Abstract
A gap exists between the growth of active germplasm collections and their effective use because of the size of these collections and the limitations of financial resources, skilled labor, and information on the traits of each accession conserved in germplasm banks. To overcome this, the development of core collections (CCs) is of significant importance. In this study, we established the CCs of lima bean (Phaseolus lunatus) accessions from the Germplasm Bank of Phaseolus at the Universidade Federal do Piauí based on two selection strategies and performed validation based on nuclear microsatellite (ncSSR) data. Two CCs were established from 395 accessions and characterized by 11 phenotypic traits of the seed. These CCs were defined using proportional and logarithmic sampling strategies, each consisting of 79 accessions. A set of 13 ncSSRs was used to validate and estimate the genetic diversity. The estimates of parameters (means, variances, and amplitudes of variation), kurtosis, skewness, and average variability retention index indicated no differences between the two CCs when compared with the entire collection (EC). However, the comparison between the accessions that did not coincide with the two CCs through ncSSRs showed differences. The logarithmic CC showed higher expected heterozygosity, mean number of alleles, and number of private alleles (0.521, 4.308, and 30, respectively) than those of the proportional CC strategy (0.352, 2.154, and 2, respectively), indicating greater genetic diversity. Thus, the logarithmic CC proved to be more suitable for conservation purposes and should be further examined in the establishment of thematic collections of lima beans.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The lima bean (Phaseolus lunatus L.) is an important species of the Fabaceae family distributed worldwide (Baudet 1977). It provides an alternative source of proteins, fibers, and mineral elements for the growing population (Seidu et al. 2018). Studies conducted by Chacón-Sánchez and Martínez-Castillo (2017) reported that wild lima bean is classified into three major gene pools according to genomic data: two Mesoamerican (MI and MII) and one Andean (AI) gene pool. The possible existence of another Andean gene pool, the AII gene pool restricted to the Andes in central Colombia, has been proposed, although this has not been confirmed.
Genetic variability in the species largely relates to the process of global dispersion, which requires adaptation to different environments, making it a promising crop for improving food security under the predicted scenarios of climate change in Latin America and elsewhere (García et al. 2021). Conservation of the genetic resources of the lima bean is conducted ex situ in germplasm banks.
According to Govindaraj et al. (2015), despite the high availability of genetic diversity, managing and exploring a large number of accessions in collections can be challenging. The size of the collections, presence of duplicates, limitation of financial resources, and information about the traits of each conserved accession have contributed to a disparity between the growth of these collections and their effective use (Brown 1989a; Xu et al. 2016). To overcome this, the establishment of core collections (CCs) has been proposed, which can be defined as a small set of accessions representative of the genetic variability of a germplasm collection, with minimum redundancy and appropriate for curators, breeders, and geneticists (Frankel and Brown 1984). According to Brown and Spillane (1999), the steps for selecting a CC are (1) identifying the material that will be represented, (2) determining the size of the core collection, (3) dividing the accessions used into different groups, (4) determining the number of entries per group, and (5) choosing the entries from each group that will be included in the CC. Several descriptors can be used in combination for the implementation of a CC, and the most common are passport data; geographic origin; morphological, phenotypic, biochemical, and phyto-pathological characteristics; and molecular markers (Balfourier et al. 2007; Bhattacharjee et al. 2007; Kumar et al. 2016; Martins et al. 2015).
Studies have shown the importance of seed traits in defining gene pools and estimating the genetic diversity of lima bean crops (Chacón-Sánchez and Martínez-Castillo 2017; Silva et al. 2017, 2019). Gomes et al. (2020), using 153 Brazilian accessions, proposed the use of microsatellite data for the establishment of a lima bean CC for the germplasm bank of Phaseolus at the Universidade Federal do Piauí (GBP-UFPI). However, through exchanges, the GBP-UFPI recently acquired accessions of lima beans with different origins from several countries (Andean, Mesoamerican, and South American, among others), including wild genotypes. With the incorporation of new accessions, the question arises of the need to update the core collection of lima beans in Brazil to maximize genetic diversity and ensure a balanced representation of accessions from various collection sites, with minimal redundancy. Evaluating genetic diversity differences between the core collections (CCs) and the entire collection (EC) is a useful approach for testing the effectiveness of the core collection. Various measures, such as genetic diversity parameters, have been used to evaluate and validate the CC (Meryem et al. 2023). Based on the available literature, this is the first study on a CC of this species that is compiled of the integration of geographic distribution, phenotypic, and molecular data. In this context, the main objective of this study was to construct the CC of lima bean in Brazil from the GBP-UFPI, based on two selection strategies that integrate different types of data, and to perform validation based on nuclear microsatellites (ncSSRs) to optimize this germplasm conservation.
Materials and Methods
Germplasm
This study was performed with 395 lima bean accessions from the GBP-UFPI, located in Teresina, Piauí, Brazil. All accessions were characterized based on 11 phenotypic seed descriptors in the Genetic Resources and Plant Breeding Laboratory, Plant Science Department, UFPI.
Eight quantitative and three qualitative seed traits were evaluated in the entire collection (EC) and were described using lima bean crop descriptors from the International Plant Genetic Resources Institute (IPGRI 2001). The quantitative descriptors were (a) seed area (SA, mm2), (b) seed length perimeter (SLP, mm), (c) seed length (SL, mm), (d) seed width (SW, mm), (e) seed length to width ratio (SL SW−1), (f) seed weight (100 WS, g), (g) seed thickness (ST, mm), and (h) seed thickness-to-width ratio (ST SW−1). The Smartgrain software (Tanabata et al. 2012) was used to determine the measures. Seed thickness was measured using a digital caliper. The multi-categorical qualitative descriptors evaluated were (a) seed background color (SBC), (b) standard seed color (SSC), and (c) second standard seed color (SSSC).
Procedures to Establish the Core Collection of Lima Bean
The establishment of the lima bean CC was divided into four stages (Fig. 1): (1) choice of the sample size, in which 79 accessions was considered an adequate size to be managed by the curators at a low relative cost, representing the sampling intensity of 20%; (2) use of the stratified sampling strategy, whereby the lima bean accessions were divided into four groups according to the centers of domestication and diversity: Brazilian (295 accessions), Mesoamerican (48 accessions imported from Costa Rica, El Salvador, Guatemala, Honduras, and Mexico), Andean (12 accessions imported from Colombia, Ecuador, and Peru), and Other (40 accessions imported from Azerbaijan, China, the Philippines, Ghana, Nigeria, and the USA); (3) determination of the proportion with which each group was represented, using either the proportional strategy (P), where of each group was represented in the CC in proportion to its size in the entire collection or the logarithmic strategy (L), in which the number of accessions to sample each group was proportional to the logarithm of the group size (Vasconcelos et al. 2010); and (4) selection of the accessions of each group to be included in the CC using the inverted Tocher’s method of grouping accessions with greater dissimilarity (Nascimento et al. 2018; Vasconcelos et al. 2007).
Evaluation of Candidate Core Collections with the Entire Collection
The validation process for the proportional (P) and logarithmic (L) core collections involved comparing them with the entire collection (EC) from which they were developed. Comparisons between were made of the quantitative traits using statistical parameters of amplitude, mean, variance, kurtosis, skewness, and average variability retention index (VRI), which was calculated according to Diwan et al. (1995). Comparisons of variances and means between the EC and core collections were performed using the F-test and Student’s t-test, respectively (Snedecor and Cochran 1980). The Shannon index was used to assess multi-categorical qualitative traits (Shannon and Weaver 1949).
Comparison of Core Collections Based on Molecular Data
A comparison was made of the disjoint set between the P and L (DS) CCs using nuclear microsatellites (ncSSRs). DNA was extracted from young lima bean leaflets using the modified CTAB method (Doyle and Doyle 1987) at the Genetic Diversity and Breeding Laboratory of the University of São Paulo, “Luiz de Queiroz” College of Agriculture (USP/ESALQ). DNA quality and quantity were checked by electrophoresis in 1% agarose gel with SYBR safe and compared with standard phage lambda DNA (Gomes et al. 2020).
A set of 13 ncSSRs were used: BM140, BM141, BM143, BM146, BM156, BM160, BM164, BM170, BM183, BM189, BM211, BM212 (Gaitán-Solís et al. 2002), and FF01 (Penha 2018). Simple sequence repeat (SSR) primers were synthesized using an M13 tailed in the forward primer. The ncSSRs were amplified as described by Penha (2018), and genotyping was performed using an automated DNA sequencer ABI 3500xL (Applied Biosystems). The sizes of the SSR fragments were analyzed and determined using the GeneMarker program (Applied Biosystems).
Genetic diversity was described in terms of the average number of alleles per locus (Na), percentage of polymorphic loci (%P), number of private alleles to the group (Pa), coefficient of inbreeding (F), observed heterozygosity (Ho), expected heterozygosity (He), results of principal coordinate analysis (PCoA), and analysis of molecular variance (AMOVA), using the GenAlex v. 6.5 software (Peakall and Smouse 2012).
The relationship between clusters among accessions in the DS was investigated using a dendrogram based on Rogers’ distances (1972). The dendrogram was built using the unweighted pair group method with arithmetic means (UPGMA), and the confidence of relationships was assessed with a bootstrap procedure with 1000 resamplings. MEGA software (v. 6.0) was used to edit the final tree (Tamura et al. 2013).
To assess the genetic structure existing among accessions in the DS, a Bayesian clustering algorithm was used with the program STRUCTURE 2.3.3 (Pritchard et al. 2000). The program was executed with the number of clusters varying from one to ten. For each run, 200,000 burn-in and 500,000 Monte Carlo Markov chain (MCMC) iterations were performed. The admixture model with correlated allele frequencies was used. The most likely K number of clusters was determined by Evanno et al. (2005) based on delta K, calculated using the application Structure Harvester (Earl and Vonholdt 2012).
Results and Discussion
Validation of the Representativeness of Strategies to Form the Core Collection
From the EC of 395 lima bean accessions, two CCs were established of 79 accessions each, through stratified sampling (Table 1). The number of accessions in a core collection is an important determining factor of its effective use (Brown and Spillane 1999). The size reduction allows for efforts to be concentrated on the characterization and evaluation of germplasm to form a more complete information base for this set of accessions. The size reduction also cuts costs and allows resources to be directed to other activities, such as germination tests, regeneration, and molecular analysis, facilitating user access to the germplasm collection. The previously developed lima bean CC consisted of 34 accessions (Gomes et al. 2020), whereas the CCs developed in the present study were comparatively higher because of the larger number of accessions in the initial germplasm collection.
The proportion with which each group was represented was based on the P and L sampling strategies, in which 49 accessions coincided. The 30 accessions that differentiated the proportional CC were all from the Brazil group, which had the highest number of accessions. However, the L strategy allowed for the groups with fewer accessions, the Mesoamerican, Andean, and Other groups, to be more represented in the CC. Thus, the L strategy decreased the weight given to large groups compared to that of the P strategy (Oliveira et al. 2010).
Once established, each CC was investigated for its ability to retain the EC variability. Table 2 summarizes the comparison between EC and CCs. When comparing the EC with the CCs regarding the values of amplitudes and means, no significant differences were observed for the means of the traits, except for seed thickness in the logarithmic CC. The P and L strategies maintained similar values to the EC but were unequal in variance (variance of the EC < variance of the CC) and kurtosis (kurtosis of the EC > kurtosis of the CC), which indicates differences between the distributions. According to Van Hintum et al. (2000), maintaining genetic representativeness in the core collection implies the maintenance of the mean and amplitude, and expansion of phenotypic variance among individuals in the core collection in relation to the EC, which occurred in the two established CCs. The same pattern was observed in comparisons with the average variability retention index, whereby strategy P (99.32%) did not differ from strategy L (99.62%). Sampling is efficient when the average variability retention index is greater than 80% (Frankel and Brown 1984). Regarding the Shannon index, it was observed that for the qualitative traits related to seed color, the established CCs (P and L) showed values similar to or close to those of the entire collection (EC), which indicated that a good portion of the variability related to seed colors was maintained (Table 3). In summary, these two strategies were efficient in retaining variability when considering phenotypic traits.
Comparison Between Core Collections Based on Molecular Data
The use of molecular markers, which directly reflect variation at the genetic level, is appropriate because each accession chosen to make up the CC is based on its allelic potential, regardless of its phenotypic expression. However, methodologies based on molecular markers are expensive, labor-intensive, and require knowledge and equipment that are not always available (Mahmoodi et al. 2019). Therefore, the use of molecular markers only for the validation of CCs established from large germplasm collections through phenotypic traits reduces costs and ensures the selection of the best CC strategy.
The 13 SSR markers analyzed in the comparison of the accessions that did not coincide in the P and L CCs generated 58 alleles, with an average of 4.46 alleles per locus, ranging from two (BM146, BM164, BM189, and FF01) to nine alleles (BM140) (Table 4). The mean values of observed heterozygosity (Ho) were generally lower than those of expected heterozygosity (He), except for the BM160 locus, resulting in high inbreeding coefficients (F). Martínez-Castillo et al. (2006) also found an excess of heterozygotes in the BM160 locus in wild lima bean populations in the Yucatan Peninsula, Mexico. The results of low Ho values were expected since lima bean is a mixed species with a predominance of autogamy (Penha et al. 2017). According to Burle et al. (2010), in autogamous species such as Phaseolus beans, low Ho values are expected because most of the loci are homozygotes. In lima beans, low Ho values were also observed by Chacón-Sánchez and Martínez-Castillo (2017), when comparing domesticated accessions with their wild ancestors, with means of Ho = 0.00 and He = 0.25. Silva et al. (2019) found low Ho (ranging from 0 to 0.065, with a mean of 0.025) in 183 lima bean landraces collected from different regions of Brazil. Andueza-Noh et al. (2015) also found low Ho (0.040), but high He (0.667), along with a high F (0.943), in 155 domesticated and wild lima bean accessions.
Comparisons of the estimates of genetic diversity (%P, Na, and He) revealed that the values for the non-coincident accessions of the logarithmic CC were higher than those of the proportional CC (Table 5). These results indicate greater genetic diversity in the logarithmic CC. The explanation lies in the fact that the non-coincident accessions of proportional CC were all from the Brazil group; hence, they are Brazilian and domesticated accessions. Conversely, the non-coincident accessions of the logarithmic CC were part of the Mesoamerican, Andean, and Other origin groups and had a domesticated and wild biological state. The high value of F (0.808) found in strategy P (Table 5) is justified by the founding effect and the fact that they were all domesticated accessions, which was not the case with strategy L.
The logarithmic CC had a higher number of private alleles than the proportional CC (Table 5), which demonstrates that strategy L reduces the weight given to groups with a large number of accessions that may have a higher level of genetic redundancy. This information can be used to increase the probability of sampling rare alleles that confer broad or localized adaptation. According to Brown (1989b), strategy L is superior to strategy P if the degree of genetic variation between and within a group is unknown. If stratification is successful in concentrating the alleles of interest, then the logarithmic scale is a good way to recover alleles present in different distribution patterns. In the case of unsuccessful stratification, the two strategies (P and L) are equally appropriate and slightly less efficient than simple random sampling.
The greater genetic diversity of the logarithmic CC was also represented by the PCoA since it showed a greater distribution of the accessions in the first two coordinates (Fig. 2), which accounted for 41.76% of the variation and allowed for the observation of an overlap of some accessions belonging to logarithmic CC with the accessions selected for proportional CC. This result suggests that the logarithmic CC contains much of the genetic diversity contained in the proportional CC. Analysis of the accessions in the proportional and logarithmic CCs using the UPGMA method (Fig. 3) showed clusters similar to the PCoA. The accessions of the logarithmic CC were distributed throughout the dendrogram; that is, there were representatives of the accessions of the logarithmic CC in almost all groups formed by the analysis. In contrast, the proportional CC accessions were divided into two distinct clusters.
UPGMA dendrogram constructed with the distances of Rogers (1972), illustrating the clustering of accessions that did not coincide in the proportional CC (blue rhombuses) and logarithmic CC (orange squares), based on the variation of 13 microsatellite loci
AMOVA showed that only 8% of the variation occurred between the CCs, indicating that most of the genetic variability was maintained in the CCs. Thus, most of the variations were within each CC (Table 6).
The likely number of genetic clusters was K = 2, suggesting the use of the ΔK method (Fig. 4), showing the formation of two main groups. However, a small peak was also observed at K = 5. The representation of the bar graph for K = 2 and K = 5 of the Bayesian analysis of STRUCTURE (Fig. 5) showed a complex ancestry relationship, which may be partly caused by the gene flow that occurs freely between wild and domesticated accessions, evidenced mainly in the accessions of the logarithmic CC since the proportional CC contained only Brazilian and domesticated accessions. Considering K = 2 (Fig. 5) in the proportional CC, the formation of two distinct genetic groups was observed. Gomes et al. (2020), when analyzing the structure of genetic variability through Bayesian analysis among 153 Brazilian bean accessions, also found different genetic groups. Logarithmic CC can be considered the preferred collection for the conservation and characterization of lima beans since it showed greater genetic variability than proportional CC.
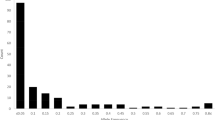
Graphic representation of the optimal K value using 13 SSR loci of the accessions that did not coincide in the proportional (P) and logarithmic (L) CCs obtained by the program STRUCTURE inferred using the criterion ΔK of Evanno et al. (2005)
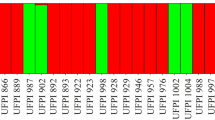
Bar graph in K = 2 and K = 5 of the Bayesian analysis obtained by the program STRUCTURE, showing association coefficients for accessions that did not coincide in the proportional (P) and logarithmic (L) CCs. Accessions are ordered according to the CC to which they belong. Different colors represent distinct genetic groups
Overall, we report that the established lima bean core collection provided by this study will be essential for improving conservation, the use of genetic resources, and the efficiency of the breeding program for the species. The demand for information and seeds from the accessions that make up the CC will likely increase. Given this perspective, procedures for regeneration and multiplication must be adopted. Moreover, it is worth highlighting that the constitution of the CC should not be considered permanent; instead, it should be dynamic, evolving with the evaluation of new accessions and the generation of new information. Future studies should involve the establishment of thematic core collections, such as those focusing on drought tolerance and pathogen resistance, in the face of climate change.
Conclusion
The present study is the first to use molecular diversity data in conjunction with phenotypic data and geographic distribution to establish a CC for lima beans. The small size of the CC is advantageous for field studies and mapping of genomic associations. Logarithmic CC can provide access to genetically diverse and agronomically important germplasms, which can be useful for broadening the genetic basis of the crop. The use of data of different types should be prioritized when available to establish the core collection. The morpho-agronomic data had low efficiency in establishing differences between the two CCs but were fundamental for the initial selection of accessions. Logarithmic CC proved to be more suitable for conservation purposes and should be examined in detail in the establishment of thematic core collections for the species.
Availability of Data and Materials
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Andueza-Noh RH, Martínez-Castillo J, Chacón-Sánchez MI (2015) Domestication of small-seeded lima bean (Phaseolus lunatus L.) landraces in Mesoamerica: evidence from microsatellite markers. Genetica 143:657–669. https://doi.org/10.1007/s10709-015-9863-0
Balfourier F, Roussel V, Strelchenko P, Exbrayat-Vinson F, Sourdille P, Boutet G, Koenig J, Ravel C, Mitrofanova O, Beckert M, Charmet G (2007) A worldwide bread wheat core collection arrayed in a 384-well plate. Theor Appl Genet 114:1265–1275. https://doi.org/10.1007/s00122-007-0517-1
Baudet JC (1977) The taxonomic status of the cultivated types of lima bean (Phaseolus lunatus L.). Tropical Grain Legume 7:29–30
Bhattacharjee R, Khairwal IS, Bramel PJ, Reddy KN (2007) Establishment of a pearl millet [Pennisetum glaucum (L.) R. Br.] core collection based on geographical distribution and quantitative traits. Euphytica 155:35–45. https://doi.org/10.1007/s10681-006-9298-x
Brown AHD (1989a) Core collections: a practical approach to genetic resources management. Genome 31:818–824
Brown AHD (1989b) The case for core collections. In: Brown AHD, Frankel OH, Marshall DR, Williams JT (eds) The use of plant genetic resources. Cambridge University, Cambridge, pp 136–156
Brown AHD, Spillane C (1999) Implementing core collections – principles, procedures, progress, problems and promise. In: Hodgking T (ed) Johnson RC. Core collections for today and tomorrow, IPGRI, Rome pp, pp 1–9
Burle ML, Fonseca JR, Kami JA, Gepts P (2010) Microsatellite diversity and genetic structure among common bean (Phaseolus vulgaris L.) landraces in Brazil, a secondary center of diversity. Theor Appl Genet 121:801–813. https://doi.org/10.1007/s00122-010-1350-5
Chacón-Sánchez MI, Martínez-Castillo J (2017) Testing domestication scenarios of lima bean (Phaseolus lunatus L.) in mesoamerica: insights from genome-wide genetic markers. Front Plant Sci 8:1551. https://doi.org/10.3389/fpls.2017.01551
Diwan N, McIntosh MS, Bauchan GR (1995) Methods of developing a core collection of annual Medicago species. Theor Appl Genet 90:755–761
Doyle JJ, Doyle JL (1987) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Earl DA, vonHoldt BM (2012) Structure harvester: a website and program for visualizing structure output and implementing the Evanno method. Conservation Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Frankel OH, Brown AHD (1984) Plant genetic resources today: a critical appraisal. In Holden JHW, Williams JT (ed) Crop genetic resources: conservation and evaluation, Allen & Unwin, London,. pp 249–257
Gaitán-Solís E, Duque MC, Edwards KJ, Tohme J (2002) Microsatellite repeats in common bean (Phaseolus vulgaris): isolation, characterization, and cross-species amplification in Phaseolus ssp. Crop Sci 42:2128–2136. https://doi.org/10.2135/cropsci2002.2128
Garcia T, Duitama J, Zullo SS, Gil J, Ariani A, Dohle S, Palkovic A, Skeen P, Bermudez-Santana CI, Debouck DG, Martínez-Castillo J, Gepts P, Chacón-Sánchez MI (2021) Comprehensive genomic resources related to domestication and crop improvement traits in Lima bean. Nat Commun 12:702. https://doi.org/10.1038/s41467-021-20921-1
Gomes RLF, Costa MF, Alves-Pereira A, Bajay MM, Viana JPG, Valente SES, Lopes ACA, Zucchi MI, Pinheiro JB (2020) A lima bean core collection based on molecular markers. Sci Agric 77(2):e20180140. https://doi.org/10.1590/1678-992X-2018-0140
Govindaraj M, Vetriventhan M, Srinivasan M (2015) Importance of genetic diversity assessment in crop plants and its recent advances: an overview of its analytical perspectives. Genetics Research International 2015:431487. https://doi.org/10.1155/2015/431487
IPGRI (2001) Descritores para Phaseolus lunatus (feijão-espadinho). International Plant Genetic Resources Institute, 51p
Kumar S, Ambreen H, Variath MT, Rao AR, Agarwal M, Kumar A, Goel S, Jagannath A (2016) Utilization of molecular, phenotypic, and geographical diversity to develop compact composite core collection in the oilseed crop, safflower (Carthamus tinctorius L.) through maximization strategy. Front Plant Sci 7:1554. https://doi.org/10.3389/fpls.2016.01554
Mahmoodi R, Dadpour MR, Hassani D, Zeinalabedini M, Vendramin E, Micali S, Nahandi FZ (2019) Development of a core collection in Iranian walnut (Juglans regia L.) germplasm using the phenotypic diversity. Sci Hortic 249:439–448. https://doi.org/10.1016/j.scienta.2019.02.017
Martínez-Castillo J, Zizumbo-Villarreal D, Gepts P, Delgado-Valerio P, Colunga-GarcíaMarín P (2006) Structure and genetic diversity of wild populations of lima bean (Phaseolus lunatus L.) from the Yucatan Peninsula. Mexico Crop Sci 46:1071–1080. https://doi.org/10.2135/cropsci2005.05-0081
Martins FA, Silva DJH, Carneiro PCS (2015) Establishing a core collection from the integration of morphoagronomical, phytopathological and molecular data. Rev Ciênc Agron 46(4):836–845. https://doi.org/10.5935/1806-6690.20150072
Meryem M, Younes A, Leila M, Mohammed A, Salwa EAT, Abdelkarim FM (2023) Development of a core collection for Tetraclinis articulata using ISSR markers and maximization strategy. Plant Mol Biol Rep 41(3):427–439. https://doi.org/10.1007/s11105-023-01372-y
Nascimento HL, Gomes RLF, Lopes ACA, Almeida RC, Carvalho LCB (2018) Selection of representative genotypes of lima bean (Phaseolus lunatus L.) from Brazil. Annual Report of the Bean Improvement Cooperative 61:201–202
Oliveira MF, Nelson RL, Geraldi IO, Cruz CD, Toledo JFF (2010) Establishing a soybean germplasm core collection. Field Crop Res 119:277–289. https://doi.org/10.1016/j.fcr.2010.07.021
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research - an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Penha JS, Lopes ACA, Gomes RLF, Pinheiro JB, Assunção-Filho JR, Silvestre EA, Viana JPG, Martínez-Castillo J (2017) Estimation of natural outcrossing rate and genetic diversity in Lima bean (Phaseolus lunatus L. var. lunatus) from Brazil using SSR markers: implications for conservation and breeding. Genet Resour Crop Evol 64:1355–1364. https://doi.org/10.1007/s10722-016-0441-9
Penha JS (2018) Diversidade genética, domesticação e plasticidade fenotípica de feijão- fava (Phaseolus lunatus L.). PhD Thesis, Universidade de São Paulo
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959 (PMID: 10835412)
Rogers JS (1972) Measures of genetic similarity and genetic distance. In: Studies in genetics VII. University of Texas, Austin
Seidu KT, Osundahunsi OF, Osamudiamen PM (2018) Nutrients assessment of some lima bean varieties grown in southwest Nigeria. Int Food Res J 25:848–853
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Silva RNO, Burle ML, Pádua JG, Lopes ACA, Gomes RLF, Martínez-Castillo J (2017) Phenotypic diversity in lima bean landraces cultivated in Brazil, using the Ward-MLM strategy. Chil J Agric Res 77:35–40. https://doi.org/10.4067/S0718-58392017000100004
Silva RNO, Lopes ACA, Gomes RLF, Pádua JG, Burle ML (2019) High diversity of cultivated lima beans (Phaseolus lunatus) in Brazil consisting of one Andean and two Mesoamerican groups with strong introgression between the gene pools. Genet Mol Res 18(4):gmr18441. https://doi.org/10.4238/gmr18441
Snedecor GW, Cochran WG (1980) Statistical Methods. Iowa State University, Ames
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30(12):2725–2729. https://doi.org/10.1093/molbev/mst197
Tanabata T, Shibaya T, Hori K, Ebana K, Yano M (2012) SmartGrain: High-throughput phenotyping software for measuring seed shape through image analysis. Plant Physiol 160(4):1871–1880. https://doi.org/10.1104/pp.112.205120
Van Hintum TJL, Brown AHD, Spillane C, Hodgkin T (2000) Core collections of plant genetic resources. IPGRI Technical Bulletin, International Plant Genetic Resources Institute, Rome
Vasconcelos ES, Cruz CD, Bhering LL, Ferreira A (2007) Estratégias de amostragem e estabelecimento de coleções nucleares. Pesq Agropec Bras 42(4):507–514. https://doi.org/10.1590/S0100-204X2007000400008
Vasconcelos ES, Cruz CD, Silva DJH, Bhering LL (2010) Tamanho de coleção original, métodos de agrupamento e amostragem para obtenção de coleção nuclear de germoplasma. Pesq Agropec Bras 45(12):1448–1455. https://doi.org/10.1590/S0100-204X2010001200016
Xu Y, Chen C, Ji D, Xu K, Xie X, Xie C (2016) Developing a core collection of Pyropia haitanensis using simple sequence repeat markers. Aquaculture 452:351–356. https://doi.org/10.1016/j.aquaculture.2015.11.016
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, methodology and analysis were performed by R.C.A., L.C.B.C., A.A.P., V.B.S., M.I.Z., J.B.P., J.M.C., A.C.A.L. and R.L.F.G. The first draft of the manuscript was written by R.C.A. and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethic Approval and Consent to Participate
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Message
• For the first time, a lima bean core collection was established with molecular and phenotypic data.
• The logarithmic method is better than the proportional one for establishing the lima bean core collection.
• Morpho-agronomic data were fundamental for the selection of lima bean accessions.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
da Costa Almeida, R., Carvalho, L.C.B., Alves-Pereira, A. et al. Establishment and Molecular Validation of a Lima Bean (Phaseolus lunatus) Core Collection in Brazil. Plant Mol Biol Rep (2024). https://doi.org/10.1007/s11105-024-01486-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11105-024-01486-x