early and accurate diagnosis is a critical first step in mitigating losses caused by plant diseases. An incorrect diagnosis can lead to improper management decisions, such as selection of the wrong chemical application that could potentially result in further reduced crop health and yield. In tomato, initial disease symptoms may be similar even if caused by different pathogens, for example early lesions of target spot (TS) caused by the fungus Corynespora cassicola and bacterial spot (BS) caused by Xanthomonas perforans. In this study, hyperspectral imaging (380–1020 nm) was utilized in laboratory and field (collected by an unmanned aerial vehicle; UAV) settings to detect both diseases. Tomato leaves were classified into four categories: healthy, asymptomatic, early and late disease development stages. Thirty-five spectral vegetation indices (VIs) were calculated to select an optimum set of indices for disease detection and identification. Two classification methods were utilized: (i) multilayer perceptron neural network (MLP), and (ii) stepwise discriminant analysis (STDA). Best wavebands selection was considered in blue (408–420 nm), red (630–650 nm) and red edge (730–750 nm). The most significant VIs that could distinguish between healthy leaves and diseased leaves were the photochemical reflectance index (PRI) for both diseases, the normalized difference vegetation index (NDVI850) for BS in all stages, and the triangular vegetation index (TVI), NDVI850 and chlorophyll index green (Chl green) for TS asymptomatic, TS early and TS late disease stage respectively. The MLP classification method had an accuracy of 99%, for both BS and TS, under field (UAV-based) and laboratory conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fresh-market tomato plants face challenges from numerous diseases in Florida production. Foliar diseases on fresh-market tomato, specifically TS and BS, are intensively managed in Florida through cultural and chemical means (Sharma and Bhattarai 2019; Onofre et al. 2019). Target spot (TS) is a serious disease problem on tomato probably due to its ability to survive over season between crops on plant debris and weed hosts, but also by becoming increasingly resistant to registered fungicides (Shazia et al. 2018; MacKenzie et al. 2018). Bacterial spot (BS) disease on tomato is a chronic problem in Florida’s warm and humid environment, and reduces plant vigor and marketable yield (Liao et al. 2019). Target spot on tomato can decrease yield by 33% (Pernezny et al. 1996) causing an estimated $3.5 million loss to Florida tomato growers (MacKenzie et al. 2018). Both diseases affect leaves, stems and fruit, thereby adversely affecting tomato yields. Growers are reliant on the timely application of fungicides and bactericides for management of these diseases. One of the big issues that growers face is the similarity of early disease symptoms of TS and BS as expressed by tomato. Particularly when lesions are small, additional expertise is needed to diagnose and identify each pathogen (Pernezny and Raid 2001).
Bacterial leaf spot is caused by four species of Xanthomonas (Potnis et al. 2015) with X. perforans being the most common species occurring in Florida tomato. Leaves infected with BS disease will initially show small, dark, irregular water-soaked lesions and eventually the leaves may turn yellowish in color and the plant can be defoliated. Leaves, stems and fruit are susceptible. Fruits are especially susceptible in the immature stage of development. In the late stage, the lesions expand and increase from approximately 3.2 to 6.3 mm in diameter (Zitter 1985).
Target spot disease is caused by the fungus Corynespora cassiicola. This disease is widespread throughout North, South and Central America, Africa, the Caribbean, Asia and Europe (Singh and Allen 1979). It is considered a major foliar disease in the tomato production industry (Pernezny et al. 2002). The fungus attacks the older leaves at the plant base, then moves up to the young leaves rapidly. The fungal disease can eventually defoliate the whole plant. The first symptoms on leaves appear as tiny lesions in an irregular shape (> 1 mm) and may have a yellow margin. In the late disease stage, the lesions increase up to 10 mm in size and have ring-shaped outlines. The spots spread throughout the plants causing the leaves to turn yellow and necrotic. Small lesions can occur on the fruit as well.
Rapid and accurate identification of a plant pathogen is essential for implementing timely disease management tactics (Ampatzidis et al. 2017; Cruz et al. 2017). Scouting of plants for disease detection is an essential component of Integrated Pest Management. Visual scouting is labor-intensive, expensive, requires a level of expertise in pest identification, and the observations may be subjective in terms of disease identification (Cruz et al. 2019). Accurate disease identification at the beginning of an outbreak is essential for the selection of effective active ingredients in spray treatments and to reduce the dosage required with significant environmental and economic benefits. Tomato diseases are currently detected in the field by visual scouting of plants and fruits. However, diagnosis based on visual symptoms is difficult due to the inability to differentiate among similar foliar symptoms of diseases. Initially, symptoms for these diseases may appear alike requiring additional confirmatory tests in a lab that could delay the diagnosis by several days. A misidentification could result in the application of non-effective chemical treatment. When favorable environmental conditions for disease development exist, preventative fungicide applications are not sufficient for disease management and control alone. Thus, timing of fungicides and/or bactericides is critical.
An automated system to detect and identify disease symptoms that assesses the entire field would increase the actual area scouted, and thus increase the likelihood of identifying disease hot spots, reduce the man-hours needed for scouting and increase the precision of disease identification (Luvisi et al. 2016). An automated system can generate the spatial distribution of a disease outbreak for targeting precise management tactics. The tomato industry would benefit from an automated method for early detection to assist growers in making timely management decisions and to limit disease spread. The rapid development of new technologies provides a unique opportunity for developing intelligent agricultural systems for precision applications (Ampatzidis et al. 2016, 2018) and food traceability (Ampatzidis et al. 2012; Ampatzidis and Vougioukas 2009; Galvez et al. 2018). Technological advances in computer vision, mechatronics, artificial intelligence and machine learning have enabled the development and implementation of remote sensing technologies for plant symptom identification and management. These technologies can be used to distinguish between a variety of similar insects (Partel et al. 2019a), disease symptoms (Cruz et al. 2019; Hariharan et al. 2019), and weeds (Partel et al. 2019b).
Several techniques based on unmanned aerial vehicles (UAV) have been developed for plant phenotyping and disease detection. Ampatzidis and Partel (2019) developed a UAV-based high throughput phenotyping system for citrus crops utilizing machine learning and artificial intelligence. This automated system can accurately detect and count citrus trees, measure their canopy size, count tree gaps and generate individual tree health status maps using the normalized difference vegetation index (NDVI). Ampatzidis et al. (2019) utilized this UAV-based system to evaluate citrus rootstock varieties. Smigaj et al. (2015) utilized low-cost fixed-wing UAVs equipped with two types of camera, hyperspectral in the near infrared (NIR) range and thermal, to monitor vegetation coverage in a forest. The data were collected and analyzed to monitor the spread of non-native invasive species within a stand of Scots pine trees. Calderon et al. (2013) monitored the progress of Verticillium wilt in olive trees using different types of camera mounted on a UAV (e.g., thermal, hyperspectral and multispectral cameras). They calculated chlorophyll concentration (a + b) and carotenoid blue/green/red (B/G/R) indices to monitor water stress because the flow of water is blocked by the Verticillium wilt pathogen which colonizes the vascular system that ultimately leads to water stress. Zarco-Tejada et al. (2012) utilized UAVs and hyperspectral and thermal imaging to monitor plant water stress remotely. Vivaldini et al. (2019) confirmed that the UAV-based images provided low cost data acquisition and fast scanning of large areas planted with eucalyptus, and could lead to methods to prevent outbreak of diseases to larger areas by controlling the disease expansion. Huang et al. (2019) evaluated wheat fields affected by Helminthosporium leaf blotch, in four disease development stages, to determine the infected area and to develop new target-based spraying techniques. Abdulridha et al. (2018, 2019) developed an aerial remote sensing technique for the early detection of laurel wilt in avocado plants utilizing hyperspectral and multispectral imaging and neural networks for disease identification. They evaluated several vegetation indices (VIs) for early detection of laurel wilt and achieved an overall detection accuracy of more than 95%. Dash et al. (2017) monitored forest health, by using UAVs, with higher temporal and spatial resolution than satellite images. Di Gennaro et al. (2016) developed a UAV-based technique utilizing multispectral imaging to distinguish grapevine leaf stripe disease in asymptomatic stages with high accuracy. Several other studies developed UAV-based techniques for plant health monitoring (Thomas et al. 2018; Albetis et al. 2017; Mahlein 2016; Salami et al. 2014).
Lu et al. (2018) used a spectroradiometric device for tomato disease detection in laboratory conditions and they recommended the need of developing a remote sensing technique to detect diseases in the field. There is limited study on disease detection in tomato in field conditions by using UAV-based hyperspectral imaging; the current study attempted to cover this gap. To achieve this goal, a remote sensing system was developed to early detect and distinguish two major Florida tomato diseases, BS and TS. The objectives of this study were to: (i) utilize hyperspectral imaging to develop disease detection techniques to distinguish BS and TS symptoms on plants at different stages of disease development under indoor (laboratory) and outdoor (UAV-based) conditions; and (ii) select the optimal spectral bands and VIs for distinguishing between BS and TS symptoms on tomato in both indoor and outdoor (UAV-based) conditions.
Materials and methods
Experimental plot design
The field experiment (26° 27′ 36.9″ N and 81° 26′ 24.2″ W) was conducted at the University of Florida’s Southwest Florida Research and Education Center (SWFREC), Immokalee, USA. Six-week-old tomato seedlings ‘FL47’ were transplanted on 5 Sep 2018 into Immokalee fine sand. Guidelines, established by the University of Florida/IFAS, were followed for land preparation, fertility, irrigation, weed management and insect control. Beds were 0.81 m wide with 1.83 m centers covered with white polyethylene film. Each plot consisted of ten plants spaced 0.46 m apart within a 4.57 m row with 3.05 m between each plot (Figs. 1d and 2d). The experiment was divided into two blocks and plots within each block received only one treatment of inoculation with either TS or BS and untreated.
Tomato leaves inoculated with the fungus, C. cassicola, causing TS disease in three developmental stages: a asymptomatic stage post-inoculation; b early stage; and c late stage (this image presents an example of spectral data collection on six RoI); and d Tomato plants infected with TS in the SWFREC field
Tomato plants inoculation
Bacterial spot disease
Plants were inoculated with X. perforans races 3 and 4 on 17 Oct 2018. Bacteria were grown in 25 ml of Difco Nutrient Broth (NB) overnight on a shaker incubator and transferred to 500 ml NB and incubated as before for 24 h. The bacterial suspension was adjusted to 106 CFU ml−1 and applied to tomato plants to run-off using a hand pump sprayer. Lesions of BS disease were confirmed by re-isolation of the bacteria onto Difco Nutrient Agar to verify presence of bacteria.
Target spot disease
Plants in plots were inoculated with C. cassiicola on 15 Oct 2018. Cultures of CC #19 and CC# 20 (kindly provided by Gary Vallad) were grown for 14 days on ¼ Potato dextrose agar + rifampicin and ampicillin (MilliporeSigma, St. Louis, MO 63103, USA). Plates were flooded with sterile water and fungi were scraped from surface. The suspension containing mycelium and spores was filtered through three layers of cheesecloth and adjusted to approximately 104 spores’ ml−1 in sterile water. The inoculum was applied with a hand pump sprayer and plants were sprayed to runoff. Confirmation of TS lesions was through microscopic examination of the lesions for presence of spores.
Tomato leaf sampling
The leaves were collected over time during Oct (18, 23) and Nov (1, 6, 9, 15) 2018 and analyzed by an indoor hyperspectral sensing system (Fig. 3a). The UAV data were collected during Oct/Nov 2018, by a UAV-based hyperspectral sensing system (Fig. 3b). The leaves were collected by plant pathologists to avoid any confusion in sampling collection. Leaves were collected in several disease development stages; for example, in the asymptomatic stage (leaves without any sign of lesions), early disease development stage (leaves with small lesions appearing slightly), and in the late stage (leaves with clear lesions). Figures 1 and 2 show the TS and BS symptoms on tomato leaves. The TS late stage showed the leaves twisted and chlorotic from coalescing lesions containing concentric, target-like rings (Schlub et al. 2007). The symptoms of BS in the late stage showed the spots start as dark green, and then gradually becoming purple and gray with black centers. The BS disease may result in starting tinny and cracked leaf tissue. Healthy leaves, without any symptoms, were obtained and inspected visually to confirm absence of lesions.
Hyperspectral data collection
Indoor hyperspectral data collection
All the collected samples were scanned using a benchtop hyperspectral imaging system (Fig. 3), Pika L 2.4 (Resonon Inc., Bozeman MT, USA) equipped with a 23 mm lens which has a spectral range of 380–1020 nm, 281 spectral channels, 15.3º field of view and a spectral resolution of 2.1 nm. Resonon hyperspectral imagers (RHI) are line-scan imagers (also referred to as push-broom imagers). The system also consists of a linear stage assembly which is moved by a stage motor. There are regulated lights placed above the linear stage to create optimal conditions for performing the scans. The hyperspectral imaging system was placed such that the distance from the lens to the linear stage was 0.5 m. The lights were also at the same level as the lens on a parallel plane. All the scans were performed using the Spectronon Pro (Resonon Inc., Bozeman MT, USA) software, which was connected to the camera system using a USB cable. Before performing the scans of the leaves, dark current noise was removed using the software and then the camera was calibrated by using a white tile (reflectance reference), provided by the manufacturer of the camera system, placed in the same conditions as in where the scans were to be performed. The regions of interest (RoIs) were chosen manually by randomly selecting six spectral scans from each leaf to avoid any bias. The number of pixels for each RoI was about 800–900 included representative leaf tissue (symptomatic and asymptomatic) in order to collect data from the whole leaf (Fig. 2c). The detached leaves were kept in container (cooler) and delivered to the lab within 15 min. Leaves were scanned immediately after receiving from the field. Each scan consisted of 4–6 leaves based upon their size such that they fit in the total scan size. After collecting each scan, the spectral data of the leaves were collected using a post-processing data analysis software (Spectronon Pro, Resonon Inc., Bozeman MT, USA). Several areas containing the symptoms on the leaves were selected using the selection tool and the spectrum was generated. In case of healthy and asymptomatic stages, several random spots on leaves were selected. Once the spectra were generated, the reflectance data were exported from the software in the form of excel sheets using the export option.
Outdoor hyperspectral data collection
Hyperspectral data were collected using an unmanned aerial vehicle (UAV) (Matrice 600 Pro Hexacopter, DJI, Shenzen, China) and the same imaging system, Resonon Pika L 2.4 (Fig. 3b); the same hyperspectral camera was used indoors and outdoors after replacing lenses covering the same spectral range. The UAV-based imaging system included: (i) Pika L 2.4 hyperspectral camera (Resonon, Bozeman, MT, USA); (ii) visible-near infrared (V-NIR) objective lenses for the Pika L camera with a focal length of 17 mm, field of view (FOV) of 17.6 degrees and instantaneous field of view (IFOV) of 0.71 mrad, (iii) a Global Navigation Satellite System (GNSS) (Tallysman 33-2710NM-00-3000, Tallysman Wireless Inc., Ontario, Canada)/Inertial Measurement Unit (IMU) (Ellipse N, SGB Systems S.A.S., France) flight control system for multi-rotor aircraft to record sensor position and orientation; (iv) a hyperspectral data analysis software (Spectronon Pro, Resonon, Bozeman, MT, USA) with an ability to correct the GNSS/IMU data using a geo-rectification plugin. Data were collected at 30 m above the ground with a speed of 1.5 m s−1. RoIs were selected randomly, 3–4 pixel for each region, to avoid background interactions. Six RoIs for each plant were chosen; forty tomato plants were chosen in this study (Fig. 4).
After the hyperspectral data were acquired, the maps and images were analyzed by using the Spectronon Pro software. The spatial resolution of the hyperspectral data collected was 0.1 m. The RoIs were selected manually (and randomly) for each plant, and multispectral scans were performed to ensure that the entire canopy was covered spectrally (Fig. 4). The RoIs were then exported as a text file and processed using the SPSS software (SPSS 13.0, Inc., Chicago; Microsoft Corp., Redmond, WA, USA).
Vegetation indices
The similarity of initial symptoms of TS and BS makes it difficult to distinguish between the lesions of these two diseases visually, therefore 35 VIs were calculated in this study to select the appropriate VI for disease detection at each disease development stage (Table 1). To obtain better results and to compare the classification accuracies of all VIs, each disease development stage (asymptomatic stage, early stage and late or advanced development stage) was analyzed individually. Some vegetation indices are associated with chlorophyll absorbances such as the chlorophyll index green (Chl green) (Vincini et al. 2007) and the anthocyanin reflectance index (ARI) (Gitelson et al. 2001), and could be potentially used for detecting abnormalities and diseases. The most common VI utilized in remote sensing is the normalized difference vegetation index (NDVI); when the value of this index is significantly low, it usually indicates vegetation stress caused by diseases, nutrient deficiencies or other factors. In general, plant diseases can cause changes in chlorophyll, reduce plant water content and create water stress, as well as affect canopy structure and pigment concentration. The 35 VIs, selected in this study, target to detect changes in plant’s chlorophyll, water content and pigment concentration (among other factors).
Data analysis
Three parameters were utilized to explore in which wavelength, significant differences occurred in order to distinguish diseases and disease development stages: (i) spectral difference value which was calculated by subtracting the mean reflectance value of BS and TS infected leaves from the mean reflectance value of healthy leaves at each wavelength; (ii) sensitivity value, which was calculated as the mean reflectance value of diseased leaves divided by the mean reflectance value of healthy leaves (at each wavelength); (iii) linear correlation coefficient (r), which visualizes the intensity of each spectrum band.
SPSS software (SPSS Statistics ver.13.0, IBM Corporation, Aramonk, NY, USA) was utilized in this study for hyperspectral analysis. The classifications were between putative healthy leaves, BS and TS in different diseases stages (asymptomatic, early and late disease stage) by using the spectral reflectance measurements for each stage. The analyses were performed independently for the asymptomatic, early and late stage spectral measurements, as well as in a data set composed of all the reflectance data in order to select the best bands for each development stage.
Classification methods
Two classification methods were used in this study: Multilayer Perceptron (MLP), and Stepwise Discriminate analysis (STDA).
Multilayer perceptron (MLP)
The most popular classification method in artificial neural networks (ANN) architecture is the MLP, which is extensively used for explaining issues related to data classification. The MLP networks are qualified using back propagation learning algorithms (Foody 2004). MLP classification is a very common method because it provides a precise classification prediction. The perceptron computes a single output from multiple real-valued inputs by forming a linear combination according to its input weights, and then putting the output through some non-linear activation function (Das and Sengur 2010). MLP is a controlled network, so it needs a desired response to be qualified. In this study, input data were determined as training 70%, testing 20% and holdout 10%; so, the training of data set taken approximately from 2/3 of the whole data and the residual has been taken as test and holdout sets. The activation functions under investigation were TS and BS in the laboratory and field conditions. The input data included spectral data of healthy versus asymptomatic, early and late development stages for both diseases. Several studies have utilized the MLP method to detect and classify diseases in different crops (Tamouridou et al. 2018; Lowe et al. 2017; Barbedo 2013).
Stepwise discriminate analysis (STDA)
In this study, healthy, TS and BS with different categories of disease development stages (asymptomatic, early and late stage) were used as input data. STDA was chosen to build a model to best predict in which group a dataset (spectral data) belong. In order to improve the analysis, a backwards process was used to eliminate the variables with low contribution to the prediction of group membership. Therefore, the effective discriminant analysis would keep only the variables that contribute the highest to the discrimination between clusters (Reynolds et al. 1988). The F-value is utilized as indicator for entering and removing value from the prediction model. If the F value for a variable is statistically significant in the discrimination group, that means that the variable contributes to the estimation of group participation (Huberty 1984). In this study, the training dataset was 70%, and the testing data set was 30%. Pydipati et al. (2006) and Burks et al. (2000) used STDA in agricultural applications, especially in plant disease detection.
Results and discussion
Spectral reflectance analysis of the BS disease in laboratory conditions
The spectral signature of the healthy plants showed high absorption of photosynthetic pigment in the visible range, which caused the peak region around 550 nm. It can also be noticed that the spectral reflectance values were high in the near-infrared range (NIR). These patterns were consistently maintained in spectral signatures of all healthy plants therefore, the spectral signature of healthy plants was used as a basis to compare with the spectral signatures of the diseased plants. Figure 5a shows the average of spectral signature of healthy and BS affected plants in three different stages (asymptomatic, early and late development stages). In the visible range (400–700) nm, the spectral signatures of healthy and asymptomatic plants have a similar pattern. The peak reflectance values of the spectral signatures in the visible range were 13.6% for healthy and 13.35% for early and asymptomatic stages. In BS late stage, the spectral reflectance was higher, when compared to the other stages, in the green band (495–570 nm) and in the red band (550–690 nm). In NIR range (700–900 nm), the spectral reflectance of late stage displayed lower value and shifted down, while the curve of asymptomatic stage showed higher reflectance than the other stages (Fig. 5a).
The differences in the sensitivity values for different stages of BS can be clearly noticed in the visible range (447–693 nm). The maximum sensitivity value was at 693 nm in the BS late stage; the minimum value was recorded in the asymptomatic stage. In NIR range (700–900 nm), the sensitivity values were very close to each other (Fig. 5b).
Differences between healthy and BS diseased plants were noticed in early, asymptomatic and late stages in the NIR band. The maximum spectral difference value was at around 1000 nm for all stages. In the visible range, a negative value was recorded in the late stage, between 501 and 718 nm (Fig. 5c). Hence, the spectral difference values (like the sensitivity values) can be used to distinguish the three disease development stages.
The correlation coefficient for asymptomatic and early stage were completely identical in both visible and NIR range (both produce an identical curve, which cannot be distinguished in Fig. 5d). Figure 5d shows some differences between the early/asymptomatic stages and late stage in 735 nm and 813 nm.
Spectral reflectance analysis of the TS disease in laboratory conditions
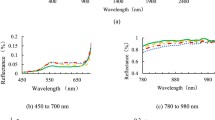
The spectral signature of TS disease lesions showed a higher peak value in the late stage and it was consistently higher when compared to the other disease development stages in the green (495–570 nm) and red range (550–690 nm) (Fig. 6a). In early stage, the spectral signature displayed a lower peak value than other stages, but the spectral reflectance value in green range was almost the same as a healthy plant. In NIR (700–900 nm), the spectral signature of asymptomatic plants shows a lower reflectance value than other disease development stages, while the late stage has a higher reflectance value than others. This increment was most distinctive in the visible range (550–690 nm) and between 787 and 900 nm in NIR range (Fig. 6a). These differences in NIR range can be used for early disease detection, as disease detection is crucial to be executed when no symptoms are visible (early stage) and proceed with spraying to avoid a disease outbreak. In late TS stages, the color of the leaves turned to yellow and brown in some areas. This would lead to reduction of chlorophyll content, increase of the lesion size, and increase of the darkness of the lesion in leaf structure; all these factors might affect the light absorption in visible range especially in red range and scatter the light in the NIR range.
From the sensitivity values (Fig. 6b), the three disease development stages of TS can be distinguished in the spectral area of 472–726 nm; especially the asymptomatic/early with the late stages. The sensitivity values of the early and asymptomatic stages showed a difference in 520–620 nm (Fig. 6b). In NIR range, there were significant differences between all stages.
The spectral difference values varied based on the severity of TS (Fig. 6c). In the asymptomatic stage, there were no changes occurring in leaf pigment, or it was not detectable by these methods. The most significant differences started in the red edge range when the lesions developed, as the lesion size increased, and leaves developed dark necrotic spots. At this moment, the light scattered depending on the quantity of leaf cell damage (Wang et al. 2008). Due to senescent tissue and the accumulation of brown and reddish-brown pigments, the spectral reflectance increased in red and red edge range (600–700 nm) for the TS disease when compared to the spectral signature of healthy tomato leaves. The correlation coefficient values of asymptomatic, early and late stages were similar (similar curves in Fig. 6d).
UAV-based disease detection
Figure 7a shows the UAV-based spectral signature of healthy, TS and BS affected plants (measured in the field at the flowering growth stage). In the visible range, the spectral reflectance value of healthy plants was higher than other disease development stages, with a peak value of 15.8% at 556 nm (green band). The spectral reflectance value of healthy plants was lower than the other disease development stages in NIR range; the spectral signature of all the stages intersected at 746 nm (red edge). In the NIR (700–1000 nm) range, the spectral reflectance value of BS was higher than the healthy and the TS disease-affected plants. The upper portion (crown) of the tomato canopy showed few symptoms of the disease. Therefore, disease detection by using UAV at 30 m height is considered as a challenging task. The leaves at the bottom portion were showing a higher level of disease severity. The leaves in the upper portion of a plant were considered as an asymptomatic stage, because they did not have any visual symptoms. All plants were already infected, but most leaves looked visually in a good shape, except a few leaves that showed early symptoms. The TS disease starts in the bottom part of the plant and gradually infects the other parts of plant. The severity of disease affects the photosynthesis absorption of the plants especially in blue and red range, so the spectral signature would be varied based on progress of the disease (Mahlein et al. 2010). Abdulridha et al. (2016) and Jacquemoud and Baret (1990) presented that the pigment amount and the leaf structure would be the essential factors that might affect the light reflectance or scattering the light of a leaf. Figure 7b shows the sensitivity values of BS and TS diseases; the highest sensitivity value was at 408 nm for both diseases. In the NIR, the sensitivity value increased at 933 nm for both diseases, but the BS was slightly higher than TS. Figure 7c illustrates the spectral difference values between healthy and BS, and healthy and TS. The maximum peak value was recorded for TS at 730 nm; then, the values reduced dramatically (negative values). The highest value of the correlation coefficient (r = 0.87) was at 408 and 978 nm, while it showed a negative value at 876 nm (Fig. 7d). All these parameters (Fig. 7) can be used to identify both diseases and distinguish them from healthy plans in the field especially in lower canopy of the plants. Critical wavelengths must be used, based on Fig. 7, for accurate disease detection.
Classification results
Lab-based analysis
Bacterial spot disease detection
Table 2 and Fig. 8 present the most significant spectral reflectance bands having higher classification weight values and can be used to distinguish between healthy and BS symptomatic plants for the indoor conditions. The MLP method recorded high classification accuracy in asymptomatic and early stages of 97% and 98%, respectively. The STDA method recorded 98% accuracy in the laboratory-based analysis, on late disease development stage. The MLP method was selected to be the best classification method because it had higher values than STDA in most categories. The most significant band (with high disease detection accuracies) was the 412 nm (100%). The highest classification accuracies (100%) for healthy versus BS asymptomatic, early and late stages were found at 1003, 426.9 and 688 nm, respectively. The most significant bands to distinguish between healthy and diseased plants were located in the red and blue bands (412, 416, and 426.9 nm). In the asymptomatic stage, the best bands to detect the disease were in the NIR with more than 1000 nm.
Target spot disease detection
Best bands that were selected to distinguish the asymptomatic, early and late disease development stages were the 382 nm (100%), 966.8 nm (100%), and 675.4 nm (100%). The bands that were selected for TS detection were in the visible and NIR range. The classification method with the highest accuracy was the MLP (99%) for indoor conditions. The MLP classification rates in the asymptomatic, early and late disease development stages (98%, 98% and 99% respectively) were higher than the STDA rates (Fig. 8a). The highest classification value of 99% was recorded at TS late stage.
Classification and best bands selection for the TS and BS diseases
The MLP classification method for distinguishing between TS and BS diseases achieved 92%, 83% and 84% accuracy for asymptomatic, early and late stages, respectively. The STDA methods recorded lower classification rates in all stages (88%, 80% and 82% in the three disease stages) (Table 2 and Fig. 8a). The most significant bands were selected in the red and red edge range. It was impossible to identify the two disease lesion types visually, since initial disease symptoms are very similar for both diseases (e.g., lesion color).
Field (UAV)-based analysis
The results of the UAV-based classification and the best bands selection were similar to the lab-based results. The best bands selected for TS and BS diseases were in blue range and red edge range (412–759 nm) (Table 2; Field-base). More specifically, the most significant bands with high classification weight values for the TS disease at the UAV-based analysis were the 416 nm (100%) and 737 nm (98%) (Table 2), and for the BS the 412 nm (100%) and the 416 nm (99%).
The MLP and STDA methods have similar overall classification rates for detecting BS (98%) and TS (97% for MLP and 96% for STDA) (Fig. 8b). It is to be noticed that both the lab and field analyses achieved similar disease detection accuracies, even though they had different procedure to be followed (e.g., the lab experiments were conducted in optimal conditions, in contrast to the field conditions).
Most significant VIs for detecting BS and TS diseases
Lab-based analysis
Thirty-five VIs, related to different physiological plant parameters, were calculated for each disease in different stages. Based on the progression of the disease development stage, some indices are more suitable than others for disease detection. The NDVI850 accurately distinguished healthy plants from BS asymptomatic, early, and late development disease stages. Most spectral changes occurred in the visible range, especially in the red band. After inoculation of tomato plants with X. perforans, the spectral reflectance value started to change from the asymptomatic stage up until the late stage of disease infection, when the necrosis increased dramatically with the formation of dark lesions, which would affect the pigment value (Franke et al. 2005; Devadas et al. 2009). The photosynthesis process would be also affected, by the reduction of the chlorophyll content; therefore, the Chl green was also able to distinguish BS affected from healthy plants. Franke et al. (2005) indicated that NDVI and chlorophyll indices are able to distinguish the rust infection in individual wheat leaves spots on the basis of associated changes in photosynthetically-active biomass.
The triangle vegetation index (TVI, the NDVI850 and the Chl green gave the highest classification weight values to distinguish the TS in asymptomatic, early and late development stage respectively (Table 3; Lab-based). The TVI indicates the radiant energy absorption of chlorophyll (De Tomás et al. 2012). Shi et al. (2018) selected the TVI to be as one of the VIs to differentiate between diseased rice from healthy rice in the various stages of the disease development. Some VIs could be used to detect both BS and TS diseases and distinguish them from healthy plants; for example, the NDVI850 and the Chl green. Both diseases affect photosynthetic pigment absorption and chlorophyll content. In TS late stage, the most significant VIs for disease detection were the NDVI, Chl green and WI. The water index (WI) was also utilized by Naidu et al. (2009) to detect grapevine leaf roll, associated with virus-3 in wine grape cultivars; the results showed that the WI was the most significant VI to distinguish between healthy and non-healthy wine grapes.
Field (UAV)-based analysis
The most suitable VIs for distinguishing between healthy and TS or BS affected plants in the UAV-based analysis were the PRI and Chl green (Table 3; Field-base). The most significant VI that could distinguish between healthy plants and BS and TS infected plants was the PRI in field condition. The PRI is sensitive to variations in carotenoid colors (e.g. xanthophyll pigments) in live vegetation. Carotenoid colors concentration is indicative of photosynthetic light use efficiency, or the rate of carbon dioxide uptake by foliage per unit energy absorbed of chlorophyll content. Therefore, any change in chlorophyll concentration will cause a difference in the value of PRI (Filella et al. 2018). Due to this sensitivity of the PRI, it could be used to distinguish a diseased plant from a healthy plant and identify clear differences between them. Furthermore, the PRI represents changes in the xanthophyll cycle and can detect changes in yellow or brown carotenoid plant pigments that cause the chlorosis of leaf color (Sukhova and Sukhov 2018).
Conclusion
The objective of this study was to detect and distinguish between TS and BS infected tomato plants at different disease development stages in both laboratory conditions, by using a benchtop hyperspectral system, and in the field, by using a UAV-based hyperspectral imaging technique. The spectral signatures of both diseases under three different disease development stages showed significant differences (in spectral reflectance values). TS and BS diseases have very similar symptoms and it is very difficult to distinguish between the two diseases visually in initial stages of disease development. Satisfactory results were obtained in the laboratory and field (UAV-based technique) conditions to detect both diseases in asymptomatic, early and late stages. The MLP method had a higher classification value than the STDA method; the MLP classification accuracies were very high (97–99%) for all stages. The disease detection accuracy utilizing VIs was varied, based on disease severity. The best VIs for detecting BS were the PRI, NDVI, WI and Chl green for both indoor and outdoor conditions. The most significant VIs for detecting TS were the PRI, TVI, NDVI850, WI and Chl green for both indoor and outdoor conditions.
References
Abdulridha, J., Ampatzidis, Y., Ehsani, R., & de Castro, A. (2018). Evaluating the performance of spectral features and multivariate analysis tools to detect laurel wilt disease and nutritional deficiency in avocado. Computers and Electronics in Agriculture, 155, 203–211. https://doi.org/10.1016/j.compag.2018.10.016.
Abdulridha, J., Ehsani, R., Abd-Elrahma, A., & Ampatzidis, Y. (2019). A remote sensing technique for detecting laurel wilt disease in avocado in presence of other biotic and abiotic stresses. Computers and Electronics in Agriculture, 156, 549–557. https://doi.org/10.1016/j.compag.2018.12.018.
Abdulridha, J., Ehsani, R., & de Castro, A. (2016). Detection and differentiation between laurel wilt disease, phytophthora disease, and salinity damage using a hyperspectral sensing technique. Agriculture-Basel, 6(4), 13. https://doi.org/10.3390/agriculture6040056.
Albetis, J., Duthoit, S., Guttler, F., Jacquin, A., Goulard, M., Poilve, H., et al. (2017). Detection of flavescence doree grapevine disease using unmanned aerial vehicle (UAV) multispectral imagery. Remote Sensing, 9(4), 308. https://doi.org/10.3390/rs9040308.
Ampatzidis, Y., De Bellis, L., & Luvisi, A. (2017). iPathology: Robotic applications and management of plants and pant diseases. Sustainability, 9(6), 1010. https://doi.org/10.3390/su9061010.
Ampatzidis, Y., Kiner, J., Abdolee, R., & Ferguson, L. (2018). Voice-controlled and wireless solid set canopy delivery (VCW-SSCD) system for mist-cooling. Sustainability, 10(2), 421. https://doi.org/10.3390/su10020421.
Ampatzidis, Y., & Partel, V. (2019). UAV-based high throughput phenotyping in citrus utilizing multispectral imaging and artificial intelligence. Remote Sensing, 11(4), 410. https://doi.org/10.3390/rs11040410.
Ampatzidis, Y., Partel, V., Meyering, B., & Albrecht, U. (2019). Citrus rootstock evaluation utilizing UAV-based remote sensing and artificial intelligence. Computers and Electronics in Agriculture, 164, 104900. https://doi.org/10.1016/j.compag.2019.104900.
Ampatzidis, Y., Tan, L., Haley, R., & Whiting, M. D. (2016). Cloud-based harvest management information system for hand-harvested specialty crops. Computers and Electronics in Agriculture, 122, 161–167. https://doi.org/10.1016/j.compag.2016.01.032.
Ampatzidis, Y. G., & Vougioukas, S. G. (2009). Field experiments for evaluating the incorporation of RFID and barcode registration and digital weighing technologies in manual fruit harvesting. Computers and Electronics in Agriculture, 66(2), 166–172. https://doi.org/10.1016/j.compag.2009.01.008.
Ampatzidis, Y. G., Whiting, M. D., Scharf, P. A., & Zhang, Q. (2012). Development and evaluation of a novel system for monitoring harvest labor efficiency. Computers and Electronics in Agriculture, 88, 85–94. https://doi.org/10.1016/j.compag.2012.06.009.
Babar, M. A., Reynolds, M. P., Van Ginkel, M., Klatt, A. R., Raun, W. R., & Stone, M. L. (2006). Spectral reflectance to estimate genetic variation for in-season biomass, leaf chlorophyll, and canopy temperature in wheat. Crop Science, 46(3), 1046–1057. https://doi.org/10.2135/cropsci2005.0211.
Barbedo, J. G. A. (2013). Digital image processing techniques for detecting, quantifying and classifying plant diseases. Springerplus, 2, 660. https://doi.org/10.1186/2193-1801-2-660.
Barnes, J. D., Balaguer, L., Manrique, E., Elvira, S., & Davison, A. W. (1992). A reappraisal of the use of DMSO for the extraction and determination of chlorophylls-A and chlorophylls-B in lichens and higher-plants. Environmental and Experimental Botany, 32(2), 85–100. https://doi.org/10.1016/0098-8472(92)90034-Y.
Bausch, W. C., & Duke, H. R. (1996). Remote sensing of plant nitrogen status in corn. Transactions of the ASAE, 39(5), 1869–1875.
Blackburn, G. A. (1998). Spectral indices for estimating photosynthetic pigment concentrations: A test using senescent tree leaves. International Journal of Remote Sensing, 19(4), 657–675. https://doi.org/10.1080/014311698215919.
Broge, N. H., & Leblanc, E. (2001). Comparing prediction power and stability of broadband and hyperspectral vegetation indices for estimation of green leaf area index and canopy chlorophyll density. Remote Sensing of Environment, 76(2), 156–172. https://doi.org/10.1016/s0034-4257(00)00197-8.
Burks, T. F., Shearer, S. A., & Payne, F. A. (2000). Classification of weed species using color texture features and discriminant analysis. Transactions of the ASAE, 43(2), 441–448.
Calderon, R., Navas-Cortes, J. A., Lucena, C., & Zarco-Tejada, P. J. (2013). High-resolution airborne hyperspectral and thermal imagery for early, detection of verticillium wilt of olive using fluorescence, temperature and narrow-band spectral indices. Remote Sensing of Environment, 139, 231–245. https://doi.org/10.1016/j.rse.2013.07.031.
Chappelle, E. W., Kim, M. S., & McMurtrey, J. E. (1992). Ration analysis of reflectance spectra (RARS)—An algorithm for the remote estimation concentration of chlorophyll-a, chlorophyll-b, and carotenoid soybean leaves. Remote Sensing of Environment, 39(3), 239–247. https://doi.org/10.1016/0034-4257(92)90089-3.
Cruz, A., Ampatzidis, Y., Pierro, R., Materazzi, A., Panattoni, A., De Bellis, L., et al. (2019). Detection of grapevine yellows symptoms in Vitis vinifera L. with artificial intelligence. Computers and Electronics in Agriculture, 157, 63–76. https://doi.org/10.1016/j.compag.2018.12.028.
Cruz, A. C., Luvisi, A., De Bellis, L., & Ampatzidis, Y. (2017). X-FIDO: An effective application for detecting olive quick decline syndrome with deep learning and data fusion. Frontiers in Plant Science, 8, 1741. https://doi.org/10.3389/fpls.2017.01741.
Das, R., & Sengur, A. (2010). Evaluation of ensemble methods for diagnosing of valvular heart disease. Expert Systems with Applications, 37(7), 5110–5115. https://doi.org/10.1016/j.eswa.2009.12.085.
Dash, J. P., Watt, M. S., Pearse, G. D., Heaphy, M., & Dungey, H. S. (2017). Assessing very high resolution UAV imagery for monitoring forest health during a simulated disease outbreak. ISPRS Journal of Photogrammetry and Remote Sensing, 131, 1–14. https://doi.org/10.1016/j.isprsjprs.2017.07.007.
De Tomás, A., Nieto, H., Guzinski, R., Mendiguren, G., Sandholt, I., & Berline, P. (2012). In multi-scale approach of the surface temperature/vegetation index triangle method for estimating evapotranspiration over heterogeneous landscapes. EGU General Assembly, 101, 131–138.
Devadas, R., Lamb, D. W., Simpfendorfer, S., & Backhouse, D. (2009). Evaluating ten spectral vegetation indices for identifying rust infection in individual wheat leaves. Precision Agriculture, 10(6), 459–470. https://doi.org/10.1007/s11119-008-9100-2.
Di Gennaro, S. F., Battiston, E., Di Marco, S., Facini, O., Matese, A., Nocentini, M., et al. (2016). Unmanned aerial vehicle (UAV)-based remote sensing to monitor grapevine leaf stripe disease within a vineyard affected by esca complex. Phytopathologia Mediterranea, 55(2), 262–275. https://doi.org/10.14601/Phytopathol_Mediterr-18312.
Filella, I., Zhang, C., Seco, R., Potosnak, M., Guenther, A., Karl, T., et al. (2018). A MODIS photochemical reflectance index (PRI) as an estimator of isoprene emissions in a temperate deciduous forest. Remote Sensing, 10(4), 557. https://doi.org/10.3390/rs10040557.
Foody, G. M. (2004). Supervised image classification by MLP and RBF neural networks with and without an exhaustively defined set of classes. International Journal of Remote Sensing, 25(15), 3091–3104. https://doi.org/10.1080/01431160310001648019.
Franke, J., Menz, G., Oerke, E. C., & Rascher, U. (2005). Comparison of multi- and hyperspectral imaging data of leaf rust infected wheat plants. In G. D. U. Manfred Owe (Ed.), SPIE-volume 5976 remote sensing for agriculture, ecosystems, and hydrology VII (p. 59761D). Washington, USA: SPIE - The International Society for Optical Engineering.
Galvez, J. F., Mejuto, J. C., & Simal-Gandara, J. (2018). Future challenges on the use of blockchain for food traceability analysis. TrAC, Trends in Analytical Chemistry, 107, 222–232. https://doi.org/10.1016/j.trac.2018.08.011.
Gamon, J. A., Penuelas, J., & Field, C. B. (1992). A narrow-waveband spectral index that tracks diurnal changes in photosynthetic efficiency. Remote Sensing of Environment, 41(1), 35–44. https://doi.org/10.1016/0034-4257(92)90059-s.
Gitelson, A. A., Gritz, Y., & Merzlyak, M. N. (2003). Relationships between leaf chlorophyll content and spectral reflectance and algorithms for non-destructive chlorophyll assessment in higher plant leaves. Journal of Plant Physiology, 160(3), 271–282. https://doi.org/10.1078/0176-1617-00887.
Gitelson, A. A., Kaufman, Y. J., Stark, R., & Rundquist, D. (2002). Novel algorithms for remote estimation of vegetation fraction. Remote Sensing of Environment, 80(1), 76–87. https://doi.org/10.1016/s0034-4257(01)00289-9.
Gitelson, A. A., & Merzlyak, M. N. (1996). Signature analysis of leaf reflectance spectra: Algorithm development for remote sensing of chlorophyll. Journal of Plant Physiology, 148(3–4), 494–500.
Gitelson, A. A., Merzlyak, M. N., & Chivkunova, O. B. (2001). Optical properties and nondestructive estimation of anthocyanin content in plant leaves. Photochemistry and Photobiology, 74(1), 38–45. https://doi.org/10.1562/0031-8655(2001)074%3c0038:opaneo%3e2.0.co;2.
Haboudane, D., Miller, J. R., Pattey, E., Zarco-Tejada, P. J., & Strachan, I. B. (2004). Hyperspectral vegetation indices and novel algorithms for predicting green LAI of crop canopies: Modeling and validation in the context of precision agriculture. Remote Sensing of Environment, 90(3), 337–352. https://doi.org/10.1016/j.rse.2003.12.013.
Haboudane, D., Miller, J. R., Tremblay, N., Zarco-Tejada, P. J., & Dextraze, L. (2002). Integrated narrow-band vegetation indices for prediction of crop chlorophyll content for application to precision agriculture. Remote Sensing of Environment, 81(2–3), 416–426. https://doi.org/10.1016/s0034-4257(02)00018-4.
Hariharan, J., Fuller, J., Ampatzidis, Y., Abdulridha, J., & Lerwill, A. (2019). Finite difference analysis and bivariate correlation of hyperspectral data for detecting laurel wilt disease and nutritional deficiency in avocado. Remote Sensing, 11(15), 1748. https://doi.org/10.3390/rs11151748.
Huang, H. S., Deng, J. Z., Lan, Y. B., Yang, A. Q., Zhang, L., Wen, S., et al. (2019). Detection of helminthosporium leaf blotch disease based on UAV imagery. Applied Sciences-Basel, 9(3), 558. https://doi.org/10.3390/app9030558.
Huberty, C. J. (1984). Issues in the use and interpretation of discriminant-analysis. Psychological Bulletin, 95(1), 156–171. https://doi.org/10.1037//0033-2909.95.1.156.
Hunt, E. R., Jr., & Rock, B. N. (1989). Detection of changes in leaf water content using near- and middle-infrared reflectances. Remote Sensing of Environment, 30(1), 43–54.
Jacquemoud, S., & Baret, F. (1990). Prospect—A model of leaf optical-properties spectra. Remote Sensing of Environment, 34(2), 75–91. https://doi.org/10.1016/0034-4257(90)90100-z.
Jordan, C. F. (1969). Derivation of leaf area index from quality of light on the forest floor. Ecology, 50, 663–666.
Liao, Y. Y., Strayer-Scherer, A. L., White, J., Mukherjee, A., De la Torre-Roche, R., Ritchie, L., et al. (2019). Nano-magnesium oxide: A novel bactericide against copper-tolerant Xanthomonas perforans causing tomato bacterial spot. Phytopathology, 109(1), 52–62. https://doi.org/10.1094/phyto-05-18-0152-r.
Lowe, A., Harrison, N., & French, A. P. (2017). Hyperspectral image analysis techniques for the detection and classification of the early onset of plant disease and stress. Plant Methods, 13, 80. https://doi.org/10.1186/s13007-017-0233-z.
Lu, J. Z., Ehsani, R., Shi, Y. Y., de Castro, A. I., & Wang, S. (2018). Detection of multi-tomato leaf diseases (late blight, target and bacterial spots) in different stages by using a spectral-based sensor. Scientific Reports, 8, 2793. https://doi.org/10.1038/s41598-018-21191-6.
Luvisi, A., Ampatzidis, Y. G., & De Bellis, L. (2016). Plant pathology and information technology: Opportunity for management of disease outbreak and applications in regulation frameworks. Sustainability, 8(8), 831. https://doi.org/10.3390/su8080831.
MacKenzie, K. J., Sumabat, L. G., Xavier, K. V., & Vallad, G. E. (2018). A review of corynespora cassiicola and its increasing relevance to tomato in Florida. Plant Health Progress, 19, 303–309. https://doi.org/10.1094/PHP-05-18-0023-RV.
Mahlein, A. K. (2016). Plant disease detection by imaging sensors—Parallels and specific demands for precision agriculture and plant phenotyping. Plant Disease, 100(2), 241–251. https://doi.org/10.1094/pdis-03-15-0340-fe.
Mahlein, A. K., Steiner, U., Dehne, H. W., & Oerke, E. C. (2010). Spectral signatures of sugar beet leaves for the detection and differentiation of diseases. Precision Agriculture, 11(4), 413–431. https://doi.org/10.1007/s11119-010-9180-7.
Merton, R. (1998). Monitoring community hysteresis using spectral shift analysis and the red-edge vegetation stress index. JPL Airborne Earth Science Workshop. NASA, Jet Propulsion Laboratory, Pasadena, California, USA
Metternicht, G. (2003). Vegetation indices derived from high-resolution airborne videography for precision crop management. International Journal of Remote Sensing, 24(14), 2855–2877. https://doi.org/10.1080/01431160210163074.
Naidu, R. A., Perry, E. M., Pierce, F. J., & Mekuria, T. (2009). The potential of spectral reflectance technique for the detection of Grapevine leafroll-associated virus-3 in two red-berried wine grape cultivars. Computers and Electronics in Agriculture, 66(1), 38–45. https://doi.org/10.1016/j.compag.2008.11.007.
Onofre, R. B., Rebello, C. S., Mertely, J. C., & Peres, N. A. (2019). First report of target spot caused by Corynespora cassiicola on strawberry in North America. Plant Disease, 103(6), 1412. https://doi.org/10.1094/pdis-01-19-0057-pdn.
Partel, V., Kakarla, C., & Ampatzidis, Y. (2019a). Development and evaluation of a low-cost and smart technology for precision weed management utilizing artificial intelligence. Computers and Electronics in Agriculture, 157, 339–350. https://doi.org/10.1016/j.compag.2018.12.048.
Partel, V., Nunes, L., Stansly, P., & Ampatzidis, Y. (2019b). Automated vision-based system for monitoring Asian citrus psyllid in orchards utilizing artificial intelligence. Computer and Electronics in Agriculture, 162, 328–336.
Penuelas, J., Baret, F., & Filella, I. (1995). Semiempirical indexes to assess carotenoids chlorophyll-a ratio from leaf spectral reflectance. Photosynthetica, 31(2), 221–230.
Penuelas, J., Filella, I., Biel, C., Serrano, L., & Save, R. (1993). The reflectance at the 950–970 nm region as an indicator of plant water status. International Journal of Remote Sensing, 14(10), 1887–1905.
Penuelas, J., Gamon, J. A., Fredeen, A. L., Merino, J., & Field, C. B. (1994). Reflectance indexes associated with physiological-changes in nitrogen-limited and water-limited sunflower leaves. Remote Sensing of Environment, 48(2), 135–146. https://doi.org/10.1016/0034-4257(94)90136-8.
Penuelas, J., Pinol, J., Ogaya, R., & Filella, I. (1997). Estimation of plant water concentration by the reflectance water index WI (R900/R970). International Journal of Remote Sensing, 18(13), 2869–2875. https://doi.org/10.1080/014311697217396.
Pernezny, K., Datnoff, L. E., Mueller, T., & Collins, J. (1996). Losses in fresh-market tomato production in Florida due to target spot and bacterial spot and the benefits of protectant fungicides. Plant Disease, 80(5), 559–563. https://doi.org/10.1094/pd-80-0559.
Pernezny, K., & Raid, R. N. (2001). Occurrence of bacterial leaf spot of Escarole caused by Pseudomonas cichorii in the Everglades agricultural area of Southern Florida. Plant Disease, 85(11), 1208. https://doi.org/10.1094/pdis.2001.85.11.1208b.
Pernezny, K., Stoffella, P., Collins, J., Carroll, A., & Beaney, A. (2002). Control of target spot of tomato with fungicides, systemic acquired resistance activators, and a biocontrol agent. Plant Protection Science, 38(3), 81–88.
Potnis, N., Timilsina, S., Strayer, A., Shantharaj, D., Barak, J. D., Paret, M. L., et al. (2015). Bacterial spot of tomato and pepper: Diverse Xanthomonas species with a wide variety of virulence factors posing a worldwide challenge. Molecular Plant Pathology, 16(9), 907–920. https://doi.org/10.1111/mpp.12244.
Pydipati, R., Burks, T. F., & Lee, W. S. (2006). Identification of citrus disease using color texture features and discriminant analysis. Computers and Electronics in Agriculture, 52(1–2), 49–59. https://doi.org/10.1016/j.compag.2006.01.004.
Raun, W. R., Solie, J. B., Johnson, G. V., Stone, M. L., Lukina, E. V., Thomason, W. E., et al. (2001). In-season prediction of potential grain yield in winter wheat using canopy reflectance. Agronomy Journal, 93(1), 131–138. https://doi.org/10.2134/agronj2001.931131x.
Reynolds, C. F., Kupfer, D. J., Houck, P. R., Hoch, C. C., Stack, J. A., Berman, S. R., et al. (1988). Reliable discrimination of elderly depressed and demented patients by electroencephalographic sleep data. Archives of General Psychiatry, 45(3), 258–264.
Roujean, J. L., & Breon, F. M. (1995). Estimating par absorbed by vegetation from bidirectional reflectance measurements. Remote Sensing of Environment, 51(3), 375–384. https://doi.org/10.1016/0034-4257(94)00114-3.
Salami, E., Barrado, C., & Pastor, E. (2014). UAV flight experiments applied to the remote sensing of vegetated areas. Remote Sensing, 6(11), 11051–11081. https://doi.org/10.3390/rs61111051.
Schlub, R., L, Smith, L., J, Datnoff, L., E, & Pernezny, K. (2007). An overview of target spot of tomato caused by Corynespora cassiicola. ll International Symposium on Tomato Disease SHS Acta Horticulturae, p. 808.
Sharma, S., & Bhattarai, K. (2019). Progress in developing bacterial spot resistance in tomato. Agronomy-Basel, 9(1), 26. https://doi.org/10.3390/agronomy9010026.
Shazia, A., Khan, S. M., Khan, M. F., Hameed, U., et al. (2018). Antifungal activity of different systemic fungicide against Fusarium oxysporum f. sp. Lycopersici associated with tomato wilt and emergence of resistance in pathogen. Pakistan Journal of Phytopathology, 30(2), 169–176.
Shi, Y., Huang, W. J., Ye, H. C., Ruan, C., Xing, N. C., Geng, Y., et al. (2018). Partial least square discriminant analysis based on normalized two-stage vegetation indices for mapping damage from rice diseases using PlanetScope datasets. Sensors, 18(6), 1901. https://doi.org/10.3390/s18061901.
Singh, S. R., & Allen, D. J. (1979). Cowpea pests and diseases (Vol. 2). Ibadan, Nigeria: International Institute of Tropical Agriculture.
Smigaj, M., Gaulton, R., Barr, S. L, & Suárez, J. C. (2015). UAV-borne thermal imaging for forest health monitoring: Detection of disease induced canopy temperature increase. The International Archives of the Photogrammetry, Remote Sensing and Spatial Information Sciences, (Vol. XL-3/W3).
Sukhova, E., & Sukhov, V. (2018). Connection of the photochemical reflectance index (PRI) with the photosystem II quantum yield and nonphotochemical quenching can be dependent on variations of photosynthetic parameters among investigated plants: A meta-analysis. Remote Sensing, 10(5), 771. https://doi.org/10.3390/rs10050771.
Tamouridou, A. A., Pantazi, X. E., Alexandridis, T., Lagopodi, A., Kontouris, G., & Moshou, D. (2018). Spectral identification of disease in weeds using multilayer perceptron with automatic relevance determination. Sensors, 18(9), 2770. https://doi.org/10.3390/s18092770.
Thomas, S., Kuska, M. T., Bohnenkamp, D., Brugger, A., Alisaac, E., Wahabzada, M., et al. (2018). Benefits of hyperspectral imaging for plant disease detection and plant protection: A technical perspective. Journal of Plant Diseases and Protection, 125(1), 5–20. https://doi.org/10.1007/s41348-017-0124-6.
Vincini, M., Frazzi, E., & D’Alessio, P. (2007). Comparison of narrow-band and broad-band vegetation indexes for canopy chlorophyll density estimation in sugar beet. In J. V. Stafford (Ed.), Precision agriculture ‘07: Proceedings of the 6th European Conference on Precision Agriculture (pp. 189–196). Wageningen, The Netherlands: Wageningen Academic Publishers.
Vivaldini, K. C. T., Martinelli, T. H., Guizilini, V. C., Souza, J. R., Oliveira, M. D., Ramos, F. T., et al. (2019). UAV route planning for active disease classification. Autonomous Robots, 43(5), 1137–1153. https://doi.org/10.1007/s10514-018-9790-x.
Wang, F.-M., Huang, J.-F., Xu, J.-F., & Wang, X.-Z. (2008). Wavebands selection tor rice information extraction based on spectral bands inter-correlation. Spectroscopy and Spectral Analysis, 28(5), 1098–1101.
Zarco-Tejada, P. J., Gonzalez-Dugo, V., & Berni, J. A. J. (2012). Fluorescence, temperature and narrow-band indices acquired from a UAV platform for water stress detection using a micro-hyperspectral imager and a thermal camera. Remote Sensing of Environment, 117, 322–337. https://doi.org/10.1016/j.rse.2011.10.007.
Zitter, T. A. (1985). Bacterial disease of tomato. Cooperative Extension. New York: Cornell University. Fact sheet page: 735.50.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Abdulridha, J., Ampatzidis, Y., Kakarla, S.C. et al. Detection of target spot and bacterial spot diseases in tomato using UAV-based and benchtop-based hyperspectral imaging techniques. Precision Agric 21, 955–978 (2020). https://doi.org/10.1007/s11119-019-09703-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11119-019-09703-4