Abstract
Aims
Agromining aims to improve the fertility of naturally metal-rich soils by extracting metals, such as nickel (Ni), using hyperaccumulator plants. Ultramafic soils are characterized by low fertility levels, limiting hyperaccumulator yields. Here, we characterize the potential benefits for phytoextraction efficiency of co-cropping a Ni-hyperaccumulator (Odontarrhena chalcidica) and a legume (Vicia sativa), following a two-year field experiment.
Methods
A two-year field experiment was set up in an ultramafic zone in North-West Spain. Three treatments were tested: co-cropping, fertilized control with ammonium nitrate and non-fertilized control.
Results
Over the 2 years, co-cropping increased O. chalcidica’s biomass by 24% and 403% compared to fertilized and non-fertilized controls, respectively. Moreover, co-cropping had higher Ni-yields for both years, while fertilization had a negative effect on soil parameters. A non-metric multidimensional scaling analysis of the operational taxonomic units showed that the soil bacterial diversity changed over time. Soil exchangeable Ni and organic carbon influenced the phyla’s relative abundance. Metabolic genes were dominant and their relative abundances increased over time with co-cropping.
Conclusion
Pluriannual co-cropping of a hyperaccumulator with a legume improved both hyperaccumulator and Ni yields. In contrast, mineral fertilization was shown to be detrimental to some soil microbial parameters. Thus, ameliorating agromining by replacing mineral fertilizers would combine an eco-efficient strategy with sustainable metal recovery.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Agromining aims to set up a new type of cropping systems employed either in degraded or naturally metal-rich soils such as ultramafic soils (Morel 2013). The main goals are to extract metals from the soil through the implementation of innovative cropping systems involving hyperaccumulator plants. It has been well-received by the public and its major advantage remains its low cost compared to conventional methods of soil decontamination or mining (Chaney et al. 2018). Yet agromining can be limited by low plant biomass productivity and by limited availability of soil metals (van der Ent et al. 2015; Benizri and Kidd 2018), and by the inherent nutrient deficiencies of ultramafic soils. In the case of nickel (Ni), the feasibility of agromining from ultramafic soils has been clearly demonstrated (Li et al. 2003; Bani et al. 2015). However, cropping systems need further research in order to optimize Ni-agromining ie. to improve plant biomass, Ni-yields and ultramafic soils’ quality.
In order to improve the development of hyperaccumulator plants and finally optimize the efficiency of metal extraction, various strategies have been developed. Indeed, several studies have suggested that the accumulation of metals by hyperaccumulator plants is influenced by their rhizospheric microflora that influence the biogeochemical cycling of soil metals (Mengoni et al. 2001; Abou-Shanab et al. 2003; Durand et al. 2016; Benizri and Kidd 2018). Numerous works have shown the positive effect of mineral fertilization on the biomass production of Ni-hyperaccumulator plants such as Odontarrhena spp. (formerly genus Alyssum section Odontarrhena, Španiel et al. 2015). The addition of organic amendments such as manure or grape and apple pomace is known to improve soil quality and structure, as well as the nutrients’ bioavailability and can stimulate the soil’s biological activity (Bernal et al. 2007; Martínez-Fernández et al. 2014; Álvarez-López et al. 2016).
Some authors have tested co-cropping using different hyperaccumulators(Lucisine et al. 2014; Rue et al. 2015). Lucisine et al. (2014) found that co-cropping different hyperaccumulator plants promoted the bioavailability of metals in the soil and modified the genetic and phenotypic structures of the rhizosphere bacterial communities.
Conversely, only a few studies have focused on the combination of hyperaccumulator plants with non-metal hyperaccumulator ones and have shown an improvement in the hyperaccumulator growth and an increase in heavy metal phytoavailability, thereby increasing remediation efficiency (Gove et al. 2002; Wu et al. 2007; Jiang et al. 2010; Wei et al. 2011; Gao et al. 2010, 2012). Among companion plants, legumes have frequently been used (Pan et al. 2008; Liu et al. 2011; Jiang et al. 2009, 2015; Saad et al. 2016, 2018a; Zu et al. 2017. Moreover, co-cropping legumes with hyperaccumulators improved soil quality, increased soil porosity, reduced its apparent density, increased aggregate stability and decreased the resistance to deep root penetration (Saad et al. 2018b). As for conventional agriculture, legumes are able to fix nitrogen from the air, thanks to the presence of symbiotic N2-fixing bacteria (De Antoni et al. 2015). Furthermore, these new agromining cropping systems can lower the risk of nitrogen leaching and any consequent underground water pollution.
Based on a field experiment in an ultramafic outcrop in NW Spain, we previously showed, only after 1 year, that the introduction of a legume into a Ni agromining system improved both plant biomass and Ni yields (Saad et al. 2018c). The objective of this work was to evaluate the performance of Odontarrhena chalcidica when co-cropped with a legume, Vicia sativa, on an ultramafic soil, within an agromining system and particularly to observe whether two consecutive years of this cropping system could further increase, along time, the hyperaccumulator biomass and the Ni yields. We hypothesized that legumes could be of a particular interest for agromining systems where nitrogen availability is often limited, especially since ultramafic soils show macronutrient deficiencies. In this study, biomass productivity and Ni yields were assessed through time, as well as the evolution of soil physicochemical, biological characteristics. In addition, this study brought new findings on the temporal variation of the genetic and functional diversity of bacterial communities in the rhizosphere of the hyperaccumulator.
Materials and methods
Experimental site
A field experiment was carried out for 2 years near the village of Eidián, Pontevedra (Galicia, North-West Spain; N 42°49′55,08” W 8°00′14,60″). This is an abandoned agricultural area colonized by vegetation such as Erica scoparia L., which is typical of the ultramafic soils in the region. In the past, this area was one of important agricultural activity, but which has since been abandoned and colonized by a typical vegetation of the serpentine soil in the region (Erica scoparia and Ulex europaeus). In the spring of 2015, the existent vegetation was removed and the soil was then ploughed before sowing plants. The soil was ploughed again, after the first harvest, in the spring of 2016.
Soil characteristics and experimental design
The soil physicochemical properties were determined by the Soil Analysis Laboratory of INRA (Arras, France). The soil was a Leptic Phaeozem (Magnesic) (IUSS Working Group WRB 2014). The soil contained 17.5, 30.6 and 51.9%, clay, silt and sand, respectively, had a C/N ratio of 13.9, an Mg/Ca ratio of 2.79 and an available phosphorus content (Olsen P) of 23 mg kg−1. Soil pH was 5.76 and the total Ni content was 861 mg Ni kg−1. For each year and before sowing plants, all the plots were amended with gypsum at a rate of 4.5 tons per hectare in order to improve the Ca/Mg ratio. Soil P and K contents were also improved by the amendments of 122.5 kg P ha−1 and 156.2 kg K ha−1 (0–52-34 NPK). Fertilizers were mixed into the first 10 cm of the topsoil by a tractor. The legume seeds (V. sativa var. Prontivesa) were provided by Semillas Batlle (http://www.semillasbatlle.es, Spain) and the seeds of O. chalcidica came from an Albanian population and were collected near Pogradec (39°47′17,5”N, 21°25′19,1″E, Albania) in August 2014. Legume seeds were sown at the rate of 6 g per m2 and O. chalcidica seedlings were transplanted at a density of 40 plants per plot (equivalent to 4 plants per m2).
The experimentation was laid out following a randomized complete block design with 4 blocks and each plot measuring 10 m2 (5 × 2 m). The treatments that were tested each year were: the co-cropping treatment named “CoC” (cropping the legume at the same time as O. chalcidica), the fertilized control treatment named “FCon” (control of O. chalcidica with two 60-kg N ha−1 repeated fertilization inputs, in the form of ammonium nitrate powder dissolved in water) and the non-fertilized control treatment “NFCon” (cropping O. chalcidica without fertilization). The plots were spaced 50 cm apart. A plastic film was buried vertically 50 cm deep to delimit the fertilized plots and to prevent horizontal N losses through sub-surface run-off and contamination of adjacent plots.
Due to its slow growth rate, O. chalcidica seeds were sown on germination plates. The plates were put in a greenhouse for 18 weeks before transplanting the seedlings into the field in the September of each year. At the same time, the legume seeds were sown for the co-cropping (“CoC”) treatment. For these treatments, legume shoots were harvested after 4 months of culture each year. The aerial parts of the legume were harvested, then dried and crushed in the laboratory. The dried and crushed legume biomass was then incorporated into the field topsoil. These steps were repeated each year. Nitrogen fertilization was applied twice at a rate of 60 kg N ha−1 for the treatment “FCon” (in March and April of each year).
Plant analyses
Nutrient content and Ni concentration in the dried and crushed biomass of the legume were analyzed before being incorporated into the topsoil of field (Supplementary Table 1). O. chalcidica shoots were harvested at the flowering stage (May of 2016 and 2017). Then, the shoots were oven-dried at 70 °C for 72 h and their dry weights recorded. Subsamples (0.5 g) of dry and ground shoot tissue were acid-digested at 95 °C in 2.5 ml of concentrated HNO3 and 5 ml of H2O2 (30%). The final solutions were filtered (0.45 μm DigiFILTER, SCP science, Canada) and completed to 25 ml with deionized water. The Ni concentration in the solution was measured with an Inductively Coupled Plasma-Atomic Emission Spectrometer (ICP-AES, Liberty II, Varian). The total C and N in the shoots were analyzed by combustion at 900 °C with a CHNS analyzer (vario MICRO cube, Elementar Analysensysteme GmbH). Plant quality controls from the International Soil-Analytical Exchange of WEPAL were used for these analyses.
Soil analyses
Soil physicochemical analyses
At each harvest, 200 g of fresh rhizosphere soil (obtained from the rootball of 5 plants from each plot) were collected and sieved (5 mm), and stored at 4 °C before being brought to France for analysis. Half of the fresh soil samples were dried at 40 °C for the physicochemical analyses and the other half was kept at 4 °C for the microbial analyses. Two grams of fresh rhizosphere soil were frozen at −80 °C for further molecular analyses.
Soil moisture was determined by heating subsamples to 105 °C until a constant weight was attained. Available Ni in soil samples was extracted with a DTPA–TEA solution (0.005 M Diethylene Triamine Pentaacetic Acid, DTPA, 0.01 M CaCl2, 0.1 M triethanolamine, pH 7.3), according to Lindsay and Norvell (1978) and the [Ni] in solutions was measured with an ICP-AES. The CEC and exchangeable cations were measured according to international ISO standard 23,470. Soil pH was measured using a pH meter in a soil–water suspension (soil:water ratio = 1:5, v:v). Total and organic C and N were quantified with a CHNS analyzer. Soil quality controls from the International Soil-Analytical Exchange of WEPAL were used for these analyses.
Soil microbial analyses
Microbial biomass carbon (MBC) and nitrogen (MBN) in the soil were determined according to Jenkinson and Ladd (1981) and Brookes et al. (1985). Urease activity was measured according to Tabatabai and Bremner (1972). Arylsulfatase, β-glucosidase and alkaline phosphatase activities were determined according to the modified protocol used by Dick et al. (2013).
Genomic DNA was extracted from soil samples using the FastDNA™ SPIN kit for Soil (MP Biomedicals™, France) in accordance with the manufacturer’s protocol. DNA solutions concentrations were first measured with a spectrophotometer (SmartSpec Plus spectrophotometer, BIO-RAD) and then adjusted to 1.66 ng μl−1 with sterile ultra-pure water, using a robot (epMotion P5073, Eppendorf) in 96-well microplates (MicroAmp® Optical 96-Well Reaction plate). The 16S rRNA gene libraries were constructed according to Klindworth et al. (2013). Amplicons were prepared according to a modified protocol described by Goux et al. (2016). The sequences obtained were de-multiplexed, quality-trimmed and assigned to OTUs at 97% similarity with the FROGS pipeline (http://frogs.toulouse.inra.fr/). Taxonomy affiliation was performed using the Silva database (Silva.nr_v132, https://www.arb-silva.de/). This sequencing data project has been deposited at the DDBJ/EMBL/GenBank under the accession reference KBZD00000000. The version described in this paper is the first version, KBZD01000000. Alpha and Beta diversities and the graphical representation NMDS (Non-metric MultiDimensional Scaling) were studied using QIIME software (Quantitative Insights Into Microbial Ecology, version 1.8.0). Relative abundances were calculated using XLSTAT software (XLSTAT version Ecology 18.07, http://www.xlstat.com).
The metabolic functions of the OTUs were predicted using the Tax4Fun package (Aßhauer et al. 2015), which transforms the SILVA based OTUs into a taxonomic KEGG profile (Kyoto Encyclopedia of Genes and Genomes) organisms (fctProfiling = T), normalized by the 16S rRNA copy number (normCopyNo = T).
Statistical analyses
Statistical parametric analyses were performed on plant parameters and metagenomic data (One-way ANOVA, normality tests and K-sample comparison). Furthermore, all the soil parameters studied were submitted to PCA. Redundancy analysis (RDA) was performed between the soil physicochemical characteristics and the relative abundancy of the bacterial phyla. These statistical analyses were carried out on XLSTAT software (XLSTAT version Ecology 18.07, http://www.xlstat.com). For all tests, differences were considered statistically significant if p value <0.05. A multivariate Regression Tree (MRT, De’Ath 2002) was constructed using the R package “mvpart” 1.6–2 with default sets to understand the correlation between the relative abundance of the major phyla and the soil physicochemical parameters. This method performs hierarchical dichotomous clustering of community data by selecting soil parameters that maximize the homogeneity within group samples.
Results
Potential shoot biomass, Ni, C and N yields
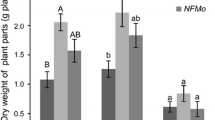
First year results of plant parameters were obtained from Saad et al. (2018c). For the second year, we measured again the shoot biomass, Ni, C and N yields in order to assess its variation along time. For the first year, the potential shoot biomass of O. chalcidica from the treatments of “CoC” and “FCon” were higher than “NFCon” (827, 884 and 160 kg ha−1 respectively, Saad et al. 2018c). Same trends were observed for the second year: 2013, 1625 and 400 kg ha−1 for “CoC”, “FCon” and “NFCon” respectively (Fig. 1). At the same time, no significant difference was detected between “CoC” and “FCon” treatments for each year. “CoC” and “FCon” treatments showed 143% and 84% augmentations for the second year in comparison with the yields of the first year. For the second year, “CoC” had the highest potential biomass of all the treatments. The same trends were shown for the potential shoot Ni yields, where “CoC” and “FCon” had the respective Ni yields of 7.8 and 5.8 kg ha−1 for the first year (Saad et al. 2018c), and 13.6 and 9.9 kg ha−1 for the second year. As in the case of the shoot biomass yield, “NFCon” had a lower potential Ni yield for both years (1.3 and 4 kg ha−1, respectively for the first and the second year). Results of major and minor elements are presented in the Supplementary Table 2.
Potential biomass, Ni, C and N yields of the shoots (kg ha−1) of O. chalcidica. “CoC”, “FCon” and “NFCon” correspond to co-cropping, fertilized control and non-fertilized control, respectively. “Y1” and “Y2” correspond to first and second year of cultivation. For each plant parameter, bars represent means ± standard error. Values for the same plant parameter followed by the same letter are not significantly different at p ≤ 0.05 (n = 4)
Over time, “CoC” and “FCon” treatments showed a significant increase in potential shoot C yield, when comparing the first year (Saad et al. 2018c) to the second year (corresponding to 292 vs. 871 kg C ha−1 for “CoC” and 316 vs. 679 kg C ha−1 for “FCon”). For the potential shoot N yield, a clear and significant improvement was shown for “FCon”, when comparing the 2 years of cultivation (21 vs. 41 kg N ha−1) and had the highest value presented by any of the treatments. However, “CoC-Y2” had a significantly higher shoot N yield than that of “NFCon-Y2” (23 vs. 5 kg N ha−1). In addition, over time, “NFCon” did not show any significant improvement for either C or N potential yields and had the lowest values when compared to the other treatments, for either year of cultivation.
Soil physicochemical and microbial analyses
First year soil parameters results (except High-throughput 16S rRNA amplicon sequencing results) were obtained from Saad et al. (2018c). The results presented in Fig. 2 were obtained by submitting the various soil physicochemical and microbial parameters studied to a Principal Component Analysis (PCA). Results of the soil physicochemical and microbial parameters are presented in Supplementary Table 3. Axis 1 explains 39% of the total variability and separates “FCon-Y1” from the other treatments of the first year’s harvest (Fig. 2a). This first principal component is strongly correlated with both soil total carbon concentration (C) and soil organic carbon concentration (Corg). This suggests that these two parameters vary together. This component can be viewed as a measure of the carbon status of the soil. The second component (Axis 2) corresponds to the level of the bioavailable nickel concentration. Along this Axis, which explains 21% of the total variability, first year treatments (“CoC-Y1”, “FCon-Y1” and “NFConY1”) are clearly separated from the second-year treatments (“CoC-Y2”, “FCon-Y2” and “NFConY2”). The soils from the “FCon-Y1” treatment are characterized by negative correlations with almost all microbial and soil physicochemical parameters. Treatments from the second-year cultivation are negatively correlated with the soil DTPA-extractable Ni concentration (Ni-DTPA) and phosphatase microbial activity (Phos) (Fig. 2b). These results are in agreement with the correlations observed between soil physicochemical and microbial parameters presented in the Supplementary Table 4.
Principal Component Analysis (PCA) generated from soil parameters measured for each cultivation treatment. a Points represent the coordinated means of different treatments (“CoC”, “FCon” and “NFCon” correspond to co-cropping, fertilized control and non-fertilized control, respectively. “Y1” and “Y2” correspond to first and second years of cultivation) and the standard error of four replicate samples. b Soil microbial and physico-chemical parameters involved in the discrimination of samples. Soil parameters are abbreviated as the following: microbial biomass carbon (MBC), microbial biomass nitrogen (MBN), soil arylsulfatase activity (Aryl), soil urease activity (Urease), soil β-glucosidase activity (β-glucosidase), soil phosphatase activity (Phos), soil total carbon concentration (C), soil total nitrogen concentration (N), soil organic carbon concentration (Corg), soil organic nitrogen concentration (Norg), soil pH (pH), soil Cation Exchange Capacity (CEC), bioavailable nickel concentration extracted with Diethylene Triamine Pentaacetic Acid (Ni-DTPA) and soil exchangeable Ni concentration (Niex)
High-throughput 16S rRNA amplicon sequencing results
First year DNA soil extracts (stored at −80 °C) were re-sequenced at the same time as the second year samples. Alpha diversity for the soil samples was studied using the Shannon index, equitability, Chao1 and Simpson evenness (Table 1). The Shannon index for the “FCon” treatment was significantly lower than any of the second-year cultivation treatments (10.19 vs. 10.43 and 10.39 respectively for “FCon-Y2” vs. “CoC-Y2” and “NFCon-Y2”). “CoC” treatment showed a higher equitability value than that of the “FCon” treatment (0.911 for “CoC-Y2”, “0.896” for FCon-Y2”), thus indicating that the bacterial community for “CoC-Y2” tended to have more homogeneous OTU proportions than “FCon-Y2”. The Chao1 was significantly higher for “CoC-Y2” and “NFCon-Y2” than any treatment for both years. Moreover, the Chao1 for “CoC-Y2” was significantly higher than “CoC-Y1”. The Simpson evenness did not vary significantly between treatments whatever the year considered.
Relative bacterial abundance at the phyla and subphyla level is represented in Fig. 3. A total of 3015 OTUs were found for the treatments as a whole and were affiliated within 15 different phyla and subphyla (“Other” phylum regroups all the phyla with a relative abundance <1%). Proteobacteria phylum, which includes the subphyla of γ-Proteobacteria, α-Proteobacteria and δ-Proteobacteria, was well-represented for all the treatments. The relative abundance of Gemmatimonadetes increased significantly with time for the “CoC” treatment (6.51 and 8.81% for the first and second year of cultivation, respectively). A significant increase was observed for the “FCon” treatment with time concerning the relative abundance of α-Proteobacteria (8.91 and 11.30%, for Y1 and Y2, respectively) and Planctomycetes (FCon-Y1: 2.47% and FCon-Y2: 3.22%). The relative abundance of Verrucomicrobia also increased significantly from first and second year for the “CoC” (0.81 and 1.68%) and “FCon” treatments (1.20 and 1.80%). In addition, the relative abundance of Patescibacteria significantly increased with time, whatever the treatment. No significant differences were found for Bacteroidetes, Actinobacteria, Chloroflexi, δ-Proteobacteria and Nitrospirae. The relative abundance of the γ-Proteobacteria subphylum decreased significantly during the second year (21.40, 21.39 and 21.90% for “CoC-Y2”, “FCon-Y2” and “NFCon-Y2”, respectively), when compared with the relative abundances observed for the first year of cultivation (25.95, 25.26 and 26.10% for “CoC-Y1”, “FCon-Y1” and “NFCon-Y1”, respectively). The same trend was found for the Rokubacteria, which displayed a significant decrease for all the treatments, except for the fertilized one (“FCon”), for which no significant difference was observed between the 2 years of cultivation. The relative abundance of Fibrobacteres was only found to decrease significantly between the 2 years for the fertilized treatment (1.59% and 0.35% for “FCon-Y1” and “FCon-Y2”, respectively). Concerning Acidobacteria, only a decrease in the relative abundance for the “FCon-Y2” treatment (16.74%) can be observed when compared to “NFCon-Y2” (20.52%).
Relative abundance of bacterial subphyla and phyla identified in all samples (%). “Others” refer Cyanobacteria, Elusimicrobia, Entotheonellaeota, FCPU426, Firmicutes, Latescibacteria, Multi-affiliation, Omnitrophicaeota, Spirochaetes, Tenericutes, WPS-2 and WS2. “CoC”, “FCon” and “NFCon” correspond to co-cropping, fertilized control and non-fertilized control treatments, respectively. “Y1” and “Y2” correspond to first and second year of cultivation. Means ± standard error followed by the same letter are not significantly different according to Duncan’s test at p ≤ 0.05 (n = 4)
The NMDS (Non-metric Multidimensional Scaling) graphical representation at OTU level allowed a comparison of the treatments (Fig. 4) based on the unweighted phylogenetic criteria and on sequence alignment (sequences not influenced by their number). Treatments from the second year of cultivation were clearly separated along Axis 1 (NMDS1) from those of the first year. In addition, the “FCon” treatment, whatever the year of cultivation, was separated from all other treatments along Axis 2 (NMDS2) and this was clearer in the case of the first year.
Non-metric multidimensional scaling: distribution of the treatments according to their bacterial community. Points represent the coordinate means of different treatments (“CoC”, “FCon” and “NFCon” correspond to co-cropping, fertilized control and non-fertilized control treatments, respectively. “Y1” and “Y2” correspond to first and second year of cultivation). Bars correspond to standard error (based on four replicates for each treatment)
The functional potential of the bacterial community metagenome profile was evaluated using the Tax4Fun approach. Based on the predicted metagenomes, six of the Level 1 KEGG Orthology (KO) family genes were found (Fig. 5). These metabolism-related genes clearly dominated the overall functional structure of the soil bacterial community. The metabolism-related KEGG pathways showed a significant increase with time in the case of the “CoC” treatment (affecting KEGG Level 2 pathways: carbohydrates metabolism, data not shown). With time, a significant negative impact of the mineral fertilization treatments was detected for those genes related to the genetic information process (affected KEGG Level 2 pathways: translation, replication and repair and transcription, data not shown). Concerning the genes related to the environmental information processing, “CoC-Y1” showed a higher significant percentage than “FCon-Y1” and “NFCon-Y1”. In contrast, mineral fertilization showed a significant positive impact with time on this KO group (affecting KEGG Level 2 pathways: membrane transport, data not shown). Regarding the genes predicted relating to the cellular processes pathways, a significant decrease was showed with time in the case of the “CoC” treatment (affecting KEGG level 2 pathways: cell motility).
Gene profiles of bacterial community in O. chalcidica rhizosphere predicted in all treatments (%). “CoC”, “FCon” and “NFCon” correspond to co-cropping, fertilized control and non-fertilized control treatments, respectively. “Y1” and “Y2” correspond to first and second year of cultivation. Means ± standard error followed by the same letter are not significantly different according to Duncan’s test at p ≤ 0.05 (n = 4)
Global analyses
A redundancy analysis (RDA) was performed between soil physicochemical characteristics and the relative abundancy at the bacterial phylum level (>10%) for all treatments (“CoC”, “FCon” and “NFCon” from Y1 and Y2), (Fig. 6). Axis 1 explains 53.73% of the total variability separating second year treatments (negative abscises) from those of the first year (positive abscises). This Axis is strongly correlated with soil bioavailable Ni concentration (Ni-DTPA) and a greater relative abundance of the γ-Proteobacteria subphylum and Bacteroidetes phylum. Along Axis 2, which explains 22.73% of the total variability, “FCon-Y1” treatment is strongly separated from other treatments from the first year of cultivation. “NFCon-Y2” treatment (negative ordinates) was found to be clearly separated along Axis 2 from all the other treatments, whatever the year of cultivation. This Axis appears to be correlated with soil pH. Spearman correlations (p < 0.05) were measured between the relative abundance of the bacterial phyla and the soil physicochemical characteristics for all the treatments. Only significant correlations were retained. γ-Proteobacteria subphylum and Bacteroidetes phylum were more abundant in the rhizosphere soils collected during the first year of cultivation and particularly in the case of “CoC-Y1”, when compared to those of the second year. γ-Proteobacteria was positively correlated to soil Ni-DTPA (R = 0.51) and negatively correlated to soil’s total nitrogen content, CEC and Niex (R = −0.51, R = −0.71 and R = −0.57, respectively). Actinobacteria, Gemmatimonadetes, α-Proteobacteria and Acidobacteria were more abundant in the rhizosphere soils of the second-year treatments. Moreover, the relative abundance of the Gemmatimonadetes was visibly linked to the rhizosphere soils of “CoC-Y2” treatment and was positively correlated to soil CEC (R = 0.56) and exchangeable Ni (Niex) (R = 0.46) and negatively correlated to soil Ni-DTPA (R = − 0.61). α-Proteobacteria subphylum was positively correlated to soil Niex (R = 0.50).
Redundancy Analysis (RDA) performed between soil physico-chemical characteristics and relative abundancy of bacterial subphyla and phyla for planted treatments. Points ± standard error represent the coordinate means of different treatments (“CoC”, “FCon” and “NFCon” correspond to co-cropping, fertilized control and non-fertilized control treatments, respectively. “Y1” and “Y2” correspond to first and second year of cultivation) based on 4 points for each treatment (4 replicates). Coordinates of soil parameters were multiplied by a factor of 4 in order to be clearer on the RDA graph. Abbreviations: soil total carbon concentration (C), soil total nitrogen concentration (N), soil organic carbon concentration (Corg), soil organic nitrogen concentration (Norg), soil pH (pH), soil Cation Exchange Capacity (CEC), bioavailable nickel concentration extracted with Diethylene Triamine Pentaacetic Acid (Ni-DTPA) and soil exchangeable Ni concentration (Niex)
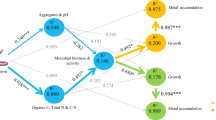
A multivariate regression tree (MRT) was performed to reveal those environmental factors which most affected the bacterial community composition (Fig. 7). This analysis provided a tree with three terminal nodes based on exchangeable Ni (Niex) and soil organic carbon (Corg) which together explain 41.5% of the standardized abundance variance. Niex was the most influential parameter on the phyla relative abundance. All samples collected during the first year were characterized by low Niex levels (<1.18 mg kg−1) and were separated from those of the second year (higher Niex values). Concerning samples from the first year, Corg was identified as the second major environmental factor which affected the bacterial community composition. “CoC-Y1” (Corg >3.18 g kg−1) was separated from other treatments (Corg <3.18 g kg−1). γ-Proteobacteria and Acidobacteria were the most abundant phyla along the tree and showed a decrease when Niex exceeded 1.18 mg kg−1. Actinobacteria and α-Proteobacteria showed little variation along the tree. Gemmatimonadetes abundance increased with the increase of Niex and decreased when Corg was higher than 3.18 g kg−1. Bacteroidetes were shown to be favored with the increase of Corg at low Niex levels.
Multivariate regression tree of the relation between the relative abundance of the major subphyla and phyla and the soil physico-chemical parameters. The bar plots show the mean relative abundance of each phylum at the terminal nodes. Abbreviations: soil organic carbon concentration (Corg) and soil exchangeable Ni concentration (Niex)
Discussion
After 2 years of cultivation, O. chalcidica from the “CoC” treatment had the highest potential biomass and Ni yields of all the treatments and reached 2013 kg ha−1 and 13.6 kg ha−1, respectively. When a Brassicaceae has been co-cropped with a legume, an increased biomass and N-accumulation have previously been observed by many authors (Banikm et al. 2000; Andersen et al. 2005; Szumigalski and Van Acker 2005). While improving crop productivity, co-cropping the Brassicaceae with the legume can also reinforce its competitive ability against weeds (Szumigalski and Van Acker 2005). In addition, the introduction of legumes to conventional agro-ecosystems has already shown its use in improving soil organic matter and structure. Consequently, crop productivity has been seen to be enhanced due to enriched soil nutrient levels and deep soil exploration (Teasdale et al. 1991; Fisk et al. 2001; Sarrantonio and Gallandt 2003). Biomass yields obtained from the “FCon” and “NFCon” treatments confirm those of the field experiments carried out in Albania by Bani et al. (2015). For their field trials, the fertilization rate was the same as in this study and contributed to a strong increase in O. chalcidica’s biomass compared to the non-fertilized plots (2300 kg ha−1 vs. 272.2 kg ha−1). In addition, a significant increase in Ni yield was observed when the nitrogen fertilizers were used in the first-year trial (Bani et al. 2015). Lower yields were obtained in our study than those obtained by Bani et al. (2015). This could be explained by the differences in both the Spanish and Albanian climate and soil characteristics. We should not forget either that the soil bioavailable Ni concentration (DTPA-extractable Ni) of the site studied was around 41 mg Ni kg−1, whereas in Albanian studied soils, this reached more than 120 mg Ni kg−1. Moreover, a significant increase in potential shoot C yields were observed for both “CoC” and “FCon” treatments over time. This was related to the obtention of better biomass of the hyperaccumulator plant for these two treatments. Shoot N yields were only improved for the “FCon” treatment over time. N has been proven to be a limiting factor for an optimal crop yield (Harker et al. 2012). This explains why, in the case of “FCon” treatment, N chemical fertilization increased O. chalcidica’s shoot N yield.
Principal Component Analysis revealed a negative impact of the “FCon” treatment on soil parameters. Being the preferred nitrogen source for most bacteria and fungi, nitrogen fertilizers used at high rates can negatively affect soil microorganisms (Marzluf 1997; Omar and Ismail 1999; Geisseler and Scow 2014). Over time, soil microbial phosphatase activity decreased for all the treatments. In our study, soil was amended with 122.5 kg P ha−1 each year before plantation in order to improve P content. This was in accordance with Allison and Vitousek (2005) who showed that phosphatase activity was declined in response to phosphate additions. Moreover, over time, low soil DTPA-extractable Ni levels were obtained for the “CoC” and “FCon” treatments.. Echevarria et al. (1998) showed that the DTPA-extractable Ni in the soil, is the soluble soil Ni fraction that is most likely to be absorbed by hyperaccumulator roots.
Over recent years, many reports have employed Next Generation Sequencing to reveal a fundamental and new understanding of the rhizomicrobiome structure and diversity (Metzker 2010; Mendes et al. 2013; Knief 2014; Yasir et al. 2015; Lopez et al. 2017), but very little is known about the microbial diversity associated with the rhizosphere of hyperaccumulator plants. However, the hyperaccumulators’ rhizosphere bacterial community has been recognized to influence plant growth and development in metal-rich soils, by modifying metal mobility and bio-availability (Reeves and Adigüzel 2008; Sessitsch et al. 2013). Of the 27 phyla identified in the studied soils, 15 had relative abundances exceeding 1%. Whatever the treatment, the bacterial phyla identified in O. chalcidica rhizosphere are commonly encountered as dominant taxa in soils (Rastogi et al. 2010). Proteobacteria was the dominant phylum in the hyperaccumulator rhizosphere, as observed in many other soil types, including those that are multi-contaminated (Cr, Zn and Pb) (Gołębiewski et al. 2014), naturally metal-rich soils (Lopez et al. 2019), agricultural soils (Yang et al. 2017) or even forest soils (Uroz et al. 2010). The bacteria belonging to this phylum have been defined as copiotrophic (Lienhard et al. 2014) and are known to prefer carbon-rich environments, such as rhizospheres (Yang et al. 2017). However, the relative abundance of the γ-Proteobacteria subphylum decreased significantly for the second year compared with the first year of cultivation. This bacterial phylum is known for its capacity to rehabilitate brownfields by bioleaching the heavy metals (Yang et al. 2016). In addition, bacterial species of this phylum can tolerate high Ni concentrations in metal-rich soils (Idris et al. 2006). Our results showed that the γ-Proteobacteria subphylum is strictly linked to the soil Ni-DTPA in the RDA analysis. The decrease in the soil Ni-DTPA for the second year of cultivation could explain the relative abundance reduction of the γ-Proteobacteria subphylum in the soil. In fact, the decrease in soil bioavailable Ni concentrations could favor the growth of other non-tolerant bacterial communities and increase their competition with the Ni-tolerant ones. Other notable phyla were in order of abundance: Acidobacteria, Bacteroidetes and Actinobacteria. These results confirmed those obtained from contaminated soils (Kim et al. 2006). Indeed, in soil, Acidobacteria constitute on average 20% of all bacteria (Naether et al. 2012) and this observation is in accordance with our results (19%). Among the previously-known environmental factors that correlate to Acidobacteria abundance in soils, pH is the most prominent (Jones et al. 2009) as confirmed by our redundancy analysis (Fig. 6). Moreover, we observed that Acidobacteria’s relative abundance decreased with time for the fertilized treatment. In fact, even if Acidobacteria plays a crucial role in the C cycle due to its ability to degrade complex plant-derived polysaccharides, such as cellulose and lignin (Ward et al. 2009), we can hypothesize that the bacterial communities were negatively influenced by the mineral fertilization and were more dependent on N addition than on decomposing plant materials. These conditions reduced the relative abundance of C-dependent bacteria in the soil such as Acidobacteria. In contrast, the relative abundance of Gemmatimonadetes increased significantly with time for the “CoC” treatment. This phylum is known to be a ubiquitous polyphosphate-accumulating bacteria (Zhang et al. 2003). In fact, legumes are known to solubilize phosphorus through soil acidification resulting from the secretion of large amounts of protons in the rhizosphere soil (Hinsinger 2001; Yan et al. 2002). The significant number of protons in the soil was confirmed both by the RDA and the multivariate regression tree analyses, where this phylum was positively correlated to soil CEC and exchangeable Ni. Consequently, in these soil conditions, Gemmatimonadetes were favored in the soils of the treatment including the legume. A significant increase was detected with time for the “FCon” treatment concerning the relative abundance of α-Proteobacteria and Planctomycetes. In fact, the α-Proteobacteria subphylum is known to use ammonia and nitrate as its sole nitrogen source (Madigan et al. 1984). In addition, Planctomycetes species have been observed in environments in all trophic states, with some reports of higher numbers occurring in eutrophic and polluted waters (Staley et al. 1980). Furthermore, Planctomycetes have large genomes, which is a feature of copiotrophs that prefer nutrient-rich environment (Lauro et al. 2009). The relative abundance of Verrucomicrobia increased significantly with time for the “CoC” and “FCon” treatment. This phylum is known to be favored by high nutrient availabilities (Haukka et al. 2006) and the increase in its relative abundance could be related to the decomposition of the legume organic matter (in the case of co-cropping) and the addition of the mineral fertilization (in the case of the fertilized treatment). Recently, the three candidate phyla, Parcubacteria, Microgenomates and Gracilibacteria, have been grouped into the Patescibacteria superphylum (Rinke et al. 2013; Hedlund et al. 2014). The Patescibacteria superphylum showed a significant increase for all the treatments with time. Patescibacteria sequences were first reported in groundwater and sediments of anoxic aquatic environments (Elshahed et al. 2005; Youssef et al. 2011; Wrighton et al. 2012). Nevertheless, prospective metagenomic analyses have since established that this phylum has a widespread environmental distribution including the maize rhizosphere (Correa-Galeote et al. 2016). Concerning the Fibrobacteres phylum, its relative abundance decreased with time for the fertilized treatment. In fact, this phylum is known to decompose plant material and utilize cellulose as a carbon source (Qi et al. 2008). As we previously hypothesized, the bacterial communities could be negatively influenced by the mineral fertilization and were more dependent on N addition than on decomposing plant materials. Consequently, this could induce a decrease in the relative abundance in the soil of C-dependent bacteria such as the Fibrobacteres.
The NMDS showed that the treatments of the first-year cultivation were separate from those of the second year, confirming that the soil bacterial diversity was modified. In addition, the “FCon” treatment showed a clear separation from the other treatments for both years: underlining that this treatment induced a particular bacterial community. The same trends were shown for the RDA analysis. Alpha-diversity indexes of the soil samples confirmed these observations. Indeed, the Shannon index decreased significantly with time for the “FCon” treatment. Moreover, the Chao1 was lower for the “FCon” treatment for both years of cultivation. Soil microbial diversity has been shown to decrease after long-term application of NPK chemical fertilizers (Postma-Blaauw et al. 2012; Sun et al. 2015). Moreover, it has been recently shown that the mineral fertilization can decrease the soil bacterial diversity after just 1 year of cultivation (Liang et al. 2020). Conversely, previous studies have revealed that organic amendments, such as compost or manure, improved the soil bacterial diversity. Indeed, Zhang et al. (2015) showed that addition of livestock manure, straw or green manure enhance albic paddy soil nutrients, enzyme activities and affect positively the microbial biomass and structure. In the same way, Sun et al. (2015) showed, among typical lime concretion black soils subjected to 30 years of NPK fertilization, that the use of pig or cow manure improved soil bacterial diversity. In our study, the co-cropping treatment improved the soil bacterial diversity. As proofed by recent studies (Benizri and Amiaud 2005; Gao et al. 2012), the coexistence of different plant species induced a variety of rhizodeposits (Zak et al. 2003), thus generating a better bacterial diversity with a range of functional microbial groups (Wardle et al. 2004; Benizri and Amiaud 2005; Gao et al. 2010). Enhancing soil bacterial diversity is known to improve soil nitrogen and carbon cycles. Indeed, Griffiths et al. (2000, 2001) showed a positive effect of the soil microbial diversity on mineralization of complex carbon sources. Moreover, Saad et al. (2018a) showed that co-cropping O. chalcidica with Vicia sativa ameliorated the soil aggregate stability and the soil particules size. A better structured soil allows a deep development of the plant root system resulting in an enhanced plant nutrition (Passioura 1991).
Based on the predicted metagenomes using the Tax4Fun approach, genes belonging to metabolism were identified as the major gene families at the Level 1 KO groups. Our results confirm recent studies where the metabolism-related functions were found in great abundance in the rhizospheres of hyperaccumulator plants (Lin et al. 2013; Lopes et al. 2016; 2019). In addition, we found that carbohydrate metabolism increased significantly with time for the co-cropping treatment. This could be related to the fact that the complex compounds, present in organic matter (i.e. incorporated legume residues) in the case of the co-cropping treatment, are recognized as a major C and N source for the bacterial activity. The latter would activate complex enzyme systems in order to degrade and utilize these compounds (Kögel-Knabner 2002). In addition, it is theorized that the increase in plant growth creates a positive feedback, which increases root exudates for bacterial metabolism (Mahoney et al. 2017). Moreover, we can hypothesize that, after 2 years of cultivation and the legume introduction, the soil was better structured due to a greater presence of micro-aggregates (Saad et al. 2018a, b), thereby enhancing soil bacterial metabolism. With time, a significant negative impact of the mineral fertilization (“FCon”) was detected on the percentages of the translation, replication and repair and transcription gene families. The co-cropping treatment showed a higher significant percentage for those genes related to environmental information pathways than the other treatments of the first year of cultivation. In contrast, mineral fertilization showed a significant increase with time for this KO group and especially for the membrane transport category. In addition, a significant decrease with time was shown for the cellular process pathways in the case of the co-cropping treatment (affected KEGG Level 2 pathways: cell motility). These results could be explained by the fact that membrane transport and cell motility could permit the bacteria to interact with their surroundings and react to chemical gradients generated by rhizodeposits and other signals in the rhizosphere (Somers et al. 2004). On one hand, the addition of the mineral fertilization could be a possible reason for the increase in the percentage of the bacterial membrane transport category. Since membrane transporters play an important role in many different aspects of bacterial physiology, this results in facilitating both the import of nutrients and the extrusion of toxins and antimicrobial compounds (Davidson and Chen 2004; Lin et al. 2013). On the other hand, the enrichment of the soil with legume residues in the case of co-cropping could generate a stable and locally rich-environment for the bacteria, thus minimizing their need to be mobile in order to search for soil nutrients through bacterial chemotaxis (Somers et al. 2004).
Even if an increasing number of studies have attempted to characterize how microbial distribution patterns respond to plants and environmental factors, to our knowledge, few studies have ever investigated the influence of physicochemical factors and their modifications with time, on the rhizosphere bacterial community of hyperaccumulator plants growing on ultramafic soils (Pardo et al. 2018; Visioli et al. 2018). The multivariate regression tree showed that Niex was the most influential parameter on the phyla’s relative abundance with all first-year samples characterized by low Niex levels (<1.18 mg kg−1) and separate from those of the second year with a higher Niex. Nevertheless, the presence of metals in soils can induce changes in the structure and diversity of the soil bacterial communities, as evidenced by numerous studies (Sandaa et al. 1999; Mengoni et al. 2001; Idris et al. 2004; Lopez et al. 2017).. Soil organic matter is considered as an important indicator of soil quality because of the many functions it provides. Organic matter is a source of C and N and influences the phosphorus and sulfur cycles (Carter 2002). In addition, it has the ability to complex with multivalent ions and organic compounds. Organic matter has an effect on aggregate stability, water retention and soil hydraulic properties (Carter 2002). In our study, the PCA analyses (Fig. 2) allowed for a clear differentiation between treatments from the first and those of the second year, along Axis 2. Moreover PCA showed that Corg, C and N increased with time. The application of organic fertilization (in our case, the addition of legume residues to the soil), can increase soil organic carbon and enhance the bacterial communities that are known to be involved in the decomposition of complex organic matter and soil carbon, nitrogen, and phosphorus transformations (Li et al. 2017).
Conclusions
This study showed that the introduction of a legume into a Ni agromining system improved both plant biomass and Ni yields. The results obtained from the co-cropping system clearly demonstrated the improvement of Ni yields in comparison to fertilized and non-fertilized treatments. Soil bioavailable Ni concentration was lowered with time in comparison to the soil initial concentration before the implementation of this cropping system. In addition, mineral fertilization had a negative impact on many microbial and soil physicochemical parameters. Co-cropping enhanced the soil bacterial diversity in contrast to the fertilized treatment that reduced it. Moreover, co-cropping with the legume increased the relative abundance of genes related to the bacterial metabolism. Co-cropping of O. chalcidica with the legume and incorporating the legume dry biomass into the soil, could reduce the need for fertilizers, as well as lowering the risk of nitrogen leaching and consequent underground water pollution. Further studies could be done in other regions where freezing temperatures during winter could naturally destroy the legume plant cover in order to avoid the manual incorporation of the legume biomass. In addition, more research is needed in order to understand the impact of the living legume on soil parameters prior to its incorporation into the soil. However, this work has confirmed that implementing pluriannual agromining trials on abandoned ultramafic soils can enhance the biomass and Ni yields of the hyperaccumulator plant with time. Improving agromining methods by replacing mineral fertilizers would combine an eco-efficient strategy with a sustainable metal recovery.
References
Abou-Shanab RI, Delorme TA, Angle JS, Chaney RL, Ghanem K, Moawad H, Ghozlan HA (2003) Phenotypic characterization of microbes in the rhizosphere of Alyssum murale. Int J Phytoremediation 5:367–379. https://doi.org/10.1080/15226510309359043
Allison SD, Vitousek PM (2005) Responses of extracellular enzymes to simple and complex nutrient inputs. Soil Biol Biochem 37:937–944. https://doi.org/10.1016/j.soilbio.2004.09.014
Andersen MK, Hauggaard-Nielsen H, Ambus P, Jensen ES (2005) Biomass production, symbiotic nitrogenfixation and inorganic N use in dual and tricomponent annual intercrops. Plant Soil 266:273–287. https://doi.org/10.1007/s11104-005-0997-1
Álvarez-López V, Prieto-Fernández A, Cabello-Conejo MI, Kidd PS (2016) Organic amendments for improving biomass production and metal yield of Ni-hyperaccumulating plants. Sci Total Environ 548–549:370–379. https://doi.org/10.1016/j.scitotenv.2015.12.147
Aßhauer KP, Wemheuer B, Daniel R, Meinicke P (2015) Tax4Fun: predicting functional profiles from metagenomic 16S rRNA data. Bioinformatics 31:2882–2884. https://doi.org/10.1093/bioinformatics/btv287
Bani A, Echevarria G, Sulçe S, Morel JL (2015) Improving the agronomy of Alyssum murale for extensive phytomining: a five-year field study. Int J Phytoremediat 6514:117–127. https://doi.org/10.1080/15226514.2013.862204
Banikm P, Sasmal T, Ghosal P, Bagchi D (2000) Evaluation of mustard (Brassica compestris var. Toria) and legume intercropping under 1: 1 and 2: 1 row-replacement series systems. J Agron Crop Sci 185:9–14. https://doi.org/10.1046/j.1439-037X.2000.00388.x
Benizri E, Amiaud B (2005) Relationship between plant and soil microbial communities in fertilized grasslands. Soil Biol Biochem 37:2055–2064. https://doi.org/10.1016/j.soilbio.2005.03.008
Benizri E, Kidd PS (2018) The role of the rhizosphere and microbes associated with hyperaccumulator plants in metal accumulation. In: van der Ent A, Echevarria G, Baker AJM, Morel JL (Eds.), Agromining: Farming for metals - Extracting unconventional resources using plants. Springer, pp. 157–188. https://doi.org/10.1007/978-3-319-61899-9_9
Bernal MP, Clemente R, Walker DJ (2007) The role of organic amendments in the bioremediation of heavy metal-polluted soils. In: Gore RB (ed) Environmental research at the leading edge. Nova Science Publishers Inc., New York, pp 1–57
Brookes PC, Landman A, Pruden G, Jenkinson DS (1985) Chloroform fumigation and the release of soil nitrogen: a rapid direct extraction method to measure microbial biomass nitrogen in soil. Soil Biol Biochem 17:837–842. https://doi.org/10.1016/0038-0717(85)90144-0
Carter MR (2002) Soil quality for sustainable land management: organic matter and aggregation interactions that maintain soil functions. Agron J 94:38–47. https://doi.org/10.2134/agronj2002.3800
Chaney RL, Baker AJM, Morel JL (2018) The Long road to developing Agromining/Phytomining. In: van der Ent A, Echevarria G, Baker A, Morel JL (Eds) Agromining: Farming for metals - Extracting unconventional resources using plants. Springer, pp. 1–17. https://doi.org/10.1007/978-3-319-61899-9_1
Correa-Galeote D, Bedmar EJ, Fernández-González AJ, Fernández-López M, Arone GJ (2016) Bacterial communities in the rhizosphere of amilaceous maize (Zea mays L.) as assessed by pyrosequencing. Plant Sci 7:1016. https://doi.org/10.3389/fpls.2016.01016
Davidson AL, Chen J (2004) ATP-binding cassette transporters in bacteria. Annu Rev Biochem 73:241–268. https://doi.org/10.1146/annurev.biochem.73.011303.073626
De Antoni MM, Bell M, Grace PR, Scheer C, Rowlings DW, Liu S (2015) Legume pastures can reduce N2O emissions intensity in subtropical cereal cropping systems. Agric Ecosyst Environ 204:27–39. https://doi.org/10.1016/j.agee.2015.02.007
De’Ath G (2002) Multivariate regression trees: a new technique for modeling species-environment relationships. Ecology 83:1105–1117. https://doi.org/10.1890/0012-9658(2002)083[1105:mrtant]2.0.co;2
Dick LK, Jia G, Deng S, Dick RP (2013) Evaluation of microplate and bench-scale β-glucosidase assays for reproducibility, comparability, kinetics, and homogenization methods in two soils. Biol Fert Soils 49:1227–1236. https://doi.org/10.1007/s00374-013-0820-8
Durand A, Piutti S, Rue M, Morel JL, Echevarria G, Benizri E (2016) Improving nickel phytoextraction by co-cropping hyperaccumulator plants inoculated by plant growth promoting Rhizobacteria. Plant Soil 399:179–192. https://doi.org/10.1007/s11104-015-2691-2
Echevarria G, Morel JL, Fardeau JC, Leclerc-Cessac E (1998) Assessment of phytoavailability of nickel in soils. J Environ Qual 27:1064–1070. https://doi.org/10.2134/jeq1998.00472425002700050011x
Elshahed MS, Najar FZ, Aycock M, Qu C, Roe BA, Krumholz LR (2005) Metagenomic analysis of the microbial community at Zodletone spring (Oklahoma): insights into the genome of a member of the novel candidate division OD1. Appl Environ Microbiol 71:7598–7602. https://doi.org/10.1128/AEM.71.11.7598-7602.2005
Fisk JW, Hesterman OB, Shrestha A, Kells JJ, Harwood RR, Squire JM, Sheaffer CC (2001) Weed suppression by annual legume cover crops in no-tillage corn. Agron J 93:319–325. https://doi.org/10.2134/agronj2001.932319x
Gao Y, Zhou P, Mao L, Zhi Y, Zhang C, Shi W (2010) Effects of plant species coexistence on soil enzyme activities and soil microbial community structure under cd and Pb combined pollution. J Environ Sci 22:1040–1048. https://doi.org/10.1016/S1001-0742(09)60215-1
Gao Y, Miao CY, Xia J, Mao L, Wang YF, Zhou P (2012) Plant diversity reduces the effect of multiple heavy metal pollution on soil enzyme activities and microbial community structure. Front Environ Sci Eng 6:213–223. https://doi.org/10.1007/s11783-011-0345-z
Geisseler D, Scow KM (2014) Long-term effects of mineral fertilizers on soil microorganisms - a review. Soil Biol Biochem 75:54–63. https://doi.org/10.1016/j.soilbio.2014.03.023
Gołębiewski M, Deja-Sikora E, Cichosz M, Tretyn A, Wróbel B (2014) 16S rDNA pyrosequencing analysis of bacterial community in heavy metals polluted soils. Microb Ecol 67:635–647. https://doi.org/10.1007/s00248-013-0344-7
Goux X, Calusinska M, Fossépré M, Benizri E, Delfosse P (2016) Start-up phase of an anaerobic full-scale farm reactor - appearance of mesophilic anaerobic conditions and establishment of the methanogenic microbial community. Bioresour Technol 212:217–226. https://doi.org/10.1016/j.biortech.2016.04.040
Gove B, Hutchinson JJ, Young SD, Craigon J, McGrath SP (2002) Uptake of metals by plants sharing a rhizosphere with the hyperaccumulator Thlaspi caerulescens. Int J Phytoremediat 4:267–281. https://doi.org/10.1080/15226510208500087
Griffiths BS, Ritz K, Bardgett RD, Cook R, Christensen S, Ekelund F, Sørensen SJ, Bååth E, Bloem J, de Ruiter PC, Dolfing J, Nicolardot B (2000) Ecosystem response of pasture soil communities to fumigation-induced microbial diversity reductions: an examination of the biodiversity – ecosystem function relationship. Oikos 90:279–294. https://doi.org/10.1034/j.1600-0706.2000.900208.x
Griffiths BS, Ritz K, Wheatley R, Kuan HL, Boag B, Christensen S, Ekelund F, Sørensen SJ, Muller S, Bloem J (2001) An examinataion of the biodiversity-ecosystem function relationship in arable soil microbial communities. Soil Biol Biochem 33:1713–1722. https://doi.org/10.1016/S0038-0717(01)00094-3
Harker KN, O’Donovan JT, Turkington TK, Blackshaw RE, Lupwayi ZN, Smith GE, Klein-Gebbinck H, Dosdall LM, Hall LM, Willenborg CJ, Kutcher HR, Malhi SS, Vera CL, Gan Y, Lafond GP, May WE, Grant CA, Mclaren DL (2012) High-yield no-till canola production on the Canadian prairies. Can J Plant Sci 92:221–233. https://doi.org/10.4141/cjps2011-125
Haukka K, Kolmonen E, Hyder R, Hietala J, Vakkilainen K, Kairesalo T, Haario H, Sivonen K (2006) Effect of nutrient loading on bacterioplankton community composition in lake mesocosms. Microb Ecol 51:137–146. https://doi.org/10.1007/s00248-005-0049-7
Hedlund BP, Dodsworth JA, Murugapiran SK, Rinke C, Woyke T (2014) Impact of single-cell genomics and metagenomics on the emerging view of extremophile “microbial dark matter”. Extrem Life Extreme Cond 18:865–875. https://doi.org/10.1007/s00792-014-0664-7
Hinsinger P (2001) Bioavailability of soil inorganic P in the rhizosphere as affected by root-induced chemical changes: a review. Plant Soil 237:173–195. https://doi.org/10.1023/A:1013351617532
Idris R, Trifonova R, Puschenreiter M, Wenzel WW, Sessitsch A (2004) Bacterial communities associated with flowering plants of the Ni hyperaccumulator Thlaspi goesingense. Appl Environ Microbiol 70:2667–2677. https://doi.org/10.1128/AEM.70.5.2667-2677.2004
Idris R, Kuffner M, Bodrossy L, Puschenreiter M, Monchy S, Wenzel WW, Sessitsch A (2006) Characterization of Ni-tolerant methylobacteria associated with the hyperaccumulating plant Thlaspi goesingense and description of Methylobacterium goesingense sp. nov. Syst. Appl Microbiol 29:634–644. https://doi.org/10.1016/j.syapm.2006.01.011
IUSS Working Group WRB (2014) World Reference Base for soil resources 2014. International soil classification system for naming soils and creating legends for soil maps. World soil resources reports no. 106. FAO, Rome
Jenkinson DS, Ladd JN (1981) Microbial biomass in soil: measurement and turnover. Soil Biochem 5:415–471. https://doi.org/10.1016/0038-0717(91)90183-K
Jiang CA, Wu QT, Wu SH, Long XX (2009) Effect of co-cropping Sedum alfredii with different plants on metal uptake. China Environ Sci 29:985–990 [in Chinese]
Jiang CA, Wu QT, Sterckeman T, Schwartz C, Sirguey C, Ouvrard S, Perriguey J, Morel JL (2010) Co-planting can phytoextract similar amounts of cadmium and zinc to mono-cropping from contaminated soils. Ecol Eng 36:391–395. https://doi.org/10.1016/j.ecoleng.2009.11.005
Jiang CA, Wu QT, Echevarria G, Goudon R, Morel JL (2015) Biomass and metal yield of co-cropped Alyssum murale and Lupinus albus. Aust J Bot 62:159–166. https://doi.org/10.1071/BT14261
Jones RT, Robeson MS, Lauber CL, Hamady M, Knight R, Fierer N (2009) A comprehensive survey of soil acidobacterial diversity using pyrosequencing and clone library analyses. ISME J 3:442–453. https://doi.org/10.1038/ismej.2008.127
Kim J, Kang SH, Min KA, Cho KS, Lee IS (2006) Rhizosphere microbial activity during phytoremediation of diesel-contaminated soil. J Environ Sci Health Subst Environ Eng 41:2503–2516. https://doi.org/10.1080/10934520600927658
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:e1. https://doi.org/10.1093/nar/gks808
Knief C (2014) Analysis of plant microbe interactions in the era of next generation sequencing technologies. Front Plant Sci 5:216. https://doi.org/10.3389/fpls.2014.00216
Kögel-Knabner I (2002) The macromolecular organic composition of plant and microbial residues as inputs to soil organic matter. Soil Biol Biochem 34:139–162. https://doi.org/10.1016/s0038-0717(01)00158-4
Lauro FM, McDougald D, Thomas T, Williams TJ, Egan S, Rice S, DeMaere MZ, Ting L, Ertan H, Johnson J (2009) The genomic basis of trophic strategy in marine bacteria. Proc Natl Acad Sci U S A 106:15527–15533. https://doi.org/10.1073/pnas.0903507106
Li YM, Chaney R, Brewer E, Rosenberg R, Angle JS, Baker AJM, Reeves R, Nelkin J (2003) Development of a technology for commercial phytoextraction of nickel: economic and technical considerations. Plant Soil 249:107–115. https://doi.org/10.1023/A:1022527330401
Li F, Chen L, Zhang J, Yin J, Huang S (2017) Bacterial community structure after long-term organic and inorganic fertilization reveals important associations between soil nutrients and specific taxa involved in nutrient transformations. Front Microbiol 8:187. https://doi.org/10.3389/fmicb.2017.00187
Liang R, Hou R, Li J, Lyu Y, Hang S, Gong H, Ouyang Z (2020) Effects of different fertilizers on rhizosphere bacterial communities of winter wheat in the North China plain. Agronomy 10:93. https://doi.org/10.3390/agronomy10010093
Lienhard P, Terrat S, Prévost-Bouré NC, Nowak V, Régnier T, Sayphoummie S, Panyasiri K, Tivet F, Mathieu O, Levêque J, Maron PA, Ranjard L (2014) Pyrosequencing evidences the impact of cropping on soil bacterial and fungal diversity in Laos tropical grassland. Agron Sustain Dev 34:525–533. https://doi.org/10.1007/s13593-013-0162-9
Lin W, Wu L, Lin S, Zhang A, Zhou M, Lin R, Wang H, Chen J, Zhang Z, Lin R (2013) Metaproteomic analysis of ratoon sugarcane rhizospheric soil. BMC Microbiol 13:135. https://doi.org/10.1186/1471-2180-13-135
Lindsay WL, Norvell WA (1978) Development of DTPA soil test for zinc, iron, manganese and copper. Soil Sci Soc Am J 42:421–428. https://doi.org/10.2136/sssaj1978.03615995004200030009x
Liu L, Li Y, Tang J, Hu L, Chen X (2011) Plant coexistence can enhance phytoextraction of cadmium by tobacco (Nicotiana tabacum L.) in contaminated soil. J Environ Sci 23:453–460. https://doi.org/10.1016/S1001-0742(10)60430-5
Lopes LD, Perreira e Silva MC, Andreote FD (2016) Bacterial abilities and adaptation toward the rhizosphere colonization. Front Microbiol 7:1341. https://doi.org/10.3389/fmicb.2016.01341
Lopez S, Piutti S, Vallance J, Morel JL, Echevarria G, Benizri E (2017) Nickel drives bacterial community diversity in the rhizosphere of the hyperaccumulator Alyssum murale. Soil Biol Biochem 114:121–130. https://doi.org/10.1016/j.soilbio.2017.07.010
Lopez S, Goux X, Echevarria G, Calusinska M, Morel JL, Benizri E (2019) Community diversity and potential functions of rhizosphere-associated bacteria of nickel hyperaccumulators found in Albania. Sci Total Environ 654:237–249. https://doi.org/10.1016/j.scitotenv.2018.11.056
Lucisine P, Echevarria G, Sterckeman T, Vallance J, Rey P, Benizri E (2014) Effect of hyperaccumulating plant cover composition and rhizosphere-associated bacteria on the efficiency of nickel extraction from soil. Appl Soil Ecol 81:30–36. https://doi.org/10.1016/j.apsoil.2014.04.011
Madigan M, Cox SS, Stegeman RA (1984) Nitrogen-fixation and nitrogenase activities in members of the family Rhodospirillaceae. J Bacteriol 157:73–78. https://doi.org/10.1128/JB.157.1.73-78.1984
Mahoney AK, Yin C, Hulbert SH (2017) Community structure, species variation, and potential functions of rhizosphere-associated bacteria of different winter wheat (Triticum aestivum) cultivars. Front Plant Sci 8:132. https://doi.org/10.3389/fpls.2017.00132
Martínez-Fernández D, Arco-Lázaro E, Bernal MP, Clemente R (2014) Comparison of compost and humic fertilizer effects on growth and trace elements accumulation of native plant species in a mine soil phytorestoration experiment. Ecol Eng 73:588–597. https://doi.org/10.1016/j.ecoleng.2014.09.105
Marzluf GA (1997) Genetic regulation of nitrogen metabolism in the fungi. Microbiol. Mol Biol Rev 61:17–32
Mendes R, Garbeva P, Raaijmakers JM (2013) The rhizosphere microbiome: significance of plant beneficial, plant pathogenic, and human pathogenic microorganisms. FEMS Microbiol Rev 37:634–663. https://doi.org/10.1111/1574-6976.12028
Mengoni A, Barzanti R, Gonnelli C, Gabbrielli R, Bazzicalupo M (2001) Characterization of nickel-resistant bacteria isolated from serpentine soil. Environ Microbiol 3:691–698. https://doi.org/10.1046/j.1462-2920.2001.00243.x
Metzker ML (2010) Sequencing technologies – the next generation. Nat Rev Genet 11:31–46. https://doi.org/10.1038/nrg2626
Morel JL (2013) Using plants to “micro-mine” metals. In: http://www.inra.fr/en/Scientists-Students/Biomass/All-the-news/Using-plants-to-micro-mine-metals
Naether A, Foesel BU, Naegele V, Wüst PK, Weinert J, Bonkowski M, Alt F, Oelmann Y, Polle A, Lohaus G, Gockel S, Hemp A, Kalko EKV, Linsenmair KE, Pfeiffer S, Renner S, Schöning I, Weisser WW, Wells K, Fischer M, Overmann J, Friedrich MW (2012) Environmental factors affect acidobacterial communities below the subgroup level in grassland and forest soils. Appl Environ Microbiol 78:7398–7406. https://doi.org/10.1128/AEM.01325-12
Omar SA, Ismail M (1999) Microbial populations, ammonification and nitrification in soil treated with urea and inorganic salts. Folia Microbiol 44:205–212. https://doi.org/10.1007/BF02816244
Pan SW, Wei SQ, Yuan X, Cao SX (2008) The removal and remediation of phenanthrene and pyrene in soil by mixed cropping of alfalfa and rape. Agric Sci China 7:1355–1364. https://doi.org/10.1016/S1671-2927(08)60185-6
Pardo T, Rodríguez-Garrido B, Saad RF, Soto-Vázquez JL, Loureiro-Viñas M, Prieto-Fernández A, Echevarria G, Benizri E, Kidd PS (2018) Assessing the agromining potential of Mediterranean nickel-hyperaccumulating plant species at field-scale in ultramafic soils under humid-temperate climate. Sci Total Environ 630:275–286. https://doi.org/10.1016/j.scitotenv.2018.02.229
Passioura JB (1991) Soil structure and plant growth. Aust J Soil Res 29:717–728. https://doi.org/10.1071/SR9910717
Postma-Blaauw M, de Goede RGM, Bloem J, Faber JH, Brussaard L (2012) Agricultural intensification and de-intensification differentially affect taxonomic diversity of predatory mites, earthworms, enchytraeids, nematodes and bacteria. Appl Soil Ecol 57:39–49. https://doi.org/10.1016/j.apsoil.2012.02.011
Qi M, Jun HS, Forsberg CW (2008) Cel9D, an atypical 1,4–β–D–glucan glucohydrolase from Fibrobacter succinogenes: characteristics, catalytic residues, and synergistic interactions with other cellulases. J Bacteriol 190:1976–1984. https://doi.org/10.1128/JB.01667-07
Rastogi G, Osman S, Kukkadapu R, Engelhard M, Vaishampayan PA, Andersen GL, Sani RK (2010) Microbial and mineralogical characterizations of soils collected from the deep biosphere of the former Homestake gold mine, South Dakota. Microb Ecol 60:539–550. https://doi.org/10.1007/s00248-010-9657-y
Reeves RD, Adigüzel N (2008) The nickel hyperaccumulating plants of the serpentines of Turkey and adjacent areas : a review with new data. Turkish J Biol 32:143–153
Rinke C, Schwientek P, Sczyrba A, Ivanova NN, Anderson IJ, Cheng JF, Darling A, Malfatti S, Swan BK, Gies EA, Dodsworth JA, Hedlund BP, Tsiamis G, Sievert SM, Liu WT, Eisen JA, Hallam SJ, Kyrpides NC, Stepanauskas R, Rubin EM, Hugenholtz P, Woyke T (2013) Insights into the phylogeny and coding potential of microbial dark matter. Nature 499:431–437. https://doi.org/10.1038/nature12352
Rue M, Vallance J, Echevarria G, Rey P, Benizri E (2015) Phytoextraction of nickel and rhizosphere microbial communities under mono- or multispecies hyperaccumulator plant cover in a serpentine soil. Aust J Bot 63:92–102. https://doi.org/10.1071/BT14249
Saad R, Kobaissi A, Robin C, Echevarria G, Benizri E (2016) Nitrogen fixation and growth of Lens culinaris as affected by nickel availability: a pre-requisite for optimization of agromining. Environ Exp Bot 131:1–9. https://doi.org/10.1016/j.envexpbot.2016.06.010
Saad RF, Kobaissi A, Machinet G, Villemin G, Echevarria G, Benizri E (2018a) Crop rotation associating a legume and the nickel hyperaccumulator Alyssum murale improves the structure and biofunctioning of an ultramafic soil. Ecol Res 33:799–810. https://doi.org/10.1007/s11284-017-1526-4
Saad RF, Kobaissi A, Amiaud B, Ruelle J, Benizri E (2018b) Changes in physicochemical characteristics of a serpentine soil and in root architecture of a hyperaccumulating plant cropped with a legume. J Soils Sediments 18:1994–2007. https://doi.org/10.1007/s11368-017-1903-1
Saad RF, Kobaissi A, Goux X, Calusinska M, Echevarria G, Kidd P, Benizri E (2018c) Soil microbial and Ni-agronomic responses to Alyssum murale interplanted with a legume. Appl Soil Ecol 132:60–73. https://doi.org/10.1016/j.apsoil.2018.08.019
Sandaa RA, Enger O, Torsvik V (1999) Abundance and diversity of Archaea in heavy-metal-contaminated soils. Appl Environ Microbiol 65:3293–3297. https://doi.org/10.1128/AEM.65.8.3293-3297.1999
Sarrantonio M, Gallandt E (2003) The role of cover crops in North American cropping systems. J Crop Improv 8:53–74. https://doi.org/10.1300/J144v08n01_04
Sessitsch A, Kuffner M, Kidd P, Vangronsveld J, Wenzel WW, Fallmann K, Puschenreiter M (2013) The role of plant-associated bacteria in the mobilization and phytoextraction of trace elements in contaminated soils. Soil Biol Biochem 60:182–194. https://doi.org/10.1016/j.soilbio.2013.01.012
Somers E, Vanderleyden J, Srinivasan M (2004) Rhizosphere bacterial signalling: a love parade beneath our feet. Crit Rev Microbiol 30:205–235. https://doi.org/10.1080/10408410490468786
Španiel S, Kempa M, Salmerón-Sánchez E, Fuertes-Aguilar J, Mota JF, Al-Shehbaz IA, German DA, Olšavská K, Šingliarová B, Zozomová-Lihová J, Marhold K (2015) AlyBase: database of names, chromosome numbers, and ploidy levels of Alysseae (Brassicaceae), with a new generic concept of the tribe. Plant Syst Evol 301:2463–2491. https://doi.org/10.1007/s00606-015-1257-3
Staley JT, Marshall KC, Skerman VBD (1980) Budding and prosthecate bacteria from freshwater habitats of various trophic states. Microb Ecol 5:245–251. https://doi.org/10.1007/BF02020332
Sun R, Zhang XX, Guo X, Wang D, Chu H (2015) Bacterial diversity in soils subjected to long-term chemical fertilization can be more stably maintained with the addition of livestock manure than wheat straw. Soil Biol Biochem 88:9–18. https://doi.org/10.1016/j.soilbio.2015.05.007
Szumigalski A, Van Acker R (2005) Weed suppression and crop production in annual intercrops. Weed Sci 53:813–825 https://www.jstor.org/stable/4046981
Tabatabai MA, Bremner JM (1972) Assay of urease activity in soils. Soil Biol Biochem 4:479–487. https://doi.org/10.1016/0038-0717(72)90064-8
Teasdale JR, Beste CE, Potts WE (1991) Response of weeds to tillage and cover crop residue. Weed Sci 39:195–199 http://www.jstor.org/stable/4044915
Uroz S, Buée M, Murat C, Frey-Klett P, Martin F (2010) Pyrosequencing reveals a contrasted bacterial diversity between oak rhizosphere and surrounding soil. Environ Microbiol Rep 2:281–288. https://doi.org/10.1111/j.1758-2229.2009.00117.x
van der Ent A, Baker AJM, Reeves RD, Chaney RL, Anderson CWN, Meech JA, Erskine PD, Simonnot MO, Vaughan J, Morel JL, Echevarria G, Fogliani B, Rongliang Q, Mulligan DR (2015) Agromining: farming for metals in the future? Environ Sci Technol 49:4773–4780. https://doi.org/10.1021/es506031u
Visioli G, Sanangelantoni AM, Conti FD, Bonati B, Gardi C, Menta C (2018) Above and belowground biodiversity in adjacent and distinct serpentine soils. Appl Soil Ecol 133:98–103. https://doi.org/10.1016/j.apsoil.2018.09.013
Ward NL, Challacombe JF, Janssen PH, Henrissat B, Coutinho PM, Wu M, Xie G, Haft DH, Sait M, Badger J, Barabote RD, Bradley B, Brettin TS, Brinkac LM, Bruce D, Creasy T, Daugherty SC, Davidsen TM, Deboy RT, Detter JC, Dodson RJ, Durkin AS, Ganapathy A, Gwinn-Giglio M, Han CS, Khouri H, Kiss H, Kothari SP, Madupu R, Nelson KE, Nelson WC, Paulsen I, Penn K, Ren Q, Rosovitz MJ, Selengut JD, Shrivastava S, Sullivan SA, Tapia R, Thompson S, Watkins KL, Yang Q, Yu C, Zafar N, Zhou L, Kuske CR (2009) Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microbiol 75:2046–2056. https://doi.org/10.1128/AEM.02294-08
Wardle DA, Bardgett RD, Klironomos JN, Setälä H, Van Der Putten WH, Wall DH (2004) Ecological linkages between aboveground and belowground biota. Science 304:1629–1633. https://doi.org/10.1126/science.1094875
Wei ZB, Guo XF, Wu QT, Long XX, Penn CJ (2011) Phytoextraction of heavy metals from contaminated soil by co-cropping with chelator application and assessment of associated leaching risk. Int J Phytoremediat 13:717–729. https://doi.org/10.1080/15226514.2010.525554
Wrighton KC, Thomas BC, Sharon I, Miller CS, Castelle CJ, VerBerkmoes NC, Wilkins MJ, Hettich RL, Lipton MS, Williams KH, Long PE, Banfield JF (2012) Fermentation, hydrogen, and sulfur metabolism in multiple uncultivated bacterial phyla. Science 337:1661–1665. https://doi.org/10.1126/science.1224041
Wu QT, Hei L, Wong JWC, Schwartz C, Morel JL (2007) Co-cropping for phyto-separation of zinc and potassium from sewage sludge. Chemosphere 68:1954–1960. https://doi.org/10.1016/j.chemosphere.2007.02.047
Yan F, Zhu YY, Müller C, Zörb C, Schubert S (2002) Adaptation of H+-pumping and plasma membrane H+ ATPase activity in proteoid roots of white lupin under phosphorus deficiency. Plant Physiol 129:50–63. https://doi.org/10.1104/pp.010869
Yang Z, Zhang Z, Chai L, Wang Y, Liu Y, Xiao R (2016) Bioleaching remediation of heavy metal-contaminated soils using Burkholderia sp. Z–90. J Hazard Mater 301:145–152. https://doi.org/10.1016/j.jhazmat.2015.08.047
Yang Y, Wang N, Guo X, Zhang Y, Ye B (2017) Comparative analysis of bacterial com- munity structure in the rhizosphere of maize by highthroughput pyrosequencing. PLoS One 12:e0178425. https://doi.org/10.1371/journal.pone.0178425
Yasir M, Azhar EI, Khan I, Bibi F, Baabdullah R, Al-Zahrani IA, Al-Ghamdi AK (2015) Composition of soil microbiome along elevation gradients in southwestern highlands of Saudi Arabia. BMC Microbiol 15:65. https://doi.org/10.1186/s12866-015-0398-4
Youssef NH, Blainey PC, Quake SR, Elshahed MS (2011) Partial genome assembly for a candidate division OP11 single cell from an anoxic spring (Zodletone spring, Oklahoma). Appl Environ Microbiol 77:7804–7814. https://doi.org/10.1128/AEM.06059-11
Zak DR, Holmes WE, White DC, Peacock AD, Tilman D (2003) Plant diversity, soil microbial communities, and ecosystem function: are there any links? Ecology 84(8):2042–2050. https://doi.org/10.1890/02-0433
Zhang H, Sekiguchi Y, Hanada S, Hugenholtz P, Kim H, Kamagata Y, Nakamura K (2003) Gemmatimonas aurantiaca gen. Nov., sp. nov., a gram-negative, aerobic, polyphosphate-accumulating micro-organism, the first cultured representative of the new bacterial phylum Gemmatimonadetes phyl. Nov. Int J Syst Evol Microbiol 53:1155–1163. https://doi.org/10.1099/ijs.0.02520-0
Zhang Q, Zhou W, Liang G, Wang X, Sun J, He P, Li L (2015) Effects of different organic manures on the biochemical and microbial characteristics of albic paddy soil in a short term experiment. PLoS One 10:e0124096. https://doi.org/10.1371/journal.pone.0124096
Zu Y, Qin L, Zhan F, Wu J, Li Y, Chen J, Wang J, Hu WY (2017) Effects of intercropping of Sonchus asper and Vicia faba on plant cadmium accumulation and root responses. Pedosphere 30:457–465. https://doi.org/10.1016/S1002-0160(17)60484-3
Acknowledgements
We would like to acknowledge the technical team of “Laboratoire Sols et Environnement” and especially Mr. Lucas Charrois and Mr. Romain Goudon for their help and support. We are thankful for the technical assistance of the Soil Microbiology Group of the Instituto de Investigaciones Agrobiológicas de Galicia (IIAG), Consejo Superior de Investigaciones Científicas (CSIC, Santiago de Compostela, Spain). We would like to thank also Mr. Manuel Gomez for his assistance and direct work in the field. This work was supported by the French National Research Agency through the national “Investissements d’avenir” program, reference ANR-10-LABX-21 - LABEX RESSOURCES21, through the “Agromine” project, reference ANR-14-CE04-0005, through the European ERA-net FACCE_SURPLUS project “AGRONICKEL: Developing Ni agromining on ultramafic land in Europe”, reference ANR-15-SUSF-0003-05, through the “Application of Nickel hyperaccumulating plants in phytomining processes” program, reference I-LINK0900 and the LIFE Environment and Resource Efficiency Programme (Life Agromine; LIFE15 ENV/FR/000512). Finally, we would like to thank Dr. Markus Puschenreiter for proofreading and commenting this paper.
Author information
Authors and Affiliations
Contributions
Ramez F. Saad carried out the experiments and wrote the manuscript. G. Echevarria, B. Rodríguez-Garrido and P. Kidd gave intellectual input and critically revised the manuscript. E. Benizri designed the experiments, provided expertise in data analyses and supervised this study.
Corresponding author
Ethics declarations
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Responsible Editor: Juan Barcelo.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This is a posthumous publication by our dear colleague Petra Kidd
Supplementary Information
ESM 1
(DOCX 33 kb)
Rights and permissions
About this article
Cite this article
Saad, R.F., Echevarria, G., Rodríguez-Garrido, B. et al. A two-year field study of nickel-agromining using Odontarrhena chalcidica co-cropped with a legume on an ultramafic soil: temporal variation in plant biomass, nickel yields and taxonomic and bacterial functional diversity. Plant Soil 461, 471–488 (2021). https://doi.org/10.1007/s11104-021-04834-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-021-04834-y