Abstract
In concert with oxygen, soil alkalinity strongly restricts the availability of iron, an essential nutrient with a multitude of functions in living organisms. In addition to its role in mitochondrial energy metabolism and as a cofactor for enzymes, in plants iron also plays key roles in photosynthesis and is required for chlorophyll biosynthesis. The ability to thrive in calcareous soils, referred to as calcicole behaviour, is the readout of an amalgam of traits of which efficient foraging of iron is a decisive factor. Recently, the well-established concept of two distinct iron uptake strategies, phylogenetically separating grasses from other land plants, was expanded by the discovery of auxiliary mechanisms that extend the range of edaphic conditions to which a species can adapt. Secretion of a tailor-made cocktail of iron-mobilising metabolites into the rhizosphere, the composition of which is responsive to a suite of edaphic and internal cues, allows survival in calcareous soils through a competitive iron acquisition strategy, which includes intricate interactions with the consortium of associated microorganisms in, on, and around the roots. This versatile, reciprocal plant-microbiome interplay affects iron mobilisation directly, but also collaterally by impacting growth, fitness, and health of the host. Here, we review the mechanisms and the multifaceted regulation of iron acquisition in plants, taking into consideration the specific constraints associated with the uptake of iron from alkaline soils. Knowledge on how plants extract iron from such soils sets the stage for a better understanding of essential ecological processes and for combatting iron malnutrition in humans.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Iron (Fe) is a crucial component of electron transport chains and a cofactor of a myriad of enzymes involved in numerous vital functions. In plants, insufficient Fe supply leads to chlorosis of young leaves and, upon prolonged exposure to inadequate Fe concentrations, severely decreased fitness. Despite the generally high abundance of Fe in soils, uptake of sufficient Fe is a challenging task for plants. In most soils, the presence of oxygen decreases the availability of Fe to levels that are insufficient to support the pivotal functions Fe is playing in plant metabolism. Besides the redox potential (Eh), free hydrogen ion concentration (pH) of the soil is a decisive factor for the phyto-availability of Fe, causing an approximately one-thousand-fold decrease of Fe activity (pFe) for each one-digit increase in pH between pH 3 and pH 8 (Schwertmann 1991). The dramatic decrease in Fe solubility, caused by rising oxygenation of the atmosphere through photosynthesis starting about 2.4 billion years ago (Kroh and Pilon 2020), forced plants that thrive under these conditions to evolve mechanisms that mobilize Fe from the pool of recalcitrant Fe oxyhydroxides in the soil. Only in the absence of oxygen, for example in waterlogged soils and at strongly acidic pH, Fe is available for plants at levels that may even exceed the demand, overrunning exclusion mechanisms and causing Fe toxicity through the formation of reactive oxygen species.
Governed by the parent material, weathering, and climate, soil pH strongly affects germination and ‘filters’ plant establishment (Fig. 1) (Wala et al. 2022). Moreover, soil pH dictates the availability of mineral nutrients and plant assemblage composition, leading to floristically distinct communities typical of alkaline or acid soils (Tansley 1917). In both natural and agricultural ecosystems, soil pH can vary widely, ranging from pH values between 3.5 (ultra-acidic soils) to 9.0 (strongly alkaline soils). While acid soils are characterized by leaching and are often associated with high levels of Al3+, Fe2+, and Mn2+ as well as limited availability of N, alkaline soils are formed when evaporation exceeds precipitation and are associated with high concentrations of Ca2+ and low available water capacity (Fig. 1) (Tsai and Schmidt 2021). Besides Fe, the availability of Zn2+ and Mn2+ is strongly restricted in soils with neutral or alkaline pH (Tyler 2003). High nitrification rates, leading to nitrate as the prevalent form of N, and an alkaline N uptake pattern where nitrate-proton co-transport causes alkalization of the apoplast, are further characteristics associated with soil alkalinity. Both acid and alkaline soils are characterised by deficiencies in plant-available phosphate.
Effects of soil pH on plant growth. The pH of the soil affects the availability of essential mineral nutrients, defines the composition of root exudates and the microbiome, and controls the concentration of potentially toxic ions such as Al3+. Alkaline pH severely restricts the availability of Fe, P, Mn, and Zn, reduces seed germination and water capacity. Many alkaline soils are characterized by high Ca and bicarbonate concentrations. The number of the element symbols reflect the availability of the respective ion. Figure was created with BioRender.com
The pronounced influence of soil pH on the composition of plant communities is a well-established ecological phenomenon (Ellenberg 1958; Chytrý et al. 2010). The factors that define calcicole-calcifuge plant strategies are the subject of a long-lasting debate over generations of plant biologists (Link 1789; Unger 1836; Grime and Hodgson 1968). Calcicole (‘chalk-loving’) behaviour is a conglomerate of traits composed of the ability to mobilize sparingly soluble mineral nutrients at high pH and to tolerate high Ca2+ and bicarbonate (HCO3−) concentrations in soils, while lack of these competences excludes calcifuge (‘chalk-fleeing’) species from habitats featuring alkaline soils. Calcifuge plants, on the contrary, are characterised by the ability to tolerate high levels of Al3+ and high free H+ activity.
For several prokaryotic and eukaryotic systems, the sensing and signalling pathways that adapt cells to the prevailing hydrogen concentration have been elucidated in detail. Sensing systems particularly for alkaline pH have been described for bacteria and yeast (Tsai and Schmidt 2021). In Saccharomyces cerevisiae, external alkalinization is perceived via the sensing protein Rim21, which forms a plasma membrane-localised complex with the chaperone Rim9 and the α-arrestin Rim8 (Fig. 2a) (Obara et al. 2012). A shift towards alkaline pH leads to a conformational change of Rim21 that allows its cytoplasmic tail to interact with Rim8, initiating recruitment of non-endosomal ESCRT (endosomal sorting complex required for transport) components to the plasma membrane complex (Xu et al. 2004; Peñalva et al. 2014). Assembly of ESCRTs set the stage for the formation of a proteolytic complex comprising Rim23, Rim20, and the protease Rim13, which cleaves the zinc finger transcription factor Rim101. Proteolytic processing of Rim101 triggers its migration to the nucleus, where it activates the expression of genes required for adaptation to alkaline pH (Hayashi et al. 2005).
External pH sensing in yeast and plants. a Sensing of alkalinity in the yeast Saccharomyces cerevisiae. Elevated external pH is perceived by a membrane-bound sensor complex and signalled into the nucleus to steer the expression of genes that adapt the cells to alkaline pH. b pH sensing in the root apoplast of A. thaliana. Low pH is sensed by protonation of a sulfotyrosine in the peptide RGF, which supports binding to its receptor RGFR and the co-receptor SERK, promoting growth of the root apical meristem. At high pH, deprotonation of aspartic and glutamic acid residues of the receptor PEPR allows binding of Pep and formation of a complex with the co-receptor BAK that activates defense responses. Based on Liu et al. (2022). Figure was created with BioRender.com
In plants, sensors for the perception of the pH in the apoplast were recently identified in A. thaliana roots (Fig. 2b) (Liu et al. 2022). At acidic pH, a sulfotyrosine of the endogenous peptide root growth factor (RGF) is protonated and forms a complex with its receptor RGFR and the co-receptor SERK, which promotes root apical meristem growth (Liu et al. 2022). The target of RGF is PLETHORA, a master regulator of root development (Aida et al. 2004; Galinha et al. 2007). At alkaline pH, the sulfotyrosine in RGF is deprotonated and the complex is destabilized. Thus, RGF acts as a bona fide pH sensor. An increase in external pH, as for example induced by pathogenic bacteria, allows the plant elicitor peptide Pep to bind its receptor PEPR and to induce pattern-triggered immunity, an interaction that is suspended at low pH due to protonation of pH-sensing aspartic and glutamic acid residues of PEPR (Liu et al. 2022). Depending on the external hydrogen ion concentration, this interlaced pH-sensing system prioritizes either growth (at acidic pH) or defense (under alkaline conditions), decisions that strongly affect the performance and fitness of the plant. Whether this system is engaged to trigger adaptive gene expression in response to alkaline conditions analogous to the Rim pathway of yeast remains, however, to be corroborated.
An estimated 25–30% of the world’s land area features soils with calcareous surface horizons. Calcareous soils are characterised by high abundance of calcium carbonate (CaCO3) in the parent material, and typically display pH values between 7.0 and 8.5. Elevated concentrations of plant-available Ca represent a crucial factor for survival in calcareous soils. The role played by Ca in ionic balance and the mechanisms by which deleterious effects of an overabundance of this nutrient are avoided vary among species. Generally, two distinct strategies are employed to cope with high soil Ca levels: 1) development of apoplastic barriers to restrict Ca uptake and translocation, or 2) tolerance to high Ca concentrations by sequestering Ca in specific cell types (Kotula et al. 2021). In some cases, elevated Ca levels can be beneficial and serve a more specific purpose. In certain calcicole species, high Ca concentrations are required to balance Fe levels. When grown on Ca-poor soils, Callisthene fasciculata Mart. (Vochysiaceae) develops necrotic spots on the leaves due to the accumulation of excess Fe (De Souza et al. 2020). Plants grown on Ca-rich soil accumulate less Fe and Zn, suggesting that Fe toxicity is restricting C. fasciculata from Ca-poor, acidic soils. On the other hand, high Ca levels can induce phosphate deficiency due to immobilisation of phosphate and precipitation of Ca phosphate (Zohlen and Tyler 2004). Efficient Ca exclusion was found to be a characteristic trait of A. thaliana demes native to calcareous soils compared to demes that are absent from such soils (Terés et al. 2019), suggesting that managing high Ca concentrations constitutes a central component in the adaptation to calcareous soils.
The elevated levels of bicarbonate (HCO3−) typical of calcareous soils pose adverse effects on growth, photosynthesis, and on the uptake of essential mineral nutrients, including Fe. The question as to whether bicarbonate has effects on the plant that are independent of the pH associated with its presence is subject of a long-standing debate. Typically, the amount of HCO3− in soils ranges from 0.4 to 3 mM, and both increases and buffers the pH of the soil. While bicarbonate appears to have a more negative impact on Fe nutrition than high pH as such, relevant results seem to be everything but equivocal across studies and species. The most common observation is a bicarbonate-induced decrease in the uptake and translocation of Fe to the leaves (Lucena et al. 2007; Alhendawi et al. 1997; Martínez-Cuenca et al. 2013; Hsieh and Waters 2016; Ding et al. 2020; Pérez-Martín et al. 2021). Translocation of Fe is possibly restricted by bicarbonate-induced alkalisation of the xylem (Wegner and Zimmermann 2004). In the moderately calcifuge species Arabidopsis thaliana, transcriptional profiling revealed that bicarbonate, but not high pH, induces a pronounced Fe deficiency response (Chen et al. 2021). It might be stated that elevated pH, bicarbonate, and Fe deficiency are inducing partly, but not always, coinciding stress responses in plants.
The corroboration of the importance of Fe-mobilising compounds, and investigations into the complex interaction of these exudates with soil components and the root microbiome, provided molecular insights into the mechanisms conferring the ability to thrive in calcareous soils, and rekindled research into the role of Fe acquisition strategies in the calcicole/calcifuge question. Here, we review recent progress on Fe uptake in general and in particular from substrate with circumneutral or alkaline pH, conditions in which the basic Fe uptake machinery is challenged or compromised, and plants are forced to employ additional mechanisms to recalibrate cellular Fe homeostasis. The elucidation of these mechanisms is not only paramount for solving persistent ecological questions; knowledge about these plant strategies is also immensely relevant from a more applied viewpoint. Yield losses and low nutritional quality of crop plants grown on calcareous soils caused by low Fe uptake efficiency makes the elucidation of the processes, players, and regulation of the mechanisms governing Fe uptake a pivotal target of research. Approximately two billion people suffer from Fe deficiency anaemia caused by insufficient diary Fe intake, often exacerbated by a mainly or entirely plant-based diet (Viteri 1998). Counteracting ‘hidden hunger’ inflicted by deficiencies of Fe and other micronutrients is one of the most urgent and challenging tasks for the community of plant biologists.
The oxymoron of Fe in soils: superabundant, but short in supply
Iron is the fourth most abundant element in soil, but, due to its low mobility in aerobic environments, the third most limiting nutrient for plants (de Mello Gabriel et al. 2021; Zuo and Zhang 2011). In soils, Fe-containing primary minerals (i.e., minerals derived from cooling magma and unaltered since) are transformed through weathering into secondary Fe minerals such as goethite, hematite, lepidocrocite, ferrihydrite, and maghemite. Under aerobic conditions, Fe2+ and Fe3+ ions released from Fe minerals are rapidly hydrolysed and precipitate either as secondary minerals, Fe hydroxides, or ferrihydrite (Cornell and Schwertmann 2003). The activity of Fe3+ (pFe3+) is dependent on soil pH and is extremely low at alkaline pH (e.g., 1024 M at pH 8.5 for Fe3+) (Lindsay and Schwab 1982), levels that are well below the requirement of plants for optimal growth. Therefore, the dissolution rate of Fe(III) oxides governs Fe supply in most soils. Besides Fe-containing minerals, organic Fe complexes, such as Fe chelated by humates and root exudates secreted by plants and microorganisms, constitute the most important pool of plant available Fe in aerobic soils. Iron-mobilizing compounds secreted by plant roots can be reducing, ligating, or acidifying and may act in an additive or synergistic manner with other soil constituents to efficiently mobilize Fe oxides in soils (Schenkeveld et al. 2016). Dynamic mobilisation of Fe by this mixture of compounds opens ‘Fe uptake windows’ that are assessable to plants for a certain period of time under a certain set of conditions. It should be stated that a realistic picture of Fe mobilization in calcareous soils can only be obtained when factoring in the presence and dynamics of Fe-binding ligands secreted by both plant roots and microorganisms. In addition, the presence of other nutrients and compounds which may dominate in alkaline soils such as Ca and HCO3− needs to be considered.
Iron acquisition from calcareous soils: mission impossible?
Uptake of Fe from calcareous soils requires an increase in Fe solubility by several orders of magnitude. To accomplish this seemingly impossible endeavour, plants have developed elaborate strategies that aid in the acquisition of Fe and in its transport to the sites of demand. Iron mobilization is governed by protonation, reduction, and chelation of Fe oxides with varying, species-dependent emphasis on particular processes (Fig. 3). Two basic, phylogenetically separated strategies to acquire Fe from the soil have been described for land plants (Römheld and Marschner 1986). In so-called strategy I plants (all spermatophytes with the exception of gramineous species), the central component of Fe acquisition is the reduction of Fe(III) chelates by an inducible plasma-membrane bound oxidoreductase, followed by uptake of the released ferrous ions. Reduction of Fe(III) is an obligatory step in species that adopted this strategy (Chaney et al. 1972). In most, but not all, strategy I species, Fe reduction is accompanied by ATPase-mediated proton extrusion, which weakens the Fe–O bond of Fe(III) oxides and leads to metal detachment (Schwertmann 1991). Secretion of Fe-mobilising compounds, such as catechol-type coumarins, is a further component of the strategy I-type Fe acquisition strategy.
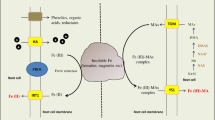
Iron uptake strategies of land plants as affected by pH. In strategy I plants, Fe is taken up as Fe2+after external reduction of Fe(III) chelates. Mobilisation of Fe is facilitated by proton extrusion and the secretion of Fe-mobilising compounds such as coumarins. Presumably, Fe(III)-coumarin complexes can be taken up as such without prior reduction of Fe via a yet to be identified coumarin importer (CI). Some species may also be capable of taking up phytosiderophores (PS) secreted by grasses via YELLOW STRIPE-LIKE (YSL) transporters. Grasses (strategy II) take up Fe after secretion of PS, which are taken up after loading with Fe3+. Mugineic acid family phytosiderophores (MAs) such as deoxymugineic acid are synthesized from nicotianamine (NA) derived from three molecules of S-adenosylmethionine (SAM) through NICOTIANAMINE SYNTHASE (NAS) via an intermediate formed by NICOTIANAMINE AMINOTRANSFERASE (NAAT) and reduction of the 3’ carbon of the intermediate via DEOXYMUGINEIC ACID SYNTHASE (DMAS). Processes that secure Fe uptake under acidic conditions and are compromised at alkaline pH are depicted on a yellow background; processes that are less affected by soil pH and operate also at alkaline pH are shown in the lower part of the figure on a blue background. Gene nomenclature refers to A. thaliana (left) and rice (Oryza sativum) (right). Broken arrow indicates a partly hypothetical mechanism. Figure was created with BioRender.com
Grasses (Poaceae) rely on the secretion of so-called phytosiderophores of the mugineic acid family that form 1:1 hexadentate complexes with Fe(III), a concept that has been referred to as strategy II (Römheld and Marschner 1986; Takagi et al. 1984). The loaded Fe(III)-phytosiderophore complex is then taken up as such, without prior reduction of ferric Fe. In grasses, biosynthesis, secretion, and uptake of phytosiderophores are increased upon Fe starvation (Takagi et al. 1984; Kobayashi and Nishizawa 2012).
In the model species A. thaliana and rice (Oryza sativum), the proteins involved in Fe uptake have been identified and functionally characterized (Fig. 3). In A. thaliana, FERRIC REDUCTASE OXIDASE 2 (FRO2) is the predominant driver of ferric reduction (Robinson et al. 1999), although coumarins (and other, less well-defined root exudates; see below) are participating in this response to a degree defined by the compound, the media pH, and the Fe status of the plant. Uptake of Fe2+ ions released from the chelator is mediated by IRON-REGULATED TRANSPORTER 1 (IRT1) (Eide et al. 1996; Vert et al. 2002). IRT1-mediated Fe uptake is accompanied by the influx of secondary substrates, in particular Mn2+ and Zn2+, which are sequestered into the vacuole by a suite of transporters such as MTPa2, ZIP9, and MTPc3. In addition, IRT1 supports the uptake of harmful metals travelling as stowaways such as Co2+ and Cd2+ (Arrivault et al. 2006). It was suggested that IRT1 acts as a transceptor, sensing the cytoplasmatic concentration of non-Fe metals that bind to a histidine-rich stretch of the protein, and trigger its degradation via CIPK23-mediated phosphorylation and IDF1-mediated lysine-63 polyubiquitination (Dubeaux et al. 2018). Proton extrusion in response to Fe starvation is mediated by the P-type ATPase AHA2 (Santi and Schmidt 2009). FRO2, IRT1, and AHA2 co-localize to the plasma membrane, organised in a tripartite Fe uptake complex (Martín-Barranco et al. 2020).
A magic potion for calcicole plants
Conspicuously, the Fe(II)- and 2-oxoglutarate-dependent dioxygenase F6′H1 was identified as putative component of the IRT1 interactome in a co-immunoprecipitation approach using a functional IRT1-mCitrine fusion protein as bait (Martín-Barranco et al. 2020). F6’H1 catalyses the first committed step in the biosynthesis of coumarins, the conversion of feruloyl-CoA to 6′-hydroxyferuloyl-CoA. In A. thaliana roots, F6’H1 massively accumulates upon Fe starvation, together with proteins involved in the general phenylpropanoid pathway (Lan et al. 2011). Similarly, transcriptomic surveys of Fe-deficient roots found genes involved in the coumarin biosynthetic pathway strongly induced, including the ABC-type transporter PDR9/ABCG37, which was later shown to be critical for the secretion of coumarins (Fourcroy et al. 2014; Yang et al., 2010; Rodríguez-Celma et al. 2013a, b; Ziegler et al. 2017; Kim et al. 2019). Mutations in F6’H1 are detrimental for growth on alkaline substrates (Rodríguez-Celma et al. 2013b; Schmid et al. 2014). While these observations circumstantiate that coumarin production is increased in roots of Fe-deficient plants, it remained unclear which of the compounds are critical for the mobilisation of Fe. The phytoalexin scopoletin was found to accumulate in Fe-deficient A. thaliana roots (Lan et al. 2011; Rosenkranz et al. 2021). However, scopoletin is not – or to a limited extent—able to mobilise Fe (Schmid et al. 2014; Rajniak et al. 2018), albeit there is some debate as to whether this holds true in soil (Baune et al. 2020). The formation of scopoletin from 6-hydroxyferuloyl-CoA can occur spontaneously aided by light, but in roots, in particular under Fe-deficient conditions, the reaction is mediated by a member of the BAHD-acyltransferase family, COUMARIN SYNTHASE (COSY) (Vanholme et al. 2019). SCOPOLETIN 8-HYDROXYLASE (S8H) converts scopoletin into the catechol fraxetin (7,8-dihydroxy-6-methoxycoumarin), which appears to be the dominant Fe-mobilising coumarin at neutral and alkaline pH (Siwinska et al. 2018; Tsai et al. 2018). Resembling bacterial catechol-type siderophores (Hider and Kong 2010), fraxetin features two adjacent hydroxyl groups that efficiently bind and mobilize Fe from Fe(III)-oxides by chelation and reduction (Sisó-Terraza et al. 2016a; Tsai et al. 2018). Recently, Rajniak et al. (2018) identified sideretin (5-hydroxyfraxetin) as a further catecholic coumarin produced by Fe-deficient plants, synthesized through hydroxylation of fraxetin via the cytochrome P450 enzyme CYP82C4. Fraxetin, sideretin, and esculetin (6,7-dihydroxycoumarin) are the major players that enable A. thaliana (and very likely other species) to extract Fe from calcareous soils. Coumarins are stored as glucosides in the vacuole and are deglycosylated by β-glucosidases such as BGLU42 before secretion (Stringlis et al. 2018).
Coumarins: is that all there is?
Besides coumarins, Fe-deficient plants release various kinds of low-molecular-weight compounds into the rhizosphere, including organic acids, amino acids, sugars, flavins, and flavonoids, which can act as nutrient sources for microbes, growth promoters, chemoattractants, phytoalexins, chelators, acidifiers, detoxifiers, or growth inhibitors (Susín et al. 1993; 1994; Schmidt 1999; Jin et al. 2007; Cesco et al. 2010; Dakora and Phillips 2002; Mimmo et al. 2014; Astolfi et al. 2020). Some species, such as Beta vulgaris or Medicago truncatula, secrete flavins when subjected to Fe deficiency (Susín et al. 1993), a response that was thought to be mutually exclusive to the exudation of coumarins (Rodríguez-Celma et al. 2013a, b), although this does not always seem to be the case (Lefèvre et al. 2018). Notably, flavins and coumarins appear to be functionally analogous in promoting the reductive dissolution of scarcely soluble Fe(III) compounds (Rodríguez-Celma et al. 2013b; Sisó-Terraza et al. 2016b). At present, it is unclear why some plant species prioritize flavin secretion while others rely on the exudation of coumarins. Crucially, secretion of riboflavin is favoured by low pH, whereas fraxetin is more abundant in exudates from plants grown on alkaline media (Susín et al. 1993; Sisó-Teraza et al. 2016a). This difference is related to the secretion but not to the dissolution of Fe; both fraxetin and riboflavin can mobilize Fe at pH values prevailing in calcareous soils (Shi et al. 2013; Tsai et al. 2018). Of note, in contrast to B. vulgaris and M. truncatula, Medicago scutellata secretes flavins under both acidic and alkaline conditions (Gheshlaghi et al. 2021), possibly indicating a specific adaptation of Medicago scutellata to alkaline soils.
As a word of caution, it should be mentioned that the secretion of Fe-mobilising compounds determined under controlled conditions may not adequately reflect the exudation rates in natural substrates. Only limited information is available regarding the concentration of exudates in the rhizosphere, but a recent study suggests that coumarin levels in soil may be lower than those assayed in sterile agar-based media or hydroponics (Sarashgi et al. 2021). Moreover, under more natural conditions, scopoletin was found to dominate over catecholic coumarins (Rosenkranz et al. 2021). These observations are raising the question as to whether coumarins are in fact the magic potion conferring calcicole traits. However, coumarin-deficient f6’h1 mutants are unable to grow in calcareous soils (Schmid et al. 2014; Rosenkranz et al. 2021), an observation that corroborates the importance of Fe-mobilising root exudates for survival in such soils. High reactivity with soil components, which restricts the action of catecholic coumarins to the proximity of the roots, could provide an alternative explanation for their reduced detection in the soil solution. We will discuss other roles and peculiarities of coumarins later on in this review.
Phytosiderophores: evolution’s best offer?
The chelation-based Fe uptake strategy of grasses is less affected by media pH than reduction-based Fe acquisition and works efficiently in calcareous soils (Fig. 3). Secretion of phytosiderophores is mediated by TRANSPORTER OF MUGINEIC ACID FAMILY PHYTOSIDEROPHORES 1 (TOM1), first identified in rice and barley (Nozoye et al. 2011). Uptake of the Fe(III)-phytosiderophore complex into root cells is mediated by the high-affinity plasma membrane transporter YELLOW STRIPE 1 (YS1) (Curie et al. 2001) and YELLOW STRIPE 1-LIKE proteins, i.e., OsYSL15 in rice (Fig. 1) (Lee et al. 2009). Phytosiderophores are also critically involved in the long-distance transport of Fe. In rice phloem, the primary form of Fe is Fe(III)-deoxy-mugineic acid (Nishiyama et al. 2012); xylem saps of barley, maize, and rice were shown to contain mugineic acid and 2-deoxy-mugineic acid (Alam et al. 2001; Kawai et al. 2001; Ariga et al. 2014). Mugineic acid biosynthesis is initiated by the formation of nicotianamine (NA) via trimerization of S-adenosyl methionine catalysed by NICOTIANAMINE SYNTHASE (NAS), followed by an amino group transfer to form a 3-keto intermediate, a step that is mediated by NICOTIANAMINE AMINOTRANSFERASE (NAAT) (Higuchi et al. 1999; Kanazawa et al. 1995). Reduction of the 3’ carbon of the intermediate via DEOXYMUGINEIC ACID SYNTHASE (DMAS) yields deoxymugineic acid (Bashir et al. 2006). The various mugineic acid family phytosiderophores are then formed by subsequent steps in a species- and cultivar-dependent manner (Ma et al. 1995).
The ability to thrive in calcareous soils appears to be tightly correlated with the amount of secreted MA. Rice plants transformed with barley genes encoding enzymes of the MA biosynthetic pathway showed higher tolerance to low Fe availability in calcareous soils than untransformed plants (Suzuki et al. 2008; Gómez-Galera et al. 2012; Banakar et al. 2017). Similarly, in wild grasses, phytosiderophore secretion was higher in calcicole than in calcifuge species, suggesting that grasses of the latter group are excluded from calcareous soil due to their low Fe acquisition efficiency (Gries and Runge 1992; 1995). Notably, calcicoles not only secreted more siderophores, they also had lower metabolic requirements for Fe. Factoring in the latter trait, an up to 50 times higher Fe efficiency of calcicole compared to calcifuge grass taxa was estimated (Gries and Runge 1995). Thus, this dual adaptation of high MA secretion and low Fe demand seems to be decisive for the ability to colonize calcareous soils in grass species. It should be noted, however, that—similar to coumarins secreted by strategy I species—the complex interaction between soil characteristics, nutrient status of the plant, and phytosiderophore secretion may lead to an overestimation of the concentration of phytosiderophores under experimental conditions relative to concentrations determined in situ (Oburger et al. 2014).
In rice, IRT-type transporters are induced upon Fe starvation in addition to phytosiderophore-mediated Fe uptake (Ishimaru et al. 2006), a process that was thought to have evolved as a consequence of the cultivation of rice plants in paddy fields, where high concentrations of Fe2+ are readily available for IRT1-mediated Fe uptake. OsIRT orthologs are, however, also responsive to Fe deficiency in wild Oryza species, suggesting species-dependent evolution of this strategy preceding rice domestication (Wairich et al. 2019). Notably, as a further strategy I component, rice plants secrete phenolic compounds such protocatechuic acid and caffeic acid through a transporter designated PEZ2 (Bashir et al. 2011).
Inter-organ signalling orchestrates root Fe uptake
The tomato (Solanum lycopersicum) mutant T3238fer harbours a recessive mutation that renders it incapable to mount any of the typical Fe deficiency responses (Brown et al. 1971; Ling et al. 1996). The mutation was later mapped to the transcription factor SlFER/SlbHLH85 (Ling et al. 2002). SlFER was the first identified constituent of a surprisingly complex signalling cascade, comprising a suite of bHLH proteins that keep the cellular concentration of Fe balanced by positively and negatively regulating proteins involved in Fe uptake from the soil (reviewed in Liang 2022; Gao et al. 2019; Gao and Dubos 2021; Riaz and Guerinot 2021). The A. thaliana homolog of SlFER is FIT (FER-LIKE IRON DEFICIENCY-INDUCED TRANSCRIPTION FACTOR; bHLH29) (Jakoby et al. 2004; Colangelo and Guerinot 2004; Yuan et al. 2005). FIT forms heterodimers with the clade Ib bHLH proteins bHLH38, bHLH39, bHLH100, and bHLH101 to regulate genes mediating key processes of the strategy I Fe uptake system, i.e., FRO2, IRT1, AHA2, F6ʹH1, S8H, and CYP82C4 (Fig. 3) (Yuan et al. 2008; Wang et al. 2013). Within these heterodimers, clade Ib proteins confer DNA binding, while FIT possess transcription activation activity to interdependently regulate Fe uptake (Cai et al. 2022).
FIT and its homologs are exclusively expressed in root cells, necessitating information from above-ground to adequately adjust Fe uptake rates. The photosystems in chloroplasts are the strongest sink for Fe and are, most probably, the sites from which the demand for Fe is communicated to regulate its uptake by roots. Iron deficiency appears to be first sensed in the phloem of leaf cells, supporting this concept (Khan et al. 2018). Shoot-to-root communication that steers Fe uptake was demonstrated by various experimental setups. Altering the demand for and the supply with Fe by cooling the root zone while exposing the shoots to warmer air triggers an increase in root Fe uptake to compensate for the decreased transport of Fe to the leaves (Schmidt and Steinbach 2000). Similar observations were made with split-root plants, where one half of the root system is exposed to Fe while the other half is grown in Fe-deplete media (Schmidt et al. 1996; Vert et al. 2003; Tabata et al. 2022). In this set-up, the Fe demand of the shoots must be provided by the Fe-replete half of the root system, tuning uptake rates to compensate for the lack of Fe supply by the other half of the root system. Conspicuously, the response to Fe starvation is more pronounced in the Fe-supplied root portion, a pattern that seems to be advantageous for efficient mining of Fe. These results are indicative of an intricate interplay between local and systemic signals and unambiguously suggest one or—more likely—a suite of positive and negative signals that migrate between different plant parts to orchestrate root Fe uptake.
The nature of the long-distance signal(s) remains largely enigmatic. Iron, loaded into the phloem of leaf cells by oligopeptide transporters (OPTs), and IRONMAN (IMA) peptides have been suggested as respective negative and positive regulators, but the observed pattern appears to be more complex. AtOPT3 and its rice homolog OsOPT7 play critical roles in Fe homeostasis (Stacey et al. 2008; Mendoza-Cózatl et al. 2014; Bashir et al. 2015). AtOPT3 is preferentially expressed on the plasma membrane of phloem cells in the leaf vasculature and required for shoot-to-root transport of Fe. Similar to what was observed for transgenic A. thaliana lines overexpressing IMA, mutations in OPT3 lead to a constitutive upregulation of the root Fe deficiency responses, while shoot cells show the transcriptional signature of Fe toxicity (Grillet et al. 2018; Hirayama et al. 2018; Khan et al. 2018). In A. thaliana, shoot-specific expression of OPT3 rescued the opt3 phenotype (Mendoza-Cózatl et al. 2014). Supplying opt3 roots with Fe, however, did not decrease the constitutively elevated root reductase activity of the mutant, supporting a concept in which the perception of phloem Fe by root cells is critical for proper Fe sensing.
In an attempt to further clarify this matter, Tabata et al. (2022) undertook a transcriptomic analysis of split-root halves of A. thaliana plants exposed to either Fe-free or Fe-replete media. Besides FRO2 and IRT1, this survey found in particular genes involved in coumarin secretion to be up-regulated in the Fe-exposed half of the root system, a response that was absent in septuple ima7x mutants with minimal IMA activity. Reciprocal grafting experiments with wild-type plants and IMA3 overexpressing lines showed that IMA3 is required for systemic signalling, a route which includes the transcription factor MYB72 (Tabata et al. 2022). MYB72 and its homolog MYB10 were previously shown to be critical for the survival in alkaline soil (Palmer et al. 2013). Notably, the expression of IMA3 (≈80-fold), MYB72 (> 400-fold), and S8H (≈190-fold) is dramatically induced by bicarbonate treatment (Chen et al. 2021), indicative of a massive increase in the production and—most likely—secretion of coumarins in the presence of bicarbonate. In line with the upregulation of Fe uptake genes in the Fe-supplied split-root half, local resupply of Fe to cucumber roots caused a boost of Fe uptake in the Fe-resupplied portion of the roots, followed by rapid translocation to leaves and the other, Fe-free grown part of the root system (Valentinuzzi et al. 2020). Together, these observations are rather consistent with a root-shoot–root communication route than with simple shoot-to-root signalling. Employment of such a circuit is supported by the observation that separated roots can perceive biotic stimuli such as the local colonialization of microbial communities, and drive systemic exudation of metabolites by unshared areas of the root system (Korenblum et al. 2020).
The transcription factor ELONGATED HYPOCOTYL 5 (HY5) was recently introduced as novel actor in Fe-related long-distance communication. In tomato plants, phyB-dependent accumulation and subsequent binding of shoot-derived SlHY5 to SlFER orchestrate downstream Fe deficiency responses (Guo et al. 2021). HY5 is phloem mobile and able to migrate from shoots to roots (Chen et al. 2016), making this protein well-suited for conveying information between plant parts. In A. thaliana, HY5 was found to bind to the promotor of the putative Fe sensor BRUTUS (BTS) to regulate Fe-responsive genes, supporting a role of HY5 in long-distance Fe signalling (Mankotia et al. 2022). The expression of genes involved in Fe uptake, in particular those encoding proteins mediating the production of coumarins (i.e., F6’H1, S8H, CYP82C4, PDR9, and BGLU42) was compromised in hy5 mutants. When grown in calcareous soil, homozygous hy5 mutants are more chlorotic than wild-type plants, underlining the importance of HY5 for coumarin-mediated Fe acquisition (Mankotia et al. 2022). It is still unclear though as to how specificity of HY5 action is achieved, since HY5 is also involved in the regulation of the uptake of N, P, and Cu (Chen et al. 2016; Sakuraba et al. 2018; Zhang et al. 2014).
Transcripts of the cation/H+ exchanger (CAX) family member MdCAX3 were recently shown to represent further means of inter-organ communication (Hao et al. 2022). In apple (Malus domestica), MdCAX3 mRNA migrates from shoots to the roots, where it is translated and activated by the molecular chaperone MdCXIP1. MdCAX3 pumps Zn2+ ions into the vacuole and prevents MdIRT1 from being degraded by the elevated Zn2+ levels typically observed in roots of Fe-deficient plants. Degradation of AtIRT1 by secondary substrates such as Zn2+, possibly to prevent the accumulation of toxic levels of secondary non-Fe metal substrates, was reported for A. thaliana roots (Dubeaux et al. 2018). In addition, MdCAX3 aids in Fe acquisition by activating plasma membrane ATPases to acidify the apoplast.
Consistent with Fe being sensed in the vasculature, communication of the shoot Fe status to the roots is impaired in A. thaliana mutants defective in the plasma membrane-bound NA-metal transporters YSL1 and YSL2, which are predominantly expressed in the leaf vasculature (Kumar et al. 2017). In contrast to the constitutively upregulated Fe deficiency responses of opt3 roots, the ysl1 ysl2 mutant does not—or to a lesser extent—mount such responses when subjected to Fe-deficient conditions (Kumar et al. 2017). Homozygous ysl1 ysl2 double mutant plants are able to induce a partial Fe-deficiency response in their leaves, suggesting that it is not the sensing of Fe that is compromised. It appears that in ysl1 ysl2 plants Fe is not efficiently removed from the veins, resulting in elevated Fe levels in the vasculature. A probable scenario explaining the phenotype of ysl1 ysl2 involves OPT3, which may transport enough Fe from the vasculature to the roots to signal Fe sufficiency despite compromised lateral transport of, which in turn triggers a partial Fe deficiency response in the leaves (Kumar et al. 2017).
Crucially, the expression of both AtIMA1 and AtYSL1 in shoots is affected by compromised trimethylation of lysine 27 of histone 3 (H3K27me3) (Park et al. 2020), suggesting that post-translational histone modifications add a further layer to the complexity of inter-organ Fe signalling. Differential expression and differential H3K27me3 deposits were observed for AtIMA1 and AtbHLH39 in response to phosphate starvation, which, indirectly, alters the Fe status of the plants (Yen et al. 2017). Histone methylation may be more globally changed (but less well-explored) in response to alterations of the Fe status (Yen et al. 2017; Singh et al. 2020). The picture that emerges from these examples implies a rather complex interplay between different signals involved in inter-organ communication, an elaborate network which has only begun to be explored.
Peculiarities of Fe acquisition contribute to calcicole behaviour
Elevated root reductase activity, introduced by transgenic approaches or caused by natural variation, appears to confer chlorosis resistance to plants growing in calcareous soils. Reduction of ferric Fe is the rate-limiting step in Fe uptake (Grusak et al. 1990), making strategies to improve Fe efficiency through elevated root reductase activity a promising approach. Several studies appear to support this concept. For example, heterologous expression of the A. thaliana FRO2 gene ameliorated chlorosis symptoms in soybean (Vasconcelos et al. 2006). Furthermore, non-coding allelic variation in the FRO2 gene among natural A. thaliana accessions was found to be associated with root length, FRO2 expression, and root reduction activity, with accessions having higher reductase activity performing better in calcareous substrate (Satbhai et al. 2017). However, chlorosis-resistant species, cultivars, or demes are not generally characterized by high ferric chelate activity (see below and de la Guardia and Alcántara 2002; Terés et al. 2019; Wang et al. 2022).
A major constraint for strategy I plants inhabiting calcareous soils is the pronounced inhibition of root-mediated ferric chelate reduction at neutral or alkaline pH (Susín et al. 1996). Repression of FRO2 activity under such conditions is caused by decreased gene expression (Santi and Schmidt 2009; Tsai and Schmidt 2021) and—more importantly—decreased reduction rates in vivo, possibly caused by repulsion of (negatively charged) Fe(III) chelates by (also negatively charged) pectic polysaccharides in the cell wall. The slightly acidic pH optimum of root Fe reduction is maintained by net proton efflux supported by P-type ATPases, a process that is efficiently buffered in calcareous soils and renders this component of the strategy I-type Fe acquisition system rather inefficient. Moreover, expression of AtAHA2 (but not that of the housekeeping AtAHA1) is reduced at elevated pH (Santi and Schmidt 2009). Engineering a Fe(III) chelate reductase for better performance at high pH revealed that an amino acid substitution at position 312 is critical for the activity of the yeast reductase FRE1 under alkaline conditions (Ishimaru et al. 2006; Oki et al. 2004). When expressed in rice under control of the OsIRT1 promoter, the transgene—referred to as reconstructed FRE1 (refre1)—conferred higher Fe chelate reduction rates, increased Fe uptake, elevated grain yield, and improved tolerance to low Fe availability in calcareous soils (Ishimaru et al. 2007). The positive outcome of this approach raises the question as to why no such pH-insensitive reductase has been evolved in calcicole plant species. The answer may lie in an energetically unfavourable outcome of such an endeavour; since high redox activity may come with the penalty of low affinity (Km) for the substrate. In a comparison of Plantago species differing in their ecological distribution pattern, maximal velocity (Vmax) of the Fe chelate reductase was found to be an insufficient proxy for the efficiency of Fe uptake among the species under study. Iron uptake efficiency was rather associated with the Km of the reaction (Schmidt and Fühner 1998), suggesting that substrate availability is a decisive factor for the tolerance to low Fe availability. Notably, olive trees, adapted to Mediterranean calcareous soils and highly tolerant to low available Fe, do not increase their ferric reductase activity upon Fe starvation (de la Guardia and Alcántara 2002). Thus, another possible explanation for the absence of a pH-insensitive reductase in calcicole species may lie in the fact that other solutions are simply more efficient.
Iron chelating ligands extend the pH range for efficient Fe uptake
While the elucidation of the signalling cascades leading to adaptive changes in gene expression in plants is still pending, it has been observed that short-term shifts in media pH caused remarkably rapid and pronounced changes in the transcriptome of plants (Lager et al. 2010; Payá-Milans et al. 2017; Pérez-Martín et al. 2021). Similarly, transferring Fe-deficient A. thaliana plants from slightly acidic pH to media with circumneutral pH and restricted Fe availability altered the expression of a large subset of genes (Tsai and Schmidt 2020). Most of the genes that were differentially expressed in response to the change in pH are not generally responsive to the Fe regime, suggesting that pH- and Fe-signalling run largely separate courses. However, in some remarkable cases, pH- and Fe-signalling circuits appear to merge to prioritise specific phenotypic readouts. An example of such intertwining of signalling pathways emerged from the above-mentioned survey, which revealed a pronounced repression of the gene encoding the sideretin-producing enzyme CYP82C4 at elevated pH, a response that was later confirmed in other studies (Fig. 4) (Gautam et al. 2021). By contrast, expression of S8H, mediating the preceding formation of fraxetin, was robustly increased by high pH (Tsai and Schmidt 2020; Gautam et al. 2021; Chen et al. 2021). This pH-dependent control of gene activity results in the production of different coumarins at different external hydrogen concentrations: while acidic pH supports the biosynthesis of sideretin, elevated pH favours the formation of its less oxidised homolog fraxetin (Fig. 4). The oppositional regulation of the two genes seems, at first sight, to be counterintuitive, given the fact that both genes are controlled by the transcription factor FIT and are highly induced (at acidic pH) by Fe deficiency. Ecologically, however, a pH-dependent prioritization of specific coumarins appears to be highly advantageous. Sideretin is unstable at high pH and less efficient in mobilizing Fe at high pH than fraxetin (Rajniak et al. 2018; Tsai et al. 2018; Sisó-Terraza et al. 2016b; Gautam et al. 2021). Fraxetin, on the other hand, is critical for survival on substrates with high pH and limited Fe availability (Tsai et al. 2018). Without pH-dependent modulation of gene expression, the coumarin biosynthetic pathway would inevitably lead to the biosynthesis of sideretin, which does not support Fe uptake under alkaline conditions. Thus, the pH-dependent regulation of the pathway allows for efficient Fe mining under a wide range of edaphic conditions.

Adapted from Tsai and Schmidt (2021). Figure was created with BioRender.com
pH-dependent production of coumarins in response to external pH. Acidic media pH supports the biosynthesis of sideretin via CYP82C4. At circumneutral or alkaline pH, a putative pH signal interferes with the default gene expression pattern, supporting the expression of S8H while repressing CYP82C4. This differential regulation of the last and penultimate steps of the coumarin pathway supports the production of fraxetin, which is more efficient in mobilizing Fe under such conditions.
Growth, chlorophyll, and Fe content correlated with fraxetin content among A. thaliana accessions differing in performance on high pH/low Fe media, corroborating the supposition that fraxetin is a major component of calcicole behaviour (Tsai et al. 2018). In support of this finding, the decisive trait for the adaptation of an A. thaliana deme to carbonate-rich soil was a higher secretion of catechol-type coumarins such as esculetin and fraxetin relative to plants native to Ca-poor soil (Terés et al. 2019). Notably, a deme adapted to carbonate-rich soil exhibited lower root reductase activity than a carbonate‐sensitive deme, suggesting that the higher reductase activity of plants from the non-adapted population reflected a lower Fe status rather than an adaptive trait (Terés et al. 2019). Thus, secretion of coumarins (or other Fe-mobilising compounds) appears to be a more promising strategy for efficient Fe mining at elevated pH than increased root reduction activity. Importantly, secretion of coumarins can – at least to a certain extent or under certain conditions – bypass the reductive Fe uptake system, enabling plants to take up Fe-coumarin complexes without prior reduction of Fe (Fig. 3). Re-uptake of secreted coumarins is energetically favourable and provides a competitive advantage over other organisms that may otherwise profit from the secreted compounds. Coumarins taken up from the rhizosphere could be glycosylated and stored in the vacuole before undergoing a new circle of de-glycosylation, secretion, and chelation. Evidence for such a ‘merry-go-round’ of coumarins derived from the observation that coumarin-deficient A. thaliana f6’h1 mutants contain fraxetin when provided exogenously, indicating that roots can indeed take up catecholic coumarins (Robe et al. 2021a, b, c). Such re-uptake of Fe(III)-coumarin complexes is a plausible scenario which awaits further support. A transporter for Fe(III)-chelates has, however, not yet been identified.
The uptake of Fe(III) coumarin complexes by non-gramineous plants resembles the promotive effect observed during intercropping. Here, strategy I plants such as peanut (Arachis hypogaea) profit from phytosiderophores secreted by grasses that are taken up via AhYSL1 or AhYSL3.1 transporters, the expression of which is induced in intercropped peanut plants (Xiong et al. 2013; Guo et al. 2014). Notably, these two YSL proteins were shown to complement the yeast fet3fet4 double mutant (defective in high- and low-affinity Fe-uptake systems) supplied with Fe(III)- deoxymugineic acid (Xiong et al. 2013). It thus appears that strategy I plants are able to hijack phytosiderophores with an auxiliary Fe uptake system, which may have a wide range of suitable substrates (Fig. 3). The next question to be asked is as to whether strategy I plants can produce and secrete phytosiderophores of the MA family and have thus adopted components of the strategy II system just as rice (and possibly other strategy II species) employs traits typical of strategy I plants. In support of this supposition, olive (Olea europaea) trees were found to contain endogenous 2’-deoxymugineic acid and contigs (i.e., contiguous representations of a genomic region) coding for MA biosynthesis enzymes such as NAS, NAAT, and DMAS as well as for YS1 transporters (Suzuki et al. 2016). Moreover, MA derivatives were identified in root exudates of Fe-deficient tomato plants in an untargeted metabolomic survey (Astolfi et al. 2020). The assumption that tomato roots can indeed produce phytosiderophores was supported by the presence of putative SlTOM1 orthologous genes in the tomato genome. Phytosiderophores were also detected in exudates of grapevine (Marastoni et al. 2020) and in the metal hyper-tolerant species Nicotiana thyrsiflora and Puya sp. (Kińska et al. 2022). It appears from these observations that the mechanisms for Fe uptake are not as strictly separated by phylogenetic borders as previously suggested. However, the reciprocal adoption of non-typical Fe acquisition mechanisms seems to represent auxiliary, non-inducible systems that may be critical under specific conditions, but are not as equally important as the canonical Fe uptake strategy of the respective group. Moreover, it remains to be unequivocally demonstrated that MA-type phytosiderophores are secreted and taken up by strategy I plants and whether this mechanism is of significant physiological importance for this group of plants.
The plant-microbiome holobiont
The term holobiont refers to an amalgamated organism composed of microbial species and the eukaryotic host, where a suite of microbes harboured within and on the outer surface of the root provide additional functionality (Meyer-Abich 1943; Margulis and Fester 1991; Vandenkoornhuyse et al. 2015; Theis et al. 2016; Baedke et al. 2020). The interaction between host and microbiome can be viewed as a facilitator that provides the plant with additional genes, a concept that has been referred to as ‘the powerhouse of the adjustment to local conditions’ (Vandenkoornhuyse et al. 2015).
Plant–microbe interactions are of particular importance for the uptake of Fe. The rhizosphere is strongly influenced by carbon-rich molecules derived from or secreted by plant roots, compounds that can be both detrimental or beneficial for the microbes. Rhizosphere microorganisms such as bacteria, fungi, archaea, viruses, and oomycetes may profit from the host by using the various kinds of rhizodeposits (i.e., material lost from plant roots) such as exudates, nutrients, mucilage, and volatiles as an energy source. Iron acquisition by the host, on the other hand, may be augmented by bacterial siderophores through one or more of the following mechanisms: 1) direct uptake of the Fe-siderophore complex, 2) limiting the growth of pathogens and, consequently, reduced competition for Fe in the rhizosphere, and 3) by stimulation of the host Fe uptake system by microbes. Thus, the host affects the activity, composition, and functional plasticity of the microbiome, and the microbiome affects growth, Fe status, and fitness of the plant (Fig. 5).
Plant-microbiome interactions relevant to Fe acquisition. Root exudates such as coumarins can be beneficial to soil bacteria or repel plant pathogens. The microbiome in the rhizosphere can support the host’s Fe uptake by stimulation of plant Fe uptake, by repressing the growth of pathogenic bacteria and thus reducing the competition for soil Fe, and by hijacking bacterial siderophores. Beneficial microbes can further strengthen the plant’s defence by inducing systemic resistance. Coumarins secreted by the plant improve Fe uptake by providing substrate for the reducing Fe uptake system and via uptake of the Fe(III) coumarin complex as such. PS; phytosiderophores. Figure was created with BioRender.com
Cross-species hijacking of siderophores in the soil
Bacteria produce a wide range of Fe-scavenging siderophores, which play important roles in ecosystem functioning and participate in a variety of ecological processes such as pathogen competition, Fe cycling in soils, cross-kingdom signalling, and plant growth promotion (Trivedi et al. 2020). Crucially, siderophore-producing species appear to be enriched in the rhizosphere relative to the bulk soil (Jin et al. 2010; Coleman-Derr et al. 2016; Bulgarelli et al. 2015; Xu et al. 2018). Bacterial siderophores mobilize Fe in the soil and could provide a bio-available source of soluble Fe for plants, a mechanism that can become critical in soils in which other sources of Fe are firmly limited. In principle, plants may assimilate Fe from bacterial siderophores either by reductive splitting of Fe(III) chelates and subsequent uptake of the released ferrous ion, direct absorption of siderophore-Fe complexes as such, or after ligand exchange followed by uptake of the plant-borne Fe-chelate. While the exact mechanism by which plants profit from the abundance of bacterial siderophores remain to be elucidated, a suite of studies supports the view that plants indeed efficiently highjack Fe bound to bacterial siderophores.
Unequivocal evidence for such a mechanism derived from an investigation by Vansuyt et al. (2007), showing that A. thaliana plants can take up the bacterial siderophore pyoverdine. The authors demonstrated that supplementation of the media with the bacterial siderophore rescued the A. thaliana irt1 mutant, whereas substrates of the FRO2/IRT1 uptake system such as FeEDTA were ineffective. Pyoverdine was immunologically detected in plant roots, excluding other mechanisms such as ligand exchange to account for the observation. A transcriptomic survey conducted later revealed that application of pyoverdine in its Fe-free structure (apo-pyoverdine) induces the expression of Fe uptake genes and represses defense-related genes (Trapet et al. 2016), indicating wide-ranging effects of the bacterial siderophore on growth and immunity. Uptake of Fe chelators of bacterial origin has been suggested earlier for siderophores produced by the plant growth-promoting bacterium Pseudomonas sp. strain GRP3 and mung bean (Vigna radiata L. Wilzeck) (Sharma et al. 2003), and since then for several bacterial-plant systems such as Azospirillum brasilense and cucumber (Pii et al. 2015), Pseudomonas strain sp. SP3 and apple (Gao et al. 2022), and P. illinoisensis and peanut plants (Liu et al. 2017). While the the transporter(s) facilitating the uptake of bacterial siderophores remain to be identified, the potential importance of the highly diverse cocktail of Fe-siderophore complexes of bacterial origin for the mobilization of Fe is immense. More than 500 different bacterial siderophores with hydroxamate, catecholate, and α-hydroxycarboxylate functional groups have been identified, (Hider and Kong 2010), suggesting niche specificity and functional diversity of the various siderophores in metal acquisition (Baars et al. 2016). Thus, the potential ability for plants to utilise Fe pools of poor mobility is extended to a great extent, allowing for survival in stands with extremely poor Fe availability.
Another way by which commensal bacteria improve plant Fe acquisition is via stimulation of the host’s Fe uptake system. Inoculation of rice plants with the endophytic Streptomyces hygroscopicus OsiSh-2 improved the Fe status of its host by exploiting the highly efficient Fe uptake system of the bacterium in which siderophore secretion was increased during rice colonisation (Cao et al. 2021). Moreover, OsiSh-2 increased the host’s Fe acquisition by up-regulation plant-specific processes. Interestingly, OsiSh-2 inoculation specifically induced the strategy I-type Fe uptake system in rice such as root-mediated ferric reduction and media acidification, while phytosiderophore production was down-regulated. These observations suggest that the bacterium supports Fe acquisition through two different modes: by providing Fe-loaded bacterial siderophores and by inducing specific components of the host Fe uptake mechanism (Cao et al. 2021).
Conspicuously, secretion of siderophores is associated with the loss of substantial energy resources, making it attractive to highjack Fe and siderophores produced by other organisms. Factually, such siderophore piracy is supported by the evolution of uptake pathways for siderophores of foreign origin (Lee et al. 2012; Harrington et al. 2015), improving Fe uptake efficiency of the cheater and attenuating that of the producer. Other studies show that siderophore piracy is, in fact, bidirectional. In the grass Brachypodium distachyon, genes involved in phytosiderophore production were upregulated when plants were cultured with the soil bacterium Pseudomonas fluorescens SBW25::gfp/lux (SBW25), but the abundance of secreted phytosiderophores was lower compared to media without the bacterium (Boiteau et al. 2021). By contrast, the presence of B. distachyon suppressed pyoverdine production by the bacterium. These observations suggest that microbes can induce phytosiderophore production of their host and intercept the alien Fe complexes as a Fe source.
The effects of microbial-derived siderophores on the host are, however, not restricted to the scenarios outline above. A further mechanism is tied to the competition for Fe among plants, pathogens, and beneficial bacteria. By assessing the ability of siderophores produced by various rhizosphere bacteria to suppress or promote the growth of the pathogenic bacterium Ralstonia solanacearum, it was found that pathogen-suppressive microbes produce siderophores that the pathogen cannot take up, resulting in plant protection (Gu et al. 2020). Thus, bacterial siderophores can improve plant growth indirectly and govern microbiome function in natural ecosystems.
Root exudates shape the microbiome
Coumarin secretion by plants not only mobilises Fe in the soil, but also considerably modulates the structure of microbial assemblages (Stringlis et al. 2018, 2019; Voges et al. 2019; Harbort et al. 2020; Hou et al. 2021). By employing a synthetic microbial community (SymCom), Voges et al. (2019) demonstrated that secretion of catecholic coumarins by plant roots triggered a beneficial shift of the microbial community, repressing proliferation of the Pseudomonas strain sp. Root329 but not that of the host-beneficial Pseudomonas simiae WCS417; plants harbouring mutations in genes that compromise the production or secretion of catecholic coumarins such as f6’h1 and pdr9 were ineffective (Voges et al. 2019). Notably, root colonization by WCS417 stimulates the host’s Fe uptake, leading to improved growth and increased Fe levels in both roots and shoots (Verbon et al. 2019). Volatile organic compounds (VOCs) from beneficial Pseudomonas can induce AtMYB72 by an AtOPT3- and Fe-status-independent, photosynthesis-related shoot-to-root signal (Zamioudis et al. 2015; Verbon et al. 2019), suggesting that beneficial bacteria can induce processes downstream of and controlled by MYB72 such as the secretion of coumarins and systemic resistance. Thus, coumarins secreted by plants can alter bacterial communities, which in turn induce coumarin secretion via MYB72 in a feed forward manner (Fig. 5). The mechanisms by which microbes induce MYB72 expression are not yet fully resumed, but at least in the case of Trichoderma VOCs this process appears to be associated with a rapid burst of nitric oxide (Pescador et al. 2022), a mechanism that may apply to other bacteria-host systems.
Microbiota populations are not stable and subject to rapid evolution driven by environmental constraints. Plant-antagonistic Pseudomonas protegens bacteria were shown to evolve into mutualists within six plant growth cycles due to improved competition for root exudates and a higher tolerance to the root-derived phytoalexin scopoletin, indicative of a fast plant-mediated ‘ecological filtering’ (Li et al. 2021). Surprisingly, the new mutualists were able to induce the expression of MYB72 in A. thaliana, which promoted the production of coumarins and plant growth (Li et al. 2021). Thus, plants cannot only recruit specific beneficial bacteria, but can also alter the functionality of the associated microbiota (Fig. 5).
The catechol fraxetin appears to be of particular importance for plant-microbiome-communication and critical for commensal-mediated plant growth stimulation (Harbort et al. 2020). A surprising finding was the observation that commensals support the growth of the coumarin-deficient f6ʹh1 mutant. The concentration of fraxetin required to rescue the chlorotic phenotype displayed by f6ʹh1 plants when grown in calcareous substrate was significantly lower in the presence of commensals compared to that required for ameliorating the f6’h1 phenotype in commensal-free media (Harbort et al. 2020), suggesting a boost of the host’s Fe uptake system by a yet unexplored mechanism not related to coumarin secretion by the host.
Plants growing in calcareous soils are generally undersupplied with multiple mineral nutrients and may have more complex effects on the microbiome. Unsurprisingly, deficiencies in other mineral nutrients such as P or imbalances in Fe/phosphate supply can alter the composition of root exudates and, subsequently, microbiome composition (Chutia et al. 2019; Tang et al. 2022). In response to phosphate starvation, A. thaliana roots secrete immune-related RAPID ALKALINIZATION FACTOR (RALF) peptides to suppress plant immunity via the receptor kinase FERONIA, which shifts the microbiome towards colonization by bacteria that induce phosphate starvation genes and mitigate phosphate deficiency (Tang et al. 2022). Thus, the plant nutritional status is critical for the composition of the microbiome, which, in turn, support the acquisition of limiting nutrients.
Notably, the communication between the microbiome and plants is not strictly confined to roots. In A. thaliana, suboptimal light conditions perceived by leaves were shown to alter root bacterial communities, which support growth of the host, but decrease its foliar defences. Thus, root commensals can trigger and steer also above-ground stress responses in plants, indicating that microbiota-root-shoot communication circuits are drivers of the fitness and phenotypic plasticity of the holobiont (Hou et al. 2021; Stassen et al. 2021). Such signalling circuits might be widespread and crucial for the prioritization of both below- and above-ground plant stress responses.
Conclusions
Much has been learned regarding the mechanisms and regulation of Fe uptake over the past decade. The acknowledgement of the importance of Fe-mobilising exudates of both plant and microbial origin, which extend the range of edaphic conditions over which Fe uptake is supported, has revealed the tremendous plasticity by which plants can respond to edaphic cues. The plethora of interactions between the root microbiome and the host has added another facet to the ways and means by which plants take up Fe, an aspect that has not yet been explored comprehensively. It appears that plants function as an integrated continuum with the microbiome and ‘outsource’ parts of the Fe acquisition tasks to profit from soil bacteria. The mutual exchange between plants and microbiota is not restricted to the trade of goods such as Fe and organic compounds, but contributes to—or even determines—plant fitness in calcareous soils. It should also be stated that the ability to thrive in such soils represents a multifactorial blend of physiological readouts comprising various facets. Ecological factors, such as competition, ecological amplitudes, and habitat preferences, further complicates the picture. It seems obvious, however, that the efficiency by which Fe is mined from alkaline soils is one of the strongest determinants for conferring calcicole behaviour. Fitting plants to a given soil, either by traditional breeding methods or via transgenic approaches, requires knowledge regarding the traits conferring Fe efficiency on alkaline substrates, knowledge that aids in understanding functionally of natural ecosystems and in securing nutritional quality of edible plant parts.
References
Aida M, Beis D, Heidstra R, Willemsen V, Blilou I, Galinha C, Nussaume L, Noh YS, Amasino R, Scheres B (2004) The PLETHORA genes mediate patterning of the Arabidopsis root stem cell niche. Cell 119(1):109–120. https://doi.org/10.1016/j.cell.2004.09.018
Alhendawi RA, Römheld V, Kirkby EA, Marschner H (1997) Influence of increasing bicarbonate concentrations on plant growth, organic acid accumulation in roots and iron uptake by barley, sorghum, and maize. J Plant Nutrition 20(12):1731–1752. https://doi.org/10.1080/01904169709365371
Alam S, Kamei S, Kawai S (2001) Metal micronutrients in xylem sap of iron-deficient barley as affected by plant-borne, microbial, and synthetic metal chelators. Soil Sci Plant Nutr 47(1):149–156. https://doi.org/10.1080/00380768.2001.10408377
Ariga T, Hazama K, Yanagisawa S, Yoneyama T (2014) Chemical forms of iron in xylem sap from graminaceous and non-graminaceous plants. Soil Sci Plant Nutr 60(4):460–469. https://doi.org/10.1080/00380768.2014.922406
Arrivault S, Senger T, Krämer U (2006) The Arabidopsis metal tolerance protein AtMTP3 maintains metal homeostasis by mediating Zn exclusion from the shoot under Fe deficiency and Zn oversupply. Plant J 46(5):861–879. https://doi.org/10.1111/j.1365-313X.2006.02746.x
Astolfi S, Pii Y, Mimmo T, Lucini L, Miras-Moreno MB, Coppa E, Violino S, Celletti S, Cesco S (2020) Single and combined Fe and S deficiency differentially modulate root exudate composition in tomato: A double strategy for Fe acquisition? Int J Mol Sci 21(11):4038. https://doi.org/10.3390/ijms21114038
Baune M, Kang K, Schenkeveld WD, Kraemer SM, Hayen H, Weber G (2020) Importance of oxidation products in coumarin-mediated Fe (hydr) oxide mineral dissolution. Biometals 33(6):305–321. https://doi.org/10.1007/s10534-020-00248-y
Bashir K, Inoue H, Nagasaka S, Takahashi M, Nakanishi H, Mori S, Nishizawa NK (2006) Cloning and characterization of deoxymugineic acid synthase genes from graminaceous plants. J Biol Chem 281(43):32395–32402. https://doi.org/10.1074/jbc.M604133200
Bashir K, Ishimaru Y, Shimo H, Kakei Y, Senoura T, Takahashi R, Sato Y, Sato Y, Uozumi N, Nakanishi H, Nishizawa NK (2011) Rice phenolics efflux transporter 2 (PEZ2) plays an important role in solubilizing apoplasmic iron. Soil Sci Plant Nutr 57(6):803–812. https://doi.org/10.1080/00380768.2011.637305
Bashir K, Ishimaru Y, Itai RN, Senoura T, Takahashi M, An G, Oikawa T, Ueda M, Sato A, Uozumi N, Nakanishi H (2015) Iron deficiency regulated OsOPT7 is essential for iron homeostasis in rice. Plant Mol Biol 88(1):165–176. https://doi.org/10.1007/s11103-015-0315-0
Banakar R, Alvarez Fernandez A, Abadía J, Capell T, Christou P (2017) The expression of heterologous Fe (III) phytosiderophore transporter HvYS 1 in rice increases Fe uptake, translocation and seed loading and excludes heavy metals by selective Fe transport. Plant Biotechnol J 15(4):423–432. https://doi.org/10.1111/pbi.12637
Baedke J, Fábregas-Tejeda A, Nieves Delgado A (2020) The holobiont concept before Margulis. J Exp Zool Part B 334(3):149–155. https://doi.org/10.1002/jez.b.22931
Baars O, Zhang X, Morel FM, Seyedsayamdost MR (2016) The siderophore metabolome of Azotobacter vinelandii. Appl Environ Microbiol 82(1):27–39. https://doi.org/10.1128/AEM.03160-15
Boiteau RM, Markillie LM, Hoyt DW, Hu D, Chu RK, Mitchell HD, Pasa-Tolic L, Jansson JK, Jansson C (2021) Metabolic interactions between Brachypodium and Pseudomonas fluorescens under controlled iron-limited conditions. mSystems 6(1):e00580-20. https://doi.org/10.1128/mSystems.00580-20
Brown JC, Chaney RL, Ambler JE (1971) A new tomato mutant inefficient in the transport of iron. Physiol Plant 25(1):48–53. https://doi.org/10.1111/j.1399-3054.1971.tb01086.x
Bulgarelli D, Garrido-Oter R, Münch PC, Weiman A, Dröge J, Pan Y, McHardy AC, Schulze-Lefert P (2015) Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17(3):392–403. https://doi.org/10.1016/j.chom.2015.01.011
Chaney RL, Brown JC, Tiffin LO (1972) Obligatory reduction of ferric chelates in iron uptake by soybeans. Plant Physiol 50(2):208–13. https://doi.org/10.1104/pp.50.2.208
Chytrý M, Danihelka J, Axmanová I, Božková J, Hettenbergerova E, Li CF, Rozbrojova Z, Sekulová L, Tichý L, Vymazalova M, Zelený D (2010) Floristic diversity of an eastern Mediterranean dwarf shrubland: the importance of soil pH. J Veg Sci 21(6):1125–1137. https://doi.org/10.1111/j.1654-1103.2010.01212.x
Cornell RM, Schwertmann U (2003) The iron oxides: structure, properties, reactions, occurrences, and uses (Vol. 2, p. 71). Weinheim: Wiley-vch. https://doi.org/10.1515/CORRREV.1997.15.3-4.533
Chen J, Li X, Ye X, Guo P, Hu Z, Qi G, Cui F, Liu S (2021) An S-ribonuclease binding protein EBS1 and brassinolide signaling are specifically required for Arabidopsis tolerance to bicarbonate. J Exp Bot 72(4):1449–1459. https://doi.org/10.1093/jxb/eraa524
Chen X, Yao Q, Gao X, Jiang C, Harberd NP, Fu X (2016) Shoot-to-root mobile transcription factor HY5 coordinates plant carbon and nitrogen acquisition. Curr Biol 26(5):640–646. https://doi.org/10.1016/j.cub.2015.12.066
Chutia R, Abel S, Ziegler J (2019) Iron and phosphate deficiency regulators concertedly control coumarin profiles in Arabidopsis thaliana roots during iron, phosphate, and combined deficiencies. Front Plant Sci 10:113. https://doi.org/10.3389/fpls.2019.00113
Cesco S, Neumann G, Tomasi N, Pinton R, Weisskopf L (2010) Release of plant-borne flavonoids into the rhizosphere and their role in plant nutrition. Plant Soil 329(1):1–25. https://doi.org/10.1007/s11104-009-0266-9
Curie C, Panaviene Z, Loulergue C, Dellaporta SL, Briat JF, Walker EL (2001) Maize yellow stripe1 encodes a membrane protein directly involved in Fe (III) uptake. Nature 409(6818):346–349. https://doi.org/10.1038/35053080
Colangelo EP, Guerinot ML (2004) The essential basic helix-loop-helix protein FIT1 is required for the iron deficiency response. Plant Cell 16(12):3400–3412. https://doi.org/10.1105/tpc.104.024315
Cai Y, Yang Y, Ping H, Lu C, Lei R, Li Y, Liang G (2022) Why FIT and bHLH Ib interdependently regulate Fe-uptake. bioRxiv 2022.02.12.480172. https://doi.org/10.1101/2022.02.12.480172
Coleman-Derr D, Desgarennes D, Fonseca-Garcia C, Gross S, Clingenpeel S, Woyke T, North G, Visel A, Partida-Martinez LP, Tringe SG (2016) Plant compartment and biogeography affect microbiome composition in cultivated and native Agave species. New Phytol 209(2):798–811. https://doi.org/10.1111/nph.13697
Cao L, Gao Y, Yu J, Niu S, Zeng J, Yao Q, Wang X, Bu Z, Xu T, Liu X, Zhu Y (2021) Streptomyces hygroscopicus OsiSh-2-induced mitigation of Fe deficiency in rice plants. Plant Physiol Biochem 158:275–283. https://doi.org/10.1016/j.plaphy.2020.11.013
Ding W, Clode PL, Lambers H (2020) Effects of pH and bicarbonate on the nutrient status and growth of three Lupinus species. Plant Soil 447(1):9–28. https://doi.org/10.1007/s11104-019-03980-8
Dubeaux G, Neveu J, Zelazny E, Vert G (2018) Metal sensing by the IRT1 transporter-receptor orchestrates its own degradation and plant metal nutrition. Mol Cell 69(6):953–964. https://doi.org/10.1016/j.molcel.2018.02.009
Dakora FD, Phillips DA (2002) Root exudates as mediators of mineral acquisition in low-nutrient environments. Food security in nutrient-stressed environments: exploiting plants’ genetic capabilities, 201-213. https://doi.org/10.1007/978-94-017-1570-6_23
de Mello Gabriel GV, Pitombo LM, Rosa LM, Navarrete AA, Botero WG, do Carmo JB, de Oliveira LC (2021) The environmental importance of iron speciation in soils: evaluation of classic methodologies. Environ Monit Assess 193(2):1–9. https://doi.org/10.1007/s10661-021-08874-w
de Souza MC, Williams TC, Poschenrieder C, Jansen S, Pinheiro MH, Soares IP, Franco AC (2020) Calcicole behaviour of Callisthene fasciculata Mart., an Al-accumulating species from the Brazilian Cerrado. Plant Biol 22(1):30–37. https://doi.org/10.1111/plb.13036
de la Guardia MD, Alcántara E (2002) A comparison of ferric-chelate reductase and chlorophyll and growth ratios as indices of selection of quince, pear and olive genotypes under iron deficiency stress. Plant Soil 241(1):49–56. https://doi.org/10.1023/A:1016083512158
Eide D, Broderius M, Fett J, Guerinot ML (1996) A novel iron-regulated metal transporter from plants identified by functional expression in yeast. PNAS 93(11):5624–5628. https://doi.org/10.1073/pnas.93.11.5624
Ellenberg H (1958) Bodenreaktion (einschließlich Kalkfrage). In Die Mineralische Ernährung der Pflanze/Mineral Nutrition of Plants (pp. 638–708). Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-94729-2_23
Fourcroy P, Sisó-Terraza P, Sudre D, Savirón M, Reyt G, Gaymard F, Abadía A, Abadia J, Álvarez-Fernández A, Briat JF (2014) Involvement of the ABCG 37 transporter in secretion of scopoletin and derivatives by Arabidopsis roots in response to iron deficiency. New Phytol 201(1):155–167. https://doi.org/10.1111/nph.12471
Galinha C, Hofhuis H, Luijten M, Willemsen V, Blilou I, Heidstra R, Scheres B (2007) PLETHORA proteins as dose-dependent master regulators of Arabidopsis root development. Nature 449(7165):1053–1057. https://doi.org/10.1038/nature06206
Gautam CK, Tsai HH, Schmidt W (2021) IRONMAN tunes responses to iron deficiency in concert with environmental pH. Plant Physiol 187(3):1728–1745. https://doi.org/10.1093/plphys/kiab329
Gao F, Robe K, Gaymard F, Izquierdo E, Dubos C (2019) The transcriptional control of iron homeostasis in plants: a tale of bHLH transcription factors? Front Plant Sci 10:6. https://doi.org/10.3389/fpls.2019.00006
Gao F, Dubos C (2021) Transcriptional integration of plant responses to iron availability. J Exp Bot 72(6):2056–2070. https://doi.org/10.1093/jxb/eraa556
Gao B, Chai X, Huang Y, Wang X, Han Z, Xu X, Wu T, Zhang X, Wang Y (2022) Siderophore production in Pseudomonas sp. strain SP3 enhances iron acquisition in apple rootstock. J Appl Microbiol. https://doi.org/10.1111/jam.15591
Gheshlaghi Z, Luis-Villarroya A, Álvarez-Fernández A, Khorassani R, Abadía J (2021) Iron deficient Medicago scutellata grown in nutrient solution at high pH accumulates and secretes large amounts of flavins. Plant Sci 303:110664. https://doi.org/10.1016/j.plantsci.2020.110664
Guo Z, Xu J, Wang Y, Hu C, Shi K, Zhou J, Xia X, Zhou Y, Foyer CH, Yu J (2021) The phyB-dependent induction of HY5 promotes iron uptake by systemically activating FER expression. EMBO Rep 22(7):e51944. https://doi.org/10.15252/embr.202051944
Guo X, Xiong H, Shen H, Qiu W, Ji C, Zhang Z, Zuo Y (2014) Dynamics in the rhizosphere and iron-uptake gene expression in peanut induced by intercropping with maize: role in improving iron nutrition in peanut. Plant Physiol Biochem 76:36–43. https://doi.org/10.1016/j.plaphy.2013.12.019
Gu S, Wei Z, Shao Z, Friman VP, Cao K, Yang T, Kramer J, Wang X, Li M, Mei X, Xu Y (2020) Competition for iron drives phytopathogen control by natural rhizosphere microbiomes. Nat Microbiol 5(8):1002–1010. https://doi.org/10.1038/s41564-020-0719-8
Gómez-Galera S, Sudhakar D, Pelacho AM, Capell T, Christou P (2012) Constitutive expression of a barley Fe phytosiderophore transporter increases alkaline soil tolerance and results in iron partitioning between vegetative and storage tissues under stress. Plant Physiol Biochem 53:46–53. https://doi.org/10.1016/j.plaphy.2012.01.009
Gries D, Runge M (1992) The ecological significance of iron mobilization in wild grasses. J Plant Nutr 15(10):1727–1737. https://doi.org/10.1080/01904169209364434
Gries D, Runge M (1995) Responses of calcicole and calcifuge poaceae species to iron-limiting conditions. Bot Acta 108(6):482–489. https://doi.org/10.1111/j.1438-8677.1995.tb00525.x
Grillet L, Lan P, Li W, Mokkapati G, Schmidt W (2018) IRON MAN is a ubiquitous family of peptides that control iron transport in plants. Nat Plants 4(11):953–963. https://doi.org/10.1038/s41477-018-0266-y
Grime JP, Hodgson JG (1968) Investigation of the ecological significance of lime-chlorosis by means of largescale comparative experiments. In Brit Ecol Soc Symp 9 67–99
Grusak MA, Welch RM, Kochian LV (1990) Does iron deficiency in Pisum sativum enhance the activity of the root plasmalemma iron transport protein? Plant Physiol 94(3):1353–1357. https://doi.org/10.1104/pp.94.3.1353
Hayashi M, Fukuzawa T, Sorimachi H, Maeda T (2005) Constitutive activation of the pH-responsive Rim101 pathway in yeast mutants defective in late steps of the MVB/ESCRT pathway. Mol Cell Biol 25(21):9478–9490. https://doi.org/10.1128/MCB.25.21.9478-9490.2005
Hao P, Lv X, Fu M, Xu Z, Tian J, Wang Y, Zhang X, Xu X, Wu T, Han Z (2022) Long‐distance mobile mRNA CAX3 modulates iron uptake and zinc compartmentalization. EMBO Rep e53698. https://doi.org/10.15252/embr.202153698
Harrington JM, Duckworth OW, Haselwandter K (2015) The fate of siderophores: antagonistic environmental interactions in exudate-mediated micronutrient uptake. Biometals 28(3):461–472. https://doi.org/10.1007/s10534-015-9821-4
Harbort CJ, Hashimoto M, Inoue H, Niu Y, Guan R, Rombolà AD, Kopriva S, Voges MJ, Sattely ES, Garrido-Oter R, Schulze-Lefert P (2020) Root-secreted coumarins and the microbiota interact to improve iron nutrition in Arabidopsis. Cell Host Microbe 28(6):825–837. https://doi.org/10.1016/j.chom.2020.09.006
Hsieh EJ, Waters BM (2016) Alkaline stress and iron deficiency regulate iron uptake and riboflavin synthesis gene expression differently in root and leaf tissue: implications for iron deficiency chlorosis. J Exp Bot 67(19):5671–5685. https://doi.org/10.1093/jxb/erw328
Higuchi K, Suzuki K, Nakanishi H, Yamaguchi H, Nishizawa NK, Mori S (1999) Cloning of nicotianamine synthase genes, novel genes involved in the biosynthesis of phytosiderophores. Plant Physiol 119(2):471–480. https://doi.org/10.1104/pp.119.2.471
Hirayama T, Lei GJ, Yamaji N, Nakagawa N, Ma JF (2018) The putative peptide gene FEP1 regulates iron deficiency response in Arabidopsis. Plant Cell Physiol 59(9):1739–1752. https://doi.org/10.1093/pcp/pcy145
Hider RC, Kong X (2010) Chemistry and biology of siderophores. Nat Prod Rep 27(5):637–657. https://doi.org/10.1039/b906679a
Hou S, Thiergart T, Vannier N, Mesny F, Ziegler J, Pickel B, Hacquard S (2021) A microbiota–root–shoot circuit favours Arabidopsis growth over defence under suboptimal light. Nat Plants 7(8):1078–1092. https://doi.org/10.1038/s41477-021-00956-4
Ishimaru Y, Suzuki M, Tsukamoto T, Suzuki K, Nakazono M, Kobayashi T, Wada Y, Watanabe S, Matsuhashi S, Takahashi M, Nakanishi H (2006) Rice plants take up iron as an Fe3+-phytosiderophore and as Fe2+. Plant J 45(3):335–346. https://doi.org/10.1111/j.1365-313X.2005.02624.x
Ishimaru Y, Kim S, Tsukamoto T, Oki H, Kobayashi T, Watanabe S, Matsuhashi S, Takahashi M, Nakanishi H, Mori S, Nishizawa NK (2007) Mutational reconstructed ferric chelate reductase confers enhanced tolerance in rice to iron deficiency in calcareous soil. Proc Natl Acad Sci USA 104(18):7373–7378. https://doi.org/10.1073/pnas.0610555104
Jakoby M, Wang HY, Reidt W, Weisshaar B, Bauer P (2004) FRU (BHLH029) is required for induction of iron mobilization genes in Arabidopsis thaliana. FEBS Lett 577(3):528–534. https://doi.org/10.1016/j.febslet.2004.10.062
Jin CW, You GY, He YF, Tang C, Wu P, Zheng SJ (2007) Iron deficiency-induced secretion of phenolics facilitates the reutilization of root apoplastic iron in red clover. Plant Physiol 144(1):278–285. https://doi.org/10.1104/pp.107.095794
Jin CW, Li GX, Yu XH, Zheng SJ (2010) Plant Fe status affects the composition of siderophore-secreting microbes in the rhizosphere. Ann Bot 105(5):835–841. https://doi.org/10.1093/aob/mcq071
Kawai S, Kamei S, Matsuda Y, Ando R, Kondo S, Ishizawa A, Alam S (2001) Concentrations of iron and phytosiderophores in xylem sap of iron-deficient barley plants. Soil Sci Plant Nutr 47(2):265–272. https://doi.org/10.1080/00380768.2001.10408390
Kanazawa K, Higuchi K, Nishizawa NK, Fushiya S, Mori S (1995) Detection of two distinct isozymes of nicotianamine aminotransferase in Fe-deficient barley roots. J Exp Bot 46(9):1241–1244. https://doi.org/10.1093/jxb/46.9.1241
Kińska K, Cruzado-Tafur E, Parailloux M, Torró L, Lobinski R, Szpunar J (2022) Speciation of metals in indigenous plants growing in post-mining areas: Dihydroxynicotianamine identified as the most abundant Cu and Zn ligand in Hypericum laricifolium. Sci Total Environ 809:151090. https://doi.org/10.1016/j.scitotenv.2021.151090
Kroh GE, Pilon M (2020) Regulation of iron homeostasis and use in chloroplasts. Int J Mol Sci 21(9):3395. https://doi.org/10.3390/ijms21093395
Kotula L, Clode PL, Ranathunge K, Lambers H (2021) Role of roots in adaptation of soil-indifferent Proteaceae to calcareous soils in south-western Australia. J Exp Bot 72(4):1490–1505. https://doi.org/10.1093/jxb/eraa515
Kobayashi T, Nishizawa NK (2012) Iron uptake, translocation, and regulation in higher plants. Annu Rev Plant Biol 63:131–52. https://doi.org/10.1146/annurev-arplant-042811-105522
Kim SA, LaCroix IS, Gerber SA, Guerinot ML (2019) The iron deficiency response in Arabidopsis thaliana requires the phosphorylated transcription factor URI. Proc Natl Acad Sci USA 116(50):24933–24942. https://doi.org/10.1073/pnas.1916892116
Khan MA, Castro-Guerrero NA, McInturf SA, Nguyen NT, Dame AN, Wang J, Bindbeutel RK, Joshi T, Jurisson SS, Nusinow DA, Mendoza-Cozatl DG (2018) Changes in iron availability in Arabidopsis are rapidly sensed in the leaf vasculature and impaired sensing leads to opposite transcriptional programs in leaves and roots. Plant Cell Environ 41(10):2263–2276. https://doi.org/10.1111/pce.13192
Korenblum E, Dong Y, Szymanski J, Panda S, Jozwiak A, Massalha H, Meir S, Rogachev I, Aharoni A (2020) Rhizosphere microbiome mediates systemic root metabolite exudation by root-to-root signaling. Proc Natl Acad Sci USA 117(7):3874–3883. https://doi.org/10.1073/pnas.1912130117
Kumar RK, Chu HH, Abundis C, Vasques K, Rodriguez DC, Chia JC, Huang R, Vatamaniuk OK, Walker EL (2017) Iron-nicotianamine transporters are required for proper long distance iron signaling. Plant Physiol 175(3):1254–1268. https://doi.org/10.1104/pp.17.00821
Lan P, Li W, Wen TN, Shiau JY, Wu YC, Lin W, Schmidt W (2011) iTRAQ protein profile analysis of Arabidopsis roots reveals new aspects critical for iron homeostasis. Plant Physiol 155(2):821–834. https://doi.org/10.1104/pp.110.169508
Lager ID, Andréasson OL, Dunbar TL, Andreasson E, Escobar MA, Rasmusson AG (2010) Changes in external pH rapidly alter plant gene expression and modulate auxin and elicitor responses. Plant Cell Environ 33(9):1513–1528. https://doi.org/10.1111/j.1365-3040.2010.02161.x
Lee S, Chiecko JC, Kim SA, Walker EL, Lee Y, Guerinot ML, An G (2009) Disruption of OsYSL15 leads to iron inefficiency in rice plants. Plant Physiol 150(2):786–800. https://doi.org/10.1104/pp.109.135418
Lee W, van Baalen M, Jansen VA (2012) An evolutionary mechanism for diversity in siderophore-producing bacteria. Ecol Lett 15(2):119–125. https://doi.org/10.1111/j.1461-0248.2011.01717.x
Lefèvre F, Fourmeau J, Pottier M, Baijot A, Cornet T, Abadía J, Álvarez-Fernández A, Boutry M (2018) The Nicotiana tabacum ABC transporter NtPDR3 secretes O-methylated coumarins in response to iron deficiency. J Exp Bot 69(18):4419–4431. https://doi.org/10.1093/jxb/ery221
Li E, de Jonge R, Liu C, Jiang H, Friman VP, Pieterse CM, Bakker PA, Jousset A (2021) Rapid evolution of bacterial mutualism in the plant rhizosphere. Nat Commun 12(1):1–13. https://doi.org/10.1038/s41467-021-24005-y
Liang G (2022) Iron uptake, signaling, and sensing in plants. Plant Commun 14:100349. https://doi.org/10.1016/j.xplc
Link HF (1789) Florae Göttingensis specimen, sistens vegetabilia saxo calcareo propria: Diss. Inaug. Botan.
Lindsay WL, Schwab AP (1982) The chemistry of iron in soils and its availability to plants. J Plant Nutr 5(4–7):821–840
Liu L, Song W, Huang S, Jiang K, Moriwaki Y, Wang Y, Men Y, Zhang D, Wen X, Han Z, Chai J, Guo H (2022) Extracellular pH sensing by plant cell-surface peptide-receptor complexes. Cell S0092–8674(22)00915–1. https://doi.org/10.1016/j.cell.2022.07.012
Lucena C, Romera FJ, Rojas CL, García MJ, Alcántara E, Pérez-Vicente R (2007) Bicarbonate blocks the expression of several genes involved in the physiological responses to Fe deficiency of Strategy I plants. Functional Plant Biol 34(11):1002–1009. https://doi.org/10.1071/FP07136
Ling HQ, Pich A, Scholz G, Ganal MW (1996) Genetic analysis of two tomato mutants affected in the regulation of iron metabolism. Mol Gen Genet MGG 252(1):87–92. https://doi.org/10.1007/BF02173208
Ling HQ, Bauer P, Bereczky Z, Keller B, Ganal M (2002) The tomato fer gene encoding a bHLH protein controls iron-uptake responses in roots. Proc Natl Acad Sci USA 99(21):13938–13943. https://doi.org/10.1073/pnas.212448699
Liu D, Yang Q, Ge K, Hu X, Qi G, Du B, Liu K, Ding Y (2017) Promotion of iron nutrition and growth on peanut by Paenibacillus illinoisensis and Bacillus sp. strains in calcareous soil. Braz J Microbiol 48:656–670. https://doi.org/10.1016/j.bjm.2017.02.006
Ma JF, Shinada T, Matsuda C, Nomoto K (1995) Biosynthesis of phytosiderophores, mugineic acids, associated with methionine cycling. J Biol Chem 270(28):16549–16554. https://doi.org/10.1074/jbc.270.28.16549
Margulis L, Fester R (1991) Symbiosis as a source of evolutionary innovation: speciation and morphogenesis. MIT Press, Cambridge, Massachusetts, USA
Martínez-Cuenca MR, Legaz F, Forner-Giner MÁ, Primo-Millo E, Iglesias DJ (2013) Bicarbonate blocks iron translocation from cotyledons inducing iron stress responses in Citrus roots. J Plant Physiol 170(10):899–905. https://doi.org/10.1016/j.jplph.2013.01.012
Martín-Barranco A, Spielmann J, Dubeaux G, Vert G, Zelazny E (2020) Dynamic control of the high-affinity iron uptake complex in root epidermal cells. Plant Physiol 184(3):1236–1250. https://doi.org/10.1104/pp.20.00234
Mankotia S, Singh D, Monika K, Meena H, Meena V, Yadav R, Pandey AK, Satbhai SB (2022) Elongated Hypocotyl 5 (HY5) regulates BRUTUS (BTS) to maintain iron homeostasis in Arabidopsis thaliana. bioRxiv. https://doi.org/10.1101/2022.04.26.489524
Marastoni L, Lucini L, Miras-Moreno B, Trevisan M, Sega D, Zamboni A, Varanini Z (2020) Changes in physiological activities and root exudation profile of two grapevine rootstocks reveal common and specific strategies for Fe acquisition. Sci Rep 10(1):1–12. https://doi.org/10.1038/s41598-020-75317-w
Mendoza-Cózatl DG, Xie Q, Akmakjian GZ, Jobe TO, Patel A, Stacey MG, Song L, Demoin DW, Jurisson SS, Stacey G, Schroeder JI (2014) OPT3 is a component of the iron-signaling network between leaves and roots and misregulation of OPT3 leads to an over-accumulation of cadmium in seeds. Mol Plant 7(9):1455–1469. https://doi.org/10.1093/mp/ssu067
Meyer-Abich A (1943) I. Das typologische Grundgesetz und seine Folgerungen für Phylogenie und Entwicklungsphysiologie. Acta Biotheor 7(1):1–80. https://doi.org/10.1007/BF01603792
Mimmo T, Del Buono D, Terzano R, Tomasi N, Vigani G, Crecchio C, Pinton R, Zocchi G, Cesco S (2014) Rhizospheric organic compounds in the soil–microorganism–plant system: their role in iron availability. Eur J Soil Sci 65(5):629–642. https://doi.org/10.1111/ejss.12158
Nishiyama R, Kato M, Nagata S, Yanagisawa S, Yoneyama T (2012) Identification of Zn–nicotianamine and Fe–2’-deoxymugineic acid in the phloem sap from rice plants (Oryza sativa L.). Plant Cell Physiol 53(2):381–390. https://doi.org/10.1093/pcp/pcr188
Nozoye T, Nagasaka S, Kobayashi T, Takahashi M, Sato Y, Sato Y, Uozumi N, Nakanishi H, Nishizawa NK (2011) Phytosiderophore efflux transporters are crucial for iron acquisition in graminaceous plants. J Biol Chem 286(7):5446–5454. https://doi.org/10.1074/jbc.M110.180026
Obara K, Yamamoto H, Kihara A (2012) Membrane protein Rim21 plays a central role in sensing ambient pH in Saccharomyces cerevisiae. J Biol Chem 287(46):38473–38481. https://doi.org/10.1074/jbc.M112.394205
Oburger E, Gruber B, Schindlegger Y, Schenkeveld WD, Hann S, Kraemer SM, Wenzel WW, Puschenreiter M (2014) Root exudation of phytosiderophores from soil-grown wheat. New Phytol 203(4):1161–1174. https://doi.org/10.1111/nph.12868
Oki H, Kim S, Nakanishi H, Takahashi M, Yamaguchi H, Mori S, Nishizawa NK (2004) Directed evolution of yeast ferric reductase to produce plants with tolerance to iron deficiency in alkaline soils. Soil Sci Plant Nutr 50(7):1159–1165. https://doi.org/10.1080/00380768.2004.10408589
Payá-Milans M, Nunez GH, Olmstead JW, Rinehart TA, Staton M (2017) Regulation of gene expression in roots of the pH-sensitive Vaccinium corymbosum and the pH-tolerant Vaccinium arboreum in response to near neutral pH stress using RNA-Seq. BMC Genomics 18(1):1–16. https://doi.org/10.1186/s12864-017-3967-0
Palmer CM, Hindt MN, Schmidt H, Clemens S, Guerinot ML (2013) MYB10 and MYB72 are required for growth under iron-limiting conditions. PLoS Genet 9(11):e1003953. https://doi.org/10.1371/journal.pgen.1003953
Park EY, Tsuyuki KM, Parsons EM, Jeong J (2020) PRC2-mediated H3K27me3 modulates shoot iron homeostasis in Arabidopsis thaliana. Plant Signaling Behav 15(9):1784549. https://doi.org/10.1080/15592324.2020.1784549
Pérez-Martín L, Busoms S, Tolrà R, Poschenrieder C (2021) Transcriptomics reveals fast changes in salicylate and jasmonate signaling pathways in shoots of carbonate-tolerant Arabidopsis thaliana under bicarbonate exposure. Int J Mol Sci 22(3):1226. https://doi.org/10.3390/ijms22031226
Peñalva MA, Lucena-Agell D, Arst Jr HN (2014) Liaison alcaline: Pals entice non-endosomal ESCRTs to the plasma membrane for pH signaling. Curr Opin Microbiol 22:49–59. https://doi.org/10.1016/j.mib.2014.09.005
Pescador L, Fernandez I, Pozo MJ, Romero-Puertas MC, Pieterse CM, Martínez-Medina A (2022) Nitric oxide signalling in roots is required for MYB72-dependent systemic resistance induced by Trichoderma volatile compounds in Arabidopsis. J Exp Bot 73(2):584–595. https://doi.org/10.1093/jxb/erab294
Pii Y, Penn A, Terzano R, Crecchio C, Mimmo T, Cesco S (2015) Plant-microorganism-soil interactions influence the Fe availability in the rhizosphere of cucumber plants. Plant Physiol Biochem 87:45–52. https://doi.org/10.1016/j.plaphy.2014.12.014
Rajniak J, Giehl RF, Chang E, Murgia I, von Wirén N, Sattely ES (2018) Biosynthesis of redox-active metabolites in response to iron deficiency in plants. Nat Chem Biol 14(5):442–450. https://doi.org/10.1038/s41589-018-0019-2
Robe K, Conejero G, Gao F, Lefebvre-Legendre L, Sylvestre-Gonon E, Rofidal V, Hem S, Rouhier N, Barberon M, Hecker A, Gaymard F, Izquierdo E, Dubos C (2021) Coumarin accumulation and trafficking in Arabidopsis thaliana: a complex and dynamic process. New Phytol 229(4):2062–2079. https://doi.org/10.1111/nph.17090
Rodríguez-Celma J, Pan I, Li WD, Lan PD, Buckhout TJ, Schmidt W (2013a) The transcriptional response of Arabidopsis leaves to Fe deficiency. Front Plant Sci 4:276. https://doi.org/10.3389/fpls.2013.00276
Rodríguez-Celma J et al (2013b) Mutually exclusive alterations in secondary metabolism are critical for the uptake of insoluble iron compounds by Arabidopsis and Medicago truncatula. Plant Physiol 162(3):1473–1485. https://doi.org/10.1104/pp.113.220426
Römheld V, Marschner H (1986) Evidence for a specific uptake system for iron phytosiderophores in roots of grasses. Plant Physiol 80(1):175–180. https://doi.org/10.1104/pp.80.1.175
Robinson NJ, Procter CM, Connolly EL, Guerinot ML (1999) A ferric-chelate reductase for iron uptake from soils. Nature 397(6721):694–697. https://doi.org/10.1038/17800
Rosenkranz T, Oburger E, Baune M, Weber G, Puschenreiter M (2021) Root exudation of coumarins from soil-grown Arabidopsis thaliana in response to iron deficiency. Rhizosphere 17:100296. https://doi.org/10.1016/j.rhisph.2020.100296
Riaz N, Guerinot ML (2021) All together now: regulation of the iron deficiency response. J Exp Bot 72(6):2045–2055. https://doi.org/10.1093/jxb/erab003
Robe K, Conejero G, Gao F, Lefebvre-Legendre L, Sylvestre-Gonon E, Rofidal V, Hem S, Rouhier N, Barberon M, Hecker A, Gaymard F (2021b) Coumarin accumulation and trafficking in Arabidopsis thaliana: a complex and dynamic process. New Phytol 229(4):2062–2079. https://doi.org/10.1111/nph.17090
Robe K, Stassen M, Chamieh J, Gonzalez P, Hem S, Santoni V, Dubos C, Izquierdo E. (2021c). Uptake of Fe-fraxetin complexes, an IRT1 independent strategy for iron acquisition in Arabidopsis thaliana. bioRxiv 2021c.08.03.454955. https://doi.org/10.1101/2021.08.03.454955
Sakuraba Y, Kanno S, Mabuchi A, Monda K, Iba K, Yanagisawa S (2018) A phytochrome-B-mediated regulatory mechanism of phosphorus acquisition. Nat Plants 4(12):1089–1101. https://doi.org/10.1038/s41477-018-0294-7
Sarashgi A, Puschenreiter M, Baune M, Paffrath V, Oburger E, Giehl RFH, Rosenkranz T (2021) Does the exudation of coumarins from Fe-deficient, soil-grown Brassicaceae species play a significant role in plant Fe nutrition? Rhizosphere 19:100410. https://doi.org/10.1016/j.rhisph.2021.100410
Santi S, Schmidt W (2009) Dissecting iron deficiency-induced proton extrusion in Arabidopsis roots. New Phytol 183(4):1072–1084. https://doi.org/10.1111/j.1469-8137.2009.02908.x
Satbhai SB, Setzer C, Freynschlag F, Slovak R, Kerdaffrec E, Busch W (2017) Natural allelic variation of FRO2 modulates Arabidopsis root growth under iron deficiency. Nat Commun 8(1):1–10. https://doi.org/10.1038/ncomms15603
Schwertmann U (1991) Solubility and dissolution of iron oxides. Plant Soil 130(1):1–25. https://doi.org/10.1007/BF00011851
Schmid NB, Giehl RF, Döll S, Mock HP, Strehmel N, Scheel D, Kong X, Hider RC, von Wirén N (2014) Feruloyl-CoA 6’-Hydroxylase1-dependent coumarins mediate iron acquisition from alkaline substrates in Arabidopsis. Plant Physiol 164(1):160–172. https://doi.org/10.1104/pp.113.228544
Schmidt W (1999) Mechanisms and regulation of reduction-based iron uptake in plants. The New Phytol 141(1):1–26. https://doi.org/10.1046/j.1469-8137.1999.00331.x
Schmidt W, Steinbach S (2000) Sensing iron—A whole plant approach. Ann Bot 86(3):589–593. https://doi.org/10.1006/anbo.2000.1223
Schmidt W, Boomgaarden B, Ahrens V (1996) Reduction of root iron in Plantago lanceolata during recovery from Fe deficiency. Physiol Plant 98(3):587–593. https://doi.org/10.1111/j.1399-3054.1996.tb05715.x
Schmidt W, Fühner C (1998) Sensitivity to and requirement for iron in Plantago species. New Phytol 138(4):639–651. https://doi.org/10.1046/j.1469-8137.1998.00156.x
Schenkeveld WD, Wang Z, Giammar DE, Kraemer SM (2016) Synergistic effects between biogenic ligands and a reductant in Fe acquisition from calcareous soil. Environ Sci Technol 50(12):6381–6388. https://doi.org/10.1021/acs.est.6b01623
Shi Z, Zachara JM, Wang Z, Shi L, Fredrickson JK (2013) Reductive dissolution of goethite and hematite by reduced flavins. Geochim Cosmochim Acta 121:139–154. https://doi.org/10.1016/j.gca.2013.05.039
Siwinska J, Siatkowska K, Olry A, Grosjean J, Hehn A, Bourgaud F, Meharg AA, Carey M, Lojkowska E, Ihnatowicz A (2018) Scopoletin 8-hydroxylase: a novel enzyme involved in coumarin biosynthesis and iron-deficiency responses in Arabidopsis. J Exp Bot 69(7):1735–1748. https://doi.org/10.1093/jxb/ery005
Sisó-Terraza P, Rios JJ, Abadía J, Abadía A, Álvarez-Fernández A (2016a) Flavins secreted by roots of iron-deficient Beta vulgaris enable mining of ferric oxide via reductive mechanisms. New Phytol 209(2):733–745. https://doi.org/10.1111/nph.13633
Sisó-Terraza P, Luis-Villarroya A, Fourcroy P, Briat JF, Abadía A, Gaymard F, Abadía J, Álvarez-Fernández A (2016b) Accumulation and secretion of coumarinolignans and other coumarins in Arabidopsis thaliana roots in response to iron deficiency at high pH. Front Plant Sci 7:1711. https://doi.org/10.3389/fpls.2016.01711
Singh DP, Singh V, Shukla R, Sahu P, Prabha R, Gupta A, Sarma BK, Gupta VK (2020) Stage-dependent concomitant microbial fortification improves soil nutrient status, plant growth, antioxidative defense system and gene expression in rice. Microbiol Res 239:126538. https://doi.org/10.1016/j.micres.2020.126538
Stacey MG, Patel A, McClain WE, Mathieu M, Remley M, Rogers EE, Gassmann W, Blevins DG, Stacey G (2008) The Arabidopsis AtOPT3 protein functions in metal homeostasis and movement of iron to developing seeds. Plant Physiol 146(2):589–601. https://doi.org/10.1104/pp.107.108183
Stassen MJ, Hsu SH, Pieterse CM, Stringlis IA (2021) Coumarin communication along the microbiome–root–shoot axis. Trends Plant Sci 26(2):169–183. https://doi.org/10.1016/j.tplants.2020.09.008
Stringlis IA, Yu K, Feussner K, de Jonge R, Van Bentum S, Van Verk MC, Berendsen RL, Bakker PA, Feussner I, Pieterse CM (2018) MYB72-dependent coumarin exudation shapes root microbiome assembly to promote plant health. Proc Natl Acad Sci USA 115(22):E5213–E5222. https://doi.org/10.1073/pnas.1722335115
Stringlis IA, De Jonge R, Pieterse CM (2019) The age of coumarins in plant–microbe interactions. Plant Cell Physiol 60(7):1405–1419. https://doi.org/10.1093/pcp/pcz076
Susín S, Abian J, Sanchez-Baeza F, Peleato ML, Abadia A, Gelpi E, Abadia J (1993) Riboflavin 3 ‘-and 5 ‘-sulfate, two novel flavins accumulating in the roots of iron-deficient sugar beet (Beta vulgaris). J Biol Chem 268(28):20958–20965. https://doi.org/10.1016/S0021-9258(19)36879-6
Susín S, Abián J, Peleato ML, Sánchez-Baeza F, Abadía A, Gelpi E, Abadía J (1994) Flavin excretion from roots of iron-deficient sugar beet (Beta vulgaris L.). Planta 193(4):514–519. https://doi.org/10.1007/BF02411556
Susín S, Abadia A, González-Reyes JA, Lucena JJ, Abadia J (1996) The pH requirement for in vivo activity of the iron-deficiency-induced “Turbo” ferric chelate reductase. A comparison of the iron-deficiency-induced iron reductase activities of intact plants and isolated plasma membrane fractions in sugar beet. Plant Physiol 110(1):111–123. https://doi.org/10.1104/pp.110.1.111
Suzuki M, Nozoye T, Nagasaka S, Nakanishi H, Nishizawa NK, Mori S (2016) The detection of endogenous 2’-deoxymugineic acid in olives (Olea europaea L.) indicates the biosynthesis of mugineic acid family phytosiderophores in non-graminaceous plants. Soil Sci Plant Nutr 62(5–6):481–488. https://doi.org/10.1080/00380768.2016.1230724
Suzuki M, Morikawa KC, Nakanishi H, Takahashi M, Saigusa M, Mori S, Nishizawa NK (2008) Transgenic rice lines that include barley genes have increased tolerance to low iron availability in a calcareous paddy soil. Soil Sci Plant Nutr 54(1):77–85. https://doi.org/10.1111/j.1747-0765.2007.00205.x
Sharma A, Johri BN, Sharma AK, Glick BR (2003) Plant growth-promoting bacterium Pseudomonas sp. strain GRP3 influences iron acquisition in mung bean (Vigna radiata L. Wilzeck). Soil Biol Biochem 35(7):887–894. https://doi.org/10.1016/S0038-0717(03)00119-6
Tang J, Wu D, Li X, Wang L, Xu L, Zhang Y, Xu F, Liu H, Xie Q, Dai S, Coleman-Derr D (2022) Plant immunity suppression via PHR1-RALF-FERONIA shapes the root microbiome to alleviate phosphate starvation. EMBO J 41(6):e109102. https://doi.org/10.15252/embj.2021109102
Tansley AG (1917) On competition between Galium saxatile L. (G. hercynicum Weig.) and Galium sylvestre Poll. (G. asperum Schreb.) on different types of soil. J. Ecol 173-179. https://doi.org/10.2307/2255655
Takagi SI, Nomoto K, Takemoto T (1984) Physiological aspect of mugineic acid, a possible phytosiderophore of graminaceous plants. J Plant Nutr 7(1–5):469–477. https://doi.org/10.1080/01904168409363213
Tabata R, Kamiya T, Imoto S, Tamura H, Tabata M, Hirayama T, Tsukagoshi H, Tanoi K, Suzuki T, Hachiya T, Sakakibara H (2022) Systemic regulation of iron acquisition by Arabidopsis in environments with heterogeneous iron distributions. Plant Cell Physiol. https://doi.org/10.1093/pcp/pcac049
Terés J, Busoms S, Perez Martín L, Luís-Villarroya A, Flis P, Álvarez-Fernández A, Tolrà R, Salt DE, Poschenrieder C (2019) Soil carbonate drives local adaptation in Arabidopsis thaliana. Plant Cell Environ 42(8):2384–2398. https://doi.org/10.1111/pce.13567
Tsai HH, Schmidt W (2021) The enigma of environmental pH sensing in plants. Nat Plants 7(2):106–115. https://doi.org/10.1038/s41477-020-00831-8
Tsai HH, Rodríguez-Celma J, Lan P, Wu YC, Vélez-Bermúdez IC, Schmidt W (2018) Scopoletin 8-hydroxylase-mediated fraxetin production is crucial for iron mobilization. Plant Physiol 177(1):194–207. https://doi.org/10.1104/pp.18.00178
Tsai HH, Schmidt W (2020) pH-dependent transcriptional profile changes in iron-deficient Arabidopsis roots. BMC Genomics 21(1):1–11. https://doi.org/10.1186/s12864-020-07116-6
Tyler G (2003) Some ecophysiological and historical approaches to species richness and calcicole/calcifuge behaviour—contribution to a debate. Folia Geobot 38(4):419–428. https://doi.org/10.1007/BF02803249
Theis KR, Dheilly NM, Klassen JL, Brucker RM, Baines JF, Bosch TC, Cryan JF, Gilbert SF, Goodnight CJ, Lloyd EA, Sapp J (2016) Getting the hologenome concept right: an eco-evolutionary framework for hosts and their microbiomes. Msystems 1(2):e00028-e116. https://doi.org/10.1128/mSystems.00028-16
Trivedi P, Leach JE, Tringe SG, Sa T, Singh BK (2020) Plant–microbiome interactions: from community assembly to plant health. Nat Rev Microbiol 18(11):607–621. https://doi.org/10.1038/s41579-020-0412-1
Trapet P, Avoscan L, Klinguer A, Pateyron S, Citerne S, Chervin C, Mazurier S, Lemanceau P, Wendehenne D, Besson-Bard A (2016) The Pseudomonas fluorescens siderophore pyoverdine weakens Arabidopsis thaliana defense in favor of growth in iron-deficient conditions. Plant Physiol 171(1):675–693. https://doi.org/10.1104/pp.15.01537
Unger F (1836) Über den Einfluss des Bodens auf die Verteilung der Gewächse: nachgewiesen in der Vegetation des nordöstlichen Tirols. Bei Rohrmann und Schweigerd.
Vanholme R, Sundin L, Seetso KC, Kim H, Liu X, Li J, De Meester B, Hoengenaert L, Goeminne G, Morreel K, Haustraete J (2019) COSY catalyses trans–cis isomerization and lactonization in the biosynthesis of coumarins. Nat Plants 5(10):1066–1075. https://doi.org/10.1038/s41477-019-0510-0
Valentinuzzi F, Pii Y, Carlo P, Roberto T, Fontanella MC, Beone GM, Astolfi S, Mimmo T, Cesco S (2020) Root-shoot-root Fe translocation in cucumber plants grown in a heterogeneous Fe provision. Plant Sci 293:110431. https://doi.org/10.1016/j.plantsci.2020.110431
Vasconcelos M, Eckert H, Arahana V, Graef G, Grusak MA, Clemente T (2006) Molecular and phenotypic characterization of transgenic soybean expressing the Arabidopsis ferric chelate reductase gene, FRO2. Planta 224(5):1116–1128. https://doi.org/10.1007/s00425-006-0293-1
Vandenkoornhuyse P, Quaiser A, Duhamel M, Le Van A, Dufresne A (2015) The importance of the microbiome of the plant holobiont. New Phytol 206(4):1196–1206. https://doi.org/10.1111/nph.13312
Vansuyt G, Robin A, Briat JF, Curie C, Lemanceau P (2007) Iron acquisition from Fe-pyoverdine by Arabidopsis thaliana. Mol Plant-Microbe Interact 20(4):441–447. https://doi.org/10.1094/MPMI-20-4-0441
Vert G, Grotz N, Dédaldéchamp F, Gaymard F, Guerinot ML, Briat JF, Curie C (2002) IRT1, an Arabidopsis transporter essential for iron uptake from the soil and for plant growth. Plant Cell 14:1223–1233. https://doi.org/10.1105/tpc.001388
Vert GA, Briat JF, Curie C (2003) Dual regulation of the Arabidopsis high-affinity root iron uptake system by local and long-distance signals. Plant Physiol 132(2):796–804. https://doi.org/10.1104/pp.102.016089
Verbon EH, Trapet PL, Kruijs S, Temple-Boyer-Dury C, Rouwenhorst TG, Pieterse CM (2019) Rhizobacteria-mediated activation of the Fe deficiency response in Arabidopsis roots: impact on Fe status and signaling. Front Plant Sci, 909. https://doi.org/10.3389/fpls.2019.00909
Viteri FE (1998) A new concept in the control of iron deficiency: community-based preventive supplementation of at-risk groups by the weekly intake of iron supplements. Biomed Environ Sci 11(1):46–60
Voges MJ, Bai Y, Schulze-Lefert P, Sattely ES (2019) Plant-derived coumarins shape the composition of an Arabidopsis synthetic root microbiome. Proc Natl Acad Sci USA 116(25):12558–12565. https://doi.org/10.1073/pnas.1820691116
Wala M, Kołodziejek J, Wilk T (2022) Acidity and availability of aluminum, iron and manganese as factors affecting germination in European acidic dry and alkaline xerothermic grasslands. PeerJ 10:e13255. https://doi.org/10.7717/peerj.13255
Wairich A, de Oliveira BH, Arend EB, Duarte GL, Ponte LR, Sperotto RA, Ricachenevsky FK, Fett JP (2019) The combined strategy for iron uptake is not exclusive to domesticated rice (Oryza sativa). Sci Rep 9(1):1–17. https://doi.org/10.1038/s41598-019-52502-0
Wang N, Cui Y, Liu Y, Fan H, Du J, Huang Z, Yuan Y, Wu H, Ling HQ (2013) Requirement and functional redundancy of Ib subgroup bHLH proteins for iron deficiency responses and uptake in Arabidopsis thaliana. Mol Plant 6(2):503–513. https://doi.org/10.1093/mp/sss089
Wang B, Wei H, Chen Z, Li Y, Zhang WH (2022) Carbonate-induced chemical reductants are responsible for iron acquisition in strategy I wild herbaceous plants native to calcareous grasslands. Plant Cell Physiol 63(6):770–784. https://doi.org/10.1093/pcp/pcac038
Wegner LH, Zimmermann U (2004) Bicarbonate-induced alkalinization of the xylem sap in intact maize seedlings as measured in situ with a novel xylem pH probe. Plant Physiol 136(3):3469–3477
Xu W, Smith FJ Jr, Subaran R, Mitchell AP (2004) Multivesicular body-ESCRT components function in pH response regulation in Saccharomyces cerevisiae and Candida albicans. Mol Biol Cell 15(12):5528–5537. https://doi.org/10.1091/mbc.e04-08-0666
Xu J, Zhang Y, Zhang P, Trivedi P, Riera N, Wang Y, Liu X, Fan G, Tang J, Coletta-Filho HD, Cubero J (2018) The structure and function of the global citrus rhizosphere microbiome. Nat Commun 9(1):1–10. https://doi.org/10.1038/s41467-018-07343-2
Xiong H, Kakei Y, Kobayashi T, Guo X, Nakazono M, Takahashi H, Nakanishi H, Shen H, Zhang F, Nishizawa NK, Zuo Y (2013) Molecular evidence for phytosiderophore-induced improvement of iron nutrition of peanut intercropped with maize in calcareous soil. Plant Cell Environ 36(10):1888–1902. https://doi.org/10.1111/pce.12097
Yang TJ, Lin WD, Schmidt W (2010) Transcriptional profiling of the Arabidopsis iron deficiency response reveals conserved transition metal homeostasis networks. Plant Physiol 152(4):2130–2141. https://doi.org/10.1104/pp.109.152728
Yen MR, Suen DF, Hsu FM, Tsai YH, Fu H, Schmidt W, Chen PY (2017) Deubiquitinating enzyme OTU5 contributes to DNA methylation patterns and is critical for phosphate nutrition signals. Plant Physiol 175(4):1826–1838. https://doi.org/10.1104/pp.17.01188
Yuan YX, Zhang J, Wang DW, Ling HQ (2005) AtbHLH29 of Arabidopsis thaliana is a functional ortholog of tomato FER involved in controlling iron acquisition in strategy I plants. Cell Res 15(8):613–621. https://doi.org/10.1038/sj.cr.7290331
Yuan Y, Wu H, Wang N, Li J, Zhao W, Du J, Wang D, Ling HQ (2008) FIT interacts with AtbHLH38 and AtbHLH39 in regulating iron uptake gene expression for iron homeostasis in Arabidopsis. Cell Res 18(3):385–397. https://doi.org/10.1038/cr.2008.26
Zamioudis C, Korteland J, Van Pelt JA, van Hamersveld M, Dombrowski N, Bai Y, Hanson J, Van Verk MC, Ling HQ, Schulze-Lefert P, Pieterse CM (2015) Rhizobacterial volatiles and photosynthesis-related signals coordinate MYB 72 expression in Arabidopsis roots during onset of induced systemic resistance and iron-deficiency responses. Plant J 84(2):309–322. https://doi.org/10.1111/tpj.12995
Zhang H, Zhao X, Li J, Cai H, Deng XW, Li L (2014) MicroRNA408 is critical for the HY5-SPL7 gene network that mediates the coordinated response to light and copper. Plant Cell 26(12):4933–4953. https://doi.org/10.1105/tpc.114.127340
Ziegler J, Schmidt S, Strehmel N, Scheel D, Abel S (2017) Arabidopsis transporter ABCG37/PDR9 contributes primarily highly oxygenated coumarins to root exudation. Sci Rep 7(1):1–11. https://doi.org/10.1038/s41598-017-03250-6
Zohlen A, Tyler G (2004) Soluble inorganic tissue phosphorus and calcicole–calcifuge behaviour of plants. Ann Bot 94(3):427–432. https://doi.org/10.1093/aob/mch162
Zuo Y, Zhang F (2011) Soil and crop management strategies to prevent iron deficiency in crops. Plant Soil 339(1):83–95. https://doi.org/10.1007/s11104-010-0566-0
Acknowledgements
We acknowledge the use of the BioRender software (https://biorender.com) for generating the figures of this article.
Funding
This work was funded by an Academia Sinica Investigator Award (AS-IA-111-L03) to W.S.
Author information
Authors and Affiliations
Contributions
ICVB and WS performed the literature search and wrote the manuscript. WS initiated the review, ICVB drafted the figures.
Corresponding author
Ethics declarations
Competing Interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Responsible Editor: Ismail Cakmak.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vélez-Bermúdez, I.C., Schmidt, W. Plant strategies to mine iron from alkaline substrates. Plant Soil 483, 1–25 (2023). https://doi.org/10.1007/s11104-022-05746-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-022-05746-1