Abstract
Aims
To understand the species composition, abundance and diversity of nitrogen-fixing bacteria in the Cenchrus fungigraminus rhizosphere and to screen nitrogen-fixing bacteria to study their potential role in plant growth promotion.
Methods
Soil were collected from 4 depths (G1, G2, G3 and G4) of the C. fungigraminus rhizosphere, and their physical and chemical properties were determined. The diversity and abundance of nitrogen-fixing bacteria and the nifH gene was analysed. Nitrogen-fixing bacteria were screened and selected to promote growth of C. fungigraminus seedlings.
Results
The highest diversity and abundance of nitrogen-fixing bacteria was observed in the G2 samples collected from the C. fungigraminus rhizosphere. These bacteria mainly included Proteobacteria (93.91%), Actinobacteria (0.42%), and Firmicutes (0.18%) and were significantly affected by total nitrogen, available nitrogen and soil depth. The nifH gene copy number was highest (1.56 ± 0.17 × 107 copies/g) in G2. Rhizobium pusense No. 8 and No. 28 were isolated from G1 and G2, with nitrogenase activities of 145.06 ± 4.10 and 199.78 ± 7.50 U/L. The promotion experiment revealed that the plant height, root length, and leaf length of C. fungigraminus seedlings treated with both strains significantly increased by 56.79%, 76.99% and 55.71%, and the soil moisture and total nitrogen were significantly increased compared to the control (P < 0.05). The available nitrogen, organic matter and organic carbon in soil with strain No. 28 significantly increased compared to CK.
Conclusion
Rhizobacteria in the C. fungigraminus rhizosphere play an important role as plant growth promoting rhizobacteria (PGPR). Two strains showed potential for the development of biological fertilizers.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Nitrogen is one of the most important elements affecting plant growth and development. Plants associate with microorganisms that are able to obtain molecular nitrogen from the atmosphere and convert it into ammonium nitrogen for direct use. Nitrogen-fixing bacteria are usually classified into symbiotic and nonsymbiotic bacteria (Tchan 1952). Symbiotic nitrogen-fixing microorganisms that form a symbiosis with legumes are the most efficient system for nitrogen fixation in nature and were first reported by Frank (1889) in stem- and root-nodulating bacteria. Furthermore, many rhizobial species, such as Rhizobium oryziradicis, Rhizobium rhizoryzae, Rhizobium pseudoryzae, Rhizobium oryzicola, and Rhizobium taibaishanense, have a nonsymbiotic association with crops (Zhang et al. 2015; Zhao et al. 2017). Currently, the problems regarding environmental contamination, energy shortages, food safety, etc., threaten sustainable agriculture. There is a need to reduce the use of chemical fertilizers. Recent studies have demonstrated that the host plant and its developmental stages play an important role in shaping the rhizosphere microbial community structure (Chaparro et al. 2014; Hou and Babalola 2013). The rhizosphere microbiome is mainly derived from plant roots and has been shown to enhance plant growth and plant stress tolerance under certain environmental conditions, such as soil type, soil nutrients, plant cultivar, climate change and anthropogenic activities (Igiehon and Babalola 2018; Vejan et al. 2016). Such bacteria are generally referred to as plant growth-promoting rhizobacteria (PGPR) (Lugtenberg and Kamilova 2009). PGPR have been shown to enhance plant growth by direct or indirect mechanisms such as biological nitrogen fixation, phosphate solubilization, siderophore production, rhizosphere engineering, and phytohormone production (Chabot et al. 1996). According to Nakkeeran et al. (2005) and Swarnalakshmi et al. (2020), ideal PGPR should possess high rhizosphere competence, enhance plant growth capabilities, display a broad spectrum of action, be safe for the environment, be compatible with other rhizobacteria, and be tolerant to heat, UV radiation, and oxidizing agents. PGPR have been shown to promote the growth of many crops, including maize (Ferrarezi et al. 2022), wheat (Majeed et al. 2015) and rice (Barraquio et al. 1997). Rhizobia are employed as PGPR to increase plant growth through phosphate solubilization, the fixation of nitrogen, and the production of ACC deaminase, siderophores and indole-3-acetic acid (Cavite et al. 2021).
Nitrogen-fixing genes (nif) encode nitrogen-fixing enzymes that catalyse nitrogen fixation. Approximately 149 nitrogen-fixing genes, including nifH, nifD, nifK, nifE, nifN and nifB, have been identified (Santos et al. 2012). The nifH gene encodes a key enzyme involved in nitrogen fixation, i.e., dinitrogenase, which is highly conserved and widely distributed among nitrogen-fixing bacteria. The nifH gene expression in plants at different growth stages was consistent with changes in the composition of nitrogen-fixing bacteria (Lin et al. 2021). Average annual rainfall and temperature were reported to be the main factors affecting changes in endophytic nitrogen-fixing bacteria in different regions (Jia et al. 2020). High-throughput sequencing technology is able to overcome the limitation of microbiological studies based on traditional pure culture and can be used to quantitatively analyse samples of noncultivable, dominant, and rare bacteria and can comprehensively and accurately reflect flora composition and abundance (Monteiro et al. 2016).
Cenchrus fungigraminus Z. X. Lin & D. M. Lin & S. R. Lan sp. Nov. (Pennisetum giganteum Z. X. Lin or Giant grass has been used before) is a new species and perennial C4 plant of the family Gramineae with high yield and arid adaptability (Lin et al. 2022). It was collected in South Africa and has been widely used in environmental management (Zhou et al. 2021), in the production of edible and medicinal mushrooms (Su et al. 2022), and as fodder (Liu et al. 2018), yielding a good economic benefit for rural areas. Currently, C. fungigraminus is grown in 31 provinces in China, including Inner Mongolia, Qinghai, Hainan, Ningxia, Guizhou, Sichuan and Fujian, and in 106 countries worldwide (Lin and Lin 2018). The total nitrogen content of C. fungigraminus may exceed its total nitrogen uptake from soil and fertilizer, presumably due to the presence of the endophytic nitrogen-fixing bacteria. Klebsiella and Bradyrhizobium that were shown to be the main nitrogen-fixing bacteria present in the roots and stems of C. fungigraminus in Hainan Province (Lin et al. 2019). In this study, we evaluated the diversity and abundance of nitrogen-fixing bacteria in the rhizosphere of C. fungigraminus in Hainan and screened the main PGPR to explore their potential effect on C. fungigraminus seedlings and soil. We hypothesized that the diversity and abundance of nitrogen-fixing bacteria in the C. fungigraminus rhizosphere would be significantly affected by soil depth and some strains may have the ability to promote plant growth as PGPR. Our results provide a theoretical basis for the screening and development of PGPR from the C. fungigraminus rhizosphere as biological fertilizers.
Materials and methods
Sample collection and processing
C. fungigraminus was planted in 2017 at the Danzhou Demonstration Base (22 m × 37 m) of the National Engineering Research Center of Juncao Technology in Hainan Province, People’s Republic of China (109°53’ E and 19°50’ N). In total, 4 groups of soil samples were collected in 2021 from the C. fungigraminus surface soil (G1), rhizosphere soil (1–5 mm from the plant roots; these samples were carefully shaken to collect the soil attached to the roots) (G2), 10 cm depth from the main root of C. fungigraminus (G3) and 20 cm depth from the main root of C. fungigraminus (G4). According to Li et al. (2020) and Xomphoutheb et al. (2020), G2 is rhizosphere soil, while G1, G3 and G4 are nonrhizosphere soil samples. Six natural soil replicates of each group were randomly collected from the demonstration base using a clean soil auger and placed in clean, labelled sample bags. Each sample was divided into two parts; one part was stored at 4 °C until physical and chemical analyses were performed. The other part was stored at -80 °C until further analysis. The collected soil samples were ground and sieved through a 2 mm sieve. DNA extraction from the soil samples was performed by a Soil DNA Isolation Kit (Invitrogen, USA) following the manufacturer’s instructions. The integrity of the extracted DNA was determined by 1% (w/v) agarose gel electrophoresis with SYBR Safe (Invitrogen, USA). The concentration of DNA was estimated by a Nanodrop spectrophotometer (Thermo, USA). The soil used for the plant promotion experiment was obtained from the National Engineering Research Center of Juncao Technology.
High-throughput sequencing and diversity analysis
The phylogeny of the nifH gene is significantly consistent with the presence of specific16S rRNA, with highly conserved sequences, abundant data information, and variable regions; this gene can be used to indicated the existence of nitrogen-fixing bacteria in samples and to reveal the relationship between the community structure of nitrogen-fixing bacteria and the environment (Cantera et al. 2004; Terakado-Tonooka et al. 2008). An amplified fragment of nifH, a functional gene for nitrogen fixation, was selected for subsequent high-throughput sequencing. The primer sequences were nifH-F: 5′-TGCGAYCCSAARGCBGACTC-3′ and nifH-R: 5′-ATSGCCATCATYTCRCCGGA-3′. The polymerase chain reaction (PCR) conditions were predenaturation at 98 °C for 5 min, denaturation at 98 °C for 30 s, and annealing at 55 °C for 30 s, for a total of 30 cycles, and then extension at 72 °C for 5 min. The target fragments were approximately 360 bp in size and were recovered using a gel extraction kit after 1% agarose gel electrophoresis of the amplified products for high-throughput sequencing by Shanghai Personalbio Technology Co., Ltd. (Lin et al. 2021).
The nifH fragments were sequenced and then analysed using the QIIME2 (v2.13.4) analysis platform. The quality of the reads and the effect of splicing were filtered by quality control, and the samples were effectively distinguished according to the sequence at the ends of the fore and aft bar-code and primer sequence to calibrate the sequence direction, namely, to optimize the data. Sequences for annotation were based on the Greengenes database (Release 13.8, http://greengenes.secondgenome.com/) (DeSantis et al. 2006). The alpha diversity was calculated at the operational taxonomic unit (OTU) level of each sample and evaluated according to the distribution of OTUs in different samples, and the appropriateness of the sequencing depth was reflected by the rarefaction curve. The matrix distance was calculated for each sample at the OTU level. Canonical correspondence analysis (CCA) is a multivariate technique used to relate the composition of a species to environmental gradients when species have a bell-shaped response curve (Molnar 1998). The CCA in this work was performed by Genescloud tools, an online platform for data analysis (https://www.genescloud.cn). At the species taxonomic composition level, the differences in species abundance and composition between different samples (groups) were further analysed by various unsupervised and supervised sorting, clustering and modelling means, combined with the corresponding statistical test methods, and attempts were made to identify marker species by linear discriminant analysis (LDA) and linear effect size (LEfSe) analysis (Segata et al. 2011; Chang et al. 2022).
Determination of the nifH gene copy number by AQ-PCR
The nifH gene copy number of soil nitrogen-fixing bacteria was measured by absolute quantification polymerase chain reaction (AQ-PCR), which generates a standard curve by using a standard with a known copy number and determines the Ct value of the unknown sample, combining the standard curve to obtain the copy number (Walker 2001). The reaction system volume was 20 µL, which included 10 µL of 2 × SYBR real-time PCR premixture, 1 µL of nifH-F (10 µmmol/L) and 1 µL of nifH-R (10 µmmol/L) as primers, 1 µL DNA template, and 7 µL of ddH2O. The reaction conditions were denaturation at 95 °C for 15 s followed by extension at 60 °C for 30 s, for a total of 40 cycles, and then collection of the fluorescence signal starting at 60 °C. AQ-PCR was performed using nifH plasmids with a concentration gradient from 10− 1 to 10− 6 as templates and a standard curve, and ddH2O was used as a negative control. The gene copy number was calculated by the following formula.
where y is the Ct value; k is the slope of the standard curve; b is the intercept of the standard curve.
Determination of soil physical and chemical characteristics
Research has demonstrated that PGPR play key roles in nutrient acquisition and assimilation and improving soil texture, nutrition, and pH, all leading to the enhancement of plant growth (Backer et al. 2018). The physical and chemical properties of the 4 groups of soil were analysed. The moisture content (WC) of the soil was determined by mass change after oven-drying at 105 °C until constant weight. The pH value was measured by a pH meter (PB-10, Sartorius, Germany) (Yang et al. 2012). The total nitrogen (TN), available nitrogen (AN), organic matter (OM) and organic carbon (OC) contents of the soil were determined following Baeumer (1974) and Schumacher et al. (2009).
Isolation and identification of nitrogen-fixing strains
The screened strains were tested for nitrogenase activity, phosphorus-dissolving capacity, ammonia production capacity and siderophores production capacity. In addition, indole tests (tryptophan in peptone was decomposed to produce indole), catalase tests (toxic hydrogen peroxide was decomposed to H2O and O2), and NO3− reduction tests (nitrate was reduced to nitrite and ammonia) were used to screen the PGPR. Each fresh soil sample (2 g) from the 4 groups was added to 18 mL ddH2O, mixed well and then diluted to 10− 5. Then, 0.1 mL of the diluted solution was inoculated into Ashby’s nitrogen-free medium and incubated at 30 °C for 24–48 h under aerobic conditions (Lin et al. 2020). The colonies were picked randomly and inoculated in Ashby’s nitrogen-free medium for screening and preservation. The nitrogenase activity of the isolated bacteria was determined by a soil nitrogenase double antibody enzyme-linked immunosorbent assay (ELISA) kit (Thermo, USA). The solution changes colour from blue to yellow, and the intensity of the colour is measured at 450 nm using a spectrophotometer. The ammonia production capacity, nitrate reduction capacity (Marschner 1995), and catalase activity (Kuscu 2019) of the representative strains were measured. The selected representative strains were inoculated into Chrome Azurol S (CAS) test medium and incubated at 30 °C for 24–48 h to observe the siderophores around the colonies. The selected representative strains were inoculated into inorganic phosphorus medium (glucose 10.0, (NH4)2SO4 0.5, yeast extract 0.5, NaCl 0.3, KCl 0.3, MgSO4 0.3, FeSO4 0.03, MnSO4 0.03, Ca3(PO4)2 5.0, agar 15.0, pH 7.0-7.5, g/L) and organic phosphorus medium (glucose 10.0, (NH4)2SO4 0.5, yeast extract 0.5, NaCl 0.3, KCl 0.3, MgSO4 0.3, FeSO4 0.03, MnSO4 0.03, lecithin 0.2, CaCO3 1.0, agar 15.0, pH 7.0-7.5, g/L) at 30 °C for 24–48 h to observe the phosphorus-solubilized circles around the colonies (Madigan and Martinko 1997).
Whole-cell protein pattern analysis by sodium dodecyl sulfate‒polyacrylamide gel electrophoresis (SDS-PAGE) offers a fairly fast and easy method of identifying a large number of strains, and it has an adequate level of taxonomic resolution at the species and subspecies levels. SDS-PAGE protein pattern analysis has been successfully used to identify various types of bacteria (Kim et al. 2010). The isolated bacteria were inoculated into Ashby’s nitrogen-free liquid medium (with 30 °C, 150 rpm shake cultured) and grown to the logarithmic phase with an OD600 value of 1.0, and the bacterial colonies were clustered by SDS-PAGE whole-cell protein electrophoresis to determine the homology of the strains (Lajudie et al. 1994). The Tanon GIS series digital gel image processing system and image analysis were used to analyse the homology of the diversity of strains.
The 16S rDNA sequence PCRs were performed as a total reaction system of 25 µL, which contained 2 × PCR TaqMix 12.5 µL, template DNA 1.0 µL, primer 27F (10 µmmol/L) 1 µL, primer 1492R (10 µmmol/L) 1 µL, and ddH2O 9.5 µL. The 16S rDNA universal primers were 27F: 5’-AGAGTTTGATCCTGGCTCAG-3’ and 1492R: 5’-GGTTACCTTGTTACGACTT-3’. The PCR thermal cycling conditions were predenaturation at 94 °C for 5 min, denaturation at 94 °C for 60 s, annealing at 60 °C for 60 s, and extension at 72 °C for 60 s, for a total of 30 cycles, followed by extension at 72 °C for 5 min and detection by 1% agar gel electrophoresis. Samples were recovered and sent to Fuzhou Boshang Biotechnology Co., Ltd. for DNA sequencing. The 16S rDNA sequences of each strain were blasted from nucleotide to nucleotide on the National Center for Biotechnology Information (NCBI) website (https://blast.ncbi.nlm.nih.gov/Blast.cgi) for comparison, and homology was established using the neighbour-joining algorithm of MEGA (v5.1) software (Saitou and Nei 1987; Tamura et al. 2004; Kumar et al. 2016).
Promotional properties of PGPR
To assay growth promotion under different PGPR solutions, three treatments were further selected to test the ability of the bacteria to promote C. fungigraminus seedling growth in pot experiments at 28–30 °C in a greenhouse (natural light). During the experiment, 15 mL of selected bacterial solution (Ashby’s nitrogen-free liquid medium, 30℃ shake cultured at 150 rpm) with a concentration of 2 × 109 cfu/mL was poured onto the C. fungigraminus seedlings near the roots. Experimental treatment-1 (T1) and experimental treatment-2 (T2) were treated with a single strain solution, and experimental treatment-3 (T3) was treated with a combined suspension with both strains (1:1, v/v). A control group (CK) was treated with the same volume of medium solution. There were 20 replicates in each group. The C. fungigraminus growth status (including the root length, root number, plant height, leaf length, and leaf number) and soil physical and chemical characteristics were observed and recorded after 5 weeks of treatment.
Data analysis
Statistical analysis of data was carried out by SPSS (v9.1) software, and the results were expressed as the means ± standard deviations (SDs). The data were statistically evaluated using a one-way analysis of variance (ANOVA) test, and values presented with different lowercase letters are significantly different at P < 0.05.
Results
Soil physiochemical properties
The physical and chemical properties of the samples collected from different locations of C. fungigraminus soil in Hainan Province are shown in Table 1. The soil pH value and contents of TN, AN, OM and OC gradually decreased with soil depth, and the contents of each parameter were lowest in G4. However, the water content increased gradually and was highest in G4.
Determination of the nifH gene copy number
The results of the genomic DNA extracted from the 24 samples and the electrophoretic amplification of nifH were detected by 1% agar gel electrophoresis (Fig. S1). The Ct value was obtained by AQ-PCR amplification of the nifH gene and was plotted as the ordinate. The standard curve equation was Y =-3.59X + 39.89, R2 = 0.999, and the amplification efficiency was 89.91% (Fig. S2). The melting curve had a single peak, indicating that there was no nonspecific amplification or primer dimer. The copy number of the nifH gene was highest in G2 (1.56 ± 0.17 × 107 copies/g) (Fig. S3).
Nitrogen-fixing bacterial population diversity
Through the prepossessing of the sequences and the removal of low-quality and ambiguous sequences, a total of 3.28 million raw sequences were obtained from the 24 samples. After initial quality filtering, noise removal and correction, 3.07 million valid sequences remained with sequence lengths ranging from 33 to 444 bp; the average sequence length was 319 bp. Based on the principle of similarity greater than 97%, the total OTU numbers of the G1, G2, G3 and G4 samples were 1650, 1821, 1704 and 1729, respectively. The richness of nitrogen-fixing bacteria in rhizosphere soil (G2) was higher than that in nonrhizosphere soil (G1, G3, G4) (Fig. S4a). Four groups of sample dilution curves with Observed_species were stable as the soil depth deepened; the sequences were constructed by QIIME2 (v2.13.4). Nonmetric multidimensional scaling (NMDS) revealed that 6 samples in each group were well clustered and indicated that the amount of sequenced data was reliable and reflected the diversity of the nitrogen-fixing bacterial communities (Fig. S4b). Alpha diversity analysis revealed the richness and diversity of the microbial communities in each group of samples. Table 2 shows that the Chao 1 index, the number of observed_species, and the Shannon index of nitrogen-fixing bacterial species were highest in G2. The diversity and abundance of the nitrogen-fixing bacterial groups were the highest in the C. fungigraminus rhizosphere soil (Table 2).
Species abundance and community structure
The overall structure of the nitrogen-fixing bacteria is shown in a circle packing chart (Carrión et al. 2019) in which the different domain, phylum, order, family, genus and species levels have different microbiota. At the phylum level, 90.93–95.98% of the bacteria belonged to Proteobacteria, with the greatest abundance occurring in G1 and the least abundance in G4. Actinobacteria accounted for 0.12–0.42% of the bacteria and had the greatest abundance in G2. Firmicutes, Cyanobacteria and Verrucomicrobia were present in small amounts in the 4 groups, with abundances of 0.09–0.34%, 0.05–0.29% and 0.06–0.14%, respectively. Spirochaetes and Chlorobi were present in G2, with abundances of 0.02% and 0.01%, respectively, but did not appear in the other groups. Unclassified clades were also present in the 4 groups with abundances of 3.66–5.35% (Fig. 1a).
Composition of the nitrogen-fixing bacterial community at different taxonomic levels in a circle packing chart. a shows the composition at the phylum level, and b shows the composition at the genus level. Each dot represents the specific taxonomic attribution of the OTU at the phylum level. The largest circle represents the level of the phyla, and the circles with different colours and sizes represent classes, orders, families, genera and species. The dots represent the top 100 most abundant OTUs, and their area is proportional to the abundance of the OTUs. Therefore, the abundance of taxa corresponding to the circle are also indicated by the area of the origin in the circle. The larger the sector area of the OTU in each group, the higher the abundance of the taxon in each group
At the genus level, the community composition and relative abundance of nitrogen-fixing bacteria in the C. fungigraminus rhizosphere soils were significantly different between different locations. The genera with the high abundance in the 4 groups included Bradyrhizobium (29.64–39.71%), Geobacter (9.35–28.99%), Azospirillum (1.75–15.15%), Desulfovium (1.75–15.15%), Desulfovibrio (2.67–5.58%), Burkholderia (1.42–6.76%), Halorhodospira (0.68–2.73%), Methylosinus (0.51–4.30%), Azotobacter (1.07–2.45%), Paraburkholderia (0.30–1.70%), Frankia (0.11–0.34%), Rhizobium (0.0023-0.01%) and unidentified bacteria (23.26–31.81%). Phyla with high abundance in G1 included Proteobacteria (95.97%), Actinobacteria (1.16%), Verrucomicrobia (0.98%), Firmicutes (0.89%), Cyanobacteria (0.61%) and other unidentified genera (3.65%). In G2, bacterial genera present in high abundance included Proteobacteria (93.91%), Actinobacteria (0.42%), Firmicutes (0.18%), Verrucomicrobia (0.06%), Cyanobacteria (0.04%), Rhizobium (0.0054%) and unidentified genera (5.35%). In group 3, Proteobacteria (90.93%), Actinobacteria (3.10%), Firmicutes (0.34%), Cyanobacteria (0.29%), and Verrucomicrobia (0.14%) were the main community components. In G4, genera with high abundance included Proteobacteria (94.10%), Actinobacteria (0.38%), Firmicutes (0.32%), Cyanobacteria (0.16%), and Verrucomicrobia (0.09%) (Fig. 1b).
As shown in Fig. 2a, the numbers of OTUs in G1, G2, G3 and G4 were 1437, 1600, 1489 and 1447, respectively, with 445 common OTUs among all groups. The numbers of unique OTUs in each group were 534 (G1), 399 (G2), 327 (G3) and 326 (G4). The overall classification of samples based on R language and pheatmap software was plotted as a heatmap, and the top 50 classification units in terms of relative abundance are shown in Fig. 2b. The colour change on the heatmap shows that the nitrogen-fixing bacterial communities of G1 and G2 had analogous compositions, while the nitrogen-fixing bacterial communities of G3 and G4 clustered together. The same result is also shown in Fig. S1b. The relative abundances of Paraburkholderia, Ruficoccus, Azonexus, Azohydromonas, Azoarcus, Azospirillum, Burkholderia and Rhizobium were higher in G1 than in the other groups, while the relative abundances of Azotobacter, Methylococcus, Rhizobium and Burkholderia were highest in G2. Azotobacter, Methylocystis, Methylosinus, Beijerinckia, and Geobacter had relatively high abundances in G3, while in G4, Ralstonia, Pelomonas, Desulfovibrio, Halorhodospira, Magnetospirillum, Frankia, and Bradyrhizobium had higher relative abundances.
a is a Venn diagram of the OTU distribution of all samples, and the number in the Venn diagram indicates the number of OTUs that were shared or unique between samples from different locations. b Heatmap of the top 50 most abundant genera (darker green indicates higher abundance, lighter yellow indicates lower abundance). c Canonical correspondence analysis (CCA) based on the nitrogen-fixing bacterial community composition and physicochemical variables
TN, AN and depth (DP) explained more than 80% of the bacterial community variation. In C. fungigraminus surface soil (G1), TN, AN, OM and OC were the main factors and were positively affected by Azospirillum, Burkholderia and Rhizobium. Compared to G1, DP was the only significant main factor and contributed nearly 40% of the total nitrogen-fixing bacterial community variation in G2, G3 and G4 (Fig. 2c). LEfSe analysis showed an abundance that was plotted based on the overall classification of the samples. With an LDA value of 3 (species diversity and richness, P < 0.001), Actinobacteria and Proteobacteria had higher abundance at the phylum level, and at the genus level, Bejerinckia, Bradyrhizobium, Methylocystis, Rhizobium and Azospirillum had higher abundance (Fig. S5). The distributions of Bejerinckia and Methylocystis in G2 were significantly higher than those of other genera, and these genera were the most abundant in this group and can be used as marker species. These results are consistent with the composition taxonomic statistics in Fig. 1.
Isolation, purification, identification and performance of PGPR
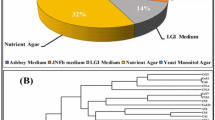
The bacterial suspension was spread on Ashby’s solid medium, and bacterial morphotypes with high colony number and vigorous growth were selected. Through microscopic examination and nitrogenase activity determination, 28 bacterial morphotypes were selected for further physiological and biochemical tests (Table 3). Of these, 7 representative strain classes were divided by SDS-PAGE electrophoresis analysis (Fig. 3a).
a SDS-PAGE electrophoretic clustering analysis of whole-cell proteins of 28 bacterial morphotypes into 7 classes; b evolutionary tree constructed by the neighbour-joining procedure in MEGA (v5.1) software. The length and bootstrap confidence values of each branch are above or below the sequence branch
Eleven representative strain determination experiments included the indole test, catalase test, methyl red test, and NO3− reduction test. The ammonia production capacity, phosphorus solubility activity and iron-producing carrier ability were also measured to determine the basic functions of the representative strains. Table 4 shows that none of the 11 selected strains had the ability to decompose glucose and tryptophan, which indicated the absence of intestinal pathogens. Strains No. 8 and No. 28 had the ability to convert the fixed nitrogen into ammonia. Both strains also displayed a positive ability to catalyse hydrogen peroxide, produce H2O and O2−, reduce NO3− to NO2− and even produce nitrogen compounds for plant growth. Strains No. 8 and No. 28 produced siderophores, and it was inferred that they may have the ability to dissolve unavailable calcium phosphate salts into bioavailable phosphate salts for plant absorption. In all, these capabilities suggest that strains No. 8 and No. 28 may have good promotion potential for plant growth as PGPR.
PGPR strains No. 8 (GenBank Accession No.: OQ119905) and No. 28 (GenBank Accession No.: OQ119906) homologous sequences were downloaded from NCBI and compared; the likelihood with known strains exceeded 97%, and the sequences belonged to Rhizobium. The 2 typical strains clustered with MK241857.1, ON358080.1, NR169377.1, and NR144599.1, and the bootstrap value exceeded 97%, which indicated that the 2 strains belonged to Rhizobium pusense (Fig. 3b). Rhizobium pusense No. 8 and Rhizobium pusense No. 28 clustered together and were screened in G1 and G2, respectively (Fig. 3b, Table 3).
C. fungigraminus seedling promotion test of PGPR
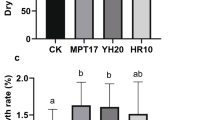
The plant height, leaf length, number of leaves, root length, and tiller number of C. fungigraminus seedlings after watering with R. pusense No. 8 bacterial solution (T1), No. 28 bacterial solution (T2), and the mixed bacterial solution (T3) are shown in Table 5; Fig. 4. Compared to the CK, the root length, height, and leaf length of C. fungigraminus in T3 significantly increased by 56.79%, 76.99% and 55.71% (P < 0.05), respectively. The AN, OM and OC of C. fungigraminus soil in T2 significantly increased by 3.09 times, 5.77 times and 5.77 times, respectively. T1, T2 and T3 significantly increased the WC and TN in the soil.
Discussion
Microorganisms have been shown to be able to form many biological communities in soil ecosystems. The formation and function of soil ecosystems at different depths are greatly influenced by microbial communities (Amundson et al. 2015). Soil depth significantly affected the abundance of dominant groups such as Actinomycetes and Proteobacteria (Li et al. 2014). The soil microbial community composition was shown to be closely correlated to the soil physical and chemical properties (Araújo et al. 2013; Wei et al. 2013) and is greatly affected by the vertical distribution of the soil texture. An important objective of research on rhizosphere microbes that fix nitrogen is to extend biological nitrogen fixation as a significant source of nitrogen for plants (Singh et al. 2020). Soil microorganisms are essential for biogeochemical cycles, colonizing plant roots, improving soil fertility and plant health and increasing crop production (Hayat et al. 2010). In the present study, the physical and chemical properties of the soil and the diversity of nitrogen-fixing bacteria from different soil locations of C. fungigraminus were compared. With increasing soil depth, the concentrations of soil TN, AN, OC and OM gradually decreased, reaching their lowest values at 20 cm depth from the rhizosphere soil. Analysis of the α diversity of nitrogen-fixing bacteria in the 4 groups showed that the Shannon index of rhizosphere soil was significantly higher than that of the other soil samples, while depth was the most significant factor, and the other groups were significantly different from surface soil in terms of the Chao1 index, Shannon index and Simpson index. Soil depth represents a strong physio-chemical gradient that greatly affects soil-dwelling microorganisms. More intensive interkingdom cooccurrence patterns were observed in the upper mineral layer (0–5 cm) than in the above organic and lower mineral soils, signifying a substantial influence of soil depth on biotic interactions (Mundra et al. 2021). Plant roots release a variety of secretions, including sugars, organic acids, amino acids, hormones, and extracellular enzymes; these substances have been shown to provide sufficient nutrition and energy for microorganisms (Zhang et al. 2004). Highly abundant genera in the four soils included Bradyrhizobium, Geobacter, Azospirillum, Desulfovibrio, Burkholderia, Halorhodospira, Methylosinus, and Azotobacter. The abundances of Paraburkholderia, Frankia and Rhizobium in rhizosphere soil were higher than those in the other groups. We also found that the diversity of the nifH gene in the rhizosphere soil was higher than that in the nonrhizosphere soil of C. fungigraminus, which was consistent with the trend observed in high-throughput sequencing results. nifH is an ideal genetic marker to detect diversity and phylogeny (Hamelin et al. 2010).
There are many species of PGPR, including Rhizobium, Bacillus, Klebsiella and Pseudomonas. They are generally capable of fixing nitrogen, producing plant growth hormones and secreting antibiotics (Kloepper et al. 1989). According to Lin et al. (2019), Klebsiella variicola mainly colonizes the endothelial layer of C. fungigraminus roots as a plant endophyte. The contents of organic matter and total nitrogen in soil are the main criteria for evaluating soil fertility and quality. It has been widely reported that Rhizobia were able to promote the growth of crops and increase soil fertility (Shota et al. 2022). In our work, 11 representative strains were isolated, and their nitrogenase activity, ammonia production capacity, siderophore production capacity and phosphorus solubilization capacity were detected. Of these, 2 strains with high nitrogenase activity and soluble phosphorus activity were selected. Furthermore, TN, AN and OM had significant effects on the nitrogen-fixing bacterial community under different treatments and had positive effects on Azospirillum, Burkholderia and Rhizobium. The similar results of CCA between the environmental factors and distribution of rhizobial genospecies showed that soil pH and the contents of total phosphorus, total potassium and total organic carbon were the main determinants of the community structure of S. davidii rhizobia (Cao et al. 2021). Two strains clustered with R. pusense, and the bootstrap value exceeded 97%. Previous studies indicated 16S rDNA sequence analysis as an authenticated technique used to study bacterial isolates at the species level (Imran et al. 2010; Alam et al. 2011). R. pusense is a gram-negative bacterium belonging to the Rhizobiaceae family. It was able to degrade glycosylated phenols and flavonoids secreted by plant roots and can produce phytohormones (Badhai et al. 2017). R. pusense MB-17 A, a PGP rhizobium isolated from mung beans, showed significant siderophore production, ammonia production and phosphorus-solubilizing ability, and the fresh weight and nodule number of mung bean were significantly increased 60 days after inoculation (Chaudhary et al. 2020). Nitrogen-fixing bacteria with phosphorus solubilizing ability can dissolve some unavailable calcium phosphate salts in the soil and convert them into bioavalable phosphorus salts for absorption and utilization by plants; Phosphorus solubilizing bacteria aslo can improve the efficiency of phosphorus absorption and utilization by plants by releasing organic acids to dissolve inorganic phosphorus in the soil, or by secreting metabolites to degrade organic phosphorus (Chauhan et al. 2017; Mehta et al. 2013). However, according to Bashan et al. (2013), we need more work on phosphorus-solubilizing ability of 2 R. pusense strains in future. Plant growth promotion could be the result of the beneficial functions of applied PGPR isolates, such as plant growth hormone production, nitrogen fixation, and P solubilization (Majeed et al. 2015). R. pusense No. 8 and R. pusense No. 28 were selected from the surface soil and rhizosphere soil in the present study, respectively, as potential soil improvers. Both strains promoted plant height, leaf length and root length growth of C. fungigraminus. Meanwhile, the growth promotion effect of the 2 strains solution was greater than that of the single strains. However, R. pusense No. 8 increased the concentrations of available nitrogen, organic matter and organic carbon in the soil more than R. pusense No. 28 and the mixture. The increase in the available nitrogen content is able to directly enhance the absorption of nitrogen by plants. Organic matter can provide nutrients for plants, promote plant growth, improve the soil structure, and improve the water retention ability of soil (Kiba and Krapp 2016). Furthermore, the inoculation of PGPR with multifunctional traits is better than inoculation of PGPR with single traits (Imran et al. 2014). The response of plants to different isolates was variable, which may be attributed to their individual traits and rhizospheric competencies, and most of the bacteria displayed good survival and plant promoting ability in the rhizosphere.
Conclusion
In summary, this foundational work examined the composition and diversity of nitrogen-fixing bacteria in C. fungigraminus rhizosphere soil, which were significantly affected by depth. R. pusense has plant growth-promoting ability as a PGPR. C. fungigraminus is becoming an important plant and is spreading worldwide. The use of effective PGPR is an opportunity for improving production in addition to maintaining soil structure and fertility. The study provides a theoretical basis for the development of plant growth-promoting rhizobacteria and the development of biological fertilizers. However, in the future, R. pusense should be applied so as to restore soil fertility, phosphorus solubilizing ability and safety to the environment, humans and animals, and the impact of long-term application on soil physical and chemical properties and microorganisms needs to be monitored and further studied.
Data availability
All authors confirm that the data supporting the findings of this study are available within the article.
References
Alam SI, Bansod S, Goel AK, Singh L (2011) Characterization of an environmental strain of Bacillus thuringiensis from a hot spring in western Himalayas. Curr Microbiol 62:547–556. https://doi.org/10.1007/s00284-010-9743-x
Amundson R, Berhe AA, Hopmans JW, Olson C, Sztein AE, Sparks DL (2015) Soil science. Soil and human security in the 21st century. Science 348(6235):1261071. https://doi.org/10.1126/science.1261071
Araújo A, Cesarz S, Leite L, Borges CD, Tsai SM, Eisenhauer N (2013) Soil microbial properties and temporal stability in degraded and restored lands of Northeast Brazil. Soil Boil Biochem 66:175–181. https://doi.org/10.1016/j.soilbio.2013.07.013
Backer R, Rokem JS, Ilangumaran G, Lamont J, Praslickova D, Ricci E, Subramanian S, Smith DL (2018) Plant growth-promoting Rhizobacteria: Context, mechanisms of action, and roadmap to commercialization of biostimulants for sustainable agriculture. Front Plant Sci 23(9):1473. https://doi.org/10.3389/fpls.2018.01473
Badhai J, Whitman WB, Das SK (2017) Draft genome sequence of Rhizobium pusense strain NRCPB10T (LMG 25623T) isolated from rhizosphere soil of chickpeas (Cicer arietinum L.) grown in India. Genome Announc 5:e00282–e00217. https://doi.org/10.1128/genomeA.00282-17
Baeumer HFK (1974) Effect of zero-tillage on organic carbon and total nitrogen content, and their distribution in different n-fractions in loessial soils. Agro Ecosyst 1:19–29. https://doi.org/10.1016/0304-3746(74)90004-3
Barraquio WL, Revilla L, Ladha JKJSN (1997) Isolation of endophytic diazotrophic bacteria from wetland rice. Plant Soil 194(1/2):15–24. https://doi.org/10.1023/A:1004246904803
Bashan Y, Kamnev AA, Bashan LED (2013) Tricalcium phosphate is inappropriate as a universal selection factor for isolating and testing phosphate-solubilizing bacteria that enhance plant growth: a proposal for an alternative procedure. Biol Fertil Soils 49(4):465–479. https://doi.org/10.1007/s00374-012-0737-7
Cantera JJL, Kawasaki H, Seki T (2004) The nitrogen-fixing gene (nifH) of Rhodopseudomonas palustris: a case of lateral gene transfer? Microbiology 150(pt 7):2237–2246. https://doi.org/10.1099/mic.0.26940-0
Cao Y, Tie D, Zhao JL, Wang XB, Yi JJ, Chai YF, Wang KF, Wang ET, Yue M (2021) Diversity and distribution of Sophora davidii rhizobia in habitats with different irradiances and soil traits in Loess Plateau area of China. Syst Appl Microbiol 44(4):126224. https://doi.org/10.1016/j.syapm.2021.126224
Carrión VJ, Perez-Jaramillo J, Cordovez V, Tracanna V, de Hollander M, Ruiz-Buck D, Mendes LW, van Ijcken WFJ, Gomez-Exposito R, Elsayed SS, Mohanraju P, Arifah A, van der Oost J, Paulson JN, Mendes R, van Wezel GP, Medema MH, Raaijmakers JM (2019) Pathogen-induced activation of disease-suppressive functions in the endophytic root microbiome. Science 366(6465):606–612. https://doi.org/10.1126/science.aaw9285
Cavite HJM, Mactal AG, Evangelista EVC, Jayvee AC (2021) Growth and yield response of upland rice to application of plant growth-promoting rhizobacteria. J Plant Growth Regul 40(2):494–508. https://doi.org/10.1007/s00344-020-10114-3
Chabot R, Antoun H, Cescas MPJP (1996) Growth promotion of maize and lettuce by phosphate-solubilizing Rhizobium leguminosarum biovar. Phaseoli Plant Soil 184(2):311–321. https://doi.org/10.1128/aem.62.8.2767-2772.1996
Chang F, He SS, Dang CY (2022) Assisted selection of biomarkers by Linear Discriminant Analysis Effect size (LEfSe) in Microbiome data. J Vis Exp 183. https://doi.org/10.3791/61715
Chaparro JM, Badri DV, Vivanco JM (2014) Rhizosphere microbiome assemblage is affected by plant development. ISME J 8(4):790–803. https://doi.org/10.1038/ismej.2013.196
Chaudhary T, Gera R, Shukla P (2020) Deciphering the potential of Rhizobium pusense MB-17a, a plant growth-promoting root endophyte, and functional annotation of the genes involved in the metabolic pathway. Front Bioeng Biotech 8:617034. https://doi.org/10.3389/fbioe.2020.617034
Chauhan A, Guleria S, Balgir PP, Walia A, Mahajan R, Mehta P, Shirkot CK (2017) Tricalcium phosphate solubilization and nitrogen fixation by newly isolated Aneurinibacillus aneurinilyticus CKMV1 from rhizosphere of Valeriana jatamansi and its growth promotional effect. Braz J Microbiol 48(2):294–304. https://doi.org/10.1016/j.bjm.2016.12.001
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72(7):5069–5072. https://doi.org/10.1128/AEM.03006-05
Ferrarezi JA, Carvalho-Estrada P, Batista BD, Aniceto RM, Tschoeke B, Andrade P, Lopes B, Bonatelli ML, Odisi EJ, Azevedo JL (2022) Effects of inoculation with plant growth-promoting rhizobacteria from the brazilian amazon on the bacterial community associated with maize in field. Appl Soil Ecol 170:104297. https://doi.org/10.1016/j.apsoil.2021.104297
Frank B (1889) Ueber die Pilzsymbiose der Leguminosen. Ber Dtsch Bot Ges 7:332–346
Hamelin J, Fromin N, Tarnawski S, Teyssier-Cuvelle S, Aragno M (2010) nifH gene diversity in the bacterial community associated with the rhizosphere of Molinia coerulea, an oligonitrophilic perennial grass. Environ Microbiol 4:477–481. https://doi.org/10.1046/j.1462-2920.2002.00319.x
Hayat R, Ali S, Amara U, Khalid R, Ahmed I (2010) Soil beneficial bacteria and their role in plant growth promotion: a review. Ann Microbiol 60(4):579–598. https://doi.org/10.1007/s13213-010-0117-1
Hou MP, Babalola OO (2013) Evaluation of plant growth promoting potential of four rhizobacterial species for indigenous system. J Cent South Univ 20:164–171. https://doi.org/10.1007/s11771-013-1472-4
Igiehon NO, Babalola OO (2018) Rhizosphere Microbiome Modulators: contributions of nitrogen fixing bacteria towards sustainable agriculture. Int J Environ Res Public Health 15(4):574. https://doi.org/10.3390/ijerph15040574
Imran A, Hafeez FY, Frühling A, Schumann P, Malik KA, Stackebrandt E (2010) Ochrobactrum ciceri sp. nov., isolated from nodules of Cicer arietinum Int J Syst Evol Micr 60(7):1548–1553. https://doi.org/10.1099/ijs.0.013987-0
Imran A, Saadalla MJA, Khan SU, Mirza MS, Malik KA, Hafeez FY (2014) Ochrobactrum sp. Pv2Z2 exhibits multiple traits of plant growth promotion, biodegradation and N-acyl-homoserine-lactone quorum sensing. Ann Microbiol 64:1797–1806. https://doi.org/10.1007/s13213-014-0824-0
Jia YL, Liao Z, Chew WF, Wang LF, Lin BS, Chen CQ, Lu GD, Lin ZX (2020) Effect of Pennisetum giganteum z.x.lin mixed nitrogen-fixing bacterial fertilizer on the growth, quality, soil fertility and bacterial community of pakchoi (Brassica chinensis L). PLoS ONE 15(2):e0228709. https://doi.org/10.1371/journal.pone.0228709
Kiba T, Krapp A (2016) Plant nitrogen acquisition under low availability: regulation of uptake and root architecture. Plant Cell Physiol 57(4):707–714. https://doi.org/10.1093/pcp/pcw052
Kim TW, Kim YH, Kim SE, Lee JH, Park CS, Kim HY (2010) Identification and distribution of Bacillus species in doenjang by whole-cell protein patterns and 16S rRNA gene sequence analysis. J Microbiol Biotechnol 20(8):1210–1214. https://doi.org/10.4014/jmb.1002.02008
Kloepper JW, Ran L, Zablotowicz RM (1989) Free-living bacterial inocula for enhancing crop productivity. Trends in Biotechnol 7:39–44. https://doi.org/10.1016/0167-7799(89)90057-7
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Kuscu ISK (2019) Changing of soil properties and urease-catalase enzyme activity depending on plant type and shading. Environ Monit Assess 191(3):178–185. https://doi.org/10.1007/s10661-019-7304-8
Lajudie PD, Willems A, Pot B, Dewettinck D, Maestrojuan G, Neyra M, Collins MD, Dreyfus B, Kersters K, Gillis M (1994) Polyphasic taxonomy of rhizobia: emendation of the genus sinorhizobium and description of Sinorhizobium meliloti comb. nov., Sinorhizobium saheli sp. nov., and Sinorhizobium teranga sp. nov. Int J Syst Evol Micr 44(4):715–733. https://doi.org/10.1099/00207713-44-4-715
Li CH, Yan K, Tang LS, Jia ZJ, Li Y (2014) Change in deep soil microbial communities due to long-term fertilization. Soil Boil Biochem 75:264–272. https://doi.org/10.1016/j.soilbio.2014.04.023
Li J, Luo ZQ, Zhang CH, Qu XJ, Chen M, Song T, Yuan J (2020) Seasonal variation in the Rhizosphere and Non-Rhizosphere Microbial community structures and functions of Camellia yuhsienensis Hu. Microorganisms 8(9):1385. https://doi.org/10.3390/microorganisms8091385
Lin ZX, Lin DM (2018) JUNCAO Science. China Agricultural Press, Beijing
Lin BS, Chen Y, Zheng SZ, Jia YL, Wang LF, Lu GD, Lin ZX (2019) Construction of nitrogen-fixing Klebsiella variicola GN02 expression vector pET28a-Lac-EGFP and its colonization of Pennisetum sinense Roxb roots. Biotechnol Biotec 33(1):1074–1084. https://doi.org/10.1080/13102818.2019.1638301
Lin BS, Zheng XT, Zheng SZ, Luo MC, Lin ZX (2020) Metabolomics analysis of ammonia secretion during the fermentation of Klebsiella variicola GN02 with highly efficient endophytic nitrogen-fixing bacteria. Appl Biochem Micro 56(4):400–411. https://doi.org/10.1134/S0003683820040109
Lin BS, Liu J, Zhang X, Weng C, Lin ZX (2021) The flora compositions of nitrogen-fixing bacteria and the differential expression of nifH gene in Pennisetum giganteum z.x.lin roots. Biomed Res Int 5568845. https://doi.org/10.1155/2021/5568845
Lin ZX, Lin DM, Liu ZJ, Lan SR (2022) Cenchrus fungigraminus Z. X. Lin & D. M. Lin & S. R. Lan sp. nov, a new species of Panicoideae (Poaceae): evidence from morphological, nuclear and plastid genome data. J Forest Environ 42(5):514–520. https://doi.org/10.13324/j.cnki.jfcf.2022.05.009
Liu YL, Mei L, Li J, Wang AN, Luo HL, Lin ZX (2018) Effect of JUNCAO-cultivated Ganoderma lucidum spent mushroom substrate-hot water extract on immune function in mice. Trop J Pharm Res 17(2):261–267. https://doi.org/10.4314/tjpr.v17i2.10
Lugtenberg B, Kamilova F (2009) Plant-growth-promoting rhizobacteria. Annu Rev Microbiol 63:541–556. https://doi.org/10.1146/annurev.micro.62.081307.162918
Madigan MT, Martinko JM (1997) Brock biology of micro-organisms. Prentice Hall. https://doi.org/10.1038/1911032b0
Majeed A, Abbasi MK, Hameed S, Imran A, Rahim N (2015) Isolation and characterization of plant growth-promoting rhizobacteria from wheat rhizosphere and their effect on plant growth promotion. Front Microbiol 6:198–207. https://doi.org/10.3389/fmicb.2015.00198
Marschner H (1995) Functions of mineral nutrients: macronutrients-sciencedirect. Mineral Nutrition of Higher Plants (2nd edn), 313–404. https://doi.org/10.1016/B978-0-08-057187-4.50014-X
Mehta P, Walia A, Chauhan A, Kulshrestha S, Shirkot CK (2013) Phosphate solubilisation and plant growth promoting potential by stress tolerant Bacillus sp. isolated from rhizosphere of apple orchards in trans himalayan region of himachal pradesh. Ann Appl Biol 163(3):430–443. https://doi.org/10.1111/aab.12077
Molnar J (1998) Paleobotanical evidence of eocene and oligocene paleoaltitudes in midlatitude western north america. GSA Bull 110(5):664–678. https://doi.org/10.1130/0016-7606(1998)110
Monteiro CC, Villegas LE, Campolina TB, Pires AC, Miranda JC, Pimenta PF, Secundino NF (2016) Bacterial diversity of the american sand fly Lutzomyia intermedia using high-throughput metagenomic sequencing. Parasit Vectors 9(1):480. https://doi.org/10.1186/s13071-016-1767-z
Mundra S, Kjønaas OJ, Morgado LN, Krabberød AK, Ransedokken Y, Kauserud H (2021) Soil depth matters: shift in composition and inter-kingdom co-occurrence patterns of microorganisms in forest soils. FEMS Microbiol Ecol 97(3):fiab022. https://doi.org/10.1093/femsec/fiab022
Nakkeeran S, Fernando WGD, Siddiqui ZA (2005) Plant Growth promoting Rhizobacteria Formulations and its scope in commercialization for the management of pests and diseases. In: Siddiqui ZA (ed) PGPR: Biocontrol and Biofertilization. Springer, Dordrecht, pp 257–296. https://doi.org/10.1007/1-4020-4152-7_10
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Santos PD, Fang Z, Mason SW, Setubal JOC, Dixon R (2012) Distribution of nitrogen fixation and nitrogenase-like sequences amongst microbial genomes. BMC Genomics 13:162–173. https://doi.org/10.1186/1471-2164-13-162
Schumacher E, Dindorf W, Dittmar M (2009) Exposure to toxic agents alters organic elemental composition in human fingernails. Sci Total Environ 407(7):2151–2157. https://doi.org/10.1016/j.scitotenv.2008.12.013
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12(6):R60
Shota K, Michiki T, Shigenobu K, Yudai I, Ryotaro H, Nahoko K, Natsumi O, Park SB, Akinori A, Makoto U, Ogawa J (2022) Characterization of regioselective glycosyltransferase of Rhizobium pusense JCM 16209T useful for resveratrol 4′-O-α-D-glucoside production. J Biosci Bioeng 134(3):213–219. https://doi.org/10.1016/j.jbiosc.2022.06.011
Singh RK, Singh P, Li HB, Song QQ, Guo DJ, Solanki MK, Verma KK, Malviya MK, Song XP, Lakshmanan P, Yang LT, Li YR (2020) Diversity of nitrogen-fixing rhizobacteria associated with sugarcane: a comprehensive study of plant-microbe interactions for growth enhancement in Saccharum spp. BMC Plant Biol 20(1):220. https://doi.org/10.1186/s12870-020-02400-9
Su DW, Su FF, Luo HL, Lin H, Lin DM, Liu PH, Lin XS, Lin ZX, Zhang LL, Lu GD (2022) Effect of different rotation systems on production and quality of Black Morel (Morchella importuna). Agronomy 12:1744–1759. https://doi.org/10.3390/agronomy12081744
Swarnalakshmi K, Yadav V, Tyagi D, Dhar DW, Kannepalli A, Kumar S (2020) Significance of plant growth promoting Rhizobacteria in grain legumes: growth promotion and crop production. Plants (Basel) 9(11):1596. https://doi.org/10.3390/plants9111596
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. PNAS 101(30):11030–11035. https://doi.org/10.1073/pnas.0404206101
Tchan YT (1952) Studies of nitrogen-fixing bacteria. ii. the presence of aerobic non-symbiotic nitrogen-fixing bacteria in soils of the sydney district. Proceedings of the Linnean Society of New South Wales
Terakado-Tonooka J, Ohwaki Y, Yamakawa H, Tanaka F, Yoneyama T, Fujihara S (2008) Expressed nifH genes of endophytic bacteria detected in field-grown sweet potatoes (Ipomoea batatas L). Microbes Environ 23(1):89–93. https://doi.org/10.1264/jsme2.23.89
Vejan P, Abdullah R, Khadiran T, Ismail S, Nasrulhaq Boyce A (2016) Role of plant growth promoting rhizobacteria in agricultural sustainability-A review. Molecules 21(5):573. https://doi.org/10.3390/molecules21050573
Walker NJ (2001) Real-time and quantitative PCR: applications to mechanism-based toxicology. J Biochem Mol Toxic 15(3):121–127. https://doi.org/10.1002/jbt.8
Wei CZ, Yu Q, Bai E, Lü XT, Li Q, Xia JY, Kardol P, Liang WJ, Wang ZW, Han XG (2013) Nitrogen deposition weakens plant-microbe interactions in grassland ecosystems. Global Change Biol 19(12):3688–3697. https://doi.org/10.1111/gcb.12348
Xomphoutheb T, Jiao S, Guo X, Mabagala FS, Sui B, Wang H, Zhao L, Zhao X (2020) The effect of tillage systems on phosphorus distribution and forms in rhizosphere and non-rhizosphere soil under maize (Zea mays L.) in Northeast China. Sci Rep 10(1):6574. https://doi.org/10.1038/s41598-020-63567-7
Yang XY, Zhang JP, Guo L, Zhao H, Zhang Y, Chen J (2012) Solvent impregnated resin prepared using ionic liquid cyphos il 104 for cr(vi) removal. T Nonferr Metal Soc 22(12):3126–3130. https://doi.org/10.1016/S1003-6326(12)61764-6
Zhang F, Shen JB, Li L, Liu XJ (2004) An overview of rhizosphere processes related with plant nutrition in major cropping systems in China. Plant Soil 260:89–99. https://doi.org/10.1023/B:PLSO.0000030192.15621.20
Zhang XX, Gao JS, Cao YH, Sheirdil RA, Wang XC, Zhang L (2015) Rhizobium oryzicola sp. nov., potential plant-growth-promoting endophytic bacteria isolated from rice roots. Int J Syst Evol Micr 65(9):2931–2936. https://doi.org/10.1099/ijs.0.000358
Zhao JJ, Zhang X, Sun L, Zhang RJ, Zhang CW, Yin HQ (2017) Rhizobium oryziradicis sp. nov., isolated from rice roots. Int J Syst Evol Micr 67:963–968. https://doi.org/10.1099/ijsem.0.001724
Zhou J, Chen SQ, Shi WJ, Rakefet DS, Li ST, Yang FL, Lin ZX (2021) Transcriptome profiling reveals the effects of drought tolerance in Giant Juncao. BMC Plant Biol 21(1):2–21. https://doi.org/10.1186/s12870-020-02785-7
Acknowledgements
We are grateful to Mr. Yongqing Wang from the Danzhou Demonstration Base of the National Engineering Research Center of Juncao Technology in Hainan Province for his help during sample collection. Many thanks to Prof. Christopher Rensing for revising the paper.
Funding
This study was supported by Special project for the protection and utilization of agricultural resources of the Department of Agriculture and Rural Affairs of Fujian Province “Research and application of key technologies for innovation and industrialized utilization of Juncao and Juncao mushroom” (KKY22001XA); Major Special Project of Fujian Province “Research and application of key technologies for innovation and industrialized utilization of Juncao” (2021NZ029009);; Open Project of Key Laboratory of Preventive Veterinary Medicine and Biotechnology for Higher Education in Fujian Province, Longyan College (2020KF01); and Doctoral Research Fund Project of Longyan College (LB20202002).
Author information
Authors and Affiliations
Contributions
Jing Li, Biaosheng Lin, Zhanxi Lin and Dongmei Lin conceived and designed the experiments; Jing Li, Yanglin Liu, Bingxin Zhou, Tingting Li and Hui Lin performed the experiments and analysed the data; Jing Li, Bingxin Zhou, Tingting Li, Biaosheng Lin and Guodong Lu performed the statistical analyses and wrote the manuscript. All authors reviewed and approved the final draft.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Responsible Editor: Luz E. Bashan.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 428 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, J., Zhou, B., Li, T. et al. Isolation of rhizobacteria from the Cenchrus fungigraminus rhizosphere and characterization of their nitrogen-fixing performance and potential role in plant growth promotion. Plant Soil 489, 405–421 (2023). https://doi.org/10.1007/s11104-023-06028-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-023-06028-0