Abstract
Purpose
Iron (Fe) deficiency seriously affects crop growth and yield. This study investigated whether and how an endophytic isolate, strain Bacillus sp. WR12, alleviates Fe deficiency in wheat (Triticum aestivum L.).
Methods
Bacillus sp. WR12 and wheat seedlings were hydroponically co-cultured for 2 weeks under Fe-deficient conditions. To evaluate the Fe deficiency alleviating potential, wheat growth parameters were compared. To elucidate possible mechanisms, the genome of WR12 was sequenced, and production of siderophores was detected. In addition, transcriptomes of wheat root were compared, and accumulation of phenolic compounds was measured.
Results
Bacillus sp. WR12 significantly increased root length and dry weight, leaf chlorophyll content and Fe content in Fe-deficient wheat seedlings. Genome analysis of WR12 and CAS agar diffusion test demonstrated that WR12 had a dhbACEBF operon and produced siderophores. Significant upregulation of this operon and more siderophore synthesis in WR12 were observed when Fe was insufficient. Transcriptomic data of roots indicated change of expression of a large number of genes in Fe-deficient wheat after WR12 inoculation. Particularly, PAL, CYP73A, COMT, F5H, HCT and CAD genes involved in phenylpropane biosynthesis were significantly up-regulated. Moreover, elevated phenol accumulation was detected in the root of wheat after inoculation with WR12 under Fe deficiency.
Conclusions
Bacillus sp. WR12 alleviates Fe deficiency in wheat and this can be partially attributed to enhanced siderophore production and phenol accumulation.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Iron (Fe) is an essential trace element and has irreplaceable functions in the growth and development of plants. As a cofactor of many enzymes and proteins, it is involved in a lot of biological processes, including photosynthesis, electron transfer, chemical transitions, mitochondrial respiration, and regulation of protein stability (Connorton et al. 2017; Zhang et al. 2019). Although abundant in the Earth’s crust, most Fe exists in the form of ferric compounds showing low solubility in soils, especially in alkaline soils that account for 30% of global land (Imsande 1998; Li and Lan 2017). This leads to very low Fe bioavailability, insufficient to satisfy the needs of plant growth. As a consequence, Fe deficiency has become one of the most widespread abiotic stresses. For crops, Fe deficiency often results in leaf chlorosis, growth retardation, and reduction in yields and quality (Sebastian et al. 2017). Moreover, Fe deficiency in crops affects dietary Fe acquisition, leading to Fe deficiency related diseases in humans (Gozzelino and Arosio 2016). According to the World Health Organization (WHO), Fe deficiency is an important global nutrition problem and seriously impacts more than 2 billion people (http://www.who.int/nutrition/topics/ida/en/). Therefore, alleviation of Fe deficiency is of great significance to both crop growth and human health.
Plant-associated microbes play vital roles in plant growth and development, and many microbes have been found to improve nutrition absorption, including Fe, and to induce abiotic stress tolerance in plants (He et al. 2020; Lata et al. 2018; Ramakrishna et al. 2020). Among these microbes, Bacillus (B.) sp. is a group of gram-positive bacteria that is commonly termed plant growth promoting bacteria (PGPB). Some species of the genus Bacillus sp. are able to promote plant growth and enhance Fe uptake. For instance, inoculation with PGPB including Bacillus altitudinis increases plant biomass, grain yield and grain Fe content in Cicer arietinum and Cajanus cajan under field conditions (Gopalakrishnan et al. 2016). Application of B. subtilis GBO3 to Manihot esculenta augments growth, photosynthesis and Fe accumulation (Freitas et al. 2015). In addition, significant increases in root length and biomass and leaves photosynthesis were observed in B. subtilis STU6 inoculated tomato under Fe-limited conditions (Zhou et al. 2019). These findings provide a possible strategy for using Bacillus sp. as inoculants to ameliorate Fe deficiency stress in crops.
As the staple food for billions of people, wheat (T. aestivum L.) supplies a large part of the daily energy and nutrients intake (Dinu et al. 2018). However, the majority of wheat lines are short of Fe. In our previous study, we obtained several endophytes belonging to the genus Bacillus sp. from the root of wheat (Sun et al. 2017; Yue et al. 2019). Considering the widespread Fe deficiency in soils and the potential of Bacillus sp. to improve Fe uptake, we wondered whether Bacillus sp. WR12, an Fe-tolerant isolate, could also ameliorate Fe deficiency in wheat. Therefore, this study was designed to: (1) assess the potential of WR12 to alleviate Fe deficiency stress in wheat seedlings using a hydroponic model; (2) evaluate the growth, siderophore production and gene expression in the siderophore biosynthesis pathway of WR12 under Fe deficiency; (3) discover the underlying mechanisms by the analyses of comparative transcriptomics of wheat root and the draft genome of WR12.
Materials and methods
Plant materials, bacteria strains and growth conditions
Seeds of wheat (cv. Zhoumai 28) were provided by Dr. Hongzhan Liu of Zhoukou Normal University. The wheat seeds were surface-sterilized using 10% sodium hypochlorite for 5 min and washed three times with sterile water (Djanaguiraman et al. 2019). Then these seeds were placed in sterile 9-cm diameter Petri dishes containing sterile water, and kept in the dark at 25 °C. After 2 days of germination, these seeds were transferred to half-strength Hoagland’s medium and grown at 25 ± 0.5 °C and 55 ± 4% relative humidity following a photoperiod of 12/12 h (light/dark), as described elsewhere (Yue et al. 2021). During plant growth, the culture solutions were renewed every other day. Bacillius sp. WR12 was cultured for 24 h at 30 °C, at 160 rpm in Luria-Bertani (LB) broth.
Co-culture of WR12 and wheat
Well-germinated wheat seeds were randomly assigned to twelve plastic basins (28 × 22 × 10 cm) each containing 1 L of half-strength Hoagland’s medium in the presence or absence of 100 μM of 2,2′-dipyridyl and 107 cfu/mL of WR12 vegetative cells. 2,2′-dipyridyl is an Fe chelator often used to create an Fe deficient condition in medium (Gaballa et al. 2008; Segond et al. 2014). The inoculation concentration of WR12 was selected based on previous researches and common sense in practice (Larcher et al. 2003; Ledea-Rodríguez et al. 2020; Sun et al. 2021). These basins were divided into four groups: group 1 without 2,2′-dipyridyl and WR12 inoculation (Fe+); group 2 without 2,2′-dipyridyl and with WR12 inoculation (Fe+ + WR12); group 3 with 2,2′-dipyridyl and without WR12 inoculation (Fe-); group 4 with 2,2′-dipyridyl and with WR12 inoculation (Fe- + WR12). Each group contained 3 replications and each replication included 30 seedlings. After two weeks, all wheat seedlings were harvested and the lengths of roots and shoots were measured. Subsequently, 20 seedlings were used for the determination of dry weight and Fe content in roots and shoots after drying for 72 h at 75 °C in an oven. 20 fresh seedlings were used to detect the content of chlorophyll and total phenolics. The roots of the remaining seedlings were frozen in liquid nitrogen and stored at −80 °C for RNA sequencing.
Detection of chlorophyll content
The content of chlorophyll in fresh leaves was detected according to a previously described method (Zhou et al. 2016). Briefly, 0.5 g of fresh leaves was homogenized with 80% acetone in darkness and centrifuged at 8000×g for 5 min. The supernatant was collected and measured at 663 nm and 645 nm using a SpectraMax i3x microplate reader (Molecular Devices, Sunnyvale, USA). The concentration of total chlorophyll (CT) was calculated based on the formulas (20.21 × A645 + 8.02 × A663) described by Arnon (1949). The chlorophyll content is presented as mg/g·fresh weight (FW).
Measurement of Fe content in wheat tissues
The digestion of roots and shoots was carried out according to a previously described method (Cuddy et al. 2013). Briefly, dried plant materials (0.1 g) were digested with 10 mL of distilled 70% nitric acid for 8 h at 120 °C. Then, 2 mL of 30% hydrogen peroxide was added to the above solutions and heated at 120 °C for 30 min. Once cool, the digested solutions were diluted to 15 mL with deionized water and filtered with 0.45 μm of micromembrane. Finally, the Fe content in the solutions was determined using a flame atomic absorption spectrophotometer (A3AFG, Persee, Beijing, China). Furthermore, Fe standard solution (1000 μg/mL in 1.0 mol/L HNO3) purchased from Macklin (Macklin, Shanghai, China) was used for calculating the standard curve. For quality control, the recovery rates of Fe in root and shoot were determined via added 60 μL of Fe standard solution during digestion. The detection limit was 5 μg/L. The recovery rates of Fe in root and shoot were 97.1% and 108.7%, respectively. Fe content in tissues is presented as μg/g·dry weight (dw).
Determination of total phenol content
The content of total phenols in roots was determined according to previously described methods with minor modifications (Azam Ansari et al. 2019; Rahimi et al. 2020). Briefly, 0.3 g of fresh root was ground in liquid nitrogen, mixed with 2 mL 95% methanol, and stood for 24 h in the dark at 4 °C. After centrifugation of 8000×g for 10 min, 0.1 mL of extract was mixed with 0.1 mL Folin–Ciocalteu reagent (Sigma-Aldrich, USA) prepared from Na2MoO4, Na2WO4, H3PO4, HCl and Li2SO4. After standing for 5 min, 0.3 mL of 10% Na2CO3 solution was added, and the final mixture was kept at room temperature for 2 h. Gallic acid was used as standard and the absorbance was recorded at 765 nm with a microplate reader in triplicate. Total phenolic content in roots is presented as mg gallic acid / g·FW.
Root RNA isolation and sequencing
The isolation and quality assurance of total RNA from root samples were performed according to our previously described method (Yue et al. 2021). Obtained RNA was then used to construct cDNA libraries using the TruSeq™ RNA sample prep kit (Illumina, San Diego, USA) and RNA sequencing was carried out in a Novaseq 6000 system (Illumina, USA), by Majorbio Biotech Co., Ltd., Shanghai, China.
Using SeqPrep and Sickle software, raw data were filtered for adapter reads, low-quality reads, higher N-ratio (> 10%) reads and < 50 bp reads. The resulting clean reads were then aligned with the wheat reference genome (version IWGSC) using TopHat2 (http://ccb.jhu.edu/software/tophat/index.shtml) and assembled using StringTie (http://ccb.jhu.edu/software/stringtie/). These raw data have been submitted to NCBI Sequence Read Archive (SRA) with the accession numbers SRR13346686 - SRR13346697.
Differentially expressed genes (DEGs) and enrichment analysis
Gene expression levels were analyzed with the transcripts per million reads (TPM) method using RSEM software (http://deweylab.github.io/RSEM/). DESeq2 software was used to assess DEGs between the control and treatment groups (Love et al. 2014). The threshold of DEGs was a false discovery rate-adjusted p value <0.05 and fold change >2.0. Enrichment analyses of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways for DEGs were conducted using Goatools and KOBAS software, respectively (Tang et al. 2015; Xie et al. 2011). A venn diagram was generated using Venny 2.1 software (https://bioinfogp.cnb.csic.es/tools/venny/index.html). Principal component analysis (PCA) analysis was performed to evaluate the relationship between biological repetitions in all groups using XLSTAT software (Bhatt and Maheshwari 2019).
Real-time quantitative PCR (qPCR) validation
Ten DEGs (5 up-regulated and 5 down-regulated) were selected for the validation of RNA-seq data by qPCR. These gene primers were designed using the online tool Primer3Plus (Supplementary Table S1). In brief, 500 ng of total RNA from RNA-seq was reverse transcribed into cDNA by HiScript® II Q RT SuperMix kit (Vazyme Biotech, Nanjing, China). The qPCR reaction was carried out in a CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad, Hercules, USA) using a 2× ChamQ™ SYBR qPCR Master Mix kit (Vazyme, China) according to the manufacturer’s protocol. The qPCR reaction protocol was as follows: 95 °C for 30 s followed by 40 cycles of 10 s at 95 °C and 30 s at 60 °C. A melting curve analysis with a temperature gradient of 0.5 °C/s from 65 °C to 95 °C was performed to confirm a single amplification product. For normalization, gapdh (gene id: TraesCS6B02G185400) was used as the reference gene because this gene was stably expressed under our experimental conditions. The relative mRNA expression was evaluated using the 2−ΔΔCt method (Livak and Schmittgen 2001).
Draft genome sequencing, annotation, and bioinformatic analyses
Strain Bacillius sp. WR12 was cultured in 200 mL of LB broth for 24 h. By centrifugation of 12,000 rpm for 10 min, bacteria cells were harvested and washed three times with sterile water. Genomic DNA was then extracted using a Bacterial DNA Extraction Kit (Tiangen Biotech, Beijing, China) in accordance with the manufacturer’s instructions. Quality and quantification analysis of the extracted DNA were performed using 1% (w/v) agarose gel electrophoresis and Nanodrop 2000c Spectrophotometer (Thermo Fisher Scientific, Waltham, USA), respectively. High quality genomic DNA was used to construct Illumina libraries and sequenced on an Illumina Hiseq Xten system (Illumina, Inc., San Diego, USA) by Beijing Abace Biology Co., Ltd.
After sequencing, quality-control of the resulting raw data was firstly conducted by PRINSEQ-lite tool to filter out adapters and low-quality reads. Using SOAPdenovo version 2.04, the remaining reads were spliced into contigs which were further assembled into scaffolds. Then, the gaps in these scaffolds were filled utilizing GapCloser v1.12. Glimmer v3.02 and Basic Local Alignment Search Tool (BLAST) were used to predict and annotate protein-coding sequences (CDSs) in the draft genome. All predicted proteins were analyzed by the Clusters of Orthologous Groups (COG, http://string-db.org/, v9.05), KEGG (http://www.genome.jp/kegg/), and GO (http://www.geneontology.org/) databases. Additionally, ribosome RNA (rRNA) and transfer RNA (tRNA) were predicted using Barrnap 0.4.2 and tRNAscan-SE v1.3.1 software, respectively. The draft genomic data of WR12 by shotgun sequencing have been submitted to GenBank with accession number JABAJR000000000 and were further annotated automatically by the NCBI prokaryotic genome annotation pipeline (PGAP).
Genomic analysis of siderophore synthesis
To find the genes related to siderophore biosynthesis, a primary search of proteins in the WR12 genomic database was conducted with the keyword “dhb” and specific “protein names” encoded by gene clusters of dhb. Subsequently, all these related genes were further identified by BLAST.
Siderophore secretion assay
Strain Bacillius sp. WR12 was inoculated into 5 mL of LB broth for 24 h, following centrifugation at 12000 rpm for 10 min. Then, the supernatant was collected for qualitative analysis of siderophores according to the method described previously with minor modification (Kalidasan et al. 2018; Schwyn and Neilands 1987). Briefly, 70 μL of cell-free-supernatant was added to a 5-mm round hole punched on the Chrome Azurol Sulphonate (CAS) agar diffusion plates and incubated at room temperature for 72 h, with ferrioxamine E (FOE) as the positive control and LB broth as the negative control. Observed orange halo zones around these holes mean the production of siderophores.
The growth, siderophore production and gene expression related to siderophores biosynthesis of WR12 under Fe deficiency
Strain Bacillius sp. WR12 was inoculated into 5 mL of LB broth with or without the chelator 2,2′-dipyridyl (100 μM, Sigma-Aldrich, USA) for 24 h and centrifuged at 12000 rpm for 5 min. The bacteria cells were collected, rinsed with sterile water, frozen in liquid nitrogen, and stored at −80 °C for RNA isolation. The supernatant was collected and used to determine siderophores.
Bacteria total RNA of WR12 was extracted using a bacteria RNA extraction kit (Vazyme Biotech, Nanjing, China). After the detection of RNA quality and concentration, total RNA was reverse transcribed and the qPCR reaction was carried out according to the method mentioned above. The relative mRNA expressions were normalized with gapdh, a stably expressed housekeeping gene in our experimental conditions, and calculated using the 2−ΔΔCt method. These gene primers are shown in Supplementary Table S2.
The measurement of siderophore content was performed according to the Arnow assay (Arnow 1937). In brief, 0.5 mL of supernatant was mixed sequentially with 50 μL of 5 M HCl, 0.5 mL of nitrite-molybdate reagent (10% NaNO2 and Na2MoO4) and 50 μL of 10 M NaOH. The absorbance was measured at 515 nm using a microplate reader (Molecular Devices, USA). The standard curve was established using different concentrations of 3,4- dihydroxybenzoic acid (3,4-DHBA). Siderophore levels are presented as μM DHBA per OD600.
Data and statistical analysis
Statistical analysis was carried out in IBM SPSS Statistics 19.0 (IBM, Armonk, USA). Data are presented as mean ± standard deviation (SD). Significant differences between control and treatment groups were analyzed by t-test and Tukey’s one-way analysis of variance (ANOVA). Values of p < 0.05 were considered as statistically significant.
Results
WR12 alleviates Fe deficiency in wheat
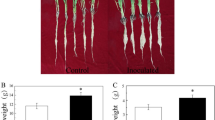
After two weeks of co-culture, WR12- inoculated wheat seedlings had similar growth and biomass to the control under the Fe sufficient condition (Fig. 1a, b). Under Fe deficiency, the growth of wheat seedlings was inhibited and leaf chlorophyll content was decreased, while WR12 alleviated these phenomena. Specifically, inoculation with WR12 significantly increased the length and dry weight in the root and the chlorophyll content in leaves compared with the uninoculated group (Fig. 1a–c). Moreover, Fe deficiency led to the decrease of Fe content in the roots and shoots, while significant increases were observed in WR12- inoculated wheat compared with WR12-free wheat (Fig. 1d).
Bacillus sp. WR12 alleviated Fe deficiency in wheat seedlings. a The length of roots and shoots; b The dry weight of roots and shoots; c Leaf chlorophyll content; d Fe contents in roots and shoots. Wheat seedlings were planted in Hoagland’s medium supplemented with or without WR12 in the presence or absence of 2,2′-dipyridyl for two weeks. Data are expressed as the means ± SD (n = 12). Different letters represented significant difference at p < 0.05
WR12 has an operon for siderophore biosynthesis and produces siderophores
By draft genome sequencing, 450 Mb of clean data were obtained. These data were assembled into 12 scaffolds. In theGO database, a total of 2942 genes were annotated, of which 1732 (58.85%) fell into the metabolic processes in the biological process category (Fig. S1). In the KEGG database, 4258 genes were annotated to 206 pathways and more than half of the pathways were related to metabolism (Fig. S2). Quality control, assembly and annotation information of this genome is shown in Supplementary Table S3. Among these metabolism pathways, a “Biosynthesis of siderophore group nonribosomal peptides” pathway (map 01053) was observed, and these genes were identified as DhbA, DhbC, DhbE, DhbB and DhbF in this pathway. Further genome mining revealed ferric uptake regulator (Fur) gene in scaffold 1 and dhbACEBF gene cluster in scaffold 2. The dhbACEBF operon was located at an area of 102,009–113,786 bp, and these five genes lengths were 786 bp, 1197 bp, 1620 bp, 939 bp and 7137 bp, respectively (Fig. 2a). Additionally, CAS agar diffusion test showed orange halo zones around these holes containing WR12-free supernatant, confirming the production of siderophores (Fig. 2b).
The dhbACEBF gene cluster for siderophore synthesis and siderophore production in Bacillus sp. WR12. a Scheme of dhbACEBF operon and siderophore synthesis pathway. After 24 h of inoculation in LB broth, WR12 was collected and its draft genome was sequenced, assembled and annotated to different databases. b Assay of siderophore production with the CAS agar. The cell free supernatant of WR12 was added to CAS agar plate and orange halo zones were observed after 72 h of incubation
WR12 secretes more siderophores under Fe deficiency
Compared with the Fe sufficient condition, WR12 grew well (84% of control) and secreted 31.5 times more siderophores under Fe deficiency (Fig. 3a, b). At transcription level, Fe deficiency induced the upregulation of dhbA, dhbB, dhbC, dhbE and dhbF genes and the downregulation of fur gene in WR12 (Fig. 3c).
Bacillus sp. WR12 secreted more siderophores under Fe deficiency. a The growth of WR12. Growth of WR12 was measured by reading OD600 after 24 h of inoculation in LB broth supplemented with or without 2,2′-dipyridyl. b Quantitative analysis of siderophores. The siderophore contents in medium from Fe sufficiency and Fe deficiency groups were determined using a microplate reader. Data are expressed as μM DHBA per OD600. c gene expression of dhbACEBF. Data are expressed as the means ± SD (n = 3). Different letters indicate significant difference at p < 0.05
WR12 induces many DEGs in wheat under Fe deficiency
RNA-seq of roots in 12 samples generated 808.3 million clean reads with Q30 > 95.9% (Table S4, 99.14% of the raw reads). These clean reads of each sample were aligned with the wheat reference genome and the mapped rate ranged from 81.52% to 90.77% (Table S5). Then, these mapped reads were assembled into 103,321 genes and 91.82% of them were successfully annotated in at least one of six databases (Table S6). PCA analysis showed that the four experimental groups were separated from each other and the three biological replicates of each experimental group gathered together, indicating effective treatment and acceptable biological replicability (Fig. 4a).
RNA-seq analyses of wheat roots. a PCA analysis between biological repetitions in all treatment groups. b Venn diagram displaying up- and down-regulated differentially expressed genes (DEGs) in Fe + vs Fe-) and Fe- vs Fe- + WR12. c and d GO and KEGG enrichment analyses of DEGs in the Fe- vs Fe- + WR12. BP, biological processes; CC, cellular components; MF, molecular functions
As shown in Fig. 4b, of the 15,881 differentially expressed genes (DEGs), 7774 were upregulated and 8107 were downregulated in Fe-deficient wheat seedlings (Fe-) in comparison with Fe-sufficient plants (Fe+). Moreover, a total of 4313 genes were differentially expressed in Fe-deficient wheat with WR12 inoculation (Fe- + WR12) compared with Fe-deficient plants (Fe-). Among these DEGs, 1898 were upregulated and 2415 were downregulated (Fig. 4b). By the GO enrichment analysis, many DEGs in Fe- vs Fe- + WR12 were related to transmembrane transport and secondary metabolic progress in the subcategory of biological processes (BP) (Fig. 4c). Moreover, KEGG enrichment analysis revealed that DEGs in Fe- vs Fe- + WR12 were significantly enriched to 9 pathways including “Phenylpropanoid biosynthesis” (map00940) (Fig. 4d). In addition, many genes associated with mugineic acid family phytosiderophores (MAs), such as nicotianamine synthase (NAS), nicotianamine aminotransferase (NAAT) and deoxymugineic acid synthase (DMAS), were consistently upregulated under Fe deficiency, whereas these genes were not altered in WR12- inoculated plants (Table S7).
WR12 activates phenylpropanoid biosynthesis and promotes phenol accumulation in wheat roots under Fe deficiency
In the pathway of phenylpropanoid biosynthesis, 72 of 81 DEGs were significantly upregulated, indicating WR12 activates the pathway. To give more detail, the expression of phenylalanine ammonia-lyase (PAL, EC 4.3.1.24), trans-cinnamate 4-monooxygenase (CYP73A, EC 1.14.14.91), caffeic acid 3-O-methyltransferase (COMT, EC 2.1.1.68), ferulate-5-hydroxylase (F5H, EC 1.14.-.-), shikimate O-hydroxycinnamoyltransferase (HCT, EC 2.3.1.133), and cinnamyl-alcohol dehydrogenase (CAD, EC 1.1.1.195) were up-regulated. Other genes, including cinnamoyl-CoA reductase (CCR, EC 1.2.1.44), 4-coumarate-CoA ligase (4CL, EC 6.2.1.12;), beta-glucosidase (EC 3.2.1.21), and peroxidase (POD, EC 1.11.1.7), were either up- or down-regulated (Fig. 5a). Further qPCR analysis of 10 selected DEGs displayed that the transcription results of the tested genes were similar to that of the RNA-seq data (Fig. 5b). In addition, total phenol content in the root was significantly decreased in Fe-deficient wheat but increased significantly in the WR12-inoculated wheat under Fe deficiency (Fig. 5c).
Bacillus sp. WR12 activated phenylpropanoid biosynthesis and promoted phenolic accumulation in wheat roots under Fe deficiency. a DEGs in the Fe- vs Fe- + WR12 enriched in phenylpropanoid biosynthesis pathway. Red boxes: significantly up-regulated genes. Blue triangles, pentagrams and circles: significantly up- and down-regulated genes. b qPCR validation of RNA-seq data in the Fe- vs Fe- + WR12. Data are expressed as the means ± SD (n = 3).c Total phenols of roots. Data are expressed as the means ± SD (n = 6). Different letters represent significant difference at p < 0.05
Discussion
The total amount of Fe in soil is more than is required by plants. However, the aerobic and alkaline conditions of soil cause soluble Fe2+ to be oxidized to insoluble Fe3+ (e.g., ferric hydroxides and ferric oxides) which is not readily available to plants (Kobayashi et al. 2019). It has been reported that the contents of free Fe3+ and Fe2+ are less than 10−15 M in well-aerated soils at physiological pH, far lower than the amount required for plant growth (10−9- 10−4 M) (Kim and Guerinot 2007). Hence, as a non-negligible abiotic stress, soil Fe deficiency adversely affects plant growth and development (Sebastian et al. 2017). Consistent with the results in previous studies, the dry biomass of the root, length of root and shoot, and content of leaf chlorophyll were significantly less in the Fe-deficient wheat seedlings than in the Fe-sufficient plants in this study. In contrast, the seedlings inoculated with WR12 showed better growth under Fe deficiency. Similar results were also observed in B. subtilis STU6-inoculated tomato and Arbuscular mycorrhizal fungi supplemented sunflower under Fe-limited conditions (Kabir et al. 2020; Zhou et al. 2019). Therefore, we suggest that Bacillus sp. WR12 has the potential to alleviate Fe deficiency stress in wheat.
Fe plays an important role in chlorophyll biosynthesis, and Fe shortage often results in chlorophyll reduction and leaf chlorosis (Roncel et al. 2016). In this study, chlorophyll synthesis was obviously inhibited in Fe-deficient wheat leaves. Conversely, WR12-inoculated plants showed higher chlorophyll content (similar to Fe sufficiency) under Fe deficiency, implying that WR12 may improve Fe uptake in Fe-deficient wheat. Furthermore, we determined Fe contents in wheat seedlings in different treatment groups and found that Fe contents were increased significantly in wheat inoculated with WR12 compared to that without WR12 under Fe deficiency. Similar results were reported in other studies. For example, Pisolithus tinctorius significantly increased Fe content and chlorophyll concentration in slash pine (Pinus elliottii) under Fe-deficient conditions (Leyval and Reid 1991), and Glomus versiforme inoculation resulted in higher chlorophyll and Fe concentrations in Fe-deficient trifoliate orange (Poncirus trifoliate) (Wang et al. 2007). These findings demonstrate that WR12 enhances Fe absorption in plants under Fe deficiency.
Gramineous plants including wheat (Strategy II) acquire Fe mainly through the synthesis and secretion of mugineic acids (MAs) (Beasley et al. 2017; Connorton et al. 2017). In the root, MAs are synthesized from S-adenosylmethionine (SAM) via a series of enzymes, such as nicotianamine synthase (NAS), nicotianamine aminotransferase (NAAT) and deoxymugineic acid synthase (DMAS) (Ma et al. 1995; Higuchi et al. 1999). Under Fe limitation, these MA-associated genes are up-regulated in plants (Kobayashi and Nishizawa 2012). Indeed, transcriptome data revealed that the transcripts of genes encoded NAS, NAAT and DMAS in Fe-deficient wheat were upregulated significantly in our study. However, these MA-associated gene expressions were not changed in Fe-deficient wheat with WR12 inoculation compared with the Fe-deficient wheat without WR12, indicating that the Fe uptake induced by WR12 inoculation was not associated with MA biosynthesis.
To find out whether WR12 produces a certain substance that promotes Fe absorption in plants, we sequenced the draft genome of WR12. By genome analyses, a gene cluster (dhbACEBF) related to siderophore synthesis was discovered, indicating that WR12 may produce siderophores, and further biochemical tests confirmed this. Siderophores are high affinity ferric iron (Fe3+) chelators synthesized and secreted by many microbes under Fe-deficient conditions (Khan et al. 2018). Many members of the genus Bacillus sp. have been confirmed to synthesize siderophores, including Bacillibactin (BB), such as B. subtilis, Bacillus licheniformis, Bacillus stratosphericus, and Bacillus megaterium (Liu et al. 2021; Bhatt and Maheshwari 2020). Bacillibactin (BB) is the major catechol-based siderophore composed of dihydroxybenzoate (DHBA), threonine and glycine (Hotta et al. 2010). Its production starts with chorismite and depends on dhbACEBF operon under ferric uptake regulator (Fur) regulation. Fur is an Fe-dependent repressor that senses intracellular Fe content and regulates the expression of genes involved in Fe acquisition (Pi and Helmann 2017). Under Fe-depleted conditions, the Fur regulon is derepressed, which results in the transcription of the dhbACEBF gene cluster and siderophore production (Chandrangsu et al. 2017). In this study, all genes in dhbACEBF operon were uniformly upregulated and siderophore production was also increased remarkably in WR12 cultured under Fe deficiency. In fact, it has been reported that siderophores promote plant growth and increase yield by enhancing the solubilization of Fe (Saha et al. 2016; Verma et al. 2021). For instance, application of Fe-fortified bacterial siderophores significantly enhanced the growth, grain yield and grain Fe content in wheat (Sharma et al. 2019). The presence of siderophores significantly increased the Fe uptake and promoted plant growth in Pteris vittate (Liu et al. 2017). Supplementation with pyoverdine, a siderophore synthesized by Pseudomonas fluorescens, resulted in the accumulation of Fe in leaves and roots of Solanum lycopersicum (Nagata et al. 2013). Hence, we suggest that the WR12-induced Fe uptake in Fe-deficient plants can be partially attributed to the production of siderophores.
In addition, KEGG analysis revealed that most of the DEGs enriched in the pathway of “phenylpropanoid biosynthesis” were up-regulated in WR12-inoculated wheat under Fe deficiency compared with under Fe-deficiency alone. The phenylpropanoid pathway starts with phenylalanine and produces thousands of secondary metabolites including phenolic compounds through a series of enzymatic reactions (Vogt 2010). Therefore, the upregulation of genes encoding PAL, CYP73A, COMT, F5H, HCT, and CAD in the present study indicated that WR12 activated this pathway under Fe deficiency. We then measured the total phenol content and observed the accumulation of phenols in Fe-deficient wheat with WR12 inoculation. Similarly, a significant increase in phenolic compounds has been observed in the Azospirillum brasilense inoculated cucumber and Paenibacillus polymyxa inoculated Arabidopsis under Fe deficiency (Pii et al. 2015; Zhou et al. 2016). Plant phenols are a group of aromatic compounds that play critical roles in response to biotic and abiotic stresses, including Fe deficiency (Jin et al. 2007a; Msilini et al. 2013). It has been reported that phenols facilitate Fe acquisition by increasing Fe solubility in soil and mediating the mobilization of root apoplastic Fe due to their reductive and chelating properties (Bashir et al. 2011; Jin et al. 2007b; Tato et al. 2013). Furthermore, phenols also influence the rhizosphere microbial community and recruit more siderophore-producing bacteria, which in turn indirectly promotes plant Fe acquisition (Jin et al. 2007b, 2010). Therefore, WR12 may exert the beneficial effect to wheat also by increasing phenol production in the root, thereby enhancing Fe absorption and alleviating Fe deficiency. The interaction between WR12 and wheat highlighted their mutualistic relationship, as indicated in this study under Fe deficiency.
In summary, the present study showed that Bacillus sp. WR12 alleviates the growth inhibition and weakened photosynthesis caused by Fe deficiency in wheat seedlings. This is because WR12 improved Fe acquisition in wheat. One potential mechanism was WR12 had an induced dhbACEBF operon and produced more siderophores under Fe deficiency. Another possibility is that WR12 enhanced Fe uptake partially by activating phenylpropanoid biosynthesis and promoting phenol secretion in the plants. Our findings support using siderophore-producing microorganisms as bio-fertilizer to improve wheat growth in response to Fe deficiency.
Abbreviations
- B :
-
Bacillus
- PGPB:
-
Plant growth promoting bacteria
- LB:
-
Luria-Bertani
- RNA-seq:
-
RNA sequencing
- DEGs:
-
Differentially expressed genes
- GO:
-
Gene ontology
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- CAS:
-
Chrome azurol sulphonate
- FOE:
-
ferrioxamine E
- DHBA:
-
Dihydroxybenzoic acid
References
Arnon D (1949) Copper enzymes in isolated chloroplasts, phenoloxidase in Beta vulgaris. Plant Physiol 24:1–15
Arnow LE (1937) Proposed chemical mechanisms for the production of skin erythema and pigmentation by radiant energy. Science 86:176
Azam Ansari M, Chung IM, Rajakumar G, A Alzohairy M, Almatroudi A, Gopiesh Khanna V, Thiruvengadam M (2019) Evaluation of polyphenolic compounds and pharmacological activities in hairy root cultures of Ligularia fischeri Turcz. f. spiciformis (Nakai). Molecules 24: 1586
Bashir K, Ishimaru Y, Shimo H, Kakei Y, Senoura T, Takahashi R, Satob Y, Uozumib N, Nakanishi H, Nishizawa NK (2011) Rice phenolics efflux transporter 2 (PEZ2) plays an important role in solubilizing apoplasmic iron. Soil Sci Plant Nutr 57:803–812
Beasley JT, Bonneau JP, Johnson AAT (2017) Characterisation of the nicotianamine aminotransferase and deoxymugineic acid synthase genes essential to strategy II iron uptake in bread wheat (Triticum aestivum L). PLoS One 12:1–18
Bhatt K, Maheshwari DK (2019) Decoding multifarious role of cow dung bacteria in mobilization of zinc fractions along with growth promotion of C. annuum L. Sci Rep 9:14232
Bhatt K, Maheshwari DK (2020) Bacillus megaterium strain CDK25, a novel plant growth promoting bacterium enhances proximate chemical and nutritional composition of Capsicum annuum L. Front Plant Sci 11:1147
Chandrangsu P, Rensing C, Helmann JD (2017) Metal homeostasis and resistance in bacteria. Nat Rev Microbiol 15:338–350
Connorton JM, Balk J, Rodríguez-Celma J (2017) Iron homeostasis in plants - a brief overview. Metallomics 9:813–823
Cuddy WS, Summerell BA, Gehringer MM, Neilan BA (2013) Nostoc, Microcoleus and Leptolyngbya inoculums are detrimental to the growth of wheat (Triticum aestivum L.) under salt stress. Plant Soil 370:317–332
Dinu M, Whittaker A, Pagliai G, Benedettelli S, Sofi F (2018) Ancient wheat species and human health: biochemical and clinical implications. J Nutr Biochem 52:1–9
Djanaguiraman M, Prasad PVV, Kumari J, Rengel Z (2019) Root length and root lipid composition contribute to drought tolerance of winter and spring wheat. Plant Soil 439:57–73
Freitas MA, Medeiros FH, Carvalho SP, Guilherme LR, Teixeira WD, Zhang H, Paré PW (2015) Augmenting iron accumulation in cassava by the beneficial soil bacterium Bacillus subtilis (GBO3). Front Plant Sci 6:596
Gaballa A, Antelmann H, Aguilar C, Khakh SK, Song KB, Smaldone GT, Helmann JD (2008) The Bacillus subtilis iron-sparing response is mediated by a Fur-regulated small RNA and three small, basic proteins. Proc Natl Acad Sci U S A 105:11927–11932
Gopalakrishnan S, Vadlamudi S, Samineni S, Kumar CVS (2016) Plant growth-promotion and biofortification of chickpea and pigeonpea through inoculation of biocontrol potential bacteria, isolated from organic soils. SpringerPlus 5:1882
Gozzelino R, Arosio P (2016) Iron homeostasis in health and disease. Int J Mol Sci 17:130
He L, Yue ZH, Chen C, Li CY, Li J, Sun ZK (2020) Enhancing iron uptake and alleviating iron toxicity in wheat by plant growth-promoting bacteria: theories and practices. Int J Agric Biol 23:190–196
Higuchi K, Suzuki K, Nakanishi H, Yamaguchi H, Nishizawa NK, Mori S (1999) Cloning of nicotianamine synthase genes, novel genes involved in the biosynthesis of phytosiderophores. Plant Physiol 119:471–480
Hotta K, Kim CY, Fox DT, Koppisch AT (2010) Siderophore-mediated iron acquisition in Bacillus anthracis and related strains. Microbiology 156:1918–1925
Imsande J (1998) Iron, sulfur, and chlorophyll deficiencies: a need for an integrative approach in plant physiology. Plant Physiol 103:139–144
Jin CW, He XX, Zheng SJ (2007a) The iron-deficiency induced phenolics accumulation may involve in regulation of Fe (III) chelate reductase in red clover. Plant Signal Behav 2:327–332
Jin CW, You GY, He YF, Tang C, Wu P, Zheng SJ (2007b) Iron deficiency-induced secretion of phenolics facilitates the reutilization of root apoplastic iron in red clover. Plant Physiol 144:278–285
Jin CW, Li GX, Yu XH, Zheng SJ (2010) Plant Fe status affects the composition of siderophore-secreting microbes in the rhizosphere. Ann Bot 105:835–841
Kabir AH, Debnath T, Das U, Prity SA, Haque A, Rahman MM, Parvez MS (2020) Arbuscular mycorrhizal fungi alleviate Fe-deficiency symptoms in sunflower by increasing iron uptake and its availability along with antioxidant defense. Plant Physiol Biochem 150:254–262
Kalidasan V, Azman A, Joseph N, Kumar S, Awang Hamat R, Neela VK (2018) Putative iron acquisition systems in Stenotrophomonas maltophilia. Molecules 23:2048
Khan A, Singh P, Srivastava A (2018) Synthesis, nature and utility of universal iron chelator - siderophore: a review. Microbiol Res 212-213:103–111
Kim SA, Guerinot ML (2007) Mining iron: iron uptake and transport in plants. FEBS Lett 581:2273–2280
Kobayashi T, Nishizawa NK (2012) Iron uptake, translocation, and regulation in higher plants. Annu Rev Plant Biol 63:131–152
Kobayashi T, Nozoye T, Nishizawa NK (2019) Iron transport and its regulation in plants. Free Radic Biol Med 133:11–20
Larcher M, Muller B, Mantelin S, Rapior S, Cleyet-Marel JC (2003) Early modifications of Brassica napus root system architecture induced by a plant growth-promoting Phyllobacterium strain. New Phytol 160:119–125
Lata R, Chowdhury S, Gond SK, White JF Jr (2018) Induction of abiotic stress tolerance in plants by endophytic microbes. Lett Appl Microbiol 66:268–276
Ledea-Rodríguez JL, Reyes-Pérez JJ, Castellanos T, Angulo C, Reynoso-Granados T, Alcaraz-Meléndez L (2020) Growth, development and quality of Moringa oleifera (Lamark) seedlings inoculated with plant growth promoting bacteria. Trop Subtropical Agroecosystems 23:74
Leyval C, Reid CPP (1991) Utilization of microbial siderophores by mycorrhizal and non-mycorrhizal pine roots. New Phytol 119:93–98
Li W, Lan P (2017) The understanding of the plant iron deficiency responses in strategy I plants and the role of ethylene in this process by omic approaches. Front Plant Sci 8:40
Liu X, Fu JW, Da Silva E, Shi XX, Cao Y, Rathinasabapathi B, Chen Y, Ma LQ (2017) Microbial siderophores and root exudates enhanced goethite dissolution and Fe/as uptake by as-hyperaccumulator Pteris vittata. Environ Pollut 223:230–237
Liu C, Ye Y, Liu J, Pu Y, Wu C (2021) Iron biofortification of crop food by symbiosis with beneficial microorganisms. J Plant Nutr:1–18
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550
Ma JF, Shinada T, Matsuda T, Nomoto K (1995) Biosynthesis of phytosiderophores, mugineic acids, associated with methionine cycling. J Biol Chem 270:16549–16554
Msilini N, Oueslati S, Amdouni T, Chebbi M, Ksouri R, Lachaâl M, Ouerghi Z (2013) Variability of phenolic content and antioxidant activity of two lettuce varieties under Fe deficiency. J Sci Food Agric 93:2016–2021
Nagata T, Oobo T, Aozasa O (2013) Efficacy of a bacterial siderophore, pyoverdine, to supply iron to Solanum lycopersicum plants. J Biosci Bioeng 115:686–690
Pi H, Helmann JD (2017) Sequential induction of Fur-regulated genes in response to iron limitation in Bacillus subtilis. Proc Natl Acad Sci U S A 114:12785–12790
Pii Y, Penn A, Terzano R, Crecchio C, Mimmo T, Cesco S (2015) Plant-microorganism-soil interactions influence the Fe availability in the rhizosphere of cucumber plants. Plant Physiol Biochem 87:45–52
Rahimi S, Talebi M, Baninasab B, Gholami M, Zarei M, Shariatmadari H (2020) The role of plant growth-promoting rhizobacteria (PGPR) in improving iron acquisition by altering physiological and molecular responses in quince seedlings. Plant Physiol Biochem 155:406–415
Ramakrishna W, Rathore P, Kumari R, Yadav R (2020) Brown gold of marginal soil: plant growth promoting bacteria to overcome plant abiotic stress for agriculture, biofuels and carbon sequestration. Sci Total Environ 711:135062
Roncel M, González-Rodríguez AA, Naranjo B, Bernal-Bayard P, Lindahl AM, Hervás M, Navarro JA, Ortega JM (2016) Iron deficiency induces a partial inhibition of the photosynthetic electron transport and a high sensitivity to light in the diatom Phaeodactylum tricornutum. Front Plant Sci 3:1050
Saha M, Sarkar S, Sarkar B, Sharma BK, Bhattacharjee S, Tribedi P (2016) Microbial siderophores and their potential applications: a review. Environ Sci Pollut Res Int 23:3984–3999
Schwyn B, Neilands JB (1987) Universal chemical assay for the detection and determination of siderophores. Anal Biochem 160:47–56
Sebastian A, Nangia A, Prasad MN (2017) Carbon-bound iron oxide nanoparticles prevent calcium-induced iron deficiency in Oryza sativa L. J Agric Food Chem 65:557–564
Segond D, Abi Khalil E, Buisson C, Daou N, Kallassy M, Lereclus D, Arosio P, Bou-Abdallah F, Nielsen Le Roux C (2014) Iron acquisition in Bacillus cereus: the roles of IlsA and bacillibactin in exogenous ferritin iron mobilization. PLoS Pathog 10:e1003935
Sharma S, Chandra S, Kumar A, Bindraban P, Saxena AK, Pande V, Pandey R (2019) Foliar application of iron fortified bacteriosiderophore improves growth and grain Fe concentration in wheat and soybean. Indian J Microbiol 59:344–350
Sun ZK, Liu K, Zhang J, Zhang Y, Xu KD, Yu DS, Wang J, Hu LZ, Chen L, Li CW (2017) IAA producing Bacillus altitudinis alleviates iron stress in Triticum aestivum L. seedling by both bioleaching of iron and up-regulation of genes encoding ferritins. Plant Soil 419:1–11
Sun ZK, Yue ZH, Liu HZ, Ma KS, Li CW (2021) Microbial-assisted wheat iron biofortification using endophytic Bacillus altitudinis WR10. Front Nutr 8:704030
Tang H, Klopfenstein D, Pedersen B, Flick P, Sato K, Ramirez F, Yunes J, Mungall C (2015) GOATOOLS: tools for gene ontology. Zenodo
Tato L, De Nisi P, Donnini S, Zocchi G (2013) Low iron availability and phenolic metabolism in a wild plant species (Parietaria judaica L.). Plant Physiol Biochem 72:145–153
Verma H, Jindal M, Rather SA (2021) Bacterial siderophores for enhanced plant growth. In: Malik AJ (ed) Handbook of research on microbial remediation and microbial biotechnology for sustainable soil. IGI Global, Hershey, pp 314–331
Vogt T (2010) Phenylpropanoid biosynthesis. Mol Plant 3:2–20
Wang MY, Xia RX, Hu LM, Dong T, Wu QS (2007) Arbuscular mycorrhizal fungi alleviate iron deficient chlorosis in Poncirus trifoliata L. Raf under calcium bicarbonate stress. J Hortic Sci Biotechnol 82:776–780
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li CY, Wei L (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39:W316–W322
Yue ZH, Shen YH, Chen YJ, Liang AW, Chu CW, Chen C, Sun ZK (2019) Microbiological insights into the stress-alleviating property of an endophytic Bacillus altitudinis WR10 in wheat under low-phosphorus and high-salinity stresses. Microorganisms 7:508
Yue ZH, Chen YJ, Chen C, Ma KS, Tian EL, Wang Y, Liu HZ, Sun ZK (2021) Endophytic Bacillus altitudinis WR10 alleviates cu toxicity in wheat by augmenting reactive oxygen species scavenging and phenylpropanoid biosynthesis. J Hazard Mater 405:124272
Zhang XZ, Zhang D, Sun W, Wang T (2019) The adaptive mechanism of plants to iron deficiency via iron uptake, transport, and homeostasis. Int J Mol Sci 20:2424
Zhou C, Guo J, Zhu L, Xiao X, Xie Y, Zhu J, Ma Z, Wang J (2016) Paenibacillus polymyxa BFKC01 enhances plant iron absorption via improved root systems and activated iron acquisition mechanisms. Plant Physiol Biochem 105:162–173
Zhou C, Zhu L, Guo J, Xiao X, Ma Z, Wang J (2019) Bacillus subtilis STU6 ameliorates iron deficiency in tomato by enhancement of polyamine-mediated iron remobilization. J Agric Food Chem 67:320–330
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 32071478), the Henan Provincial Department of Science and Technology (No. 192102310137) and the Education Department of Henan Province (No. 19A180034).
Author information
Authors and Affiliations
Contributions
Yue ZH and Sun ZK conceived the experiments. Yue ZH, Chen YJ, Hao YW, Wang CC, Zhang ZF and Sun ZK carried out the experimental work. Chen C, Liu HZ, Liu YC and Li LL analyzed the data. Yue ZH prepared the draft manuscript. Sun ZH provided all required material and finalized the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing financial interest.
Additional information
Responsible Editor: Ricardo Aroca
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 1070 kb)
Rights and permissions
About this article
Cite this article
Yue, Z., Chen, Y., Hao, Y. et al. Bacillus sp. WR12 alleviates iron deficiency in wheat via enhancing siderophore- and phenol-mediated iron acquisition in roots. Plant Soil 471, 247–260 (2022). https://doi.org/10.1007/s11104-021-05218-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-021-05218-y