Abstract
Background and aims
Litter decomposition and nitrogen metabolism are the key determinants of nutrient cycling in forest ecosystems. However, how processes and functional groups are affected by mixed litter decomposition is poorly understood. The aim of this research is to determine how the microbial functional pathways involved in nitrogen cycling varied according to the changes in the microbial community composition induced by mixing of litter.
Methods
The fallen leaf litters were collected on days 60, 150, 270, and 360 in Larix, Sassafras, and Larix/Sassafras plantations. The nitrogen properties, enzyme activities, microbial communities, and nitrogen metabolism pathways were evaluated during decomposition of three litter types.
Results
The pH, nitrate content, and organic nitrogen degradative enzyme activities of mixed litter were higher than those of Larix litter. Mixed litter promoted the abundances and potential nitrogen metabolism functions of Sphingomonas, and Janthinobacterium versus Larix litter during decomposition. The abundances of genes associated with microbial organic nitrogen degradation and assimilatory nitrate reduction differed significantly according to litter types. The abundance of microbial functional genes related to the production of ammonium was significantly higher in mixed litter than in Larix litter. Co-occurrence network analyses showed that mixed litter had a less complex but more stable microbial co-occurrence pattern versus monospecific litter. The difference of litter pH and nitrate content between mixed litter and Larix litter are determinants of changes in microbial functional potentials of nitrogen metabolism.
Conclusions
Mixing Sassafras/Larix litter would selectively modulate nitrogen metabolism related bacterial groups, together with functional pathways of organic nitrogen degradation and assimilatory nitrate reduction processes. These changes were mainly driven by litter pH and nitrate content.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Nitrogen is an essential component of all living organisms and the main nutrient limiting life on our planet. In forest ecosystems, litter decomposition is an important means of returning nitrogen to the soil. Nitrogen cycling is critical in coniferous plantations, the primary productivity of which is typically N-limited (Menge et al. 2012). In forest litter, organic decomposition is accompanied by mineralization of organic nitrogen (Cardenas et al. 2018). Decomposition of coniferous litter is limited by its low N content and high C:N ratio (Xiong et al. 2013). It is often assumed that the higher nutrient content of mixed litter versus single coniferous litter accelerates decomposition and increases nutrient mineralization (Olofsson and Oksanen 2002). Compared to such ‘single litter,’ mixed litter shows increased N release and improved material cycling (Zeng et al. 2018). However, the mechanism by which mixing of litter promotes N mineralization and return to soil is unclear.
The microbial community plays an essential role in the regulation of N cycling (Balser and Firestone 2005). The N cycle, by which the various types of N are transformed, is a collection of important biogeochemical pathways mediated by microbial communities (Galloway et al. 2004; Gruber and Galloway 2008), including N mineralization, immobilization, and various oxidation–reduction reactions. The associations between these connected functional pathways and related microbial communities may be revealed by analyzing co-occurrence patterns (Bissett et al. 2013; Cardona et al. 2016), which also enables the identification of the core microbiome involved in N metabolism. However, the relationships among the litter N properties, microbial community composition, and N-metabolism pathways are unclear (Remy et al. 2018).
Cultivation of high-quality conifer–broadleaf mixed plantations impacts microbial community function and may influence litter decomposition, nutrient release, and tree growth. In forest ecosystems, the chemical characteristics of litter, such as the carbon/nitrogen ratio (C/N) and/or nitrogen content, are used as indicators of litter quality (Aerts 1997; Strickland et al. 2009). N release from leaf litter is mainly driven by the initial tissue N concentration. There is a positive correlation between the litter decomposition rate and N content and a negative relationship between the litter decomposition rate and the initial litter C/N (Aerts et al. 2003; Semmartin et al. 2004; Garibaldi et al. 2007; Vaieretti et al. 2013). Variations in N availability and C:N ratio due to mixing of litter influence the bacterial and fungal communities (Gartner and Cardon 2004; Hättenschwiler et al. 2005; Leff et al. 2015; Pereira et al. 2019). Compared with monospecific litter, mixed litter could alter the nitrogen properties, microbial community structure and amino acid metabolism potential (Wang et al. 2019). Notably, the contents of nitrate and ammonium have significant effects on the characteristics of nitrogen metabolism and the activities of chitinase and protease in microbial communities (Geisseler et al. 2010; Petersen et al. 2012; Mooshammer et al. 2014). Litter pH was also important in litter decomposition, with accelerated litter mass loss and enhanced urease activity at high pH value (Wang et al. 2010; Lv et al. 2014). However, it is unclear how the microbial functional pathways involved in nitrogen cycling vary according to the changes in the microbial community composition induced by mixing of litter.
Microbial-mediated nitrogen metabolism has typically been studied based on analyses of functional genes using specific primers (Tang et al. 2018; Pereira et al. 2019). Microarray technologies such as GeoChip (Tu et al. 2014) are used to analyze N-metabolism pathways in a variety of environments, including forest ecosystems (Bai et al. 2013), grassland ecosystems (He et al. 2010), and acid mine drainage (Xie et al. 2011). Metagenomics approaches with genome shotgun sequencing allow assessment of the diversity of functional pathways governing key soil processes such as C-N metabolism, particularly for novel gene families not targeted by microbial ecological microarrays and/or those for which primers are not available (Zhou et al. 2015). Previous metagenomics studies have investigated nitrogen metabolism in soil ecosystems (Tu et al. 2017; Sun and Badgley 2019), however, the effects of litter mixing on the function microbial groups and N metabolism pathways are unclear.
In this study, we surveyed the major N-metabolism gene families, function microbial groups, and their responses to mixing of litter at the Changling Ridge Forest experimental site located in Jianshi County, Hubei Province, China. A total of 36 litter samples was collected from three types of plantations in 2017. Metagenome sequencing was used to survey the N-metabolism gene families important for litter decomposition. We hypothesized that multiple N-metabolism processes would be stimulated by the changed litter pH and litter nitrogen properties during the decomposition of mixed litter. In addition to characterizing the responses of bacterial and fungal N-metabolism pathways to mixing of litter, we assessed the abundances of key gene families in N metabolism as well as their taxonomic composition.
Materials and methods
Site description and sample collection
The study area was located at 109°32′–110°12′ E and 30°06′–30°54’ N at Changlinggang Forest Farm in Jianshi County, Hubei Province, China. The study area has an average elevation of 1700 m, a subtropical monsoon climate, an annual average temperature of 9 °C, a minimum temperature of −10 °C, an average maximum temperature of 22.2 °C (July), a frost-free period of about 230–290 days, and a relative humidity of 75–85%. The average annual rainfall is 1500–1800 mm. The soil is yellowish brown with a thickness of more than 1 m and a pH of 5–6 (Wang et al. 2019).
After performing a comprehensive survey of Larix kaempferi (Lamb.) Carr., Sassafras tzumu (Hemsl.) Hemsl., and Larix kaempferi/Sassafras tzumu plantations (the proportions between Sassafras and Larix were 1:1), nine independent sample plots with similar site conditions were selected. The experimental design has been described in detail by Wang et al. (2019). Larix and Sassafras are deciduous species, and their deciduous period occurs around November (Gower and Richards 1990; Song et al. 2011). There were three representative sample plots (0.06 ha each) for each plantation type, and each plot was sampled on five occasions.
We collected litter only from the sample plots and each litter sample consisted of a pool of five independent subsamples, located according to the S-shaped sampling method (Du et al. 2019). We collected fresh litter in November 2016 when major leaf pools were supplied to the soil and measured their initial nutrient concentrations. During the course of litter decomposition, we sampled the litter L-horizon on four additional occasions, at days 60 (winter), 150 (spring), 270 (summer), and 360 (autumn) of 2017 (Liu et al. 2016; Schneider et al. 2012). Due to the low microbial abundance in fresh litter, only 36 samples undergoing decomposition were selected for metagenomic sequencing. About 20 g samples were taken at each plot in each sampling date. On each sampling date, 50 g aliquots of cut leaf material from three biological replicates per sampling site were transported in a dry-ice box to the laboratory. One was frozen and stored at −80 °C for DNA extraction and metagenomics sequencing. The second was stored in a 4 °C refrigerator for determination of enzyme activities within 1 month. The third was dried at 65 °C to constant weight for determination of litter nitrogen properties.
Litter nitrogen properties and enzyme activity assays
Total C, N concentrations and pH of litter were determined as previously described (Wang et al. 2019). Nitrate-N (NO3−-N) and ammonium-N (NH4+-N) were extracted using 2 M KCl (Prescott et al. 2003), and analyzed on a continuous-flow ion auto-analyzer (Scalar SANplus Segmented Flow Analyzer, The Netherlands).
Urease activity was determined using urea (10% w/v) as the substrate by incubating the litter sample for 24 h at 37 °C and measuring the NH3 released colorimetrically at 578 nm (Hu et al. 2006). Protease activity was determined by extracting the released aromatic amino acids with trichloroacetic acid (0.92 M) followed by colorimetric assay using Folin–Ciocalteu reagent (Kandeler 1999). The results are expressed as μmol Tyr (tyrosine equivalents) g−1·h−1. Chitinase activity was determined using N-acetyl-β-D-glucosaminide as the substrate and measuring the absorbance of the reaction product, nitrophenol, at 410 nm (Kourtev et al. 2002).
DNA extraction, library construction, and metagenomic sequencing
The total DNA was extracted from the samples (10 g aliquots of cut leaf material from each plot) and used for the metagenome sequencing and further analyses. DNA was extracted from the litter samples using the EZNA® DNA Kit (Omega Bio-tek, Norcross, GA) according to the manufacturer’s protocol. DNA concentration, purity, and integrity were detected by NanoDrop2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA), TBS-380 fluorometer (Turner Biosystems, Sunnyvale, CA) and 1% agarose gel electrophoresis, respectively.
DNA was fragmented to an average size of about 300 bp using Covaris M220 (Gene Company, Shanghai, China) for paired-end library construction. Paired-end libraries were prepared using the TruSeq DNA Sample Prep Kit (Illumina, San Diego, CA). Adapters containing the full complement of sequencing primer hybridization sites were ligated to the blunt-end fragments. Paired-end sequencing was performed on the Illumina HiSeq 4000 platform (Illumina) at Majorbio Bio-Pharm Technology Co. (Shanghai, China) using the HiSeq 3000/4000 PE Cluster Kit and HiSeq 3000/4000 SBS Kit according to the manufacturer’s instructions. All the raw metagenomic datasets in this study are publicly available in the NCBI Sequence Read Achieve (SRA) database with an accession number SRP242499.
Sequence quality control and assembly
The raw reads were subject to the following treatments: the 3′-end and 5′-end of reads were trimmed to the first high-quality base using Seqprep (https://github.com/jstjohn/SeqPrep). To retain high-quality pair-end reads and single-end reads, the reads of sequence length < 50 bp and quality value <20 were excised with Sickle (https://github.com/najoshi/sickle). De Bruijn-graph-based assembler SOAPdenovo (http://soap.genomics.org.cn, version 1.06) was employed to assemble short reads. K-mers, varying from one- to two-thirds of the read length were tested for each sample. Scaffolds with length > 500 bp were retained for statistical testing; we evaluated the quality and quantity of scaffolds generated by each assembly and selected the K-mer that yielded the minimum scaffold number and the maximum N50 and N90 values. Next, scaffolds of >500 bp were extracted and broken into contigs without gaps. The contigs were used for gene prediction and annotation.
Gene prediction, taxonomy, and functional annotation
MetaGene (http://metagene.cb.k.u-tokyo.ac.jp) was used for open reading frame (ORF) prediction. The predicted ORFs with lengths >100 bp were retrieved and translated into amino acid sequences using the National Center for Biotechnology Information (NCBI) translation table (www.ncbi.nlm.nih.gov/Taxonomy/taxonomyhome.html/index.cgi?chapter = tgencodes#SG1). Removing redundancy was done with CD-HIT (www.bioinformatics.org/cd-hit) using bacterial and fungal redundant coding sequence catalogues. Sequences with 95% sequence identity (90% coverage) were considered redundant (Temu et al. 2016). Reads after quality control were mapped to representative genes with 95% identity using SOAPaligner (http://soap.genomics.org.cn), and gene abundances were evaluated. The resulting scaffolds were cut into contigs and only contigs longer than 500 bp were saved for gene prediction. BLASTP (version 2.2.28+, http://blast.ncbi.nlm.nih.gov/Blast.cgi) was used for taxonomic annotations by aligning non-redundant gene catalogs against the NCBI NR database with an e-value cutoff of 1e−5. The species abundance was calculated using the total gene abundance across all species, and the abundance profile was built based on taxonomic levels. The relative abundance was calculated by the ratio of individual specie abundance to total species abundance in each sample. KEGG pathway annotation was conducted by BLASTP search (version 2.2.28+) against the Kyoto Encyclopedia of Genes and Genomes database (www.genome.jp/keeg) with an e-value cutoff of 1e−5.
Nitrogen metabolism functional taxa and pathway reconstruction
The NR gene catalog was aligned against the Kyoto Encyclopedia of Genes and Genomes (KEGG) database (Kanehisa et al. 2004) by BLAST (Version 2.2.28+) and assigned KEGG functional annotation by KOBAAS 2.0 (Qin et al. 2010). Nitrogen pathway was reconstructed based on the annotation results of KEGG pathway ko00910 and relevant references (Tu et al. 2017).
Statistical analyses
SPSS 20.0 software was used for statistical analyses. Repeated measures analysis of variance (RMANOVA) was conducted to evaluate the significance of differences in litter nitrogen properties, enzyme activity, microbial abundance, and gene abundance according to plantation type and sampling time (Zhang et al. 2019b). The two-subject factors in the analyses were “plantation type” (Larix plantation, Sassafras plantation, and mixed plantation) and “sampling time” (60, 150, 270, and 360 days). The assumptions of statistical analyses were checked before analysis. Values were transformed to the log-scale if data did not conform to assumptions of normality. If main-effect differences were significant, pairwise post hoc comparisons of subgroup means were conducted using the least significant difference (LSD) procedure. If the interactions were significant, differences in the effects of forest type and sampling season were subjected to one-way ANOVA. Line charts were created using Origin version 9.1 (Originab, Northampton, MA). The relative abundance of functional genes was visualized as heatmaps using the R package ‘pheatmap’ (Sun and Badgley 2019). A co-occurrence network was constructed using 12 samples of each type of litter. Co-occurrence network analyses of the bacterial and fungal communities were conducted using the Python package ‘networkx’ (Mandakovic et al. 2018). To minimize pairwise comparisons and reduce network complexity, only dominant genera with relative abundance >3% were selected for network analyses (Huang et al. 2018). The most important interactions with strong positive (r > 0.7) and strong negative (r < −0.7) relationships are shown in network diagrams. The species with a high mean degree, high closeness centrality, and high betweenness centrality were collectively used to identify keystone taxa as described previously (Banerjee et al. 2018). Redundancy analyses (RDAs) and distance-based RDA (db-RDA) were performed to investigate the relationships among microbial genera, enzyme activity, functional genes, and litter nitrogen properties using the ‘vegan’ package in R (Xu et al. 2018). A p value <0.05 was considered to indicate statistical significance.
Results
Effect of mixing litter on litter nitrogen properties and enzyme activities
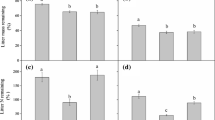
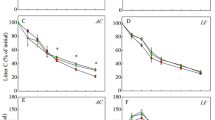
RMANOVA showed that litter C/N, nitrate and ammonium content, protease, chitinase, and urease activity were significantly different according to plantation type and sampling time (p < 0.05). In the initial stage of litter decomposition, C/N was in the order Larix litter > mixed litter > Sassafras litter (Fig. 1a). During litter decomposition, the C/N of Sassafras litter was lower than that of Larix litter before 270 days, whereas that of mixed litter was lower than that of Larix litter only at 150 and 270 days.
The ammonium content gradually decreased during Larix litter decomposition but increased in Sassafras litter and mixed litter after 150 days (Fig. 1b). There were significant differences among the three types of litter after 150 days (p < 0.05). Nitrate content generally decreased during litter decomposition, with significant differences in among the three types of litter before 270 days (p < 0.05) (Fig. 1c).
The highest urease activity was found in mixed litter, followed by Sassafras and Larix (Fig. 1d). The urease activity in mixed litter was significantly higher than in Larix litter during decomposition (p < 0.05). Significant differences in protease and chitinase activity among the three types of litter were evident before 270 days (p < 0.05) (Fig. 1e and f), and were significantly higher in mixed litter and Sassafras litter than in Larix litter (p < 0.05). Protease and chitinase activity were highest in Sassafras litter and peaked at 150 and 270 days of decomposition, respectively.
The nitrogen metabolism pathways in litter decomposition
We reconstructed the nitrogen metabolism pathways in litter decomposition based on the results of metagenomic KEGG nitrogen metabolism annotations (Fig. 2). Nitrogen metabolism involves nitrification, denitrification, dissimilatory nitrate reduction, assimilatory nitrate reduction, nitrogen fixation, and organic nitrogen degradation. To investigate changes in microbial nitrogen metabolism with litter decomposition, we analyzed the microbial taxa and genes involved in nitrogen metabolism. The results showed that nitrogen metabolism functional genes mainly came from Rhizobiales, Burkholderiales, and Sphingomonadales. The proportions of Rhizobiales involved in Organic N degradation, nitrogen fixation, assimilatory nitrate reduction, and dissimilatory nitrate reduction in Larix litter were higher than in mixed litter. While the proportions of Burkholderiales involved in assimilatory nitrate reduction, and dissimilatory nitrate reduction were higher in mixed litter than in Larix litter. The proportion of Sphingomonadales involved in Organic N degradation was higher in mixed litter than in Larix litter.
Variation in functional microbial communities and genes related to litter decomposition
To investigate changes in microbial nitrogen metabolism, we analyzed the top 10 relative abundances of taxa and functional genes related to nitrogen transformation. The results showed that Rhizobiales, Burkholderiales, and Sphingomonadales were the predominant nitrogen metabolism-related taxa during litter decomposition, irrespective of litter type (Fig. 3). With the decomposition of three types litter, the relative abundance of Burkholderiales and Sphingomonadales were decreasing over time. Notably, the relative abundance of Burkholderiales and Sphingomonadales was significantly higher in mixed litter than Larix litter before 270 days (p < 0.05). The relative abundance of Burkholderiales was significantly higher in Sassafras litter than Larix litter during decomposition (p < 0.05). While the average abundance of Rhizobiales was lower in Sassafras litter versus Larix litter during decomposition. Specially, the relative abundance of Rhizobiales was significantly lower in Sassafras litter versus Larix litter at 270 days (p < 0.05). At the genus level, Bradyrhizobium (Order Rhizobiales) was the most abundant genus during Larix litter decomposition (Fig. 4). However, the most abundant bacterial genera changed from Sphingomonas (Order Sphingomonadales) to Bradyrhizobium over time in Sassafras and mixed litter. The relative abundance of Sphingomonas was significantly higher in mixed litter than in Larix litter before 270 days, while that of Bradyrhizobium showed the opposite pattern (p < 0.05). The relative abundances of Janthinobacterium (Order Burkholderiales) were significantly higher in mixed litter than in Larix litter before 270 days (p < 0.05). Specially, Janthinobacterium and Sphingomonas abundance decreased over time, while Bradyrhizobium became more abundant.
The RMANOVA results showed that the abundances of genes associated with bacterial organic nitrogen degradation, assimilatory nitrate reduction, denitrification, and nitrification significantly differed by plantation type (p < 0.05) (Fig. 5). The abundances of genes associated with bacterial organic nitrogen degradation, dissimilatory nitrate reduction, denitrification, and nitrification significantly changed over time (p < 0.05). Notably, the abundance of bacterial organic nitrogen degradation genes was significantly higher in mixed litter than in Larix litter before 270 days of decomposition, while the genes related to assimilatory nitrate reduction showed the opposite pattern (p < 0.05). In fungi, the abundances of organic nitrogen degradation and assimilatory nitrate reduction genes significantly differed by plantation type (p < 0.05). Only the abundance of fungal denitrification genes was significantly affected by sampling time (p < 0.01). The abundance of organic nitrogen degradation genes was significantly lower in mixed litter than in Larix litter at 270 days of litter decomposition, while that of assimilatory nitrate reduction genes showed the opposite pattern (p < 0.05).
Among the functional genes in nitrogen metabolism, glutamate dehydrogenase (EC: 1.4.1.2), nitronate monooxygenase (EC: 1.13.12.16), and urease subunit alpha (EC: 3.5.1.5) were the most abundant in the bacterial community (Fig. 6a). Difference analyses showed that mixing of litter significantly increased the abundance of glutamate dehydrogenase (EC: 1.4.1.2) before 270 days of litter decomposition, but significantly decreased the abundances of ferredoxin-nitrate reductase (EC: 1.7.7.2) and ferredoxin-nitrite reductase (EC: 1.7.7.1) versus Larix litter before 150 and 270 days of litter decomposition (p < 0.05).
In the fungal community, glutamate dehydrogenase (EC: 1.4.1.2), nitrite reductase (NAD(P)H) (EC:1.7.1.4), and nitrate reductase (NAD(P)H) (EC:1.7.1.1; 1.7.1.2; 1.7.1.3) were the most abundant genes (Fig. 6b). The abundances of nitrate reductase (NAD(P)H) (EC: 1.7.1.1; 1.7.1.2; 1.7.1.3) were significantly higher in mixed litter than in Larix litter before 270 days of litter decomposition (p < 0.05). The abundance of glutamate dehydrogenase (EC: 1.4.1.2) was significantly lower in mixed litter than in Larix litter before 270 days of decomposition (p < 0.05). Collectively, the abundance of bacterial and fungal genes involved in organic nitrogen degradation and assimilatory nitrate reduction was different in mixed litter than in Larix litter.
Network analyses of microbial communities in the three types of litter
The overall networks consisted of 59 nodes with an average number of neighbors of 7.14 and a clustering coefficient of 0.65 (Fig. 7). There were significant differences among the three types of litter in terms of the bacterial and fungal communities. The network pattern of mixed litter contained more fungal classes than that of Larix litter. However, the network structure was less complex in mixed litter than in monospecific litter. And there were fewer negative linear correlated edges among microbial classes in mixed litter than in monospecific litter. Unlike Larix litter, the positive links occurred between bacterial and fungal classes in Sassafras litter and mixed litter (Fig. 7a). The abundance of the dominant genus Bradyrhizobium was positively correlated with those of Mycobacterium, Streptomyces, and Frankia in Larix litter. The abundances of the dominant genera Bradyrhizobium and Sphingomonas were negatively correlated in Sassafras litter and mixed litter (Fig. 7b and c). Unlike Bradyrhizobium, the abundance of Sphingomonas was positively correlated with that of most other genera. Network analyses showed that Mycobacterium in Larix litter was the most influential (keystone) taxon during litter decomposition. Sphingomonas was the most influential (keystone) taxon in mixed litter and Sassafras litter.
Network analyses of bacterial and fungal communities in (a)Larixlitter, (b)Sassafraslitter, and (c) mixed litter. The size of nodes represents the abundance of genera. The color of the line indicates positive and negative correlation, red: negative correlation between genera, green: positive correlation between genera
Relationships among microbial community composition, nitrogen metabolism genes, and litter nitrogen properties
The RDA results showed that the bacterial community composition was significantly affected by pH, nitrate, C/N, and protease (R2 = 0.77, p < 0.01; R2 = 0.55, p < 0.01; R2 = 0.37, p < 0.01; R2 = 0.33, p < 0.05) (Fig. 8a). Unlike Bradyrhizobium, the abundances of Sphingomonas, and Janthinobacterium were positively correlated with the pH, protease activity, and nitrate content. The fungal community was significantly affected by pH, nitrate, and protease and urease activity (R2 = 0.62, p < 0.01; R2 = 0.48, p < 0.01; R2 = 0.53, p < 0.01; R2 = 0.34, p < 0.05). The abundances of most fungal genera were positively correlated with pH, nitrate, and the protease and urease activity, and Pestalotiopsis showed the strongest such correlations (Fig. 8b).
RDA analyses of bacterial community (a), fungal community (b) composition, bacterial (c), and fungal (d) nitrogen metabolism genes in relation to litter nitrogen properties. The abbreviations of nitrogen metabolism functional genes number were presented in Table 1
The db-RDA results showed that the abundances of bacterial nitrogen metabolism genes were significantly affected by nitrate, pH, and urease activity (R2 = 0.40, p < 0.01; R2 = 0.29, p < 0.01; R2 = 0.28, p < 0.01, respectively) (Fig. 8c). The abundances of nitrite reductase (NADH) large subunit and glutamate dehydrogenase were positively correlated with nitrate content and pH, while those of nitronate monooxygenase and ferredoxin-nitrite reductase were negatively correlated with nitrate content and pH. The abundances of fungal nitrogen metabolism genes were significantly affected by pH, nitrate content, and protease activity (R2 = 0.40, p < 0.01; R2 = 0.22, p < 0.05; R2 = 0.23, p < 0.05) (Fig. 8d). The abundances of nitrate reductase (NAD(P)H), nitrite reductase (NAD(P)H), and glutamate dehydrogenase were positively correlated with pH and nitrate content.
Discussion
Nitrogen metabolism during litter decomposition is a complex process involving diverse microbial species. We focused on the known major N-metabolism processes mediated by microbial communities, namely, nitrification, denitrification, dissimilatory nitrate reduction, assimilatory nitrate reduction, nitrogen fixation, and organic nitrogen degradation. However, we did not detect anaerobic ammonium oxidation (anammox) during litter decomposition. This may be because anammox occurs in anaerobic environments, and the litter layer was not suitable for the growth and propagation of anaerobic microorganisms (Humbert et al. 2010). Although denitrification typically occurs in aerobic environments, it can also take place in anaerobic environments (Herbert 1999), as some nitrate-reducing bacteria are facultative anaerobes that possess the enzymes required for both aerobic and anaerobic respiration.
Our data indicate that each N-metabolism process was carried out by many microbial species during litter decomposition, in agreement with the highly diverse phylogenetic distribution of N-metabolism processes (Nelson et al. 2016). High microbial diversity is required to maintain active N metabolism during litter decomposition due to the limited functional redundancy of the microbial community (Philippot et al. 2013). Of the six major processes analyzed, the genes related to nitrate reduction (both assimilatory and dissimilatory) and organic nitrogen degradation showed the highest abundance; moreover, these two processes involved the greatest diversity of microbial taxa. This is likely because nitrate reduction and organic nitrogen degradation provide microorganisms with energy and nutrients (Condron et al. 2010; Moreno-Vivián et al. 1999). Compared to the other nitrogen metabolism processes, fewer taxonomic groups were related to nitrogen fixation.
In our results, Rhizobiales, Burkholderiales, and Sphingomonadales were the three major orders responsible for nitrogen metabolism processes. These taxa were reportedly the dominant species in nitrogen cycling and appeared to be N cycling generalists in that they carried genes from nitrogen metabolism pathways (Nelson et al. 2015). At the genus level, Sphingomonas members were noted for containing abundant denitrifying functional genes (Fang et al. 2018). Janthinobacterium was associated with nitrogen cycling and could utilize some amino acids (Lowell et al. 2009; Summers et al. 2013). Our results showed that the organic N degradation and nitrate reduction functional genes mainly came from Burkholderiales, and Sphingomonadales. The relative abundance of Burkholderiales and Sphingomonadales was significantly higher in mixed litter than Larix litter before 270 days. For more details, the abundances of Sphingomonas (Sphingomonasales Order), and Janthinobacterium (Burkholderiales Order) were significantly higher in mixed litter than in Larix litter, indicating that mixing of litter promoted functional potentials of nitrogen metabolism by these taxa.
Due to redundancy of microbial community functions, changes in gene abundance may be less obvious than changes in community composition (Philippot et al. 2013). The abundances of genes associated with bacterial organic nitrogen degradation, assimilatory nitrate reduction, denitrification, and nitrification significantly differed by plantation type. However, the differences in the abundances of fungal nitrogen metabolism genes according to litter type were less marked. Bacterial community composition and functional gene abundance were significantly affected by N content, while the fungal community composition was modulated by C content (Wang et al. 2019). It is possible that, compared to fungi, bacteria have a lower C/N requirement and are more sensitive to C/N changes (Keiblinger et al. 2010; Riggs and Hobbie 2016).
Microbial co-occurrence patterns enable evaluation of microbial community functions by unraveling the interactions between microbes and revealing the niche structure (Ma et al. 2018; Banerjee et al. 2018). Promotion of microbe-mediated organic degradation provides nitrogen and niches for microbial taxa, which alleviates competition and increases cooperation between microbes (Costello et al. 2012; Lin et al. 2019). The network pattern of mixed litter contained more fungal classes and fewer bacterial classes than that of Larix litter. The fewer negative linear correlated edges and less complex network structure indicated a more stable microbial co-occurrence pattern in mixed litter than in monospecific litter (Fan et al. 2018; Zhang et al. 2019a). The abundance of bacterial functional genes related to the production of ammonium was greater in mixed litter versus Larix litter. In particular, the abundance of glutamate dehydrogenase genes (related to organic nitrogen degradation) increased before 270 days of decomposition. Glutamate and ammonium were the key nitrogen donors of almost all cell biosynthesis reactions, and they were also used as the preferred nitrogen source by microbial community (Magasanik 1993; Wong et al. 2008). Degradation of organic nitrogen was the main way for microorganisms to obtain nitrogen during litter decomposition. The dominant genera Sphingomonas and Janthinobacterium were found to be rich in nitrogen metabolism genes, and could produce protease and chitinase (Byun and Blinkovsky 2004; Xiao et al. 2005; Sabri et al. 2018). Our RDA results also showed that the abundances of Sphingomonas and Janthinobacterium had positive correlations with the activities of protease and chitinase. Meanwhile, higher activities of organic nitrogen decomposing enzymes (urease, protease, and chitinase) were observed in mixed litter than Larix litter. In coniferous litter where available N and C were low, it was crucial for microbial community to regulate the activities of urease, protease, and chitinase (Geisseler et al. 2010). Therefore, the high relative abundance of the genera Sphingomonas and Janthinobacterium can reasonably explain the high organic nitrogen decomposing enzymes activities in mixed litter. The abundance of fungal functional genes involved in assimilatory nitrate reduction was higher in mixed litter versus Larix litter. For more details, the abundances of nitrate reductase (NAD(P)H) in mixed litter were higher before 270 days of decomposition. In summary, the mixed litter trended to provide more biologically available ammonium for soil N cycling during its decomposition.
Compared to Larix litter, mixed litter had a higher pH and nitrate content, which are major determinants of microbial community composition and abundance (Balser and Firestone 2005; Liu et al. 2015; Urbanová et al. 2015). Unlike Bradyrhizobium, Sphingomonas, and Janthinobacterium were positively correlated with pH and nitrate content, as reported by others (Zhalnina et al. 2015). Sphingomonas can use complex substrates and was the predominant members of the litter bacterial community during decomposition (Zeng et al. 2019). Hence, the higher pH and nitrate content of mixed litter than in Larix litter is responsible for the increased abundances of Sphingomonas, and Janthinobacterium. In contrast to Sphingomonas, the abundances of Bradyrhizobium were positively correlated with C/N, possibly due to nitrogen fixation by Bradyrhizobium (Bedmar et al. 2005), which can survive in the low-nitrogen conditions of Larix litter. The key functional genes in N cycling were highly enriched by nitrate injection (Xu et al. 2014), possibly leading to functional changes (Bowen et al. 2011). This supports our finding that higher nitrate contents were positively correlated with the abundances of Sphingomonas, Janthinobacterium, and key functional genes in nitrogen metabolism.
Conclusion
We investigated the changes in the nitrogen metabolism related microbial groups composition and functional genes during decomposition of three litter types. Our findings provide insight into how mixing of litter affects bacterial and fungal nitrogen metabolism pathways during litter decomposition. The study found that mixed litter mainly affected the abundances of genes associated with bacterial organic nitrogen degradation and assimilatory nitrate reduction. The nitrogen metabolism functional potentials of Sphingomonas, and Janthinobacterium were promoted in mixed litter versus Larix litter during decomposition. Co-occurrence network analyses showed that mixed litter had a less complex but more stable microbial pattern versus monospecific litter. The differences of microbial nitrogen metabolism potential functions among litter types were mainly driven by the litter pH and nitrate content.
References
Aerts R (1997) Climate, leaf litter chemistry and leaf litter decomposition in terrestrial ecosystems: a triangular relationship. Oikos 79:439. https://doi.org/10.2307/3546886
Aerts R, De Caluwe H, Beltman B (2003) Plant community mediated vs. nutritional controls on litter decomposition rates in grasslands. Ecology 84:3198–3208. https://doi.org/10.1890/02-0712
Bai S, Li J, He Z et al (2013) GeoChip-based analysis of the functional gene diversity and metabolic potential of soil microbial communities of mangroves. Appl Microbiol Biotechnol 97:7035–7048. https://doi.org/10.1007/s00253-012-4496-z
Balser TC, Firestone MK (2005) Linking microbial community composition and soil processes in a California annual grassland and mixed-conifer forest. Biogeochemistry 73:395–415. https://doi.org/10.1007/s10533-004-0372-y
Banerjee S, Schlaeppi K, van der Heijden MGA (2018) Keystone taxa as drivers of microbiome structure and functioning. Nat Rev Microbiol 16:567–576. https://doi.org/10.1038/s41579-018-0024-1
Bedmar EJ, Robles EF, Delgado MJ (2005) The complete denitrification pathway of the symbiotic, nitrogen-fixing bacterium Bradyrhizobium japonicum. Biochem Soc Trans 33:141–144. https://doi.org/10.1042/BST0330141
Bissett A, Brown MV, Siciliano SD, Thrall PH (2013) Microbial community responses to anthropogenically induced environmental change: towards a systems approach. Ecol Lett 16:128–139. https://doi.org/10.1111/ele.12109
Bowen JL, Ward BB, Morrison HG, Hobbie JE, Valiela I, Deegan LA, Sogin ML (2011) Microbial community composition in sediments resists perturbation by nutrient enrichment. ISME J 5:1540–1548. https://doi.org/10.1038/ismej.2011.22
Byun T, Blinkovsky A (2004) Glycyl aminopeptidase (Sphingomonas). Handbook of Proteolytic Enzymes. Elsevier, In, pp 470–471
Cardenas E, Orellana LH, Konstantinidis KT, Mohn WW (2018) Effects of timber harvesting on the genetic potential for carbon and nitrogen cycling in five north American forest ecozones. Sci Rep 8:3142. https://doi.org/10.1038/s41598-018-21197-0
Cardona C, Weisenhorn P, Henry C, Gilbert JA (2016) Network-based metabolic analysis and microbial community modeling. Curr Opin Microbiol 31:124–131. https://doi.org/10.1016/j.mib.2016.03.008
Condron L, Stark C, O’Callaghan M et al (2010) The role of microbial communities in the formation and decomposition of soil organic matter. In: Soil microbiology and sustainable crop production. Springer Netherlands, Dordrecht, pp 81–118
Costello EK, Stagaman K, Dethlefsen L et al (2012) The application of ecological theory toward an understanding of the human microbiome. Science 336(80):1255–1262. https://doi.org/10.1126/science.1224203
Du J, Niu J, Gao Z et al (2019) Catena E ff ects of rainfall intensity and slope on interception and precipitation partitioning by forest litter layer. Catena 172:711–718. https://doi.org/10.1016/j.catena.2018.09.036
Fan K, Weisenhorn P, Gilbert JA, Chu H (2018) Wheat rhizosphere harbors a less complex and more stable microbial co-occurrence pattern than bulk soil. Soil Biol Biochem 125:251–260. https://doi.org/10.1016/j.soilbio.2018.07.022
Fang W, Yan D, Wang X, Huang B, Wang X, Liu J, Liu X, Li Y, Ouyang C, Wang Q, Cao A (2018) Responses of nitrogen-cycling microorganisms to Dazomet fumigation. Front Microbiol 9:2529. https://doi.org/10.3389/fmicb.2018.02529
Galloway JN, Dentener FJ, Capone DG et al (2004) Nitrogen cycles: past, present, and future. Biogeochemistry 70:153–226. https://doi.org/10.1007/s10533-004-0370-0
Garibaldi LA, Semmartin M, Chaneton EJ (2007) Grazing-induced changes in plant composition affect litter quality and nutrient cycling in flooding Pampa grasslands. Oecologia 151:650–662. https://doi.org/10.1007/s00442-006-0615-9
Gartner TB, Cardon ZG (2004) Decomposition dynamics in mixed-species leaf litter. Oikos 104:230–246. https://doi.org/10.1111/j.0030-1299.2004.12738.x
Geisseler D, Horwath WR, Joergensen RG et al (2010) Pathways of nitrogen utilization by soil microorganisms – a review. Soil Biol Biochem 42:2058–2067. https://doi.org/10.1016/j.soilbio.2010.08.021
Gower ST, Richards JH (1990) Larixes: deciduous conifers in an Evergreen world. Bioscience 40:818–826. https://doi.org/10.2307/1311484
Gruber N, Galloway JN (2008) An earth-system perspective of the global nitrogen cycle. Nature 451:293–296. https://doi.org/10.1038/nature06592
Hättenschwiler S, Tiunov AV, Scheu S (2005) Biodiversity and litter decomposition in terrestrial ecosystems. Annu Rev Ecol Evol Syst 36:191–218. https://doi.org/10.1146/annurev.ecolsys.36.112904.151932
He Z, Xu M, Deng Y, Kang S, Kellogg L, Wu L, van Nostrand J, Hobbie SE, Reich PB, Zhou J (2010) Metagenomic analysis reveals a marked divergence in the structure of belowground microbial communities at elevated CO2. Ecol Lett 13:564–575. https://doi.org/10.1111/j.1461-0248.2010.01453.x
Herbert RA (1999) Nitrogen cycling in coastal marine ecosystems. FEMS Microbiol Rev 23:563–590. https://doi.org/10.1111/j.1574-6976.1999.tb00414.x
Hu YL, Wang SL, Zeng DH (2006) Effects of single Chinese fir and mixed leaf litters on soil chemical, microbial properties and soil enzyme activities. Plant Soil 282:379–386. https://doi.org/10.1007/s11104-006-0004-5
Huang X, Dong W, Wang H, Feng Y (2018) Role of acid/alkali-treatment in primary sludge anaerobic fermentation: insights into microbial community structure, functional shifts and metabolic output by high-throughput sequencing. Bioresour Technol 249:943–952. https://doi.org/10.1016/j.biortech.2017.10.104
Humbert S, Tarnawski S, Fromin N, Mallet MP, Aragno M, Zopfi J (2010) Molecular detection of anammox bacteria in terrestrial ecosystems: distribution and diversity. ISME J 4:450–454. https://doi.org/10.1038/ismej.2009.125
Kandeler E (1999) Xylanase, invertase and protease at the soil–litter interface of a loamy sand. Soil Biol Biochem 31:1171–1179. https://doi.org/10.1016/S0038-0717(99)00035-8
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:277–280. https://doi.org/10.1093/nar/gkh063
Keiblinger KM, Hall EK, Wanek W, Szukics U, Hämmerle I, Ellersdorfer G, Böck S, Strauss J, Sterflinger K, Richter A, Zechmeister-Boltenstern S (2010) The effect of resource quantity and resource stoichiometry on microbial carbon-use-efficiency. FEMS Microbiol Ecol 73:430–440. https://doi.org/10.1111/j.1574-6941.2010.00912.x
Kourtev P, Ehrenfeld J, Huang W (2002) Enzyme activities during litter decomposition of two exotic and two native plant species in hardwood forests of New Jersey. Soil Biol Biochem 34:1207–1218. https://doi.org/10.1016/S0038-0717(02)00057-3
Leff JW, Jones SE, Prober SM, Barberán A, Borer ET, Firn JL, Harpole WS, Hobbie SE, Hofmockel KS, Knops JM, McCulley R, la Pierre K, Risch AC, Seabloom EW, Schütz M, Steenbock C, Stevens CJ, Fierer N (2015) Consistent responses of soil microbial communities to elevated nutrient inputs in grasslands across the globe. Proc Natl Acad Sci 112:10967–10972. https://doi.org/10.1073/pnas.1508382112
Lin Y, Ye G, Kuzyakov Y et al (2019) Long-term manure application increases soil organic matter and aggregation, and alters microbial community structure and keystone taxa. Soil Biol Biochem 134:187–196. https://doi.org/10.1016/j.soilbio.2019.03.030
Liu S, Ren H, Shen L et al (2015) pH levels drive bacterial community structure in sediments of the Qiantang River as determined by 454 pyrosequencing. Front Microbiol 6. https://doi.org/10.3389/fmicb.2015.00285
Liu D, Keiblinger KM, Leitner S et al (2016) Is there a convergence of deciduous leaf litter stoichiometry, biochemistry and microbial population during decay? Geoderma 272:93–100. https://doi.org/10.1016/j.geoderma.2016.03.005
Lowell JL, Gordon N, Engstrom D, Stanford JA, Holben WE, Gannon JE (2009) Habitat heterogeneity and associated microbial community structure in a small-scale floodplain Hyporheic flow path. Microb Ecol 58:611–620. https://doi.org/10.1007/s00248-009-9525-9
Lv Y, Wang C, Jia Y et al (2014) Effects of sulfuric, nitric, and mixed acid rain on litter decomposition, soil microbial biomass, and enzyme activities in subtropical forests of China. Appl Soil Ecol 79:1–9. https://doi.org/10.1016/j.apsoil.2013.12.002
Ma B, Lv X, Cai Y et al (2018) Liming does not counteract the influence of long-term fertilization on soil bacterial community structure and its co-occurrence pattern. Soil Biol Biochem 123:45–53. https://doi.org/10.1016/j.soilbio.2018.05.003
Magasanik B (1993) The regulation of nitrogen utilization in enteric bacteria. J Cell Biochem 51:34–40. https://doi.org/10.1002/jcb.240510108
Mandakovic D, Rojas C, Maldonado J, Latorre M, Travisany D, Delage E, Bihouée A, Jean G, Díaz FP, Fernández-Gómez B, Cabrera P, Gaete A, Latorre C, Gutiérrez RA, Maass A, Cambiazo V, Navarrete SA, Eveillard D, González M (2018) Structure and co-occurrence patterns in microbial communities under acute environmental stress reveal ecological factors fostering resilience. Sci Rep 8:5875. https://doi.org/10.1038/s41598-018-23931-0
Menge DNL, Hedin LO, Pacala SW (2012) Nitrogen and phosphorus limitation over long-term ecosystem development in terrestrial ecosystems. PLoS One 7:e42045. https://doi.org/10.1371/journal.pone.0042045
Mooshammer M, Wanek W, Hämmerle I, Fuchslueger L, Hofhansl F, Knoltsch A, Schnecker J, Takriti M, Watzka M, Wild B, Keiblinger KM, Zechmeister-Boltenstern S, Richter A (2014) Adjustment of microbial nitrogen use efficiency to carbon:nitrogen imbalances regulates soil nitrogen cycling. Nat Commun 5:3694. https://doi.org/10.1038/ncomms4694
Moreno-Vivián C, Cabello P, Martínez-Luque M, Blasco R, Castillo F (1999) Prokaryotic nitrate reduction: molecular properties and functional distinction among bacterial nitrate reductases. J Bacteriol 181:6573–6584. https://doi.org/10.3846/bm.2012.131
Nelson MB, Berlemont R, Martiny AC, Martiny JBH (2015) Nitrogen cycling potential of a grassland litter microbial community. Appl Environ Microbiol 81(20):7012–7022. https://doi.org/10.1128/aem.02222-15
Nelson MB, Martiny AC, Martiny JBH (2016) Global biogeography of microbial nitrogen-cycling traits in soil. Proc Natl Acad Sci 113:8033–8040. https://doi.org/10.1073/pnas.1601070113
Olofsson J, Oksanen L (2002) Role of litter decomposition for the increased primary production in areas heavily grazed by reindeer: a litterbag experiment. Oikos 96:507–515. https://doi.org/10.1034/j.1600-0706.2002.960312.x
Pereira APA, Durrer A, Gumiere T et al (2019) Mixed Eucalyptus plantations induce changes in microbial communities and increase biological functions in the soil and litter layers. For Ecol Manag 433:332–342. https://doi.org/10.1016/j.foreco.2018.11.018
Petersen DG, Blazewicz SJ, Firestone M, Herman DJ, Turetsky M, Waldrop M (2012) Abundance of microbial genes associated with nitrogen cycling as indices of biogeochemical process rates across a vegetation gradient in Alaska. Environ Microbiol 14:993–1008. https://doi.org/10.1111/j.1462-2920.2011.02679.x
Philippot L, Spor A, Hénault C, Bru D, Bizouard F, Jones CM, Sarr A, Maron PA (2013) Loss in microbial diversity affects nitrogen cycling in soil. ISME J 7:1609–1619. https://doi.org/10.1038/ismej.2013.34
Prescott CE, Hope GD, Blevins LL (2003) Effect of gap size on litter decomposition and soil nitrate concentrations in a high-elevation spruce–fir forest. Can J For Res 33:2210–2220. https://doi.org/10.1139/x03-152
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T, Mende DR, Li J, Xu J, Li S, Li D, Cao J, Wang B, Liang H, Zheng H, Xie Y, Tap J, Lepage P, Bertalan M, Batto JM, Hansen T, le Paslier D, Linneberg A, Nielsen HB, Pelletier E, Renault P, Sicheritz-Ponten T, Turner K, Zhu H, Yu C, Li S, Jian M, Zhou Y, Li Y, Zhang X, Li S, Qin N, Yang H, Wang J, Brunak S, Doré J, Guarner F, Kristiansen K, Pedersen O, Parkhill J, Weissenbach J, MetaHIT Consortium, Bork P, Ehrlich SD, Wang J (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464:59–65. https://doi.org/10.1038/nature08821
Remy E, Wuyts K, Verheyen K et al (2018) Altered microbial communities and nitrogen availability in temperate forest edges. Soil Biol Biochem 116:179–188. https://doi.org/10.1016/j.soilbio.2017.10.016
Riggs CE, Hobbie SE (2016) Mechanisms driving the soil organic matter decomposition response to nitrogen enrichment in grassland soils. Soil Biol Biochem 99:54–65. https://doi.org/10.1016/j.soilbio.2016.04.023
Sabri NSA, Zakaria Z, Mohamad SE, Jaafar AB, Hara H (2018) Importance of soil temperature for the growth of temperate crops under a tropical climate and functional role of soil microbial diversity. Microbes Environ 33(2):144–150. https://doi.org/10.1264/jsme2.me17181
Schneider T, Keiblinger KM, Schmid E, Sterflinger-gleixner K (2012) Who is who in litter decomposition ? Metaproteomics reveals major microbial players and their biogeochemical functions 1749–1762. https://doi.org/10.1038/ismej.2012.11
Semmartin M, Aguiar MR, Distel RA et al (2004) Litter quality and nutrient cycling affected by grazing-induced species replacements along a precipitation gradient. Oikos 107:148–160. https://doi.org/10.1111/j.0030-1299.2004.13153.x
Song K, Yu Q, Shang K, Yang T, da LJ (2011) The spatio-temporal pattern of historical disturbances of an evergreen broadleaved forest in East China: a dendroecological analysis. Plant Ecol 212:1313–1325. https://doi.org/10.1007/s11258-011-9907-1
Strickland MS, Osburn E, Lauber C et al (2009) Litter quality is in the eye of the beholder: initial decomposition rates as a function of inoculum characteristics. Funct Ecol 23:627–636. https://doi.org/10.1111/j.1365-2435.2008.01515.x
Summers EA, Paoletti MG, Beggio M et al (2013) Comparative microbial community composition from secondary carbonate (moonmilk) deposits: implications for the Cansiliella servadeii cave hygropetric food web. Int J Speleol 42:181–192. https://doi.org/10.5038/1827-806X.42.3.2
Sun S, Badgley BD (2019) Changes in microbial functional genes within the soil metagenome during forest ecosystem restoration. Soil Biol Biochem 135:163–172. https://doi.org/10.1016/j.soilbio.2019.05.004
Tang Y, Yu G, Zhang X et al (2018) Changes in nitrogen-cycling microbial communities with depth in temperate and subtropical forest soils. Appl Soil Ecol 124:218–228. https://doi.org/10.1016/j.apsoil.2017.10.029
Temu T, Mann M, Räschle M, Cox J (2016) Homology-driven assembly of NOn-redundant protEin sequence sets (NOmESS) for mass spectrometry. Bioinformatics 32:1417–1419. https://doi.org/10.1093/bioinformatics/btv756
Tu Q, Yu H, He Z, Deng Y, Wu L, van Nostrand J, Zhou A, Voordeckers J, Lee YJ, Qin Y, Hemme CL, Shi Z, Xue K, Yuan T, Wang A, Zhou J (2014) GeoChip 4: a functional gene-array-based high-throughput environmental technology for microbial community analysis. Mol Ecol Resour 14:914–928. https://doi.org/10.1111/1755-0998.12239
Tu Q, He Z, Wu L et al (2017) Metagenomic reconstruction of nitrogen cycling pathways in a CO2-enriched grassland ecosystem. Soil Biol Biochem 106:99–108. https://doi.org/10.1016/j.soilbio.2016.12.017
Urbanová M, Šnajdr J, Baldrian P (2015) Composition of fungal and bacterial communities in forest litter and soil is largely determined by dominant trees. Soil Biol Biochem 84:53–64. https://doi.org/10.1016/j.soilbio.2015.02.011
Vaieretti MV, Cingolani AM, Pérez Harguindeguy N, Cabido M (2013) Effects of differential grazing on decomposition rate and nitrogen availability in a productive mountain grassland. Plant Soil 371:675–691. https://doi.org/10.1007/s11104-013-1831-9
Wang C, Guo P, Han G, Feng X, Zhang P, Tian X (2010) Effect of simulated acid rain on the litter decomposition of Quercus acutissima and Pinus massoniana in forest soil microcosms and the relationship with soil enzyme activities. Sci Total Environ 408:2706–2713. https://doi.org/10.1016/j.scitotenv.2010.03.023
Wang W, Chen D, Sun X et al (2019) Impacts of mixed litter on the structure and functional pathway of microbial community in litter decomposition. Appl Soil Ecol 144:72–82. https://doi.org/10.1016/j.apsoil.2019.07.006
Wong KH, Hynes MJ, Davis MA (2008) Recent advances in nitrogen regulation: a comparison between Saccharomyces cerevisiae and filamentous fungi. Eukaryot Cell 7:917–925. https://doi.org/10.1128/EC.00076-08
Xiao X, Yin X, Lin J, Sun L, You Z, Wang P, Wang F (2005) Chitinase genes in lake sediments of Ardley Island, Antarctica. Appl Environ Microbiol 71:7904–7909. https://doi.org/10.1128/AEM.71.12.7904-7909.2005
Xie J, He Z, Liu X, Liu X, van Nostrand J, Deng Y, Wu L, Zhou J, Qiu G (2011) GeoChip-based analysis of the functional gene diversity and metabolic potential of microbial communities in acid mine drainage. Appl Environ Microbiol 77:991–999. https://doi.org/10.1128/AEM.01798-10
Xiong Y, Fan P, Fu S et al (2013) Slow decomposition and limited nitrogen release by lower order roots in eight Chinese temperate and subtropical trees. Plant Soil 363:19–31. https://doi.org/10.1007/s11104-012-1290-8
Xu M, Zhang Q, Xia C, Zhong Y, Sun G, Guo J, Yuan T, Zhou J, He Z (2014) Elevated nitrate enriches microbial functional genes for potential bioremediation of complexly contaminated sediments. ISME J 8:1932–1944. https://doi.org/10.1038/ismej.2014.42
Xu J, Liu S, Song S et al (2018) Arbuscular mycorrhizal fungi influence decomposition and the associated soil microbial community under different soil phosphorus availability. Soil Biol Biochem 120:181–190. https://doi.org/10.1016/j.soilbio.2018.02.010
Zeng L, He W, Teng M, Luo X, Yan Z, Huang Z, Zhou Z, Wang P, Xiao W (2018) Effects of mixed leaf litter from predominant afforestation tree species on decomposition rates in the three gorges reservoir, China. Sci Total Environ 639:679–686. https://doi.org/10.1016/j.scitotenv.2018.05.208
Zeng Q, Liu Y, Zhang H, An S (2019) Fast bacterial succession associated with the decomposition of Quercus wutaishanica litter on the loess plateau. Biogeochemistry 144:119–131. https://doi.org/10.1007/s10533-019-00575-4
Zhalnina K, Dias R, de Quadros PD, Davis-Richardson A, Camargo FA, Clark IM, McGrath S, Hirsch PR, Triplett EW (2015) Soil pH determines microbial diversity and composition in the park grass experiment. Microb Ecol 69:395–406. https://doi.org/10.1007/s00248-014-0530-2
Zhang L, Adams JM, Dumont MG et al (2019a) Distinct methanotrophic communities exist in habitats with different soil water contents. Soil Biol Biochem 132:143–152. https://doi.org/10.1016/j.soilbio.2019.02.007
Zhang W, Yang K, Lyu Z, Zhu J (2019b) Microbial groups and their functions control the decomposition of coniferous litter: a comparison with broadleaved tree litters. Soil Biol Biochem. https://doi.org/10.1016/j.soilbio.2019.03.009
Zhou J, He Z, Yang Y et al (2015) High-throughput metagenomic technologies for complex microbial community analysis: open and closed formats. MBio 6:e02288–e02214. https://doi.org/10.1128/mBio.02288-14
Acknowledgements
This research was funded by The National Key Research and Development Program of China (Project no. 2017YFD0600401).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Responsible Editor: Feike A. Dijkstra.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, W., Chen, D., Zhang, Q. et al. Effects of mixed coniferous and broad-leaved litter on bacterial and fungal nitrogen metabolism pathway during litter decomposition. Plant Soil 451, 307–323 (2020). https://doi.org/10.1007/s11104-020-04523-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-020-04523-2