Abstract
Rice is highly sensitive to salt stress at flowering stage. With the objective of detection of quantitative trait loci (QTLs) in multi-environment for this stage, 180 backcross-derived lines (BC3F5) from salt tolerant donor Pokkali (AC41585) and recurrent parent IR 64 were subjected to evaluation in saline (EC = 8 dSm−1) and non-saline environments in wet season of 2014 and 2015 employing a novel phenotyping protocol. Nine multi-environmental consistent QTLs for spikelet degeneration, K+ concentration in flag leaf, stress susceptibility index for grain (SSI-Grain) and spikelet sterility (SSI-STE) on chromosomes1, 2, 3, 4 and 11 with 17–42% phenotypic variances were detected. Among several digenic epistatic interactions, one was associated with the main effect QTL (qSSI-STE-11-1) over the years. Similarly genotype × environment interaction associated with two additive QTLs, qDEG-S-2-2 and qSSI-STE-2-1 had positive effect on the resultant phenotype. Functional genes encoding calmodulin-binding protein and potassium transporter were predicted inside the consistent QTLs. Detected stable QTLs, associated markers, predicted genes and derived introgression lines with these QTLs could be utilized in future breeding programme.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salinity is now becoming a wide spread problem for rice cultivation in India and other rice-growing countries in the world (Hossain et al. 2015). Rice is mainly susceptible to salt stress at early vegetative and reproductive stages. More than 100 quantitative trait loci (QTLs) for salt stress tolerance have been mapped in all 12 chromosomes of rice, mainly for the seedling stage (Kumar et al. 2015). From Pokkali, one QTL, named Saltol, was detected and incorporated to high yielding backgrounds for imparting salt tolerance at seedling stage (Islam et al. 2012). Nevertheless, eventually, it failed to guarantee satisfactory yield under prolonged salinity stress imposed beyond the reproductive stage. Unfortunately, no significant progress has been made so far to impart reproductive stage salt tolerance in rice due to relatively more complexity of the trait, genotype × environment interaction, lack of indicator physiological traits and a well-accepted screening protocol. Research efforts have been reported to be biased towards understanding seedling stage salinity tolerance, and through the reproductive stage, salinity tolerance is equally important for reducing yield loss (Ganie et al. 2019). Robust QTLs and markers are practically unavailable for reproductive stage salinity tolerance in rice. Under salinity stress plant yield was found to be positively associated with the numbers of panicles per plant, panicle length and harvest index and negatively associated with the percentage of spikelet sterility and degeneration (Zeng et al. 2002; Surekha et al. 2008; Munns and Tester 2008; Chattopadhyay et al. 2017). Apart from that, ion homeostasis and photo-phosphorylation were also reported to play an essential role at reproductive stage salinity tolerance in rice (Hossain et al. 2015; Razzaque et al. 2017; Chattopadhyay et al. 2018). Compared to the seedling stage, a few studies reported genetic analysis and QTL identification for reproductive stage salinity tolerance (Ammar et al. 2009; Pandit et al. 2010; Hossain et al. 2015; Reza et al. 2013; Chattopadhyay et al. 2013; Kumar et al. 2015; Tiwari et al. 2016). Ammar et al. (2009) mapped 25 major QTLs through bi-parental mapping for reproductive stage salt tolerance, using salinity tolerant donor CSR27. Similarly, Pandit et al. (2010) using CSR 11 and CSR 27 mapped QTLs on chromosomes 1, 8 and 12. Later, 35 yield-related QTLs were identified using tolerant parent ‘Sadri’ for different yield attributing traits (Reza et al. 2013). From ‘Cheriviruppu’, a salt tolerant genotype, 16 QTLs for pollen fertility, Na+ concentration and flag leaf Na+:K+ ratio were detected in chromosomes 1, 7, 8 and 10 using a modified control screening protocol (Hossain et al. 2015). But none of the QTLs was stable over multi-environmental conditions. In the case of many abiotic stresses such as drought, genotype × environment interaction QTL was detected at reproductive stage as one of the important determinants which influenced the main effect QTL (Kumar et al. 2014). In spite of significant genotype × environment interaction effect in salinity stress (Chattopadhyay et al. 2018), no G × E interaction QTL was reported so far. The lack of reproducible screening protocol for this stage was a bottleneck for detection of robust QTL.
We have standardized, authenticated and employed a novel screening protocol (Chattopadhyay et al. 2018) for validation of tolerant donor, AC41585, and precise phenotyping of BC3F5 population derived from IR64 × AC41585. The present study was aimed to detect not only putative QTLs in single environment, but also stable multi-environment QTLs of the most important component traits of salinity tolerance at reproductive stage, using the novel phenotyping protocol. In addition, with the understanding of the effect of genotype × environment interaction QTLs and epistatic QTLs either in isolation or in association with the main effect QTLs, we set out to detect their presence in important component traits. Subsequent objective of the study was to predict functional genes underlaying the consistent QTLs and finally to understand the inheritance and scope of further improvement of the reproductive stage salt tolerance, a complex trait.
Materials and Methods
Plant Materials
Collected 37 ‘Pokkali’ accessions were sowing varying level of salinity tolerance at reproductive stage with 20–82% yield reduction under stress (EC = 8 dSm−1). One of them AC41585 was identified as a tolerant germplasm under salinity stress at reproductive stage as realized from its low yield reduction (< 25%) in salinized medium as compared to non-salinized condition (Chattopadhyay et al. 2013). Subsequently, we found that this accession was having high K+ uptake potential even under low K+ environment and a good Na+ excluder as supported by higher HKT expression at the reproductive stage (Chakraborty et al. 2019). Therefore, for mapping, we have used this unique germplasm which differed from many other Pokkali accessions especially for its better tolerance to salinity stress at reproductive stage. IR 64 was crossed with AC41585 and the F1 was backcrossed consecutively for three generation with recurrent parent IR 64. BC3F1 was selfed and single-seed descent method was followed to develop BC3F4 population. One hundred eighty BC3F4 lines were preliminary evaluated under salinity stress and non-stress condition in standard evaluation method (Gregorio et al. 1997). The population was normally distributed and found diverse in nature (Chattopadhyay et al. 2017). For the present experiment, 180 lines belonged that to more advanced generation (BC3F5) were employed in QTL analysis using modified precise phenotypic platform (Chattopadhyay et al. 2018).
Experimental Setup
The evaluation of mapping population was conducted at the net house which was protected against the rain by transparent shading (> 80% light transmission) in wet season 2014 and 2015 at the ICAR-National Rice Research Institute, Cuttack (20.5°N latitude and 85.83°E longitude). Precise phenotyping protocol standardized and validated in our research station (Chattopadhyay et al. 2018) was employed in the present study. In this modified setup, the composition of the growing medium ‘soil:stone (4:1)’ was modified over and above the existing soil medium (Gregorio et al. 1997). Here, 20% volume of the soil was substituted by three different sizes of gravels for stabilization and maintaining the uniformity of soil EC inside pots without changing the buffering capacity of soil. Soil medium with substituted by 20% gravels ‘soil:stone (4:1)’ has considerably higher porosity and therefore saturated much earlier than the soil alone medium by desired level (8 dSm−1) of salinity (Chattopadhyay et al. 2018). Two seedlings of 21 days old were planted in each of the perforated pot filled with fertilized soil and gravels. Three plots were taken for each of the lines under both salinized and non-salinized conditions. Pots were placed inside the plastic tub filled with water. One set of tubs was salinized mixing with required quantity of NaCl with water and water EC (ECiw) was maintained at 8 dSm−1 inside pots. A perforated PVC pipe was placed inside soil with its opening outside soil layer of each the perforated pot for continuous monitoring of soil EC and pH. Genotypes were grouped based on their maturity duration (flowering time within a 5-day interval) as detected in the previous experiments (Chattopadhyay et al. 2017). Salinity treatment with 8 dSm−1 saline water in tub was stated 14 days before booting.. The stress was continued until 7 days before grain maturity. The duration of stress at 8 dSm−1 was maintained at least for 45–50 days (Supplementary Fig. 1).
Data Recording
Data were recorded from each plant in each pot in both salinized and non-salinized environments in both wet season 2014 and 2015 of plant height (cm) (PH), days to 50% flowering (DAF), number of panicles per plant (PN), panicle length (cm) (PL), number of grains per panicle (Grain), harvest index (HI) and plant yield (g) (PY). Spikelet sterility (STE) was calculated by dividing number of unfilled grains to the total number of spikelets and reported in percentage. Vestiges of rudimentary rachis branches left on the panicle were counted as degenerated spikelets (DEG) at maturity and expressed in percentage of the total spikelets (Saha et al. 1998). The recorded data of 2 years, under saline and non-saline environments, were subjected to analysis of variances and Shapiro-Wilk test for validity of normal distribution using SPSS v. 15 software and the least significant difference (p < 0.05) was found statistically significant. To evaluate genotypes based on reaction to salinity stress, yield contributing traits and yield per se in individual environments are not enough. Salt susceptibility index (SSI) and yield stability index (YSI) for each genotype were found effective in identification of genotypes in stressed and non-stressed environments (Chattopadhyay et al. 2017). SSI is calculated using the following formula.
Stress susceptibility index (SSI) = (1 − Ysi/Ypi)/SI; SI = 1 − Ys/Yp (Fischer and Maurer 1978) (here Ysi = PY or yield attributing traits under stress, Ypi = PY or yield attributing traits under non-stress, Ys and Yp are mean yield/yield traits of all lines in this experiment under stress and normal conditions, respectively). Tolerance to salt stress at reproductive stage was also estimated through yield stability index (Bouslama and Schapaugh 1984) for yield and other important morpho-physiological traits using the following formula.
Na+ and K+ concentrations in flag leaf were detected using a flame photometer by standard procedure (Yoshida et al. 1976).
For genotype × environment interaction studies, ANOVA for mixed effect model was done considering independent variables viz. genotype, treatment (stress level) and genotype × treatment as fixed effects and environment (year), blocks within environment and genotype × environment as random effects and component traits as the response variables using PROC GLM (Federer and Wolfinger 1996; Wolfinger et al. 1997). Using PROC SGPLOT procedure of SAS 9.3 software, graphs were plotted. The t test was used to detect significant differences if any for mean component traits in multi-years.
Selection of Markers and Genotyping
Around 1 g leaf sample of each of the 180 BC3F5 lines and their parents (IR 64 and AC41585) were used for DNA extraction and purification following CTAB method (Murray and Thompson 1980). The quantity and quality of genomic DNA of each sample were determined using 0.8% agarose gel. An aliquot of 20 ng/μl diluted gDNA of each sample was used for PCR. Twelve cgSSRs markers derived from salt responsive candidate genes (Molla et al. 2015), 100 highly informative hyper-variable SSRs with repeat length range of 51–70 bp mined from rice genome (Singh et al. 2009) and 700 type I and II SSR markers were tested for polymorphism between parents. Finally, among the all tested primers, 117 were found polymorphic between IR 64 and Pokkali (AC41585) which were distributed in all 12 rice chromosomes.
The polymerase chain reaction was done in a solution (25 μl) containing 10 mMTris-HCl buffer (pH 8.2), 50 mMKCl, 1.5 mM MgCl2, 0.01% gelatine, 200 μM dNTPs, 0.2 μM primers, 1 unit Taq DNA polymerase and 40 ng of the template DNA. The amplification reaction consisted of pre-heating for 5 min at 94 °C and 36 cycles of 1 min at 94 °C (denaturation), 1 min at 53–61 °C (annealing) and 1 min at 72 °C (elongation), followed by 5 min at 72 °C (extension) in a PCR system (Ependorf make). The amplified products were separated in 2% agarose gel containing 0.5 ng mL−1 of EtBr (ethidium bromide). The separated PCR products were made visible under a UV light and photographed in Systronics Gel documentation System.
Linkage Mapping and QTL Analysis
Linkage mapping was done using ‘MAP’ option in QTL IciMapping V3 (http://www.isbreeding.net). QTL mapping was done following inclusive composite interval mapping (ICIM) using ‘BIP’ option of this software. For identification of the main effect of additive and digenic epistatic QTLs in each environment and for each trait, the ‘ICIM-ADD’ and ‘ICIM-EPI’ functions, respectively, of the software were utilized (Meng et al. 2015). We used the ‘single marker analysis’ (SMA) option for identification of significant markers (> 5% PVE) associated with traits concerned. Permutation tests (1000 permutations, 95% confidence level, 1-cM interval) were performed for each trait in each environment. Logarithm of odds (LOD) score peaks ≥ 2.5 were used to declare the presence of a putative QTL in a given genomic region. The ‘multi-environment trials’ (MET) function of the software was also utilized to determine the consensus positions for the major QTL and identification of significant additive × environment interaction effect QTLs with> 5% of the variance.
Prediction of Probable Functional Genes inside QTLs and Graphical Genotyping of Introgressed Lines
Associated genes for salinity tolerance in rice were downloaded along with their physical position from Rice Annotation Project Database (Sakai et al. 2013) and Oryzabase (Kurata and Yamazaki 2006). Functionally validated genes related to increased salinity stress tolerance were also downloaded along with their physical positions from the gene information table available in QTL Annotation Rice Online Database (Yamamoto et al. 2012). The genes located inside the QTL interval region or near to peak marker position were considered to be probable causative genes for increased salinity tolerance at reproductive stage. Functions of the identified salinity tolerance QTL-linked genes were further determined using Rice Genome Annotation Project Database (Kawahara 2013) and Rice Annotation Project Database (Sakai et al. 2013). The physical locations of markers and robust QTLs in salinity tolerant introgressed lines were represented using Graphical GenoTyping (GGT 2.0) software (Van Berloo 2008).
Results
Phenotyping of Mapping Population
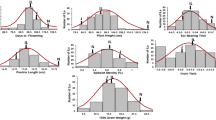
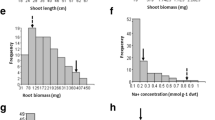
Analysis of variance (ANOVA) revealed significant differences (p < 0.01) among genotypes in mapping population for agronomical traits such as plant height (PH), days to 50% flowering (DAF), number of panicles/plant (PN), panicle length (PL), number of grains/panicle (Grain), spikelet sterility (STE), spikelet degeneration (DEG), harvest index (HI), plant yield (PY) and physiological traits such as Na+ concentration (Na), K+ concentration (K) and Na+–K+ ratio (Na–K) in flag leaf in both saline and non-saline conditions in the years 2014 and 2015. In both the years (2014 and 2015), correlation coefficient matrix (Table 1) revealed that plant yield under both the saline and non-saline conditions (PY) was positively and significantly (p < 0.01) associated with PN, Grain and HI and negatively associated with STE. In addition to that only at saline environment, PY also positively associated with PL (r = 0.286 and 0.143). Therefore, all these traits were detected as important component traits influencing plant yield and salt tolerance at flowering stage. On the other hand, low K+ concentration and high Na+ and Na–K ratio in flag leaf were significantly associated with grain sterility under salinity stress at flowering stage. PY, PL, PN, Grain, HI, DEG, STE and K under saline condition and stress susceptibility index (SSI) of these traits in both the years were normally distributed (Fig. 1). Most of the traits showed near or below unity skewness (Supplementary Table 1) which indicated their suitability for QTL analysis. Yield stability index (YSI) for plant yield for all lines were estimated over the years. A few lines such as RST-142, -4, -192, -100, -188, -123, -177, -19 and -5 were detected with transgressive segregation over tolerant parent, AC41585, for tolerance to salinity stress at reproductive stage based on high yield stability index (YSI) for grain yield. They were also high in grain yield under stress condition. Some of the lines such as RST-36, -103, -35, -94, -12, -140, -90, -118 and -191 were also detected with transgressive segregation for susceptibility over the recipient susceptible parent IR 64. They had very low yield under stress (Supplementary Table 2). Experimental plants (along with panicles in inset) belonging to two each of tolerant lines and susceptible lines along with their parents (IR64 and AC41585) raised under salinity stress (8 dSm−1) condition are displayed in Fig. 2.
Analysis of variance was done for genotype, environment and genotype × environment components. Environment was further partitioned into variance due to treatments (non-saline and saline) and years (2014 and 2015). Genotype, treatment and genotype × treatment interaction (GEI) components of variance were found significant (p < 0.001) for PN, PY, HI, DEG and STE (Table 2). But variances due to genotype × year component for all these traits were non-significant. GEI interaction plots for these traits were presented in Fig. 3. The trend lines showed that for all these traits, all environments were not parallel, indicating the presence of GEI effect. .
Genotype × treatment (1: saline, 2: non-saline) interaction for a panicle length, b panicle number, c spikelet sterility, d spikelet degeneration, e harvest index and f plant yield and genotype × year (1: 2014 and 2: 2015) interaction of of g panicle length, h panicle number, i spikelet sterility, j spikelet degeneration, k harvest index and l plant yield under four different environments of mapping population derived from IR 64/AC41585 in rice
Genotyping and Linkage Mapping
The mean frequencies of IR 64 (A) allele and AC41585 (B) allele derived by all polymorphic markers were 71.61% and 20.52%, respectively (Supplementary Table 3). The range of IR 64 genome in whole mapping population was 46.2–86.5% with an average of 68.63%. On the other hand, the genome share of the donor parent (AC41585) was 12.8–39.6% with an average of 23.56%. The average heterozygosity in this population was 5.53% (Supplementary Table 4). Among the population, the introgression line, RST-192, having highest (82%) genomic similarity with IR 64 was found tolerant as evident from high grain yield under salinity stress and high yield stability index for plant yield (0.88). Polymorphic 117 SSR markers distributed in 12 linkage groups under 12 rice chromosomes were used for linkage mapping. Map covered a genetic distance of 1235.53 cM with an average distance of 10.21 cM per marker. The highest and lowest map distances were found in chromosome 4 (192.97 cM) and chromosome 10 (52.76 cM), respectively.
Main Effect QTLs
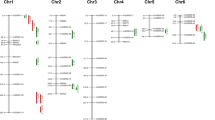
In the year 2014, 20 QTLs were identified for two traits (DEG and STE) in salinity stress condition and stress susceptibility index (SSI) of three traits (PN, STE and Grain). They were distributed in all chromosomes except chromosomes 7, 8 and 10. A total of 47 main effect QTLs were identified in 2015 (Supplementary Table 5). They were for five traits (DEG, PN, STE, Grain and K) in saline situation and stress susceptibility index (SSI) of four traits (PN, STE, Grain and HI). They were also distributed in all chromosomes except chromosomes 5 and 8 (Fig. 4). Therefore, we identified putative QTLs for salinity tolerance at reproductive stage in all chromosomes except chromosome 8. Among putative QTLs identified in the present study in 2014, nine were reproducible in nature and also were detected in 2015 (Table 3).
In 2014, two putative QTLs were found in chromosomes 4 and 12 under saline situation and another two putative QTLs were found in 2015 on chromosomes 1 and 10 for spikelet sterility (STE). On the other hand, for stress susceptibility index of this trait (SSI-STE), 14 putative QTLs were detected in 2014 on chromosomes 1, 2, 3, 4, 5, 6, 9, 11 and 12 while another 10 putative QTLs in 2015 on chromosomes 2, 3, 4 and 11. Among them, five reproducible QTLs on chromosomes 2, 3, 4 and 11 (qSSI-STE-2-1, qSSI-STE-2-2, qSSI-STE-3-1, qSSI-STE-4-3, qSSI-STE-11-1) were identified over the years. They explained 17–42% PVE with LOD score of 3.3–9 (Table 3). For Spikelet degeneration (DEG), two putative QTLs on chromosomes 2 and 4 and nine putative QTLs on chromosomes 2, 3, 4, 7, 9, 11 and 12 were found in the years 2014 and 2015, respectively, under salinity stress. Among them, two QTLs on chromosomes 2 (qDEG-2-2) and 4 (qDEG-4-1) were consistent over the years. They explained 32–37% PVE with LOD score of 4.2–6.4. In 2014, one QTL (qSSI-Grain-2-1) for stress susceptibility index for grain (SSI-Grain) was found on chromosomes 2 which almost co-localized with a QTL for DEG (qDEG-2-2). This was consistent over the years and explaining 42% PVE with LOD score of 6.2. Another six putative QTLs for SSI-Grain distributed on chromosomes 4, 6, 9, 11 and 12 and one putative QTL under stress on chromosome 2 were found in 2015. One QTL for K+ concentration in flag leaf (qK-S-1-1) found in 2015 on chromosome 1 in between markers, RM 9 and Os01g32120, explaining 37% phenotypic variation was also detected in the year 2014. Except for two QTLs for spikelet degeneration (qDEG-2-2 and qDEG-4-1), all positive alleles for salinity tolerance at reproductive stage were contributed by Pokkali (AC 41585) (Table 3).
Seven and four markers in the years 2014 and 2015, respectively, were identified through single marker analysis (Supplementary Table 6 and 7). Two of them, RM 17016 on chromosome 1 and HvSSR06-63 on chromosome 6, associated with spikelet sterility (STE) under saline condition were found consistent over the years with significant phenotypic variation (13–18%) (Supplementary Fig. 2).
Epistatic QTLs
Several significant digenic epistatic interaction loci combinations for all the traits were identified. A total of 79 significant epistatic QTLs in 2014 (Supplementary Table 8) and 314 significant epistatic QTLs in 2015 (Supplementary Table 9) for all component traits and for their stress susceptibility index were identified. Among them, six epistatic QTLs for SSI-STE were found common over the years with a range of negative (− 4%) to positive (3%) additive × additive interaction. Epistatic QTLs for SSI-STE, DEG and K in the year 2015 is presented in Fig. 5. Most of the epistatic interaction QTLs were not associated with main effect QTLs. In 2014, among 79 pairs, only five pairs of epistatic QTLs for SSI-STE were observed to be associated with main effect QTLs for this trait. An epistatic interaction was observed for QTL flanked by markers RM 9-Os01g32120 and RM17115-RM 2636 and was found to be co-located with qSSI-STE-1-1 and qSSI-STE-4-1, respectively. Another four pairs of epistatic QTLs were co-localized with main effect QTLs pairs such as qSSI-STE-1-1 and qSSI-STE-11-1, qSSI-STE-2-2 and qSSI-STE-11-1, qSSI-STE-11-1 and qSSI-STE-12-1, qSSI-STE-1-1 and qSSI-STE-12-1. They had positive and negative interactions with the main effect QTLs according to the similar or opposite sign of their additive values. Only one epistatic QTL for SSI-STE was found common over the years in association with main effect QTL (qSSI-STE-11-1). A similar sign in additive value indicated its positive effect on the main effect QTL.
Epistatic QTLs detected in Wet season 2015 for K+ concentration in flag leaf, spikelet degeneration and stress susceptibility index for spikelet sterility through analysis of a backcross-derived mapping population from IR 64/AC41585 (Note: dotted lines with yellow colour- K+ concentration, red colour—spikelet degeneration, green colour—stress susceptibility index for spikelet sterility)
Genotype × Environment Interaction QTLs
Significant interaction of genotype × environment (genotype × treatment) was found for PN, PY, HI, STE and DEG (Table 2). A total of 38 significant G × E QTLs (AE-QTLs) were identified for these traits (Table 4). A large variation in narrow sense heritability (h2) of these QTLs was found indicating their variation for inheritance. An AE-QTL for spikelet degeneration located on chromosome 7 in between RM 5436 and RM 180 showed the highest heritability (h2 = 0.992) (Table 4). Among these AE-QTLs, five for PY, seven for DEG, seven for HI, eight for PN, nine for STE (Fig. 6) and one for SSI-STE were detected in all chromosomes except chromosomes 8 and 11 with a range of 0.3 to 8.9% phenotypic variance. One AE-QTL for DEG was located on the main effect QTL (qDEG-S-2-1). Another AE-QTL for PN was also located on main effect QTL, qSSI-STE-2-1. These two additive x environment interaction QTLs with positive effects (1.3–3.5%) might enhance the phenotypic variances due to main effect QTLs (qDEG-S-2-1 and qSSI-STE-2-1), located in similar position.
Identification of Functional Genes in QTL Region
Inside qK-S-1-1 (QTL region for K+ content in flag leaf) in a 21.9-Mb region on chromosome 1, a functional gene LOC_Os01g38980.1encoding a calcium-modulating protein or calmodulin-binding protein (CaM) was found 1.15 cM apart. Two genes LOC_Os01g38980.1 and LOC_Os01g38980 encoding calmodulin-binding protein were located 2.7 cM apart from the significant associated marker, RM 17016, for STE-S located in 21.62-Mb region on chromosome 1. Inside the region of another consistent QTL qSSI-STE-2-1 in a 19.1-Mb region on chromosome 2, functional gene LOC_Os02g31910.1 encoding potassium transporter 1 was located 1.55 cM apart from the QTL peak. Apart from this functional gene, another probable functional gene Os02g33490.1 encoding pyrophosphate-energized membrane proton pump 3 was found (Supplementary Fig. 3) in this region (17–26 Mb) on chromosome 2. Around 0.5 cM apart from the another reproducible QTL peak (qSSI-STE-4-1) in a 19.83-Mb region on chromosome 4, a functional gene LOC_Os04g32920.2, encoding putative potassium transporter 1, was found. Just a 0.04-cM distance from the significant marker HvSSR06-63 for STE-S one gene LOC_Os06g45940.1 encoding HAK 13, potassium transporter 13 was located on a 27.8-Mb region on chromosome 6. Putative functional genes inside main effect QTLs and single marker analysis QTLs are presented in Table 5. Some tolerant lines (RST-100, -123, -142, -188 and -192) with higher YSI (0.83–0.91) than tolerant donor (0.75) and some susceptible lines (RST-12, -36, -90, 103 and -140) with lesser YSI (0.11–0.14) than susceptible parent IR 64 (0.19) (Supplementary Table 2) were subjected to graphical genotyping based on allelic distribution on chromosome 2. This revealed that tolerant line had AC41585 allele for the marker RM 1211 (Fig. 7) which was inside the region of a consistent QTL for SSI-STE (qSSI-STE-2-1). One tolerant line RST-192 (Fig. 8) had salinity tolerant allele from AC41585 of this QTLs. It also had another tolerant allele from AC41585 for Os01g32120 gene–based marker associated with another multi-environmental QTL qK-S-1-1 for K+ concentration in flag leaf.
Graphical genotyping of salt tolerant (RST-100, -123, -142, -188, -192) and susceptible lines (RST-12, -36, -90, -103, -140) of a backcross-derived population from IR 64/AC41585 showing allelic differences on QTL region on chromosome 2 for SSI-STE (qSSI-STE-2-1) (Note: legend A: allele from IR 64, B: allele from Pokkali (AC41585), C: heterozygote, D: missing)
Discussion
Reliable QTLs Using Unique Donor and Screening Protocol
A major bottleneck for identification of robust QTLs over the season is the ambiguity in phenotyping procedure which restricted the identification of proper donors and detection of tolerance reaction in mapping population. Prior to this experiment, we have demonstrated a novel phenotyping protocol for reproductive stage salt tolerance, which not only helped us in the detection of a reliable donor (AC41585) but also gave reproducible phenotypic information of a mapping population (Chattopadhyay et al. 2018). Tolerance level of donor parent at reproductive stage was further validated through molecular dissecting in relation to K+ uptake and its coordinated transport to flag leaf (Chakraborty et al. 2019). Reliable donor with valid salt tolerance mechanism at reproductive stage and validated high throughput phenotyping protocol made the present investigation on QTL detection more reliable than the previous occasions.
Governing Traits and Associated Novel QTLs As Well As Previously Detected Analogous QTLs
We found, the traits like higher spikelet sterility and degeneration and lower panicle and spikelet number are mainly responsible for substantial yield decline under salinity stress imposed before booting stage. In previous studies, the significant reduction of grain yield under salinity condition was found mainly due to reduction of panicle and spikelet number, increased spikelet sterility and spikelet degeneration or malformation (Zeng and Shannon 2000; Chattopadhyay et al. 2018). Many researchers reported that the set of genes and QTLs responsible for salinity tolerance at seedling and reproductive stage are different (Liu et al. 2017; Moradi and Ismail 2007). In barley, QTLs for salt tolerance at reproductive stage were detected for yield and agronomic characters (Xue et al. 2009; Eleuch et al. 2008) and stomatal and photosynthetic traits (Liu et al. 2017). In rice, several QTLs were detected for yield and yield attributing traits such as plant height, tiller number, panicle length, plant yield, biomass, pollen fertility, spikelet fertility, grain number per panicle, 1000 grain weight (Hossain et al. 2015; Reza et al. 2013) and physiological traits such as Na+ concentration and Na–K ratio in flag leaf (Hossain et al. 2015; Pandit et al. 2010; Kumar et al. 2014). In the present study, we used stress susceptibility index as an important determinant to identify the effect of component traits on tolerance reaction under stress condition at reproductive stage. We got five consistent QTLs over the seasons on chromosomes 2, 3, 4 and 11 for stress susceptibility index for spikelet sterility, which indirectly represented grain fertility (Table 3). All these identified QTLs are novel since they have not been reported earlier. In this paper, we are reporting for the first time two consistent QTLs for spikelet degeneration (qDEG-S-2-2, qDEG-S-4-3) which were supposed to be associated with low number of fertile spikelets under stress condition. Besides, we found one QTL for SSI-Grain (qSSI-Grain-2-1) (Table 3). Reduction of grain number was proposed to be due to higher accumulation of Na+ and lower K+ in floral parts and leaves which reduced the availability of glucosyle donors for starch synthetase activity in developing grains. It resulted in the accumulation of glucose on floral parts and failure of seed set (Abdullah et al. 2001). We also detected one consistent QTL for K+ concentration in flag leaf (qK-S-1-1). Near to this QTL, Saltol for seedling stage tolerance (Bonilla et al. 2002) and some other putative QTLs for reproductive stage (Hossain et al. 2015) were found. Using custom-designed array based on 6000 SNPs, Kumar et al. (2015) identified 20 loci associated with Na–K homeostasis. They found Saltol as the major salt tolerance QTL not only for seedling stage, but also for reproductive stage in relation to Na+–K+ ratio in leaves. Lower K+ concentration in plants under saline environment reduced the plant growth regulator activity caused in spikelet degeneration leading to reduction of spikelet number (Yokoyama et al. 2002). Moreover, K+ concentration was proved as an important determinant of tolerant genotypes under salinity stress at flowering stage (Chattopadhyay et al. 2018). Using 50K SNP chip, Tiwari et al. (2016) found three QTLs in between 22.4- and 26.8-Mb positions for stress susceptibility index for grain yield (SSIGY) in sodicity and salinity environment. In the similar position (25.8 Mb), we have detected two multi-environmental and pleiotropic QTLs, qSSI-Grain-2-1 and qDEG-S-2-2. Another two consistent QTLs, qSSI-STE-2-1 and qSSI-STE-2-2, were also located in between 17- and 19-Mb regions. Therefore, in the region between 17 and 26 Mb, a cluster of QTLs was detected (Table 5, Supplementary Fig. 3). Another QTL qSSI-STE-3-1 located at 34-Mb regions on chromosome 3 was found closer to a QTL for SSIGY detected by Ammar et al. (2009). Apart from this, no other multi-environmental QTL was found analogous with previously detected QTLs.
Novel Epistatic and Genotype × Environment Interaction QTLs for Salinity Tolerance at Reproductive Stage
In the present study, we could able to detect 393 epistatic QTLs for all traits and their stress susceptibility index in 2 years (Supplementary Table 8, 9). Among associated epistatic QTLs with main effect additive QTLs, only one was found common over the years (qSSI-STE-11-1). Similar sign in additive value indicated its positive effect on the main effect QTL. Although epistatic QTLs for different component traits were detected (Hossain et al. 2015; Kumar et al. 2014), multi-environmental epistatic effect on a consistent QTL was not reported earlier for salinity tolerance at reproductive stage.
In the present study, nine putative QTLs detected in one season (year 2014) were validated in the next season (year 2015) (Table 3). In comparison to previous investigations in relation to salinity tolerance at reproductive stage (Hossain et al. 2015; Reza et al. 2013; Kumar et al. 2014), this study was unique for detection of all novel QTLs which were multi-environment in nature. Generally, polygenic trait like agro-physiological parameters related to salinity stress tolerance was influenced by the environment. Therefore, inheritance of QTLs with additive effect could be influenced by additive × environment interaction variance as was observed in rice for salinity tolerance at seedling stage (Rahman et al. 2017). The effect could be either positive or negative in nature. Therefore, these additive × environment interaction QTLs even with small effect were required to detect. But we could not find such reports in earlier publication on detection of QTLs for salinity tolerance at reproductive stage in rice. We in this paper report such QTLs. G × E QTLs sharing with two additive QTLs, qDEG-S-2-1 and qSSI-STE-2-1 had synergistic effect and adding to their inheritance.
Bioinformatics Analysis Resulting in Detection of Putative Functional Genes
We found CaM functional genes (LOC_Os01g38980, LOC_Os01g38980-1/Os01g0570800) in the region of qK-S-1-1, a QTL for K+ concentration at flag leaf (Table 5). Eventually, the marker (cgSSR) present in this QTL region was derived from a Ca2+-binding regulatory gene, Os01g32120 (Molla et al. 2015; Chinpongpanich et al. 2012). This result is quite encouraging, given the known role of Ca2+ signalling and cellular Ca2+ homeostasis in salt tolerance ability of the plants. Previous studies reported that cytosolic Ca2+ signalling is indispensable for plants adaptation to salinity stress (Dodd et al. 2010). Chakraborty et al. (2016) shown that maintenance of Ca2+ homeostasis by coordinated regulation of efflux and influx of Ca2+ in the cell is related to superior K+ retention and salt tolerance ability in Brassica species under saline environment. Not only tissue K+ retention, Ca2+-mediated signalling governs at least few more concurrent signalling loops viz. salt induced signalling of CBL4 (SOS3) to activate SOS1-driven Na+ exclusion process in rice (Martinez-Atienza et al. 2007). Besides, modulation in Ca2+ signature also governs NSCC (non-selective cation channels)-mediated Na+ transport (Demidchik and Tester 2002) and prevents K+ leakage from the cell (Shabala et al. 2006; Chakraborty et al. 2018). Razzaque et al. (2017) also found that in tolerant germplasm, Horkuch, roots showed upregulation of cation transporters and constitutively expressed genes regulating membrane potential. Functional gene LOC_Os02g31910.1 (Os02g0518600) encoding potassium transporter 1 was postulated inside a consistent QTL qSSI-STE-2-1 in a 19.1-Mb region on chromosome 2. In high external Na+ condition, there prevails an apparent K+ limitation in plants due to competition of both ions in same entry points. Under such scenario, KT/KUP/HAK family of K+ transporter was reported to play an important role in uptake of K+ from K+-limited environment and in its tissue specific redistribution (Chen et al. 2015; Chen et al. 2017). HvSSR06-63 was identified as a significant marker across the environment in the region of functional gene LOC_Os06g45940.1(Os06g0671000) encoding HAK 13 (Schmidt et al. 2013) associated with spikelet sterility under salinity (Table 5, Supplementary Fig. 2) as increased K+ content and maintenance of lower Na+/K+ ratio in flag leaf and developing panicle is absolutely important for reproductive stage salt tolerance in rice (Chakraborty et al. 2019).
Conclusion
The present study detected nine novel multi-environmental additive QTLs and one multi-environmental epistatic QTL in 8 dSm−1 salinity stress during reproductive stage in rice. Among these QTLs, seven QTLs were found for such component traits, like spikelet degeneration, stress susceptibility index and spikelet sterility for which no QTL was reported previously. This investigation also reported for the first time significant genotype × environment interaction QTLs for different component traits, a few of them also positively influenced the inheritance of the main effect additive consistent QTLs such as qDEG-S-2-1 and qSSI-STE-2-1. Inside the consistent QTLs, a few functional genes such as LOC_Os01g38980, LOC_Os02g31910.1 and LOC_Os06g45940.1 were postulated. Some introgression lines with tolerant alleles from the donor parent, AC41585, were detected and they could be used as pre-breeding lines for transferring the consistent QTLs in the high yielding background.
References
Abdullah Z, Khan A, Mushtaq, Flowers TJ (2001) Causes of sterility in seed set of rice under salinity stress. J Agron Crop Sci 187:25–32. https://doi.org/10.1046/j.1439-037X.2001.00500.x
Ammar MHM, Pandit A, Singh RK, Sameena S, Chauhan MS, Singh AK, Sharma PC, Gaikwad K, Sharma TR, Mohapatra T, Singh NK (2009) Mapping of QTLs controlling Na+, K+ and Cl- ion concentrations in salt tolerant indica rice variety CSR27. J Plant Biochem Biotechnol 18:139–150
Bonilla P, Dvorak J, Mackill D, Deal K, Gregorio G (2002) RFLP and SSLP mapping of salinity tolerance genes in chromosome 1 of rice (Oryza sativa L.) using recombinant inbred lines. Philip J Agric Sci 85:68–76
Bouslama M, Schapaugh WT (1984) Stress tolerance in soybean. Part. 1: evaluation of three screening techniques for heat and drought tolerance. Crop Sci 24:933–937
Chakraborty K, Bose J, Shabala L, Shabala S (2016) Difference in root K+ retention ability and reduced sensitivity of K+-permeable channels to reactive oxygen species confer differential salt tolerance in three Brassica species. J Exp Bot 67(15):4611–4625
Chakraborty K, Basak N, Bhaduri D, Ray S, Vijayan J, Chattopadhyay K, Sarkar RK (2018) Ionic basis of salt tolerance in plants: nutrient homeostasis and oxidative stress tolerance. In Hasanuzzaman M, Fujita M, Oku H, Nahar K, Hawrylak-Nowak B (eds.) Plant nutrients and abiotic stress tolerance. Springer, Singapore, pp 325– 362. https://doi.org/10.1007/978-981-10-9044-8_14
Chakraborty K, Chattaopadhyay K, Nayak L, Ray S, Yeasmin L, Jena P, Gupta S, Mohanty SK, Swain P, Sarkar RK (2019) Ionic selectivity and coordinated transport of Na+ and K+ in flag leaves render differential salt tolerance in rice at the reproductive stage. Planta 250:1637–1653. https://doi.org/10.1007/s00425-019-03253-9
Chattopadhyay K, Nath D, Das G, Mohanta RL, Marndi BC, Singh DP, Sarkar RK, Singh ON (2013) Phenotyping and QTL-linked marker-based genotyping of rice lines with varying level of salt tolerance at flowering stage. Indian J Genet 73(4):434–437
Chattopadhyay K, Marndi BC, Sarkar RK, Singh ON (2017) Stability analysis of backcross population for salinity tolerance at reproductive stage in rice. Indian J Genet 77(1):51–58. https://doi.org/10.5958/0975-6906.2017.00007.4
Chattopadhyay K, Nayak AK, Marndi BC, Poonam A, Chakraborty K, Sarkar RK (2018) Novel screening protocol for precise phenotyping of salt-tolerance at reproductive stage in rice. Physiol Mol Biol Plants 24(6):1047–1058. https://doi.org/10.1007/s12298-018-0591-7
Chen G, Hu Q, Luo LE, Yang T, Zhang S, Hu Y, Yu L, Xu G (2015) Rice potassium transporter OsHAK1 is essential for maintaining potassium mediated growth and functions in salt tolerance over low and high potassium concentration ranges. Plant Cell Environ 38:2747–2765. https://doi.org/10.1111/pce.12585
Chen G, Liu C, Gao Z, Zhang Y, Jiang H, Zhu L, Ren D, Yu L, Xu G, Qian Q (2017) OsHAK1, a high-affinity potassium transporter, positively regulates responses to drought stress in rice. Front Plant Sci 8:1885. https://doi.org/10.3389/fpls.2017.01885
Chinpongpanich A, Limruengroj K, Phean-o-pas S, Limpaseni T, Buaboocha T (2012) Expression analysis of calmodulin and calmodulin-like genes from rice, Oryzasativa L. BMC Res Notes 5:625–625
Demidchik V, Tester M (2002) Sodium fluxes through nonselective cation channels in the plasma membrane of protoplasts from Arabidopsis roots. Plant Physiol 128:379–387
Dodd AN, Kudla J, Sanders D (2010) The language of calcium signaling. Annu Rev Plant Biol 61:593–620
Eleuch L, Jilal A, Grando S, Ceccarelli S, Schmising MVK, Tsujimoto H, Hajer A, Daaloul A, Baum M (2008) Genetic diversity and association analysis for salinity tolerance, heading date and plant height of barley germplasm using simple sequence repeat markers. J Integr Plant Biol 50:1004–1014
Federer WT, Wolfinger RD (1996) Gauss and SAS for recovering inter block and inter variety information. Technical Report Series of the Biometrics Unit, Cornell University, Ithaca
Fischer RA, Maurer R (1978) Drought resistance in spring wheat cultivars. I. Grain yield response. Aus J Agric Res 29:897–907
Ganie SA, Molla KA, Henry RJ, Bhat KV, Mondal TK (2019) Advances in understanding salt tolerance in rice. Theo Appl Genet 132(4):851–870
Gregorio G B, Senadhira D, Mendoza R D (1997) Screening rice for salinity tolerance. IRRI discussion paper series no. 22. Manila (Philippines), International Rice Research Institute, pp. 1-30
Hossain H, Rahman MA, Alam MS, Singh RK (2015) Mapping of quantitative trait loci associated with reproductive-stage salt tolerance in Rice. J Agron Crop Sci 201:17–31
Islam MR, Gregorio GB, Salam MA, Collard BCY, Singh RK, Hassan L (2012) Validation of SalTollinked markers and haplotype diversity on chromosome 1 of rice. Mol Plant Breed 3(10):103–114
Kawahara Y et al (2013) Improvement of the OryzasativaNipponbare reference genome using next generation sequence and optical map data. Rice 6(1):4
Kumar A, Dixit S, Ram T, Yadav RB, Mishra KK, Mandal NP (2014) Breeding high-yielding drought-tolerant rice: genetic variations and conventional and molecular approaches. J Exp Bot 65(21):6265–6278. https://doi.org/10.1093/jxb/eru363
Kumar V, Singh A, Amitha Mithra SV, Krishnamurthy SL, Parida SK, Jain S, Tiwari KK, Kumar P, Rao AR, Sharma SK, Khurana JP, Singh NK, Mohapatra T (2015) Genome-wide association mapping of salinity tolerance in rice (Oryza sativa). DNA Res 22:1–13. https://doi.org/10.1093/dnares/dsu046
Kurata N, Yamazaki Y (2006) Oryza base: an integrated biological and genome information database for rice. Plant Physiol 140(1):12–17
Liu X, Fan Y, Mak M, Babla M, Holford P, Wang F, Chen G, Scott G, Wang G, Shabala S, Zhou M, Chen Z-H (2017) QTLs for stomatal and photosynthetic traits related to salinity tolerance in barley. BMC Genomics 18:9. https://doi.org/10.1186/s12864-016-3380-0
Martinez-Atienza J, Jiang X, Garciablades B, Mendoza I, Zhu JK, Pardo JM, Quintero FJ (2007) Conservation of the salt overly sensitive pathway in rice. Plant Physiol 143:1001–1012
Meng L, Li H, Zhang L, Wang J (2015) QTL Ici mapping: integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J 3:269–283
Molla KA, Debnath AB, Ganie SA, Mondal TK (2015) Identification and analysis of novel salt responsive candidate gene based SSRs (cgSSRs) from rice (Oryza sativa L.). BMC Plant Biol 15(1):1
Moradi F, Ismail AM (2007) Responses of photosynthesis, chlorophyll fluorescence and ROS-scavenging systems to salt stress during seedling and reproductive stages in rice. Ann Bot 99:1161–1173
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Bio 59:651–681
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325. https://doi.org/10.1093/nar/8.19.4321
Pandit A, Rai V, Bal S, Sinha S, Kumar V, Chauhan M, Gautam RK, Singh R, Sharma PC, Singh AK, Gaikwad K, Sharma TR, Mohapatra T, Singh NK (2010) Combining QTL mapping and transcriptome profiling of bulked RILs for identification of functional polymorphism for salt tolerance genes in rice (Oryza sativa L.). Mol Genet Genomics 284:121–136. https://doi.org/10.1007/s00438-010-0551-6
Rahman MA, Bimpong IK, Bizimana JB, Pascual ED, Arceta M, Swamy MAM, Diaw F, Rahman MS, Singh RK (2017) Mapping QTLs using a novel source of salinity tolerance from Hasawi and their interaction with environments in rice. Rice 10:47. https://doi.org/10.1186/s12284-017-0186-x
Razzaque S, Haque T, Elias SM, RahmanMd S, Biswas S, Schwartz S, Ismail AM, Walia H, Juenger TE, Seraj ZI (2017) Reproductive stage physiological and transcriptional responses to salinity stress in reciprocal populations derived from tolerant (Horkuch) and susceptible (IR29) Rice. Sci Rep 7:46138. https://doi.org/10.1038/srep46138
Reza M, Mendioro MS, Diaz GQ, Gregorio GB, Singh RK (2013) Mapping quantitative trait loci associated with yield and yield components under reproductive stage salinity stress in rice (OryzaSativa). J Genet 92:433–443
Saha A, Sarkar RK, Yamagishi Y (1998) Effect of time of nitrogen application on spikelet differentiation and degeneration of rice. Bot Bull Acad Sin 39:119–123
Sakai H, Lee SS, Tanaka T, Numa H, Kim J, Kawahara Y, Wakimoto H, Yang CC, Iwamoto M, Abe T, Yamada Y, Muto A, Inokuchi H, Ikemura T, Matsumoto T, Sasaki T, Itoh T (2013) Rice Annotation Project Database (RAP-DB): an integrative and interactive database for rice genomics. Plant Cell Physiol 54(2):e6–e6
Schmidt R, Mieulet D, Hubberten HM, Obata T, Hoefgen R, Fernie AR, Fisahn J, Segundo BS, Guiderdoni E, Schippers JHM, Mueller-Roeber B (2013) Salt-responsive ERF1 regulates reactive oxygen species-dependent signaling during the initial response to salt stress in rice. Plant Cell 25(6):2115–2131
Shabala S, Demidchik V, Shabala L, Cuin TA, Smith SJ, Miller AJ, Davies JM, Newman IA (2006) Extracellular Ca2+ ameliorates NaCl-induced K+ loss from Arabidopsis root and leaf cells by controlling plasma membrane K+-permeable channels. Plant Physiol 141:1653–1665
Singh H, Deshmukh RK, Singh A, Singh AK, Gaikwad K, Sharma TR, Mohapatra T, Singh NK (2009) Highly variable SSR markers suitable for rice genotyping using agarose gels. Mol Breed 25:359–364
Surekha R, Mishra PB, Gupta SR, Rathore A (2008) Reproductive stage tolerance to salinity and alkalinity stresses in rice genotypes. Plant Breed 127:256–261
Tiwari S, Krishnamurthy SL, Kumar V, Singh B, Rao A, Mithra SV, Rai V, Singh AK, Singh NK (2016) Mapping QTLs for salt tolerance in Rice (Oryzasativa L.) by bulked segregant analysis of recombinant inbred lines using 50K SNP chip. PLoS ONE 11(4):e0153610. https://doi.org/10.1371/journal.pone.0153610
Van Berloo R (2008) GGT 2.0: versatile software for visualization and analysis of genetic data. J Hered 99:232–236
Wolfinger RD, Federer WT, Cordero-Brana O (1997) Recovering information in augmented designs, using SAS PROC GLM and PROC MIXED. Agron J 89:856–859
Xue D, Huang Y, Zhang X, Wei K, Westcott S, Li C, Chen M, Zhang G, Lance R (2009) Identification of QTLs associated with salinity tolerance at late growth stage in barley. Euphytica 169:187–196
Yamamoto E, Yonemaru JI, Yamamoto T, Yano M (2012) OGRO: the overview of functionally characterized genes in rice online database. Rice 5(1):26
Yokoyama C, Tsuda M, Hirai Y (2002) Effects of plant growth regulators on number of spikelets per panicle in Rice (Oryzasativa L.) under saline flooding conditions. Crop Physiol Cell Biol 71(3):376–382
Yoshida S, Forno DA, Cock JH, Gomez KA (1976) Laboratory manual for physiological studies of rice, 3rd edn. IRRI, Los Banos, pp 61–66
Zeng L, Shannon MC (2000) Salinity effects on the seedling growth and yield components of rice. Crop Sci 40:996–1003
Zeng L, Shannon MC, Grieve CM (2002) Evaluation of salt tolerance in rice genotypes by multiple agronomic parameters. Euphytica 127:235–245
Funding
Authors acknowledge received funding from the Director, ICAR-National Rice Research Institute, Cuttack and ICAR funded project ‘National Innovation on Climate Resilient Agriculture’, New Delhi.
Author information
Authors and Affiliations
Contributions
KC made design of the experiment and drafted manuscript. SKM, JV and BCM implemented experiments and analyse data. RKS coordinated the study. KAM and JV assisted in selection of markers and linkage mapping. AS statistically analysed the data. SR analysed molecular data. KOC and RKS assisted in preparing manuscript and revised the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key message
• Employing a population using salt tolerant donor at reproductive stage, Pokkali (AC 41585), nine multi-environmental consistent QTLs for spikelet degeneration, spikelet sterility, K+ concentration at flag leaf, etc. were discovered.
• Two genotype × environment interaction QTLs and one digenic-epistatic QTL were detected with synergistic effect on the main effect QTLs.
Supplementary Information
ESM 1
(DOC 1414 kb)
Rights and permissions
About this article
Cite this article
Chattopadhyay, K., Mohanty, S.K., Vijayan, J. et al. Genetic Dissection of Component Traits for Salinity Tolerance at Reproductive Stage in Rice. Plant Mol Biol Rep 39, 386–402 (2021). https://doi.org/10.1007/s11105-020-01257-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-020-01257-4